Abstracts
This work aimed the study of acquisition of iron from different compounds by EIEC strains. Seven serotypes were studied (O28ac.H-, O29:H-, O136:H-, O144:H-, O152:H-, O164:H- and O167:H7-), in a total of 31 strains. All serotypes were able to use distinct iron sources: hemin, hematin, hemoglobin, ovotransferin, lactoferrin, transferrin and ferritin.
EIEC; iron; iron uptake
Foi investigada a aquisição de ferro por cepas de EIEC a partir de diferentes compostos. Sete sorotipos foram estudados (O28ac.H-, O29:H-, O136:H-, O144:H-, O152:H-, O164:H- and O167:H7), totalizando 31 cepas. Foi observado que todos os sorotipos puderam usar fontes de ferro distintas: hemina, hematina, hemoglobina, ovotransferrina, transferrina, lactoferrina e ferritina.
EIEC; ferro; captação de ferro
Uptake of iron from different compounds by enteroinvasive Escherichia coli
Monique Dall´Agnol; Marina Baquerizo Martinez** Corresponding author. Mailing address: Faculdade de Ciências Farmacêuticas da Universidade de São Paulo. Av. Prof. Lineu Prestes, 580. CEP: 05508-900, São Paulo, SP, Brasil. Fax: (+5511) 813-2197. E-mail: Corresponding author. Mailing address: Faculdade de Ciências Farmacêuticas da Universidade de São Paulo. Av. Prof. Lineu Prestes, 580. CEP: 05508-900, São Paulo, SP, Brasil. Fax: (+5511) 813-2197. E-mail: mbmartin@usp.br
Departamento de Análises Clínicas e Toxicológicas, Faculdade de Ciências Farmacêuticas, Universidade de São Paulo, São Paulo, SP, Brasil
Submitted: November 27, 1998; Approved: April 20, 1999
SHORT COMMUNICATION
ABSTRACT
This work aimed the study of acquisition of iron from different compounds by EIEC strains. Seven serotypes were studied (O28ac.H-, O29:H-, O136:H-, O144:H-, O152:H-, O164:H- and O167:H7-), in a total of 31 strains. All serotypes were able to use distinct iron sources: hemin, hematin, hemoglobin, ovotransferin, lactoferrin, transferrin and ferritin.
Key words: EIEC, iron, iron uptake
The process of infectious disease depends largely upon the efficiency with which the microorganisms gain access to the host, colonize, cause pathological alterations and disseminate to a new host (5). The process of colonization involves two steps ¾ adherence and multiplication. The process of multiplication requires acquisition of growth-essential nutrients, including iron from the host (14). Iron is an essential element for living organisms. This ion is a very versatile biocatalyst (12) and this characteristic is responsible for its involvement in so many essential processes for the cells, such as breathing and ribonucleotide synthesis. Despite its importance, iron is not readily available in aquatic or terrestrial environments and in animal hosts (8).
The amount of free iron available in the human host is extremely low (10-18M) which is insufficient for bacterial growth (4). The majority of iron is found intracellularly as mioglobin, ferritin, hemosiderin and hemoglobin (12). The trace quantities of extracellular iron is bound to the glycoproteins transferrin, in the blood, and lactoferrin, in secretions and mucosal surfaces (1, 19). The ability of pathogenic microorganisms of obtaining iron from host is essential for the production of disease (5).
More commonly, enterobacteria synthesize low-molecular-weight iron-binding compounds, siderophores, and their associated outer membrane proteins receptors. There are two predominant chemical classes of siderophores in E. coli, enterobactin (fenolate) and aerobactin (hidroxamate). Siderophores have extremely high affinity for iron and are able to remove iron from glycoproteins like transferrin and lactoferrin (10).
Unlike siderophore-based systems, the utilization of host iron-binding proteins directly as source of iron requires direct contact between these proteins and the bacterial cells. Several pathogens can utilize this strategy which requires also outer membrane proteins receptors. The ability to utilize these proteins is often not the only strategy an organism has at its disposal for acquiring iron. For example, vibrios synthesize siderophores in addition to utilizing heme (19).
Several studies have been done to know which iron host compounds can support the growth of microorganisms. S. flexneri can use transferrin, lactoferrin and hemin as iron sources (18). Yersinia pestis and Y. enterocolitica can use hemin as iron source (17). Enteropathogenic E. coli (EPEC) uses hemin and hemoglobin as iron sources (9).
E. coli encloses several clones which together have evolved the ability to cause a broad spectrum of human diseases. EIEC are diarrheagenic E. coli strains which are closely related to Shigella sp. (15) However, very little is known about the iron uptake systems of enteroinvasive E. coli (EIEC) and the host sources of iron for these bacteria as well.
In the present study, a plate assay employing Luria-Bertani (LB) agar depleted of iron has been used and also with different iron sources to verify which one could support the growth of these bacteria.
A total of 31 EIEC strains, isolated from diarrheagenic patients, belonging to 7 serotypes, were tested. These strains were previously identified by biochemical and serological methods and by Serény test (13). The EIEC strains were grown in Minimal Medium broth (MM) (9) by shaking (170 rpm) at 37oC for 6 hours. The iron chelator, a, a´-dypirydil, was added to melted LB agar (triptone 10 g, yeast extract 5 g, NaCl 10 g, NaOH 1N 12.5 ml, agar 1.5%, H2O 1 L) in final concentrations of 400µM. The compounds transferrin, ovotransferrin, human and bovine lactoferrins, hemin, hematin and hemoglobin were added as iron sources in concentration of 25µg/ml (9). Bacterial suspension containing 105 CFU was inoculated as spots on the surface of LB agar supplemented with a, a´-dypirydil and the iron source. The plates were incubated at 37oC overnight for bacterial growth. The bacterial growth in LB agar and LB agar supplied with a, a´-dypirydil only were used as positive and negative controls, respectively. The iron sources and a, a´-dypirydil were purchased by Sigma Co (St. Louis, MO USA). The other reagents were purchased from DIFCO Laboratories (Detroit, Michigan USA).
The tests for enterochelin and aerobactin were done by Dr. Shelley Payne (Department of Microbiology-Univ. of Texas, Austin-USA), according to Arnow (2) and Csaky (6), respectively.
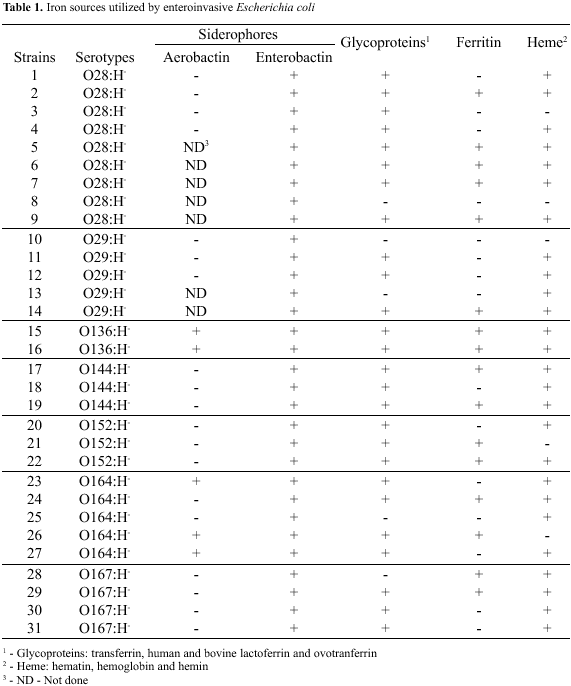
It was observed that the 7 serotypes could take iron up from distinct sources. Positive or negative results were based on visualization of bacterial growth on the media surface (Fig. 1). Heme group (hemin, hematin and hemoglobin) and/or glycoprotein group (ovotransferrin, lactoferrins and transferrin) were used by 26 strains (84%), while ferritin was used by 16 strains (52%). The strains did not show a profile of iron utilization, once there was variation in the iron source used by different serotypes, as well as by different strains of a same serotype (Table 1). The iron sources utilization was not species specific because EIEC strains could take iron up from ovotransferrin and bovine lactoferrin. The intracellular localization of ferritin can contribute to the decreasing capacity to take iron up from this compound by the siderophores. Besides, the iron must be reduced to be removed from ferritin, because without reducers, the siderophores are either ineffective for releasing iron from ferritin or they can remove it very slowly (1).
All EIEC strains produced enterobactin and some strains aerobactin (Table 1). The siderophores systems are used to obtain iron from glycoprotein group and ferritin (3; 17), however, some strains did not use one or more of these compounds as iron sources. This suggests that there can be a flaw in some steps of this process, because for the microorganisms to take iron up from siderophores, the complex iron-siderophore has to be recognized by receptors in the outer membrane and the iron has to be released by specific enzymes action (16;19).
Bacteria present in mucosal surfaces can take iron up from heme or hemoglobin released from dead epithelium cells (7). In the present work, most EIEC strains used the heme group (hemin, hematin and hemoglobin) as iron source. Stugard et al. (21) reported the presence of an outer membrane receptor for hemin (~100kDa) in Shigella flexneri and EIEC strains. Thus far it can be realized that this receptor is likely to be the mechanism used by these bacteria to take iron up from hemin and, maybe, from other heme compounds, once in S. flexneri the hemin utilization as iron source was shown to be independent of the siderophore system (11).
Being the iron transport protein present in secretions and mucosal surfaces, such as the gastrointestinal tract, the lactoferrin can be an important iron source for enteropathogens because of its localization. The iron uptake from lactoferrin can be processed either by syderophores system or by specific receptors in the outer membrane of some pathogens (17). A putative lactoferrin receptor in EIEC strains outer membrane is being investigated.
The results obtained in this work indicate that there was variation in the iron sources used by different serotypes, as well as by different strains of a same serotype. This fact might suggest that EIEC strains have a high capacity of adaptation, using the iron uptake system that requires the lowest energetic consumption. The use of several iron sources suggests that EIEC strains can use more than one mechanism for it. Therefore, the infections processes of these bacteria can be more easily achieved due to their capacity of using distinct iron sources.
ACKNOWLEDGEMENTS
Thanks to CAPES and FAPESP (94/3014-0).
RESUMO
Obtenção de ferro a partir de diferentes compostos por Escherichia coli enteroinvasora
Foi investigada a aquisição de ferro por cepas de EIEC a partir de diferentes compostos. Sete sorotipos foram estudados (O28ac.H-, O29:H-, O136:H-, O144:H-, O152:H-, O164:H- and O167:H7), totalizando 31 cepas. Foi observado que todos os sorotipos puderam usar fontes de ferro distintas: hemina, hematina, hemoglobina, ovotransferrina, transferrina, lactoferrina e ferritina.
Palavras-chave: EIEC, ferro, captação de ferro
-
1Aisen, P.; Leibman, A. Lactoferrin and transferin, a comparative study. Biochim. Biophys. Acta, 257:314-323, 1972.
-
2Arnow, L.E. Colorimetric determination of the components of 3,4-dihydroxyphenylalanine-tyrosine mixtures. J. Biol. Chem., 228:531-537, 1937.
-
3Crichton, R.R.; Charloteaux-Waubers, M. Iron transport and storage. Eur. J. Biochem., 164: 485-506, 1986.
-
4Crosa, J.H. The relationship of plasmid-mediated iron transport and bacterial virulence. Ann. Ver. Microbiol., 38: 69-89, 1984.
-
5Eichenbaum, Z.; Muller, E.; Morse, S.; Scott, J. R. Acquisition of iron from host proteins by the group A Streptococcus. Infect. Immun., 64: 5428-5429, 1996.
-
6Csaky, T. Z. On the estimation of bound hydroxylamine in biological materials. Acta Chem. Scand., 2: 450-454, 1948.
-
7Griffiths, E. The iron uptake systems of pathogenic bacteria. In: Bullen, J.J.; Griffiths, E (eds). Iron and Infection. John Wiley, Chichester, 1985, p.69-137.
-
8Guerinot, L. M. Microbial iron transport. Annu. Ver. Microbiol., 48: 743-772, 1994.
-
9Law, D.; Wilkie, K. M.; Freeman, R.; Gould, F. K. The iron uptake mechanisms of enteropathogenic Escherichia coli: the use of haem and haemoglobin during growth in an iron-limited environment. J. Med. Microbiol., 37: 15-21, 1992.
-
10Lawlor, K. M.; Daskaleros, P. A.; Robinson, R. E.; Payne, S. M. Virulence of iron transport mutants of Shigella flexneri and utilization of host iron compounds. Infect. Immun., 55: 594-599, 1987.
-
11Levine, M.M. Escherichia coli that cause diarrhea enterotoxigenic, enteropathogenic, enteroinvasive and enteroaderent. J. Infect. Dis., 155: 377-389, 1987.
-
12Litwin, M.C.; Calderwood, B.S. Role of iron in regulation of virulence genes. Clin. Microbiol., 6: 137-149, 1993.
-
13Martinez, M.B.; Whittan, T.S.; McGraw, E.A.; Trabulsi, L.R. Clonal relationship among enteroinvasive Escherichia coli. FEMS Microbiol. Letters, in press, 1999.
-
14Mietzner, T.A.; Morse, S. A.The role of iron-binding proteins in the survival of pathogenic bacteria. Annu. Rev. Nutr., 14: 471-493, 1994.
-
15Nataro, J.P.; Kaper, J.B. Diarrheagenic Escherichia coli. Clin. Microbiol. Rev., 11: 142-201, 1998.
-
16O´Brien, I.G.; Cox, B.; Gibson, F. Enterochelin hydrolysis and iron metabolism in Escherichia coli. Biochim. Biophys. Acta, 237: 537-549, 1971.
-
17Otto, B.R.; Vereij-Van, A.M.J.J.; MacLaren, D.M. Transferrins and heme-compounds as iron sources for pathogenic bacteria. Crit. Rev. Microbiol., 18: 217-233, 1992.
-
18Payne, S. M. Iron and virulence in Shigella Mol. Microbiol., 3: 1301-1306, 1989.
-
19Payne, S. M. Iron acquisition in microbial pathogenesis. Trends Microbiol., 1: 66-69, 1993.
-
20Sigel, S.P.; Payne, S.M. Effect of iron limitation on growth, siderophore production, and expression of outer membrane proteins of Vibrio cholerae. J. Bacteriol., 150: 148-155, 1982.
-
21Stugard, C.E.; Daskaleros, P.A.; Payne, S.M.A 101kDa heme-binding protein associated with congo red binding and virulence of Shigella flexneri and enteroinvasive Escherichia coli strains. Infect. Immun., 57: 3534-3539, 1989.
Publication Dates
-
Publication in this collection
06 Jan 2000 -
Date of issue
Apr 1999
History
-
Accepted
20 Apr 1999 -
Received
27 Nov 1998