Abstracts
Cryptococcus neoformans is an encapsulated yeast, etiological agent of cryptococcosis. The species is commonly associated with pigeon droppings and plant materials. The aim of the present work was to verify the presence of the yeast in pigeon droppings, and to identify the isolates obtained in serotypes and mating types (MAT). Ten samples of pigeon droppings were collected in the rural area of the city of Alfenas, Brazil. Samples were inoculated in agar Niger medium for fungal isolation and 22 isolates with characteristics of C. neoformans were obtained. The serotypes and MAT were determined by multiplex PCR using specific primers. Serotypes were also determined by using the Kit Crypto Check. Among the 22 samples evaluated, eight were identified as C. neoformans by classic identification tests. These samples were characterized as serotype A by the Kit Crypto check and as serotype A MAT alpha by the multiplex PCR. The present study reinforces the evidence that pigeon droppings are a reservoir for C. neoformans and confirms the prevalence of C. neoformans var. grubii (Aalpha) among environmental isolates. It also demonstrates that multiplex PCR is an acceptable alternative for serotype analysis because it reduces the costs for each reaction and analyses serotype and MAT simultaneously.
Cryptococcus neoformans; Serotype; Mating Type; PCR; Kit Crypto Check®
Cryptococcus neoformans é levedura encapsulada, agente etiológico da criptococose. As espécies são comumente associadas com fezes de pombos e material vegetal. O objetivo do presente trabalho foi verificar a presença de leveduras em fezes de pombos e identificar os isolados em relação aos sorotipos e "mating types". Dez amostras de fezes de pombos foram coletadas na zona rural da cidade de Alfenas, Brasil. As amostras foram inoculadas em agar Niger e 22 isolados com características de C. neoformans foram obtidos. Os sorotipos e "mating types" foram determinados pela PCR multiplex e os sorotipos foram identificados também pelo Kit Crypto Check. Dentre as 22 amostras avaliadas, oito foram identificadas como C. neoformans através dos testes clássicos. Estas amostras foram caracterizadas como sorotipo A pelo Kit Crypto check e como sorotipo A MATalfa pela PCR multiplex. O presente estudo reforça a evidência de que as fezes de pombos constituem reservatório para C. neoformans e confirma a prevalência de C. neoformans var. grubii (Aalfa) nos isolados ambientais. PCR multiplex é uma alternativa aceitável para análise do sorotipo porque reduz os custos de cada reação e analisa simultaneamente os sorotipos e "mating type".
MYCOLOGY
Serotype and mating type characterization of Cryptococcus neoformans by Multiplex PCR
Caracterização de sorotipo e "mating type" de Cryptococcus neoformans por PCR Multiplex
Vívian Gonçalves CarvalhoI; Mateus Souza TercetiI; Amanda Latercia Tranches DiasI; Claudete Rodrigues PaulaII; Juliana Pereira LyonI; Antônio Martins de SiqueiraI; Marília Caixeta FrancoI
IDepartamento de Ciências Biológicas, Universidade Federal de Alfenas, Alfenas, MG, Brazil
IILaboratório de Micologia do Instituto de Ciências Biomédicas da Universidade de São Paulo, São Paulo, SP, Brazil
Correspondence to Correspondence to: Dra Marília Caixeta Franco Departamento de Ciências Biológicas, Unifal-MG Rua Gabriel Monteiro da Silva 714 37130-000 Alfenas, MG, Brazil E-mail: mcfranco@unifal-mg.edu.br
SUMMARY
Cryptococcus neoformans is an encapsulated yeast, etiological agent of cryptococcosis. The species is commonly associated with pigeon droppings and plant materials. The aim of the present work was to verify the presence of the yeast in pigeon droppings, and to identify the isolates obtained in serotypes and mating types (MAT). Ten samples of pigeon droppings were collected in the rural area of the city of Alfenas, Brazil. Samples were inoculated in agar Niger medium for fungal isolation and 22 isolates with characteristics of C. neoformans were obtained. The serotypes and MAT were determined by multiplex PCR using specific primers. Serotypes were also determined by using the Kit Crypto Check. Among the 22 samples evaluated, eight were identified as C. neoformans by classic identification tests. These samples were characterized as serotype A by the Kit Crypto check and as serotype A MAT a by the multiplex PCR. The present study reinforces the evidence that pigeon droppings are a reservoir for C. neoformans and confirms the prevalence of C. neoformans var. grubii (Aa) among environmental isolates. It also demonstrates that multiplex PCR is an acceptable alternative for serotype analysis because it reduces the costs for each reaction and analyses serotype and MAT simultaneously.
Keywords: Cryptococcus neoformans; Serotype; Mating Type; PCR; Kit Crypto Check®.
RESUMO
Cryptococcus neoformans é levedura encapsulada, agente etiológico da criptococose. As espécies são comumente associadas com fezes de pombos e material vegetal. O objetivo do presente trabalho foi verificar a presença de leveduras em fezes de pombos e identificar os isolados em relação aos sorotipos e "mating types". Dez amostras de fezes de pombos foram coletadas na zona rural da cidade de Alfenas, Brasil. As amostras foram inoculadas em agar Niger e 22 isolados com características de C. neoformans foram obtidos. Os sorotipos e "mating types" foram determinados pela PCR multiplex e os sorotipos foram identificados também pelo Kit Crypto Check. Dentre as 22 amostras avaliadas, oito foram identificadas como C. neoformans através dos testes clássicos. Estas amostras foram caracterizadas como sorotipo A pelo Kit Crypto check e como sorotipo A MATa pela PCR multiplex. O presente estudo reforça a evidência de que as fezes de pombos constituem reservatório para C. neoformans e confirma a prevalência de C. neoformans var. grubii (Aa) nos isolados ambientais. PCR multiplex é uma alternativa aceitável para análise do sorotipo porque reduz os custos de cada reação e analisa simultaneamente os sorotipos e "mating type".
INTRODUCTION
Cryptococcus neoformans is an encapsulated yeast, etiological agent of the opportunistic mycoses, cryptococcosis. This infection manifests mainly as meningoencephalitis in immunocompromised individuals. Cryptococcosis has been responsible for great morbidity and mortality rates among patients with AIDS17.
Five distinct serotypes of C. neoformans are determined by specific antiserum: A, B, C, D and AD1,24. The occurrence is associated with ecological, morphological physiological and molecular features, epidemiology, pathogenicity and geographic distribution. The species is distributed in the varieties neoformans (serotypes A, D and A/D) and gattii (serotypes B and C). Recently, the variety grubii (serotype A) and the new species C. gattii were proposed9. The anamorphic stage of the yeast comprises two mating types (MAT), a and a, which can be crossed to produce the basidiomycetous Filobasidiella neoformans5.
Cryptococcus neoformans var. neoformans is widely distributed in nature and has been isolated from several sources. The yeast is frequently associated with pigeon droppings but has also been isolated from soil, wood in decay, fruits and vegetables4.
The aim of the present work was to identify samples of C. neoformans isolated from pigeon droppings in the rural area of the city of Alfenas, Brazil, to characterize the C. neoformans obtained in serotypes and MAT by PCR multiplex and in serotypes by the Kit Crypto Check® (Iatron, Tokyo, Japan). We also compared serotypes obtained by PCR multiples and by the Kit Crypto Check (Iatron).
MATERIAL AND METHODS
Ten samples of pigeon droppings were collected in nests from seven farms in the rural area of Alfenas city, MG, Brazil. The samples obtained were conditioned in clean and dry paper envelopes and processed immediately in the laboratory of Microbiology of the Federal University of Alfenas (Unifal-MG).
Sample processing: For the processing of the samples, we used the method proposed by SHIELDS & AJELLO (1966) modified. Four grams of each sample were solubilized under agitation for two min in 30 mL of sterile saline solution. After ten minutes, the supernatant was transferred for test tubes containing 2 mL of distilled water amended with 543 mg penicillin6.
The solution obtained was transferred on triplicate to plates containing agar Niger medium (Guizotia abyssinica - Niger seeds, 70 g; glucose, 10 g, chloramphenicol, 50 mg; biphenyl, 1 g, agar, 20 g; distilled water, 1000 mL). Samples were incubated at 28 ºC and observed for five days.
Isolation and identification of C. neoformans: Yeast-like colonies showing a cream to brown pigmentation and mucoid characteristic were isolated in pure culture in Sabouraud Dextrose Agar (Difco, laboratories, Detroit, MI, USA) and observed microscopically with Indian Ink to investigate the presence of capsule10, pigment production in agar Niger medium without biphenyl, urease production in modified Christensen's urea medium (peptone, 1 g; sodium chloride, 5 g; monopotassium phosphate, 2 g; glucose, 1 g; urea, phenol red 0.2%, 6 mL; distilled water, 1000 mL), growth at 37 ºC10, carbon source assimilation23 and non assimilation of potassium nitrate10.
Molecular characterization of C. neoformans: DNA extraction of the samples characterized as C. neoformans was made with glass beads (Sigma, St. Louis, MO, USA)2. Samples were submitted to a PCR with the aim to determine the serotype and MAT. Reference samples belong to the Culture Collection of the Biomedical Sciences Institute, University of São Paulo, São Paulo, Brazil.
The primers used for DNA amplification were based on the STE20 gene sequence1. Two PCR multiplex were performed to simultaneous amplification of serotype and MAT22. The first PCR multiplex (aAaD) was made with the primers JOHE7264/7265 and JOHE7273/7275 in the following conditions: 30 cycles of 96 ºC for one min, 61 ºC for 30s and 72 ºC for one min. The sample ICB 134 was used as positive control C(+)1 and sample ICB 170, C(+)2. The second PCR multiplex (aAaD) was made with the primers JOHE7270/7272 and JOHE7267/7268 in the following conditions 30 cycles of 96 ºC for one min, 63 ºC for 30s and 72 ºC for one min. The control for the simultaneous amplification of the bands generated by the primers 7270/7272 and 7267/7268 was obtained by the combination of the DNA of the samples ICB 163 and ICB Aa (data not shown). The primers are described on Table 1.
Serotype characterization by the commercial kit Crypto check® (Iatron): The isolates were cultivated in yeast extract-malt extract-agar (Difco) at 25 ºC. After 48h of incubation the culture was suspended in sterile physiological saline solution at McFarland scale pattern 2 (about 6 x 108 CFU/mL). A drop of each seric factor (F1, F5, F6, F7, and F8) was placed in the corresponding cycle of the agglutination glass slides, and 50 µL of the C. neoformans suspension was added over each seric factor. The glass slides were homogenized with an agitator with rotational movement at 25 rpm for two min. The slides were read by direct observation of the small clots formed, as follows: serotype A, F1 and F7; serotype B, F1 and F5; serotype C, F1 and F6; serotype D, F1 and F8; and serotype AD, F1, F7, and F8.
RESULTS
Fifty-four yeast-like colonies were isolated from the ten analyzed samples of pigeon droppings. Among these, 22 were positive for the presence of capsule and showed cream to brown pigmentation in agar Niger without biphenyl, which are characteristics of C. neoformans. According to the carbon source assimilation test the following isolates were identified: two as C. uniguttulatus, one as C. laurentii, three as C. albidus and eight as C. neoformans. So, from the 22 analyzed samples, eight of them did not belong to the genus Cryptococcus.
The eight C. neoformans isolates were obtained from two farms, being six from dropping samples of five nests in the first farm and two dropping samples of two nests in the second farm.
All of the eight isolates evaluated agglutinated with the Kit Crypto Check (Iatron) sera and were classified as C. neoformans var. grubii (serotype A).
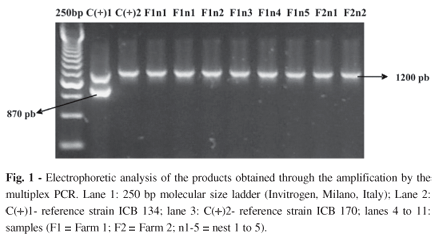
According to the multiplex PCR (Fig. 1), the C(+)1 control (ICB 134) was amplified resulting in one band of 1200 pb corresponding to the serotype A MATa alleles and another band of 870 pb related to the serotype D MAT a alleles. The amplification of C(+)2 control (ICB 170) resulting in a band of 1200 pb. The amplification of the eight C. neoformans isolates resulted in the presence of only one band, the 1200 pb one.
DISCUSSION
The isolation of C. neoformans from avian excreta has been reported by several authors3,7,16,19,20. STAIB & SCHULZ-DIETERICH (1984) concluded that the exposure to avian excreta explains at least partially the epidemiology of cryptococcosis.
REZENDE (2002) evaluated the presence of C. neoformans in pigeon and canary droppings in the urban area of the city of Alfenas and found the yeast in 11.11% of the samples analyzed. In the present work, C. neoformans could be isolated from 18.20% (eight in 44) of the samples analyzed. Similar data were obtained in other cities of Brazil and in other countries11,14,25. It has been speculated that plants in decay are the primary habitat of C. neoformans, which would explain the higher percentage of isolation in the rural area. On the other hand, this possibility does not explain the similarity with the percentage of isolation obtained in other works.
No non-specific bands were observed and no cross-reactions occurred. The produced fragments by the amplification of C(+)1 and C(+)2 controls correspond to the predicted sizes by the GeneBank (www.ncbi.nlm.nih.gov) nucleotide sequence.
The presence of 1200 pb band by the PCR multiplex allowed the identification of the variety grubii in the eight isolates.
Earlier studies had already demonstrated that MAT a was predominant among clinical and environmental isolates, despite the serotype1,8,13,24. In clinical samples, serotype A is more prevalent (77.95%) followed by serotype B (18.2%), AD (1.3%), D (0.4%) and C (0.2%)15. In the present study, serotype was determined by PCR multiplex and the Kit Crypto Check. Both methodologies showed to be easy to perform, reliable and demonstrated an acceptable concordance of results. The determination of the serotype by the Kit Crypto Check (Iatron) has also been performed by other authors1,12. BARRETO DE OLIVEIRA et al., 2004, reported that PCR multiplex with specific primers could discriminate heterozygotes isolates (AD) identified as homozygotes (A) by the Kit. In the present study, heterozygotes isolates were not identified.
Kit Crypto Check allows the quick determination of serotypes in samples of C. neoformans, however, costs are elevated when compared to PCR, and the mating type is not identified. Besides, heterozygote isolates might not be discriminated by the Kit. The present study reinforces the evidence that pigeon droppings are a reservoir for C. neoformans and confirms the prevalence of C. neoformans var. grubii (Aa) among environmental isolates. It also demonstrates that PCR multiplex is an acceptable alternative for serotype analysis because it reduces the costs for each reaction and analyses the mating type and serotype simultaneously. It represents a rapid, simple and relatively economical tool for epidemiological and virulence studies.
ACKNOWLEDGEMENTS
Our thanks go to FINEP, PIBIC/CNPq, Unifal-MG, USP for financial support.
Received: 3 August 2006
Accepted: 15 January 2007
- 1. BARRETO DE OLIVEIRA, M.T.; BOEKHOUT, T.; THEELEN, B. et al. - Cryptococcus neoformans shows a remarkable genotypic diversity in Brazil. J. clin. Microbiol., 42: 1356-1359, 2004.
- 2. BOLANO, A.; STINCHI, S.; PREZIOSI, R. et. al. - Rapid methods to extract DNA and RNA from Cryptococcus neoformans FEMS Microbiol. Lett., 1: 221-224, 2001.
- 3. EMMONS, C.W. - Saprophytic sources of Cryptococcus neoformans associated with the pigeon (Columba livia). Amer. J. Hyg., 62: 227-232, 1955.
- 4. FILIÚ, W.F.O.; WANKE, B.; AGÜENA, S.M et. al. - Cativeiros de aves como fonte de Cryptococcus neoformans na cidade de Campo Grande, Mato Grosso do Sul, Brasil. Rev. Soc. bras. Med. trop., 35: 591-595, 2002.
- 5. FRANZOT, S.P; SALKIN, I.F & CASADEVALL, A. - Cryptococcus neoformans var. grubii: separate varietal status for Cryptococcus neoformans serotype A isolates. J. clin. Microbiol., 37: 838-840, 1999.
- 6. KHOSRAVI, A.R. - Isolation of Cryptococcus neoformans from pigeon (Columba livia) droppings in northern Iran. Mycopathologia (Den Haag), 139: 93-95, 1997.
- 7. KOBAYASHI, C.C.B.A.; SOUZA, L.K.H.; FERNANDES, O.F.L. et. al. - Characterization of Cryptococcus neoformans isolated from urban environmental sources in Goiânia, Goiás State, Brazil. Rev. Inst. Med. trop. S. Paulo, 47: 203-207, 2005.
- 8. KWON-CHUNG, K.J. & BENNETT, J.T. - Distribution of alpha and alpha mating types of Cryptococcus neoformans among natural and clinical isolates. Amer. J. Epidem., 108: 337-340, 1978.
- 9. KWON-CHUNG, K.J. & VARMA, A. - Do major species concepts support one, two or more species within Cryptococcus neoformans? FEMS Yeast Res., 6: 574-587, 2006.
- 10. LACAZ, C.S.; PORTO, E. & MARTINS, J.E.C. - Micologia médica. São Paulo, Sarvier, 1991.
- 11. MONTENEGRO, H. & PAULA, C.R. - Environmental isolation of Cryptococcus neoformans var. gattii and C. neoformans var. neoformans in the city of São Paulo, Brazil. Med. Mycol., 38: 385-390, 2000.
- 12. NISHIKAWA, M.M.; LAZÉRA, M.S.; BARBOSA, G.G. et. al. - Serotyping of 467 Cryptococcus neoformans isolates from clinical and environmental sources in Brazil: analysis of host and regional patterns. J. clin. Microbiol., 41: 73-77, 2003.
- 13. OHKUSU, M.; TANGONAN, N.; TAKEO, K. et al. - Serotype, mating type and ploidy of Cryptococcus neoformans strains isolated from patients in Brazil. Rev. Inst. Med. trop. S. Paulo, 44: 299-302, 2002.
- 14. PAL, M. - First report of isolation of Cryptococcus neoformans var. neoformans from avian excreta in Kathmandu, Nepal. Rev. iberoamer. Micol., 14: 181-183, 1997.
- 15. RANDHAWA, H.S.; KOWSHIK, T. & KHAN, Z.U. - Decayed wood of Syzygium cumini and Ficus religiosa living trees in Delhi/New Delhi metropolitan area as natural habitat of Cryptococcus neoformans Med. Mycol., 41: 199-209, 2003.
- 16. REZENDE, D.G. - Isolamento de Cryptococcus neoformans de fezes de pombos e canários do município de Alfenas-MG. Alfenas, 2002. (Dissertação de Mestrado - Curso de Pós-graduação em Ciências Biológicas, Escola de Farmácia e Odontologia de Alfenas).
- 17. ROZENBAUM, R. & GONÇALVES, A.J. - Clinical epidemiological study of 171 cases of cryptococcosis. Clin. infect. Dis., 18: 369-380, 1994.
- 18. SHIELDS, A.B. & AJELLO, L. - Medium for selective isolation of Cryptococcus neoformans Science, 151: 208-209, 1966.
- 19. SOARES, M.C.B.; PAULA, C.R.; DIAS, A.L.T.; CASEIRO, M.M. & COSTA, S.O.P. - Environmental strains of Cryptococcus neoformans var. grubii in the city of Santos, SP, Brazil. Rev. Inst. Med. trop. S. Paulo, 47: 31-36, 2005.
- 20. SRIBUREE, P.; KHAYHAN, S.; KHAMWAN, C.; PANJAISEE, S. & THARAVICHITKUL, P. - Serotype and PCR-fingerprints of clinical and environmental isolates of Cryptococcus neoformans in Chiang Mai, Thailand. Mycopatologia, 158: 25-31, 2004.
- 21. STAIB, F. & SCHULZ-DIETERICH, J. - Cryptococcus neoformans in fecal matter of birds kept in cages. Control of Cryptococcus neoformans habitats. Zbl. Bakt. Mikrobiol. Hyg., 179: 179-186, 1984.
- 22. TERCETI, M.S.; CARVALHO, V.G.; DIAS, A.L.T.; PAULA, C.R. et al. - Desenvolvimento de PCR multiplex para determinação de mating type e sorotipo de Cryptococcus neoformans. In: CONGRESSO BRASILEIRO DE MICROBIOLOGIA, 23., Santos, 2005. CD Room.
- 23. WICKERHAM, L.J. & BURTON, K.A. - Carbon assimilation tests for classification of yeasts. J. Bact., 56: 363-371, 1948.
- 24. YAN, Z.; LI, X. & XU, J. - Geographic distribution of mating type alleles of Cryptococcus neoformans in four areas of the United States. J. clin. Microbiol., 40: 965-972, 2002.
- 25. YILDIRAN, S.T.; SARACLI, M.A.; GONLUM, A. & GUN, H. - Isolation of Cryptococcus neoformans var. neoformans from pigeon droppings collected throughout Turkey. Med. Mycol., 36: 391-394, 1998.
Publication Dates
-
Publication in this collection
30 Aug 2007 -
Date of issue
Aug 2007
History
-
Received
03 Aug 2006 -
Accepted
15 Jan 2007