Abstracts
The enzyme chitinase from Moniliophthora perniciosa the causative agent of the witches' broom disease in Theobroma cacao, was partially purified with ammonium sulfate and filtration by Sephacryl S-200 using sodium phosphate as an extraction buffer. Response surface methodology (RSM) was used to determine the optimum pH and temperature conditions. Four different isoenzymes were obtained: ChitMp I, ChitMp II, ChitMp III and ChitMp IV. ChitMp I had an optimum temperature at 44-73ºC and an optimum pH at 7.0-8.4. ChitMp II had an optimum temperature at 45-73ºC and an optimum pH at 7.0-8.4. ChitMp III had an optimum temperature at 54-67ºC and an optimum pH at 7.3-8.8. ChitMp IV had an optimum temperature at 60ºC and an optimum pH at 7.0. For the computational biology, the primary sequence was determined in silico from the database of the Genome/Proteome Project of M. perniciosa, yielding a sequence with 564 bp and 188 amino acids that was used for the three-dimensional design in a comparative modeling methodology. The generated models were submitted to validation using Procheck 3.0 and ANOLEA. The model proposed for the chitinase was subjected to a dynamic analysis over a 1 ns interval, resulting in a model with 91.7% of the residues occupying favorable places on the Ramachandran plot and an RMS of 2.68.
Chitinase; Moniliophthora perniciosa; kinetic characterization; purification; isoenzymes; heat stability; 3D structure; comparative modeling
A enzima quitinase de Moniliophthora perniciosa, o fungo causador da doença vassoura de bruxa em Theobroma cacao, foi parcialmente purificada por precipitação com sulfato de amônio e filtração em coluna de Sephacryl S-200 usando fosfato de sódio como tampão de extração. A metodologia de Superfície de resposta (MSR) foi usada para determinar o pH e temperatura ótimos. Foram isoladas quatro quitinases diferentes: ChitMp I, ChitMp II, ChitMp III e ChitMp IV. A ChitMp I mostrou temperatura ótima de 44-73ºC e pH ótimo de 7,0-8,4. A ChitMp II apresentou temperatura ótima entre 45-73ºC e pH ótimo de 7,0-8,4. ChitMp III apresentou temperatura ótima de 54-67ºC e pH ótimo de 7,3-8,8. ChitMp IV apresentou temperatura ótima de 60ºC e pH ótimo em 7,0. Em relação à biologia computacional a sequência primária foi feita in silico usando banco de dados do Projeto Genoma/Proteoma do M. perniciosa, obtendo-se uma sequência com 564 pb e 188 aminoácidos, que foi utilizada para a obtenção da estrutura tridimensional por metodologia de modelagem comparativa. Os modelos gerados foram submetidos à validação usando Procheck 3.0 and ANOLEA. O modelo proposto para a quitinase foi submetido à análise dinâmica por 1 ns, resultando em um modelo com 91,7% de seus resíduos em locais regiões favoráveis no gráfico de Ramachandran e RMS de 2,68.
quitinase; Moniliophthora perniciosa; caracterização cinética; purificação; isoenzimas; estabilidade térmica; estrutura 3D; modelagem comparativa
Purification, characterization and structural determination of chitinases produced by Moniliophthora perniciosa
Rafaela S. GalanteI; Alex G. TarantoII; Maria G.B. KoblitzIII; Aristóteles Góes-NetoIV; Carlos P. PirovaniV; Júlio C.M. CascardoV; Sandra H. CruzVI; Gonçalo A.G. PereiraVII ; Sandra A. de AssisI
IDepartamento de Saúde, Universidade Estadual de Feira de Santana, Km 3, BR 116 (norte), 44031-460 Feira de Santana, BA, Brasil
IILaboratório de Bioinformática, CCO, Universidade Federal de São João Del-Rei, Av. Sebastião Gonçalves Coelho, 400, Bairro Chanadour, 35501-296 Divinópolis, MG, Brasil
IIIDepartamento de Tecnologia, Universidade Estadual de Feira de Santana, Km 3, BR 116 (norte), 44031-460 Feira de Santana, BA, Brasil
IVDepartamento de Ciências Biológicas, Universidade Estadual de Feira de Santana, Km 3, BR 116 (norte), 44031-460 Feira de Santana, BA, Brasil
VLaboratório de Genômica e Expressão Gênica, DCB, Universidade Estadual de Santa Cruz, Rodovia Ilhéus-Itabuna, Km 16, 45650-000 Ilhéus, BA, Brasil, E-mail: sandraassis@uefs.br VIDepartamento de Agroindústria, Alimentos e Nutrição, Escola Superior de Agricultura Luiz de Queiroz, ESALQ, Universidade de São Paulo, Av. Pádua Dias, 11, 13418-900 Piracicaba, SP, Brasil
VIILaboratório de Genômica e Expressão, Departamento de Genética e Evolução, Instituto de Biologia, Universidade Estadual de Campinas, Caixa Postal 6109, 13083-970 Campinas, SP, Brasil
Correspondence to Correspondence to: Sandra A. de Assis E-mail: sandraassis@uefs.br
ABSTRACT
The enzyme chitinase from Moniliophthora perniciosa the causative agent of the witches' broom disease in Theobroma cacao, was partially purified with ammonium sulfate and filtration by Sephacryl S-200 using sodium phosphate as an extraction buffer. Response surface methodology (RSM) was used to determine the optimum pH and temperature conditions. Four different isoenzymes were obtained: ChitMp I, ChitMp II, ChitMp III and ChitMp IV. ChitMp I had an optimum temperature at 44-73ºC and an optimum pH at 7.0-8.4. ChitMp II had an optimum temperature at 45-73ºC and an optimum pH at 7.0-8.4. ChitMp III had an optimum temperature at 54-67ºC and an optimum pH at 7.3-8.8. ChitMp IV had an optimum temperature at 60ºC and an optimum pH at 7.0. For the computational biology, the primary sequence was determined in silico from the database of the Genome/Proteome Project of M. perniciosa, yielding a sequence with 564 bp and 188 amino acids that was used for the three-dimensional design in a comparative modeling methodology. The generated models were submitted to validation using Procheck 3.0 and ANOLEA. The model proposed for the chitinase was subjected to a dynamic analysis over a 1 ns interval, resulting in a model with 91.7% of the residues occupying favorable places on the Ramachandran plot and an RMS of 2.68.
Keywords: Chitinase, Moniliophthora perniciosa, kinetic characterization, purification, isoenzymes, heat stability, 3D structure, comparative modeling.
RESUMO
A enzima quitinase de Moniliophthora perniciosa, o fungo causador da doença vassoura de bruxa em Theobroma cacao, foi parcialmente purificada por precipitação com sulfato de amônio e filtração em coluna de Sephacryl S-200 usando fosfato de sódio como tampão de extração. A metodologia de Superfície de resposta (MSR) foi usada para determinar o pH e temperatura ótimos. Foram isoladas quatro quitinases diferentes: ChitMp I, ChitMp II, ChitMp III e ChitMp IV. A ChitMp I mostrou temperatura ótima de 44-73ºC e pH ótimo de 7,0-8,4. A ChitMp II apresentou temperatura ótima entre 45-73ºC e pH ótimo de 7,0-8,4. ChitMp III apresentou temperatura ótima de 54-67ºC e pH ótimo de 7,3-8,8. ChitMp IV apresentou temperatura ótima de 60ºC e pH ótimo em 7,0. Em relação à biologia computacional a sequência primária foi feita in silico usando banco de dados do Projeto Genoma/Proteoma do M. perniciosa, obtendo-se uma sequência com 564 pb e 188 aminoácidos, que foi utilizada para a obtenção da estrutura tridimensional por metodologia de modelagem comparativa. Os modelos gerados foram submetidos à validação usando Procheck 3.0 and ANOLEA. O modelo proposto para a quitinase foi submetido à análise dinâmica por 1 ns, resultando em um modelo com 91,7% de seus resíduos em locais regiões favoráveis no gráfico de Ramachandran e RMS de 2,68.
Palavras-chave: quitinase, Moniliophthora perniciosa, caracterização cinética, purificação, isoenzimas, estabilidade térmica, estrutura 3D, modelagem comparativa.
INTRODUCTION
Chitin, a homopolymer of N-acetyl-D-glucosamine (Glc-NAc) residues linked by β-1,4 bonds, is a common constituent in cocoons, exoskeletons and peritrophic membranes of insects, crustacean shells, and fungal cell walls (Wang et al. 2002, Schrempf 2001, Nicol 1991, Gooday 1977). Also, all organisms that contain chitin have chitinases (EC 3.2.1.14) and other chitinolytic enzymes, as these enzymes are presumably required for the morphogenesis of cell walls and exoskeletons (Patil et al. 2000, Dahiya et al. 2006). Chitinases are glycosidases; they are also found in organisms that do not contain chitin, such as plants, bacteria, and viruses. Although the physiological roles of these enzymes remain unresolved, it is clear that plant enzymes are involved in the defense against fungal pathogens and in regulating development (Merzendorfer 2006, Dahiya et al. 2006, Duo-Chuan 2006). Filamentous fungi occupy a delicate balance between biosynthesis and hydrolysis of the cell wall during the remodeling for hyphal growth, hyphal branching and septum formation, thereby requiring the involvement of chitinolytic enzymes in this process (Dahiya et al. 2006, Selvaggini et al. 2004).
Endochitinases, exochitinases and beta-N-acetylhexosamidases are the main chitinolytic enzymes, and typically several hydrolytic enzymes act synergistically to accomplish the efficient and complete biodegradation of chitin (Dahiya et al. 2005, Bormann et al. 1999).
The production of microbial chitinases has received remarkable attention in both the industrial and scientific communities, mainly because of theirs enormous potential for economically increasing the production of several useful products (Dahiya et al. 2006, Felse and Panda 1999a, b, Kim et al. 2003). These enzymes have widespread applications in the fields of medicine (Kim et al. 2003, Toharisman et al. 2005), waste management (Odds 2003, Lopes et al. 2008), biotechnology (Felse and Panda 1999a, b) and industry (Haki and Rakshit 2003). Chitinases are used for the mild degradation of chitin to prepare biologically active chitooligosaccharides and for the structural determination of the glycoprotein polysaccharide. This application of chitinase requires either commercially available individual chitinases or at least a mixture of endochitinases and exochitinases free of other proteins (Tikhonov et al. 1998). In the agricultural field, chitinases are used mainly to control phytopathogenic fungi (Felse and Panda 1999a, b, Gobbetti et al. 1999). Chitinolytic enzymes from both bacteria (Aeromonas and Serratia) and fungi (Gliocladium and Trichoderma) have been used successfully for the control of pathogenic fungi of plants (Chernin et al. 1995). Moniliophthra perniciosa, the causative agent of the witches' broom disease in Theobroma cacao, is responsible for major crop losses in South American and Caribbean cocoa plantations (Kilaru and Hasenstein 2005, Scarpari et al. 2005, Surujdeo-Maharaj and Umaharan 2004). In 1989, witches' broom disease was identified in Bahia, the leading cocoa-growing region in Brazil (Purdy and Schmidt 1996, Griffith et al. 2003). In less than 10 years, the cocoa production in Brazil decreased from 400,000 tons to 100,000 tons (Hebbar 2007). Consequently, Brazil slipped from being the third- to the fifth-largest cocoa-producing country in the world (Pereira et al. 1990). The pathogen is a hemibiotrophic basidiomycete with two distinguishable phases in its life cycle.
The concept of response surface methodology (RSM) has eased the process of development and has been of significant industrial use. Recent studies have indicated that RSM can be used to analyze the effects of different factors on proteolytic activity (Gobbetti et al. 1999) and to optimize the enzyme production (Nawani and Kapadnis 2005, Tikhonov et al. 1998).
In this study, we describe the isolation, partial purification and biochemical characterization of the chitinase from M. perniciosa. The 3D structure of the protein was elucidated by comparative modeling.
MATERIALS AND METHODS
Materials
Chitin and bovine serum albumin were purchased from Sigma Chemical Co (St Louis, MO). All the other chemicals used were of high quality analytical grade.
The microorganism used in this study was M. perniciosa (CCMB0257) from the Collection of Cultures of Microorganisms of Bahia (CCMB).
Microorganism and Enzyme Production
M. perniciosa (CCMB0257) was maintained on potato dextrose agar plates at 25ºC. To produce chitinase, M. perniciosa was grown in vegetative brooms cultivated in a special system with conditions amenable to fungi development (basidiocarps).
Extraction of the chitinase and ammonium sulphate fraction
The enzyme was extracted at 40ºC with 0.05 M sodium phosphate buffer at pH 7.0 containing 0.6 M NaCl. The ratio of M. perniciosa to the extraction buffer was 1:3 (g mL−1). The homogenate was squeezed through two layers of gauze and the extract was centrifuged at 10,000 x g for 10 min to remove the solid particles. The precipitate was discarded, and the supernatant was brought to 70% saturation by adding solid ammonium sulfate and centrifuged at 10,000 x g for 10 min after standing for 1 h. The precipitate with high PME activity was resuspended in borate acetate buffer at the ratio of 1:3 (w/v) and stored at low temperature.
The samples were desalted by ultra filtration with an Amicon membrane (10000 Da).
Measurement of Enzyme Activity
Chitinolytic activity was determined using the colloidal chitin as the substrate. A 0.2 ml portion of the enzyme extract was mixed with 0.2 ml of 1% chitin in 0.05 M phosphate buffer at pH 7.0.
The assay mixture was incubated at 50ºC and the end products of the reaction were analyzed using the DNS method (Miller 1959). One unit of chitinase activity is defined as the amount of enzymes required to release 1 µmol of GlcNAc in 1 min under the above.
Protein determination and enzyme analysis
Protein concentration was determined by the method of Bradford using bovine soroalbumin as a standard (Bradford 1976).
Chromatographic separation
Sephacryl S-200 chromatography
Following an initial round of Sephacryl S-200 (20.0 x 1.1 cm) chromatography, fractions containing chitinase activity were pooled and applied to a Sephacryl S-200 column (20.0 x 1.1 cm) previously equilibrated with ammonium sulfate buffer, pH 7.0 (the samples were subjected to a second round of chromatography). The same buffer was also used for elution, and 1.5 ml fractions were collected and assayed for chitinase activity.
Heat Stability
Chitinase samples in test tubes (selected to be equal in weight, volume and size) were incubated in water at different temperatures and for different times. After the heating process, the tubes were cooled in melting ice and the residual activity was measured. The residual activity was determined after incubation: 80ºC (0, 10, 20, 30, 40, 50 and 60 min) and 90ºC (0, 2, 4, 6, 8, 10, 12, 14, 16, 18 and 20 min).
Experimental design
A double-variable, three-level central composite design (CCD) leading to 11 sets of experiments (assays), performed in a triplicate, was used to verify the optimum pH and temperature optimum.
The STATISTICA 6.0 software (StatSoft, Tulsa, OK) was used to generate design matrix and analyze the results. Table II shows the design matrix and responses (results) obtained for the chitinase activity.
Replications and Statistical Analysis
The extractions and chromatographic isolations were repeated at least three times. Significant differences among sample means were tested.
Primary structure determination
The sequence of the chitinase of the fungus that causes witches' broom disease was obtained by in silico analysis of reads (Fig 5, Table III) deposited in the Genome Project M. perniciosa and in assembly of a contig.
Chitinase models
Chitinase models were built by threading using the Modeller program and four structures as templates and by homology comparison using the program Deep View v3.7 (Humphrey et al. 1996) and the models 1WNO and PDB 2A3E as templates (Table V), which were obtained from the PDB (Protein Data Bank) with 100% similarity with the sequence problem.
RESULTS
Purification of chitinase
Chitinase was extracted from M. perniciosa as described in Materials and Methods. The purification scheme consisted of only three steps: ammonium sulfate precipitation, gel filtration on Sephacryl S-200 and re-chromatography in gel filtration on Sephacryl S-200.
Ammonium sulfate precipitation of the enzyme resulted in a 1.63-fold increase in a specific activity. Subsequently, chitinase was eluted on a Sephacryl S-200 column equilibrated with a 50 mM sodium phosphate buffer at pH 7.0. The samples were re-eluted on a Sephacryl S-200 column. The result of this step was a 18.16-fold purification for ChitMp I2, a 7.83-fold purification for ChitMp II2, a 16.67-fold purification for ChitMp III2, and a 10.42-fold purification for ChitMp IV2 (Figure 2 and Table I)
Effect of temperature on enzyme activity
The thermal stability of the chitinase was studied by heating the samples at 80 and 90ºC for various time periods. The residual activity was measured as described in Materials and Methods.
As shown in Figure 3a, ChitMpI2 lost 10% of their initial activities after 20 min of heating at 80ºC, and were inactivated completely after 60 min of heating. Furthermore, the enzyme lost 100% of their initial activities after only 2 min of heating at 90ºC (Figure 3b).
ChitMp II2 lost 60% of its initial activity after 20 min of heating at 80ºC, and was completely inactivated after 60 min of heating (Fig. 3a). At 90ºC, the enzyme lost 100% of its initial activity after 2 min of heating (Fig. 3).
ChitMp III2 lost 60% of its initial activity after 20 min of heating at 80ºC, and was completely inactivated after 40 min of heating (Fig. 3a). At 90ºC, the enzyme lost 100% of its initial activity after 2 min of heating (Fig. 3b).
ChitMp IV2 lost 90% of its initial activity after 20 min of heating at 80ºC, and was completely inactivated after 40 min of heating (Fig. 3a). At 90ºC, the enzyme lost 95% of its initial activity after 10 min of heating (Fig. 3b).
Determination of the pH and temperature optima
Experimental design
The STATISTICA 6.0 software was used to generate design matrix and analyze the results. Table II shows the design matrix and responses (results) obtained for the chitinase activity.
The results of ChitMp I2 show that this enzyme is stable between pH 7.0 and 8.3 (Fig. 4a).
In the case of ChitMp II2, the results show that the enzyme is stable in the range of pH 7.0 - 8.4 and that the activity is higher between 45 and 73ºC (Fig. 4b).
In the case of ChitMp III2, the results show that the optimum activity is between 54 and 67ºC and a pH between 7.3 and 7.8 (Fig. 4c).
ChitMp IV2 showed optimum activity at 58 to 70ºC and a pH between 7.6 and 8.2 (Fig. 4d).
Primary structure determination (sequence)
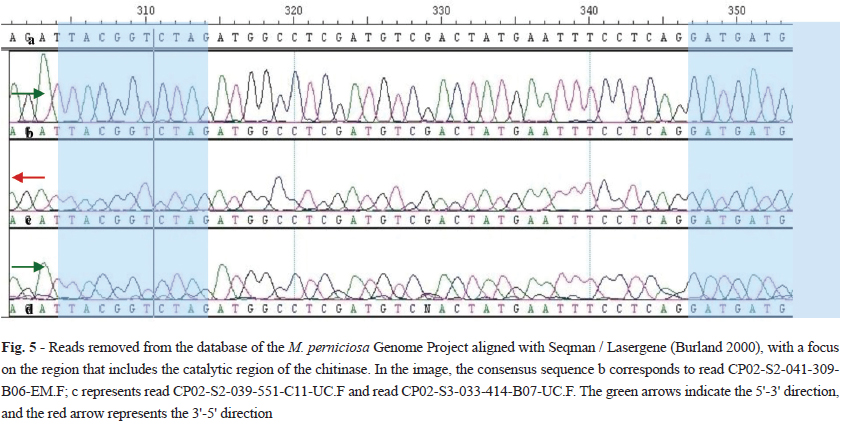
Figure 5 and Table III shows the reads removed from the database of the Genome Project M. perniciosa aligned with Seqman / Lasergene (Burland 2000), focusing on the region that includes the catalytic region of the chitinase. The fact that the same sequence is repeated in the three reads (nucleotides 310 to 343, Figure 5b, c and d) indicates that the data submitted are reliable, and all reads exhibit resolved chromatographic peaks. The low peak resolutions can interfere with the identification of nucleotides, with some nucleotides replaced by N, as in position 329 of the read CP02-S3-033-414-B07-UC.F (Fig. 5d). In the same position, reads band c show the nucleotide guanine with peaks of satisfactory quality.
This sequence was aligned to protein sequences of some fungi phylogenetically related to M. perniciosa (Lopes 2005) with high identity, particularly in the region of the catalytic site (Fig. 6, Table IV). The fragment of the sequence of chitinase has a shaped molecular mass of 21.62 kDa and a pI of 4.9, reflecting its similarity to chitinases from other fungi (Duo-Chuan 2006) and plants (Wang et al. 2002). This isoelectric point is identical to those found in fungi Verticillium lecanii (Zimm.) Viégas and Piromyces communis Gold JJ, Heath IB & Bauchop (Wang et al. 2002), and is similar to that reported by Haki and Rakshit (2003).
The sequence of the chitinase consisting of 564 bp and 188 amino acids was not complete, but it contained the conserved region described in the literature as the catalytic region of the enzyme (Synstad et al. 2004, Cederkvist et al. 2007). This region comprises the nucleotides 290 to 323 (Fig. 7).
Localization of the active site in the constructed models
Construction of three-dimensional model
The results obtained by the construction of the three-dimensional model generated by VMD 1.8.6 (Humphrey et al. 1996) show that the molecular model of chitinase has 188 residues, including 1543 atoms united by 1571 chemical bonds, with 13 hydrogen bonds at the catalytic site. Using the standard cutoff of 3.2 Ä, 22 salt bridges were found (Asp31 - Lys45, Asp99 - Lys51 and Glu130 - Arg133). Of the amino acids, 68 are hydrophobic, 47 are polar, 19 are basic and 29 are acidic. The initial model of chitinase had already been satisfactorily validated, but even so, some of the simulations and refinements introduced by molecular dynamics resulted in an updated model as shown in Table V. The model was analyzed by comparative Ramachandran charts, by PROCHECK 3.0 and shows to be improved.
Models of chitinase, built by homology and threading, have been undermined in Amber 8.0 (Case et al. 2004) with 10 sodium ions through the tool. Later, the models were subjected to refinement by 300 and 600 cycles and then were carried out molecular dynamics (MD) by 300, 1000 and 3000 ps in vacuum. For solvent, the models were subjected to refinement by 800 cycles and then were carried out molecular dynamics (MD) by 1000 and 2000 ps.
The validation results show no significant differences between the models of chitinase generated by threading and homology. The model obtained by homology, subject to refinement and 600 cycles called hqmp2, is what generated the best results in the PROCHECK validation (Table V). However, the model suggested for the chitinase of M. perniciosa was the model generated by homology with 1000 ps at 300 K in implicit solvent (Table V), which shows the Ramachandran plot, the amino acids Trp66, Ser67, Asp138, Arg151 and Ser157 are in unfavorable regions of the chart and the amino acid Lys51, Ser70, Val75, Asp138 and Asp174 in a region largely favorable, with a score of 94.7% of amino acid residues in favorable local chart Ramachandran (Fig. 8).
The effect of the quality improvement model dm1000_quit can be seen by looking at the side chains, where all parameters analyzed by PROCHECK had lower standard deviations in the chosen model when compared with side chains in the hqmp2 model (Fig. 8). The surface analysis of atomic force (Atomic Mean Force Potential - AMPF) conducted by ANOLEA (data not shown) also validated the models; however, the chosen model, dm 1000_quit, had a better AMPF and a more negative energy than other models constructed by comparative modeling, supporting the validation performed by PROCHECK. The models generated by MD 300, 1000 and 3000 ps did not show satisfactory results when validated by ANOLEA, demonstrating a positive energy (data not shown), and models generated by MD 300, 1000 and 3000 ps using the threading methodology were also not satisfactory. For these reasons, the dm1000_quit was chosen as the best model for the chitinase of M. perniciosa (Fig. 9).
The motif LDVDYEFP, corresponding to Leu98, Asp99, Gly100, Leu101, Asp102, Val103, Asp104, Tyr105, Glu106, Phe107 and Pro108 and described as the catalytic site, corresponds to residues 98-108 (Fig. 10) that participate in the formation of a β-sheet, a structural form that is easily recognized as part of a more complex system. A conserved region designed SIGG (Fig 6), corresponding to Ser62, Ile63, Gly64 and Gly65, also were found in this model it is a important represents hydrophobic domain to catalysis (Zhu et al. 2008). The final structure of the model has seven α-helices, seven β-sheets and fourteen turns. Antiparallel β-sheets can be composed of several strands that can be distributed in various parts of the sequence, and can be composed of hydrophilic (Tyr107, Asp99, Asp102, Asp104 and Glu106) or hydrophobic amino acids (Val103, Trp105 and Pro108) (Höltje et al. 2003).
Characteristics of Model generated by homology in solvent 1000 ps:
The molecule possesses 1543 atoms united by 1571 chemical bonds and 13 hydrogen bonds at the catalytic site. In VMD 1.8.6 analysis with a standard cutoff of 3.2 Ä, 22 salt bridges were found. Salt bridges and hydrogen bonds are important elements contributing to the molecular stability and substrate specificity, respectively. The overall molecular structure consists of six α-helices, two β-sheets and eight turns.
Ramachandran plot analysis showed that 91.7% of amino acid residues were found in favorable areas in the model generated in solvent after 1 ns of dynamics, resulting in an RMS of 2.68 Ä. The molecular Dynamics for 2 and 3 ns (data not shown), showed less satisfactory results than the 1 ns case. The Ramachandran plot of the initial model presents the best values as a result of the good template (50% identity).
DISCUSSION
Chitinase is one of the most important cell wall lytic enzymes; it has been purified from a number of fungal species and found to have differing physiological properties, molecular structures, and molecular weights.
In the studies made by Wang et al. (2002), the enzymes lost >50% of their initial activities after only 3 min of heating at 100ºC, and were completely inactivated after 9 min of heating.
The chitinase from Isaria japonica lost 0% of its activity at 50oC (Kawachi et al. 2001). The chitinase from Aspergillus fumigatus lost 30% of its activity after 60 minutes of heating at 55ºC (Xia et al. 2001). The chitinase from Trichoderma harzianum was inactivated at 60ºC (De La Cruz et al. 1992).
In this study, the temperature and pH optima of ChitMp I and II were similar, indicating that these enzymes act synergistically. However, ChitMp III and IV showed similar characteristics that were different from those of ChitMp I and II.
Miyashita et al. (1991) found three chitinases, ChiA, ChiB, ChiC and ChiD in S. lividans. ChiC and ChiD exhibited similar activities and generic structures, but ChiA and ChiB differ.
Felse and Panda (1999a, b) reveal that the three chitinases of T. harzianum (CHIT33, CHIT37 and CHIT42) did not result from the proteolytic breakdown of a single common chitinase, but rather were transformed from three different genes with little or no homology.
Duo-Chuan et al. (2005) found similar charac-teristics in relation to temperature (between 30 and 40ºC) and pH (4.0 - 8.0) in the two chitinases from Talaromyces flavus (Klöcker) Stolk & Samson.
The optimal temperature for many fungi is in the range between 20 and 40ºC, except for some thermophilic fungi that have maximal activities around 55 and 70ºC, such as Thermomyces lanu-ginosus Tsikl. (Guo et al. 2005) and Talaromyces emersonii Stolk (Mccormack et al. 1991). The chitinase from A. fumigatus YJ-407 retained 40% of its activity after heating at 60ºC for 30 min, and these results indicated that this enzyme was the most thermostable among the fungal chitinases reported (Xia et al. 2001).
In summary, the chitinase from M. perniciosa exists in several isoforms. One of these isoenzymes (ChitMp IV) showed high thermostability.
Compared with the modeling results, data obtained in silico from the database of the Moniliophthora perniciosa Genome Project provi-ded the region of the nucleotide sequence of the chitinase around the catalytic site of the enzyme. This region comprises 564 bp and 188 amino acids. These results were sufficient to generate a 3D model of the enzyme with 50% similarity with a model obtained from the program Deep View v3.7.
The modeled fragment includes the conserved domains SXGG (Wattanalai et al. 2004) and DxDxE (Watanabe et al. 1992, Synstad et al. 2004, Cederkvist et al. 2007), where the latter region is only found in Family 18 chitinases (Okazaki et al. 2004, Cederkvist et al. 2007). In Nomuraea rileyi (Farl.) Samson, this sequence corresponds to EDGIDIDWE (Wattanalai et al. 2004), in Rhizopus oligosporus Saito to DGFDFDIE (Takaya et al. 1998), in Trichoderma harzianum to LDGFDLDNE (Hoell et al. 2005) and in Moniliophthora perniciosa to LDGLDVDYEFP, and other conserved domains have been identified, such as SIGG (residues 62 to 65). In all these organisms, the LXXLDXDXE motif is cited as the catalytic site of the enzyme (Lu et al. 2002).
Studies by Lu et al. (2002) on Manduca sexta chitinase reveal the essential nature of residues Glu146 to Glu106 which are equivalent to Asp144 and 142 to Asp104, as well Asp 104 and 102 in the chitinase of M. perniciosa for the catalytic activity of the enzyme. Their results suggest that Glu146 is more critical than Asp142 and Asp144 to enable the participation in the first stage of the mechanism of catalysis. According to Synstad et al. (2004), a mutation in Asp142 is highly deleterious for the catalytic activity of the enzyme, and according to Lu et al. (2002), a mutation in Asp144 results in a 70% loss of enzyme activity.
From the 3D model of the chitinase and the nucleotide sequence determined by PCR, sequencing and cloning, one of the goals of future works will be to select a specific inhibitor by docking with this enzyme.
ACKNOWLEDGMENTS
This work was supported by Financiadora de Estudos e Projetos (FINEP), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and Fundação de Amparo à Pesquisa do Estado da Bahia (FAPESB). We also thank the Programa de Pós Graduação em Biotecnologia (PPGBiotec-UEFS).
Manuscript received on November 29, 2010;
accepted for publication on May 31, 2011
- AYATI M, MANDELMAN D, AGHAJARI N and HASER R. 2002. Structure catalytical domain of psycrophylic chitinase from Arthobacter, with without allosamidine. 2002. (Unpublished).
- BORMANN C, BAIER D, HÖRR I, RAPS C, BERGER J, JUNG G, SCHWARZ H. 1999. Characterization of a novel, antifungal, chitin-binding protein from streptomyces tendae that interferes with growth polarity. J Bacteriol 181: 7421-7429.
- BORTONE K, MONZINGO AF, ERNST S and ROBERTUS JD. 2002. Structure of a allosamidin complex with the Coccidioides immitis chitinase defines a role for a second acid residue in substrate-assisted mechanism. J Mol Biol 320: 293-302.
- BRADFORD MM. 1976. A rapid and Sensitive Method for the Quantitation of Microgram Quantities of Protein Utilizing the Principle of Protein-Dye Binding. Anal Biochem 72: 248-254.
- BURLAND TG. 2000. DNASTAR's Lasergene sequence analysis software. Methods Mol Biol 132: 71-91.
- CASE DA et al. 2004. Amber 8 University of California, San Francisco.
- CEDERKVIST FH, SAUA SF, KARLSEN V, SAKUDA S, EIJSINK VGH and SØRLIE M. 2007. Thermodynamic Analysis of Allosamidin Binding to a Family 18 Chitinase. Biochem 46: 12347-12354.
- CHERNIN L, ISMAILOV Z, HARAN S and CHET I. 1995. Chitinolytic Enterobacter agglomerans Antagonistic to Fungal Plant Pathogens. Appl Environ Microbiol 61(5): 1720-1726.
- DAHIYA N, TEWARI R and HOONDAL GS. 2006. Biotechnological aspects of chitinolytic enzymes: a review. Appl Microbiol Biotechnol 71: 773-782.
- DAHIYA N, TEWARI R, TIWARI RP and HOONDAL GS. 2005. Chitinase from Enterobacter sp. NRG4: Its purification, characterization and reaction pattern. Electron J Biotechnol 8: 134-145.
- DE LA CRUZ J, HIDALGO-GALLEGO A, LORA JM, BENITEZ T, PINTOR-TORO JA and LLOBELL A. 1992. A. Isolation and characterization of three chitinases from Trichoderma havzianum Eur J Biochem 206: 859-867.
- DUO-CHUAN L. 2006. Review of fungal chitinases. Mycopathologia 161: 345-360.
- DUO-CHUAN CHEN S and JING LU. 2005. Purification and partial characterization of two chitinases from the mycoparasitic fungus Talaromyces flavus Mycopathologia 159: 223-229.
- FELSE PA and PANDA T. 1999a. Regulation and cloning of microbial chitinase genes. Appl Microbiol Biotechnol 51: 141-151.
- FELSE PA and PANDA T. 1999b. Studies on applications of chitin and its derivatives. Bioprocess Eng 20: 505-512.
- GOBBETTI M, LANCIOTTI R, DE ANGELIS M, CORBO MR, MASSINI R and FOX P. 1999. Study of the effects of temperature, pH, NaCl, and aw on proteolytic and lipolytc activities of cheese-related lactic acid bacteria by quadratic response surface methodology. Enzyme Microb Technol 25: 795-809.
- GOODAY GW. 1977. Biosynthesis of the fungal walls mechanisms and implications. J Gen Microbiol 99: 1-11.
- GRIFFITH GW, NICHOLSON J, NENNINGER A, BIRCH RN and HEDGER J. 2003. Witches' broom and frosty pods two major pathogens of cacao. New Zeal J Bot 41: 423-435.
- GUO RF, LI DC and WANG R. 2005. Purification and properties of a thermophilic chitinase from thermophilic fungus Thermomyces lanuginosus Acta Microbiol Sin 45: 270-274.
- HAKI GD and RAKSHIT SK. 2003. Developments in industrially important thermostable enzymes: a review. Bioresour Technol 89: 17-34.
- HEBBAR PK. 2007. Cacao Diseases: a global perspective from an industry point of view. Phytopatology 97: 1658-1663.
- HÖLTJE HD, SIPPL W, ROGNAN D and FOLKERS G. 2003. Molecular modeling: Basic principles and applications. 2nd ed., Wiley-VCH, Germany, 430 p.
- HOELL IA, KLEMSDAL SS, VAAJE-KOLSTAD G, HORN SJ and EIJSINK VGH. 2005. Overexpression and characterization of a novel chitinase from Trichoderma atroviride strain P1. Biochim et Biophys Acta 1748: 180-190.
- Hu H, Wang G, Yang H, Zhou J, Mo L, Jin CS, Jin C and Rao Z. 2005. Crystallization and preliminary crystallo-graphic analysis of a native chitinase from the fungal pathogen Aspergillus fumigatus YJ-407 (afCHI). Acta Cryst 60: 939-940.
- HUMPHREY W, DALKE A and SCHULTEN K. 1996. "VMD - Visual Molecular Dynamics". J Mol Graphics 14: 33-38.
- KAWACHI I, FUJIEDA T, UJITA M, ISHIL Y, YAMAGISHI K, SATO H, FUNAGUMA T and HARA A. 2001. Purification and properties of extracellular chitinases from the parasitic fungus Isaria japonica J Biochem Bioengin 92(6): 544-549.
- KILARU A and HASENSTEIN KH. 2005. Development and Pathogenicity of the Fungus Crinipellis perniciosa on Interaction with Cacao Leaves. Biochem Cell Biol 95(1): 101-107.
- KIM K, YANG YJ and KIM JG. 2003. Purification and Characterization of Chitinase from Streptomyces sp. M-20. J of Biochem Mol Biol 36: 185-189.
- LOPES MA. 2005. Estudo molecular do sistema quitinolítico de Crinipellis perniciosa 64f. Dissertação (Mestre em Genética e Biologia Molecular), Universidade Estadual de Santa Cruz - Ilhéus. (Unpublished).
- LOPES MA, GOMES DS, KOBLITZ MGB, PIROVANI CP, CASCARDO JCM, GÓES-NETO A and MICHELI F. 2008. Use of response surface methodology to examine chitinase regulation in the basidiomycete Moniliophthora perniciosa Mycol Res 112: 399-406.
- LU Y, ZEN KC, MUTHUKRISHNAN S and KRAMER KJ. 2002. Site-directed mutagenesis and functional analysis of active site acidic amino acid residues D142, D144 and E146 in Manduca sexta (tobacco hornworm) chitinase. Insect Biochem Mol Biol 32: 1369-1382.
- MATSUMOTO T, NONAKA T, HASHIMOTO M, WATTANABE T and MITSUI Y. 1999. Tree-dimentional structure of catalytic domain of chitinase A1 from Bacillus circulansBacillus circulans WL-12 at a very high resolution. Proceeding of the Japan Academy Ser B 75: 269-274.
- McCORMACK J, HACKETT TJ, TUOHY MG and COUGHLAN MP. 1991. Chitinase production by Talaromyces emersonii Biotechnol Lett 13: 677-682.
- MERZENDORFER H. 2006. Insect chitin synthases: a review. J Comp Physiol B 176: 1-15.
- MILLER G. 1959. Use of dinitrosalicylic acid reagent for detection of reducing sugar. Anal Chem 31: 426-428.
- MIYASHITA K, FUJII T and SAWADA Y. 1991. Molecular cloning and characterization of chitinase genes from Streptomyces lividans 66. Jf Gen Microbiol 137: 2065- 2072.
- NAWANI NN and KAPADNIS BP. 2005. Optimization of chitinases production using statistics based experimental designs. Processes Biochemistry 40: 651-660.
- NICOL S. 1991. Life after death for empty shells. New Sci 129: 46-48.
- NOTREDAME C, HIGGINS D and HERINGA JT. 2000. Coffee: A novel method for multiple sequence alignments. J Mol Biol 302: 205-217.
- ODDS FC. 2003. Antifungal agents: their diversity and increasing sophistication. Mycologist 17: 51-55.
- OKAZAKI K, YAMASHITA Y, NODA M, SUEYOSHI N, KAMESHITA I and HAYAKAWA S. 2004. Molecular cloning and expression of the gene encoding family 19 chitinase from Streptomyces sp. J13-3. Biosci Biotechnol Biochem 68(2): 341-351.
- PATIL RS, GHORMADE V and DESHPANDE MV. 2000. Chitinolytic enzymes: an exploration. Enzyme and Microb Technol 26: 473-483.
- PEREIRA JL, RAM A, FIGUEIRED JM and ALMEIDA LCC. 1990. First occurrence of Witches' Broom disease in the principal cocoa-growing region of Brazil. Trop Agr 67: 188-189.
- PURDY LH and SHMIDT RA. 1996. Status of cacao witches'broom: biology, epidemiology, and management. Annu Rev Phytopathol 34: 573-594.
- RAO FV, ANDERSEN OA, VORA KA, DEMARTINO JA and van AALTEN DM. 2005. Methylchanthine drogs are chitinase inhibitors: investigation of inhibition and binding modes. Chem Biol 12: 973-980.
- SCARPARI LM, MEINHARDT W, MAZZAFERA P, POMELLA AWV, SCHIAVINAT MA, CASCARDO JCM and PEREIRA GAG. 2005. Biochemical changes during the development of witches'broom: the most important disease of cocoa in Brazil caused by Crinipellis perniciosa J Exp Bot 56(413): 865-877.
- SCHREMPF H. 2001. Recognition and degradation of chitin by streptomycetes. Antonie van Leeuwenhoek 79: 285-289.
- SELVAGGINI S, MUNRO CA, PASCHOUD S, SANGLARD D and GOW NAR. 2004. Independent regulation of chitin synthase and chitinase activity in Candida albicans and Saccharomyces cerevisiae Microbiol 150: 921-928.
- SURUJDEO-MAHARAJ S and UMAHARAN P. 2004. Assessment of resistance to witches'-broom disease in clonal and segregating populations of Theobroma cacao Plant Disease 88: 797-803.
- SYNSTAD B, GÄSEIDNES S, VAN AALTEN DMF, VRIEND G, NIELSEN JE and EIJSINK VGH. 2004. Mutational and computational analysis of the role conserved residues in the active site of a family 18 chitinase. Eur J Biochem 271: 253-262.
- TAKAYA N, YAMAZAKI D, HORIUCHI H, OHTA A AND TAKAGI M. 1998. Intracellular chitinase gene from Rhizopus oligosporus: molecular cloning and charac-terization, Microbiol (Reading, Engl). 144: 2647-2654.
- TIKHONOV VE, RADIGINA LA, YAMSKOV IA, NELLY D, GULYAEVA ND, ILYINA AV, ANLSIMOVA MV, VARLAMOV VP and TATARINOVA NY. 1998. Affinity purification of major chitinases produced by Streptomyces kurssanovi Enzyme Microb Technol 22: 82-85.
- TOHARISMAN A, SUHARTONO MT, SPINDLER-BARTH M, HWANG JK and PYUN YR. 2005. Purification and characterization of a thermostable chitinase from Bacillus licheniformis Mb-2. World J Microb Biot 21: 733-738.
- WANG SL, HSIAO WJ and CHANG WT. 2002. Purification and characterization of a antimicrobial chitinase extracellularly produced by Monascus purpureus CCRC31499 in a shrimp and crab shell powder medium. J Agric Food Chem 50: 2249-2255.
- WATANABE T, OYANAGI W, SUZUKI K, OHNISHI K and TANAKA H. 1992. Structure of the gene encoding chitinases D of Bacillus circulans WL-12 and possible homology of the enzyme to other prokaryotic chitinases and class II plant chitinases. J Bacteriol Jan, p. 408-414.
- WATTANALAI R, WIWAT C, BOUCIAS DG and TARTAR A. 2004 Chitinase gene of the dimorphic mycopathogen, Nomuraea rileyi J Invertebr Pathol 85: 54-57.
- XIA G, JIN C, ZHOU J, YANG S, ZHANG S and JIN C. 2001 A novel chitinases having a unique model of action from Aspergillus fumigatus YJ-407. Eur J Biochem 208: 4079-4085.
- Zhu Y, Pan J, Qiu J and Guan X. 2008. Isolation and characterization of a chitinase gene from entomopatho-genic fungus Verticillium lecanii Braz J Microbiol 39: 314-320.
Correspondence to:
Publication Dates
-
Publication in this collection
25 May 2012 -
Date of issue
June 2012
History
-
Received
29 Nov 2010 -
Accepted
31 May 2011