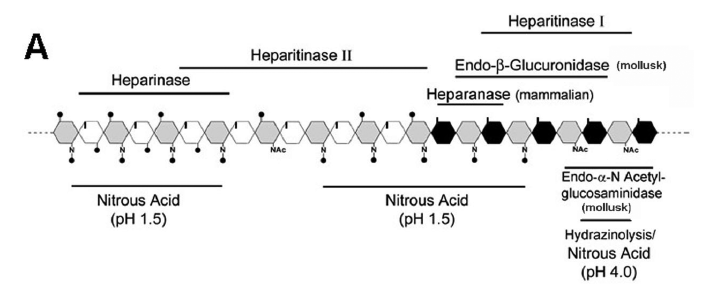
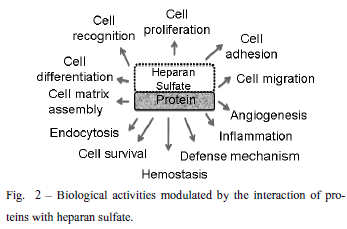
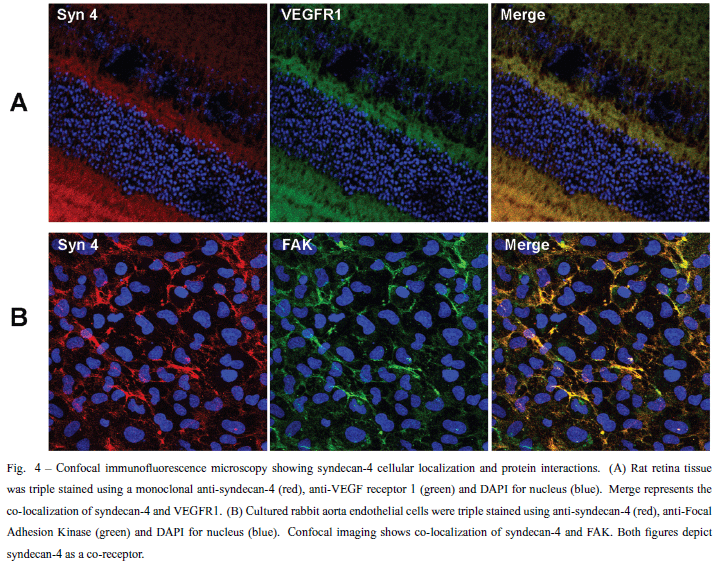
Heparan sulfate proteoglycans are ubiquitously found at the cell surface and extracellular matrix in all the animal species. This review will focus on the structural characteristics of the heparan sulfate proteoglycans related to protein interactions leading to cell signaling. The heparan sulfate chains due to their vast structural diversity are able to bind and interact with a wide variety of proteins, such as growth factors, chemokines, morphogens, extracellular matrix components, enzymes, among others. There is a specificity directing the interactions of heparan sulfates and target proteins, regarding both the fine structure of the polysaccharide chain as well precise protein motifs. Heparan sulfates play a role in cellular signaling either as receptor or co-receptor for different ligands, and the activation of downstream pathways is related to phosphorylation of different cytosolic proteins either directly or involving cytoskeleton interactions leading to gene regulation. The role of the heparan sulfate proteoglycans in cellular signaling and endocytic uptake pathways is also discussed.
glycosaminoglycans and protein interactions; growth factors; focal adhesion; extracellular matrix; cell cycle; cell proliferation