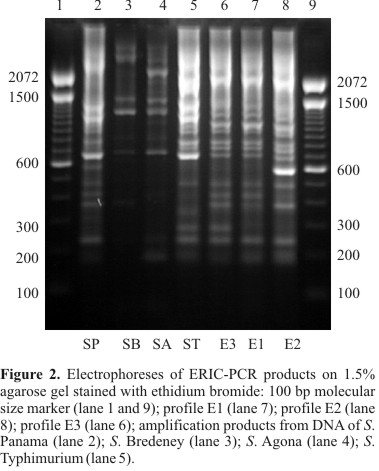
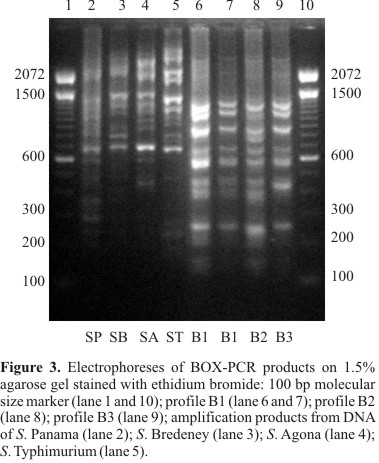
In order to study the epidemiology of Salmonella Enteritidis outbreaks and determine the source of contamination so that a recurrence can be avoided, detailed characterization is necessary. Thus, the purpose of this study was to verify whether rep-PCR was able to discriminate among Salmonella Enteritidis isolates. Phage typing, detection of virulence genes and antimicrobial resistance testing were also associated to rep-PCR results. One hundred and two S. Enteritidis isolates from broiler carcasses, food, human, pigs, poultry-related samples, and nine isolates from other countries were genotypically typed by REP-PCR, ERIC-PCR and BOX-PCR, collectively called rep-PCR. Phage typing, detection of virulence genes and antimicrobial resistance testing were also performed. Only three fingerprinting profiles were obtained with each rep-PCR method, with the majority of isolates belonging to the same profile. No relationship was observed between genotypic profile and year, place of isolation or source of infection. However, the less frequent rep-PCR profiles showed single antimicrobial resistance patterns. Although few strains isolated from swine were analyzed, different antimicrobial resistance patterns were observed. Furthermore, phage type 4 was not found in swine isolates. rep-PCR showed a lower discriminatory power as compared with antimicrobial resistance and phage typing, but the combination of genotypic and phenotypic methods was more discriminatory than any method alone, resulting in 48 different types.
Salmonella Enteritidis; rep-PCR; phage typing; antimicrobial resistance; virulence genes