Abstracts
Neosporosis is an important cause of abortion in cattle and information on their genetics and host parasite relationships are desirable. Neospora caninum samples obtained from 24 bovine fetuses from Minas Gerais, were genetically analyzed in part of the rDNA region, coding for rRNAs. Previously, brain, heart, liver, skeletal muscle, lung, kidney, spleen, thymus, lymph nodes, ovary or testis, uterus and skin of the ear were analyzed by conventional histopathology and immunohistochemistry. Of these, eight had lesions compatible in the brain and heart and two also in skeletal muscle and liver. Three brains showed antigen identification in immunohistochemistry. Brain and heart tissues were subjected to DNA extraction for PCR, whose product of 588 bp of ITS-1 region was sequenced in three samples. We obtained 96% similarity with dozens of sequences N. caninum deposited in GenBank. The phylogenetic tree showed great conservation among isolates of N. caninum in this study and those deposited in GenBank, while well-defined and specific branches were generated against other Apicomplexa. PCR for this region is useful as a diagnosis, with good analytical specificity, but the ITS-1 region is not suitable for genetic differentiation intra species because the sequences obtained were identical to the others analyzed.
Bovine; diagnosis; phylogeny; Neospora caninum; PCR
A neosporose é uma importante causa de abortos em bovinos, e informações sobre sua genética e relação parasito-hospedeiro são desejáveis. Amostras de Neospora caninum, obtidas de 24 conceptos bovinos oriundos de Minas Gerais, foram analisadas geneticamente em parte da região rDNA, codificadora de rRNAs. Previamente, cérebro, coração, fígado, músculo esquelético, pulmão, rim, baço, timo, linfonodos, ovário ou testículo, útero e pele da orelha foram analisados por histopatologia convencional e imuno-histoquímica. Dessas, oito apresentaram lesões compatíveis no encéfalo e coração e dois também no músculo esquelético e fígado. Três cérebros apresentaram identificação antigênica na imuno-histoquímica. Tecidos cerebral e cardíaco foram submetidos a extração de DNA para PCR, cujo produto de 588 pb da região ITS-1 foi sequenciado em três amostras. Obteve-se similaridade de 96% com dezenas de sequências de N. caninum depositadas no GenBank. A árvore filogenética mostrou grande conservação entre isolados de N. caninum deste estudo e aqueles depositadas no GenBank, enquanto ramos bem definidos e específicos foram gerados em relação a outros apicomplexa. A PCR para esta região é útil como diagnóstico, com boa especificidade analítica, mas a região ITS-1 não é apropriada para a diferenciação genética intraespécie, pois as sequências obtidas foram idênticas às demais analisadas.
Bovino; diagnóstico; filogenia; Neospora caninum; PCR
FULL ARTICLE
Neospora caninum in bovine fetuses of Minas Gerais, Brazil: genetic characteristics of rDNA
Neospora caninum em fetos bovinos de Minas Gerais, Brasil: características genéticas do rDNA
Domingos Sávio dos SantosI; Michele Placedino AndradeII; Mary Suzan VaraschinIII; Antônio Marcos GuimarãesIII; Christian HirschIII
IInstituto de Ensino Superior Presidente Tancredo de Almeida Neves IPTAN
IIUniversidade Federal de Lavras UFLA
IIIDepartamento de Medicina Veterinária, Universidade Federal de Lavras UFLA
Corresponding author Corresponding author: Christian Hirsch Professor do Departamento de Medicina Veterinária Universidade Federal de Lavras UFLA, Campus Universitário, s/nº, CP 3037 CEP 37200-000, Lavras, MG, Brazil e-mail: chhirsch@dmv.ufla.br
ABSTRACT
Neosporosis is an important cause of abortion in cattle and information on their genetics and host parasite relationships are desirable. Neospora caninum samples obtained from 24 bovine fetuses from Minas Gerais, were genetically analyzed in part of the rDNA region, coding for rRNAs. Previously, brain, heart, liver, skeletal muscle, lung, kidney, spleen, thymus, lymph nodes, ovary or testis, uterus and skin of the ear were analyzed by conventional histopathology and immunohistochemistry. Of these, eight had lesions compatible in the brain and heart and two also in skeletal muscle and liver. Three brains showed antigen identification in immunohistochemistry. Brain and heart tissues were subjected to DNA extraction for PCR, whose product of 588 bp of ITS-1 region was sequenced in three samples. We obtained 96% similarity with dozens of sequences N. caninum deposited in GenBank. The phylogenetic tree showed great conservation among isolates of N. caninum in this study and those deposited in GenBank, while well-defined and specific branches were generated against other Apicomplexa. PCR for this region is useful as a diagnosis, with good analytical specificity, but the ITS-1 region is not suitable for genetic differentiation intra species because the sequences obtained were identical to the others analyzed.
Keywords: Bovine, diagnosis, phylogeny, Neospora caninum, PCR.
RESUMO
A neosporose é uma importante causa de abortos em bovinos, e informações sobre sua genética e relação parasito-hospedeiro são desejáveis. Amostras de Neospora caninum, obtidas de 24 conceptos bovinos oriundos de Minas Gerais, foram analisadas geneticamente em parte da região rDNA, codificadora de rRNAs. Previamente, cérebro, coração, fígado, músculo esquelético, pulmão, rim, baço, timo, linfonodos, ovário ou testículo, útero e pele da orelha foram analisados por histopatologia convencional e imuno-histoquímica. Dessas, oito apresentaram lesões compatíveis no encéfalo e coração e dois também no músculo esquelético e fígado. Três cérebros apresentaram identificação antigênica na imuno-histoquímica. Tecidos cerebral e cardíaco foram submetidos a extração de DNA para PCR, cujo produto de 588 pb da região ITS-1 foi sequenciado em três amostras. Obteve-se similaridade de 96% com dezenas de sequências de N. caninum depositadas no GenBank. A árvore filogenética mostrou grande conservação entre isolados de N. caninum deste estudo e aqueles depositadas no GenBank, enquanto ramos bem definidos e específicos foram gerados em relação a outros apicomplexa. A PCR para esta região é útil como diagnóstico, com boa especificidade analítica, mas a região ITS-1 não é apropriada para a diferenciação genética intraespécie, pois as sequências obtidas foram idênticas às demais analisadas.
Palavras-chave: Bovino, diagnóstico, filogenia, Neospora caninum, PCR.
Introduction
Neospora caninum is an obligate intracellular parasite that causes abortions in cattle, in many countries (ANDERSON et al., 2000). It was first identified as a protozoon similar to Toxoplasma gondii in dogs with nervous symptoms, in Norway (BJERKAS et al., 1984), and only later on was it classified by Dubey et al. (1988) as a new genus and as the species N. caninum (Apicomplexa: Sarcocystidae).
Dogs - Canis lupus familiaris (McALLISTER et al., 1998), coyotes - Canis latrans (GONDIM et al., 2004) and dingoes Canis lupus dingo (KING et al., 2010) are the definitive hosts and eliminate oocysts in their feces after ingesting tissue cysts of N. caninum that are present in intermediate hosts (DUBEY et al., 2007).
The most evident clinical sign of bovine neosporosis is abortion, at any stage of gestation, but the majority of the cases occur in the sixth month of gestation (DUBEY, 2003). The diagnosis is based on detecting anti-N. caninum antibodies in adult animals and viewing suggestive lesions in the brain, heart and liver of aborted fetuses, in association with immunohistochemical, fetal serological and PCR examinations (ORTEGA-MORA et al., 2006).
Recently, several cases of bovine neosporosis have been reported in South America, including in Brazil (MOORE, 2005; GARCÍA-MELO et al., 2009).
Phylogenetic investigations on N. caninum contribute towards expanding the knowledge of the characteristics and behavior of samples of the parasite in Brazil (SANTOS et al., 2010). The gene region ITS-1 has become the target for phylogenetic studies, since this region experiences greatest mutagenic evolutionary pressure and is subject to polymorphism (PAYNE and ELLIS, 1996; AL-QASSAB et al., 2009; AL-QASSAB et al., 2010; SANTOS et al., 2010). Despite these studies, there are still few gene sequences of ITS-1 deposited in GenBank, including those originating from Brazil. The aim of the present study was to investigate occurrences of genetic groupings of N. caninum in bovine fetuses in the state of Minas Gerais, in relation to the gene region that codes for rRNA.
Material and Methods
Sixteen fetuses and eight calves of between one and 120 days of age were sent to the Veterinary Pathology Sector of the Federal University of Lavras (SPV/UFLA), between December 2007 and December 2009. The sample selection criterion was occurrences of reproductive abnormalities in dairy cattle herds, with a history of regular vaccination of calves against brucellosis and heifers and cows against the infectious bovine rhinotracheitis virus, bovine diarrhea virus and leptospirosis. As a complementary analysis, an immunohistochemical technique for parasite antigens in fetal tissues was standardized and applied. All the procedures used in this study had been approved by the Ethics Committee for Animal Use of the Federal University of Lavras (procedural no. 030/2010).
After identification, the fetus was measured using a measuring tape. The length of spine was measured from the atlanto-occipital joint to the end of the sacrum, with the aim of determining the fetal age, as recommended by Roberts (1986) and Barr et al. (1990). The age was also obtained from the service records.
Necropsies were performed at SPV/UFLA, where several types of tissue were collected for histopathological examination: central nervous system (cortex of the cerebral hemispheres, cerebellum, thalamus, rostral colliculus, caudal colliculus, cerebellar peduncles, obex and cervical medulla), heart, skeletal muscle (semitendinosus and semimembranosus), liver, lung, kidney, spleen, thymus, lymph nodes, ovary, testicle, uterus and ear skin. The placentas were not examined because only the fetuses were sent to SPV/UFLA. The tissue samples were fixed in buffered 10% formol (pH 7.2 and 0.01 M). After fixing, the material was embedded in paraffin, from which sections of thickness 5 µm were made. These were stained using the hematoxylin-eosin (HE) technique. N. caninum and lesions were investigated by means of optical microscopy.
The tissues collected were also subjected to immunohistochemical examination. For this, the streptavidin-biotin-peroxidase method was used, by means of a commercial kit (Dako LSAB + Peroxidase kit). Antigen recovery was done by means of enzymatic digestion using 0.1% trypsin (pH 7.8) for 10 minutes at 37 ºC. Subsequently, the slides were immersed in citrate buffer (pH 6.0) and irradiated in a domestic microwave oven for two minutes, at maximum power. The sections were marked using anti-N. caninum polyclonal primary antibodies, produced in goats, at a dilution of 1:2200, and anti-T. gondii (both from VRMD, PULLMAN, USA), at a dilution of 1:2400 (as a differential diagnosis), in accordance with the technique used by Mills (1992), as modified by Corbellini et al. (2002). The final staining was obtained using the chromogen DAB (3,3-diaminobenzidine, Dako). The positive control consisted of cysts of N. caninum from goats and the negative control was created by replacing the primary antibodies with beef serum.
During the necropsy, samples from the frontal lobes of the cerebral hemispheres and from the apex of the heart were collected from 24 cattle and were frozen. These samples were then subjected to DNA extraction and PCR examination. The procedures were performed in the Virology Laboratory of the Department of Veterinary Medicine of UFLA.
To obtain DNA from the samples, three different protocols were used: two based on the phenol-chloroform-isoamyl alcohol method (P:C:IA) and one on the affinity of DNA for silica, using a tissue genomic DNA extraction kit (Invisorb Spin Tissue Mini Kit, Invitek, Germany). The three protocols used the enzyme proteinase K for tissue digestion. The first protocol based on phenol was the one put forward by Sambrook (1989) (protocol A), while the second was supplied by the researcher Marcelo Fernandes Camargos (LANAGRO de Pedro Leopoldo, MG, personal communication protocol B). In addition to the difference in the initial tissue volumes (800 mg for protocol A and 250 mg for protocol B), protocol A used 100 µg.mL1 of proteinase K in the lysis reaction, followed by incubation at 50 ºC for 3 hours, with mixing of the solution from time to time. On the other hand, protocol B used 150 µg.mL1 of the enzyme in the reaction, which was incubated overnight at 60 ºC and then at 95 ºC for 10 minutes.
For PCR, the reactions were standardized to a final volume of 20 µL and were adjusted proportionally when larger volumes were used. The PCR reagents, except for water, came from the "PCR Core System I" kit (PROMEGA Corporation, USA). The reactions were standardized by means of experimenting with the MgCl2 concentrations and the alignment temperature of the primers, from data supplied by the Primer3 primer selection software (ROZEN; SKALETSKY, 1998) and BLAST software (ALTSCHUL et al., 1990), which are both available through the website of the National Center for Biotechnology Information (NCBI, 2011a).
The condition defined for PCR was an MgCl2 concentration of 1.5 mM, annealing temperature of 49 ºC for 30 seconds. The quantity of Taq DNA polymerase was 0.5 units and the DNA template was 100 ng. The concentration of primer per reaction was one 1 picomole/µL, adjusted to a final volume of 20 µL. All the PCR runs were done in a PT100 thermocycler (MJ Research Incorporated, USA), programmed in the following manner: one cycle of 95 ºC for five minutes, followed by 35 cycles of 95 ºC for 30 seconds (opening of the DNA ribbons), with primer alignment time and temperature of 30 seconds and 49 ºC, respectively. Then one cycle at 72 ºC for one minute, for extension, and concluding with a final extension period of 72 ºC for seven minutes and 4 ºC until removal.
For the PCR directed towards part of the ribosomal region of N. caninum (PCR-RR-NC), the primers Lav 1 (sense) 5'-CGGAAGGATCATTCACACG-3', which was designed in this study, and Tim 11 (antisense) 5'-CCCACTGAAACAGACGTACC-3', described by Payne and Ellis (1996), were used. The expected product from this PCR run presented 588 bp was situated between positions 4 and 592, within the region of ribosomal DNA of N. caninum. Sequence L49389 was used as the reference, as presented in Figure 3. The negative control for the PCR-RR-NC was DNA extracted from cultures of MDBK cells that had not been infected by the parasite. The positive control was standard DNA from the N. caninum sample NC-1, extracted from cultures of infected VERO cells.
To verify the PCR run, the products were subjected to electrophoresis on 1.5% agarose gel, in the presence of TAE (Tris-acetate-EDTA) running buffer solution for 1 hour at 100V. The DNA bands were viewed using ethidium bromide (0.5 µg.mL1) (Vilber Lourmat apparatus, France), and the images were capture using an L-Pix Chemi photo digitizer (Loccus Biotecnologia, Brazil), for subsequent editing and analysis.
The purified product (GenElute PCR Clean-Up kit, Sigma) was subjected to sequencing analysis for subsequent confirmation of the specificity. The sequencing process was based on the dideoxy chain termination method described by Sanger et al. (1977), using the automated sequencing apparatus available at LCBM-UFLA. The chromatogram sequences were analyzed with the aid of the Chromas software (Technelysium Pty, Australia) and were exported to Notepad files (Microsoft, USA). Confirmation of the specificity of the sequences obtained was confirmed by means of the BlastN software (ALTSCHUL et al., 1990), which is available on the NCBI website (NCBI, 2011b).
After the sequences had been edited to remove ambiguities and consensuses of the same size had been obtained, geometric analyses were undertaken with the aid of alignment through the ClustalW software (JEANMOUGIN et al., 1998). This was then fed into the MEGA4 software (KUMAR et al., 2004). For these analyses, a phylogenetic tree was constructed, obtained through the Neighbor-Joining method (SAITOU; NEI, 1987), using a bootstrap of 1000 replicates and an evolutive distance parameter, adjusted using the Kimura 2-parameter nucleotide replacement method.
Results
The 24 animals that participated in the sample were named A to X. Of these 24 cattle, eight presented microscopic lesions suggestive of infection due to N. caninum, i.e. with the presence of mononuclear inflammatory infiltrate with or without associated necrosis in several tissues. In the central nervous system (CNS), the presence of gliosis was also taken into consideration. Microscopic lesions suggestive of neosporosis were found mainly in the encephalon and heart, followed by the skeletal muscles and liver.
In the CNS, histological lesions were found throughout the encephalon and, in one animal, in the spinal cord. In the cerebral cortex, non-suppurative encephalitis characterized by mononuclear perivascular cuffs was observed in animals A and C); tissue necrosis, easily seen through tissue eosinophilia (animal B) and gliosis (animals B, A, C, R and T). Slight non-suppurative meningitis was only observed in one animal (animal A). In the cerebellum, gliosis was observed in animals A, B and T, and tissue necrosis in animals A, B and T, with mononuclear cells surrounding the necrotic area (animal A), along with slight perivascular cuffing (animals A and T). Only in one animal was a slight focus of inflammatory infiltrate observed in the grey matter of the spinal cord (animal R). In the heart, non-suppurative myocarditis characterized by multifocal areas with slight to marked mononuclear inflammatory infiltrate was observed in all the fetuses and in two of the calves. A single fetus also presented moderate non-suppurative pericarditis (animal T).
In the skeletal muscles, multifocal areas of non-suppurative myositis were observed in animals C, Q and T, and parasitic structures resembling a young cyst in skeletal fibers were observed in animal T. In the liver, multifocal areas of necrosis and mononuclear inflammatory infiltrate were observed in the portal space, in animals A and T.
The immunohistochemical marking was positive only for N. caninum and occurred in three of the eight fetuses with lesions compatible with neosporosis. Tachyzoites and cysts were marked in the encephalon of these three animals, and one of them also had marking of tachyzoites in skeletal muscles (Figure 1). The association of compatible lesions with immunohistochemical marking and PCR is demonstrated in Table 1.
Protocols based on phenol were tested because of the significantly greater tissue mass used in the extraction, considering that the silica-based kit only extracts DNA from a maximum of 40 mg of tissue. Since the parasite distribution in the infected tissues seems to be random, a greater initial tissue mass would probably increase the chance of extracting DNA from the parasite. However, protocols A and B, based on using phenol, presented low yield. Thus, it was decided to use the protocol based on DNA affinity for silica, for all the samples (Invisorb Spin Tissue Mini Kit, Invitek, Germany).
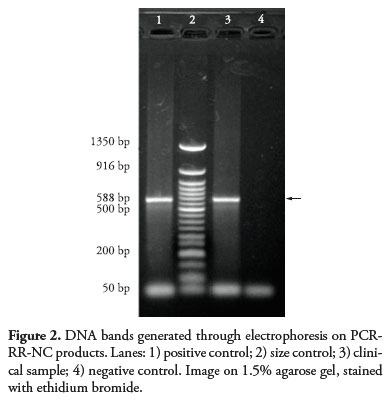
PCR corresponding to the region that codes for rRNA of N. caninum (PCR-RR-NC) was applied to 24 brain samples and 24 myocardial samples. Of these, five brain tissue samples presented positive results for the DNA of N. caninum (animals A, C, M, R and T). Figure 2 presents the image of the result from the PCR-RR-NC, which was characterized by formation of bands with 588 bp. The sequencing analysis on the purified product from these bands, which was done using the BlastN software, confirmed that sequences specific to the protozoon N. caninum were present, with a similarity rate of more than 96%.
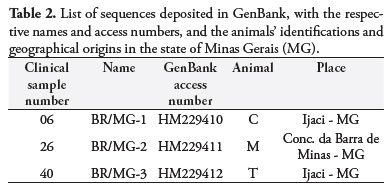
The five samples that were positive from PCR-RR-NC were again processed using this PCR, in volumes of 50 µL, and underwent subsequent extraction of the amplified product available in the bands, for direct sequencing. The result obtained, after editing to remove ambiguities and achieve consensuses, made it possible to analyze three sequences of 346 bp each, from animals C, M and T. These were deposited in GenBank under the registration numbers HM229410, HM229411 and HM229412, respectively. Table 2 presents the relationship between the sequences deposited and their geographical origin. Figure 3 represents the gene position of the DNA fragment of 346 bp, within the region that codes for rDNA of N. caninum. The sequences obtained present 100% similarity with each other and were used to feed into the MEGA4 software and generate the phylogenetic tree (Figure 4). The mutations detected among the samples obtained in the present study and in the samples available in GenBank, including the reference sequence L49389, are discriminated in Table 3.
Discussion
The histopathological findings of non-suppurative encephalitis associated with tissue necrosis and gliosis, and the myositis and myocarditis found in the bovine fetuses of this study, are in accordance with the lesions described for neosporosis (LINDSAY et al., 1999; HELMAN et al., 1998; CORBELLINI et al., 2002; DUBEY, 2003; ORTEGA-MORA et al., 2006), as were the hepatic lesions of periportal mononuclear infiltrate and foci of hepatocellular necrosis (BARR et al., 1990).
In the study by Corbellini et al. (2000), on eight bovine tissue samples with lesions compatible with N. caninum, four showed positive results from immunohistochemistry, of which one was a case in the heart and four in the brain. These results, compared with those obtained in the present study, reinforce the idea that the nervous tissue is the material of choice for diagnosing neosporosis (HELMAN et al., 1998; CORBELLINI et al., 2002; DUBEY, 2003; CABRAL et al., 2009).
With the results from this PCR run, a correlation and concordance analysis was conducted on the data obtained from histopathology and immunohistochemistry, and this is presented in Table 1. From this analysis, it could be seen that out of the eight samples with lesions, five presented amplification through PCR, which surpassed the efficiency of the immunohistochemistry, with three positive results. These findings corroborate the data presented by Baszler et al. (1999), which thus indicates that PCR is a technique of greater sensitivity than shown by immunohistochemistry, in diagnostic terms.
The percentage of intraspecies mutations was relatively low (4%) in this study. This information diverges from the findings of Payne and Ellis (1996), who stated that the ITS-1 of N. caninum was a region of polymorphism that was tolerant to gene mutations. However, in comparison with the interspecies mutations relating to the subfamily Toxoplasmatinae, the PCR-RR-NC of the present study was a highly specific diagnostic tool.
Following the study by Payne and Ellis (1996), relating to the DNA region that codes for rRNA of N. caninum, several other studies have been published. However, in specifically analyzing the ITS-1 region, few studies have demonstrated nucleotide variations (AL-QASSAB et al., 2010). Similar findings were reported by Santos et al. (2010), using sequences deposited in GenBank.
Analysis on the alignment of the sequences of N. caninum observed in this study with the other deposited in GenBank showed that equivalence was maintained. This homology between strains of N. caninum from different geographical areas is probably due to high volumes of movement and commercialization of animals from one region to another, and consequent distribution of a genetically dominant and homogenous strain.
According to Payne and Ellis (1996), the ITS-1 region of N. caninum is tolerant to mutations. However, the sequences obtained in this study referring to this region remained unchanged. This conservation of sequences probably occurred because of control over the mutagenic pressure, influenced by the immunity complex (AL-QASSAB et al., 2010). This conservation was reflected in the phylogenetic tree, which was characterized by presenting a single grouping of N. caninum, without distinguishing the geographical origin or type of host. These were also the variables obtained in the literature and comparisons are therefore possible.
Graphically, this conservation of sequences was demonstrated in the formation of the phylogenetic tree represented in Figure 4, in which branches with terminal nodes for N. caninum formed an internal node. The presence of a single branch with a terminal node, represented by the sequence of Payne and Ellis (1996), which was separated from the others because of the variations presented in Table 3, indicated that this sequence was particularly different from the others. This phylogenetic tree also promoted formation of branches to parasite species of the subfamily Toxoplasmatinae, composed of the following species: Neospora caninum, Neospora hughesi, Toxoplasma gondii and Hammondia hammondi, which were confirmed by high percentages and well defined separations of branches.
The specificity formed by the tree demonstrates that this rDNA region is important for separating parasitic protozoa of the subfamily Toxoplasmatinae that are very similar, which confirms that PCR-RR-NC is an efficient diagnostic tool, with regard to analytical specificity.
In the present study, firstly, investigation of genotype variability showed that the gene sequences were highly conserved. The lack of group formation suggests that the strains of N. caninum in cattle in the Lavras region are analogous to those in other parts of the world that have been applied in constructing the phylogenetic tree regarding the gene region ITS-1. Thus, new investigations in other gene regions of N. caninum are needed in order to confirm whether this conservation is normal in the genome of the parasite, or whether there are any gene regions that would be more appropriate for phylogenetic studies. Since knowledge of the genetic characteristics of N. caninum is still being acquired, and because this protozoon has only relatively recently been described (DUBEY et al., 1988), together with the small number of sequences deposited in GenBank, this field of research is completely open to investigation (AL-QASSAB et al., 2010).
The genus Neospora is composed of the species N. caninum and N. hughesi. In analyzing the phylogenetic tree, it can be seen that these two species have become distanced evolutionally from each other in relation to this region coding for rRNA. Only a few studies on N. hughesi are available, but this horse parasite has now been included in this genus (ÁLVAREZ-GARCÍA, 2003; TOSCAN et al., 2010). In the phylogenetic tree of the present study, formation of separate branches for the genus Neospora was observed. In fact, N. hughesi came much closer to the genera Toxoplasma and Hammondia, and this observation should be examined attentively as new gene sequences for this protozoon become available in GenBank. Thus, future studies will make it possible to better elucidate the taxonomic classification of the subfamily Toxoplasmatinae.
Acknowledgements
To the Research Support Foundation of the State of Minas Gerais (FAPEMIG), for financial assistance for carrying out this research (No. CVZ APQ-7963-5.04/07); and to Prof. Dr. Solange M. Gennari of FMVZ-USP, for donating Vero cells and the reference NC-1 sample.
Received February 21, 2011
Accepted May 10, 2011
- AL-QASSAB, S. et al. Genetic diversity amongst isolates of Neospora caninum, and the development of a multiplex assay for the detection of distinct strains. Molecular and Cellular Probes, v. 23, n. 3-4, p. 132-139, 2009.
- AL-QASSAB, S. E.; REICHEL, M. P.; ELLIS, J. T. On the Biological and Genetic Diversity in Neospora caninum. Diversity, v. 2, n. 3, p. 411-438, 2010. http://dx.doi.org/10.3390/d2030411
- ALTSCHUL, S. F. et al. Basic local alignment search tool. Journal of Molecular Biology, v. 215, n. 3, p. 403-410, 1990.
- ÁLVAREZ-GARCÍA, G. Identificación y caracterización de antígenos de "Neospora caninum" con interés inmunodiagnóstico en bovinos 2003. 302 f. Tese (Doutorado em Veterinária)-Universidad Complutense de Madrid, Madrid.
- ANDERSON, M. L.; ANDRIANARIVO, A. G.; CONRAD, P. A. Neosporosis in cattle. Animal Reproduction Science, v. 60-61, p. 417-431, 2000. http://dx.doi.org/10.1016/S0378-4320(00)00117-2
- BARR, C. B. et al. Bovine fetal encephalitis and myocarditis associated with protozoal infections. Veterinary Pathology, v. 27, n. 5, p. 354-361, 1990. PMid:2238388. http://dx.doi.org/10.1177/030098589002700508
- BASZLER, T. V. et al. Interferon-γ and interleukin-12 mediate protection to acute Neospora caninum infection in BALB/c mice. International Journal for Parasitology, v. 29, n. 10, p. 1635-1646, 1999. http://dx.doi.org/10.1016/S0020-7519(99)00141-1
- BJERKAS, I.; MOHN, S. F.; PRESTHUS, J. Unidentified cyst-forming sporozoon causing encephalomyelitis and myositis in dogs. Parasitology Research, v. 70, n. 2, p. 271-274, 1984.
- CABRAL, A. D. et al. Diagnosis of Neospora caninum in bovine fetuses by histology, immunohistochemistry, and nested-PCR. Revista Brasileira de Parasitologia Veterinária, v. 18, n. 4, p. 14-19, 2009. http://doi.editoracubo.com.br/10.4322/rbpv.01804003
- CORBELLINI, L. G. et al. Aborto bovino por Neospora caninum no Rio Grande do Sul. Ciência Rural, v. 30, n. 5, p. 863-868, 2000. http://dx.doi.org/10.1590/S0103-84782000000500021
- CORBELLINI, L. G. et al. Neosporosis as a cause of abortion in dairy cattle in Rio Grande do Sul, Southern Brazil. Veterinary Parasitology, v. 103, n. 3, p. 195-202, 2002. http://dx.doi.org/10.1016/S0304-4017(01)00600-8
- DUBEY, J. P. et al. Newly recognized fatal protozoan disease of dogs. Journal of the American Veterinary Medical Association, v. 192, n. 9, p. 1269-1285, 1988. PMid:3391851.
- DUBEY, J. P.; LINDSAY, D. S. A review of Neospora caninum and neosporosis. Veterinary Parasitology, v. 67, n. 1-2, p. 1-59, 1996. http://dx.doi.org/10.1016/S0304-4017(96)01035-7
- DUBEY, J. P. Review of Neospora caninum and neosporosis in animals. The Korean Journal of Parasitology, v. 41, n. 1, p. 1-16, 2003. PMid:12666725. PMCid:2717477. http://dx.doi.org/10.3347/kjp.2003.41.1.1
- DUBEY, J. P.; SCHARES, G.; ORTEGA-MORA, L. M. Epidemiology and control of neosporosis and Neospora caninum. Clinical Microbiology Reviews, v. 20, n. 2, p. 323-367, 2007. PMid:17428888. PMCid:1865591. http://dx.doi.org/10.1128/CMR.00031-06
- GARCÍA-MELO, D. P. et al. Isolation and biological characterisation of a new isolate of Neospora caninum from an asymptomatic calf in Brazil. Acta Parasitologica, v. 54, n. 2, n. 180-185, 2009.
- GONDIM, L. F. P. et al. Coyotes (Canis latrans) are definitive hosts of Neospora caninum International Journal for Parasitology, v. 34, n. 2, p. 159-161, 2004. PMid:15037103. http://dx.doi.org/10.1016/j.ijpara.2004.01.001
- HELMAN, R. G. et al. Neosporal abortion in Oklahoma cattle with emphasis on the distribution of brain lesions in aborted fetuses. Journal of Veterinary Diagnostic Investigation, v. 10, n. 3, p. 292-295, 1998. PMid:9683083. http://dx.doi.org/10.1177/104063879801000314
- KING, J. S. et al. Australian dingoes are definitive hosts of Neospora caninum International Journal for Parasitology, v. 40, n. 8, p. 945-950, 2010. PMid:20149793. http://dx.doi.org/10.1016/j.ijpara.2010.01.008
- KUMAR, S.; TAMURA, K.; NEI, M. MEGA3: Integrated Software for Molecular Evolutionary Genetics Analysis and Sequence Alignment. Briefings in Bioinformatics, v. 5, n. 2, p.150-163, 2004. PMid:15260895. http://dx.doi.org/10.1093/bib/5.2.150
- LINDSAY, D. S.; DUBEY, J. P.; DUNCAN, R. B. Confirmation that the dog is a definitive host for Neospora caninum Veterinary Parasitology, v. 82, n. 4, p. 327-333, 1999. http://dx.doi.org/10.1016/S0304-4017(99)00054-0
- MCALLISTER, M. M. et al. Dogs are definitive hosts of Neospora caninum International Journal for Parasitology, v. 28, n. 9, p. 1473-1478, 1998. http://dx.doi.org/10.1016/S0020-7519(98)00138-6
- MILLS, B. Immunohistochemistry. In: PROPHET, E. B. et al. Laboratory Methods in Histotechnology Washington: Armed Forces Institute of Pathology, 1992. p. 247-255.
- MOORE, D. P. Neosporosis in South America. Veterinary Parasitology, v. 127, n. 2, p. 87-97, 2005. PMid:15631900. http://dx.doi.org/10.1016/j.vetpar.2004.10.001
- NATIONAL CENTER FOR BIOTECHNOLOGY INFORMATION - NCBI. Primer3 2011a. Disponível em: < http://www.ncbi.nlm.nih.gov/tools/primer-blast/>. Acesso em: 30 mar. 2011.
- NATIONAL CENTER FOR BIOTECHNOLOGY INFORMATION - NCBI. BlastN 2011b. Disponível em: < http://blast.ncbi.nlm.nih.gov/Blast.cgi?PROGRAM=blastn&BLAST_PROGRAMS=megaBlast&PAGE_TYPE=BlastSearch&SHOW_DEFAULTS=on&LINK_LOC=blasthome >. Acesso em: 30 mar. 2011.
- ORTEGA-MORA, L. M.; FERNANDEZ-GARCIA, A.; GOMEZ-BAUTISTA, M. Diagnosis of bovine neosporosis: recent advances and perspectives. Acta Parasitologica, v. 51, n. 1, p. 1-14, 2006. http://dx.doi.org/10.2478/s11686-006-0001-0
- PAYNE, S.; ELLIS, J. Detection of Neospora caninum DNA by the polymerase chain reaction. International Journal for Parasitology, v. 26, n. 4, p. 347-351, 1996. http://dx.doi.org/10.1016/0020-7519(96)00030-6
- ROBERTS, J. S. Veterinary obstetrics and genital diseases (Theriogenology) Ann Arbor: Edwards Brothers Inc, 1986. 981 p.
- ROZEN, S.; SKALETSKY, H. J. Primer3 1998. Disponível em: < http://biotools.umassmed.edu/bioapps/primer3_www.cgi #disclaimer>. Acesso em: 30 mar. 2011.
- SAITOU, N.; NEI, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Molecular Biology Evolution, v. 4, n. 4, p. 406-425, 1987.
- SAMBROOK, J. et al. Molecular cloning: a laboratory manual. New York: Cold Spring Harbor Laboratory, 1989. 1659 p.
- SANGER, F.; NICKLEN, S.; COULSON, A. R. DNA sequencing with chain-terminating inhibitors. Proceedings of the National Academy of Sciences USA, v. 74, n. 12, p. 5463-5467, 1977. http://dx.doi.org/10.1073/pnas.74.12.5463
- SANTOS, S. L. et al. Investigation of Neospora caninum, Hammondia sp., and Toxoplasma gondii in tissues from slaughtered beef cattle in Bahia, Brazil. Parasitology Research, v. 106, n. 2, p. 457-461, 2010. PMid:19943064. http://dx.doi.org/10.1007/s00436-009-1686-4
- JEANMOUGIN, F. et al. Multiple sequence alignment with ClustalX. Trends in Biochemistry Scieces, v. 23, n. 10, p. 403-405, 1998. http://dx.doi.org/10.1016/S0968-0004(98)01285-7
- TOSCAN, G. et al. Neosporose equina: ocorrência de anticorpos anti-Neospora spp. e associação entre status sorológico de éguas e de suas crias. Pesquisa Veterinária Brasileira, v. 30, n. 8, p. 641-645, 2010. http://dx.doi.org/10.1590/S0100-736X2010000800006
Corresponding author:
Publication Dates
-
Publication in this collection
21 Dec 2011 -
Date of issue
Dec 2011
History
-
Accepted
10 May 2011 -
Received
21 Feb 2011