Abstracts
Extended-spectrum β-lactamases (ESBL) in enterobacteria are recognized worldwide as a great hospital problem. In this study, 127 ESBL-producing Enterobacteriaceae isolated in one year from inpatients and outpatients at a public teaching hospital at São Paulo, Brazil, were submitted to analysis by PCR with specific primers for blaSHV, blaTEM and blaCTX-M genes. From the 127 isolates, 96 (75.6%) Klebsiella pneumoniae, 12 (9.3%) Escherichia coli, 8 (6.2%) Morganella morganii, 3 (2.3%) Proteus mirabilis, 2 (1.6%) Klebsiella oxytoca, 2 (1.6%) Providencia rettgeri, 2 (1.6%) Providencia stuartti, 1 (0.8%) Enterobacter aerogenes and 1 (0.8%) Enterobacter cloacae were identified as ESBL producers. BlaSHV, blaTEM and blaCTX-M were detected in 63%, 17.3% and 33.9% strains, respectively. Pulsed field gel eletrophoresis genotyping of K. pneumoniae revealed four main molecular patterns and 29 unrelated profiles. PCR results showed a high variety of ESBL groups among strains, in nine different species. The results suggest the spread of resistance genes among genetically different strains of ESBL-producing K. pneumoniae in some hospital wards, and also that some strongly related strains were identified in different hospital wards, suggesting clonal spread in the institutional environment.
ESBL; Enterobacteriaceae; Klebsiella pneumoniae; Pulsed Field Gel Electrophoresis; Antimicrobial resistance
Beta-lactamases de espectro estendido (ESBL) em enterobactérias são reconhecidas mundialmente como um grande problema hospitalar. Neste estudo, 127 Enterobacteriaceae produtoras de ESBL isoladas por um ano, de pacientes internados e ambulatoriais de um hospital público de ensino em São Paulo, Brasil, foram submetidas à análise pela PCR com iniciadores específicos para os genes blaSHV, blaTEM e blaCTX-M. Dos 127 isolados, 96 (75,6%) K. pneumoniae, 12 (9,3%) E. coli, 8 (6,2%) M. morganii, 3 (2,3%) Proteus mirabilis, 2 (1,6%) Klebsiella oxytoca, 2 (1,6%) Providencia rettgeri, 2 (1,6%) Providencia stuartti, 1 (0,8%) Enterobacter aerogenes e 1 (0,8%) Enterobacter cloacae foram identificados como produtores de ESBL. BlaSHV, blaTEM e blaCTX-M foram detectados em 63%, 17,3% e 33,9% das cepas, respectivamente. A genotipagem de K. pneumoniae por eletroforese em campo pulsado revelou quatro padrões moleculares principais e 29 perfis não relacionados. Os resultados da PCR demonstraram alta variedade de grupos de ESBL entre as cepas, em nove espécies diferentes. Os resultados sugerem a disseminação de genes de resistência entre cepas geneticamente diferentes de K. pneumoniae produtoras de ESBL em algumas unidades do hospital, e também que algumas cepas fortemente relacionadas foram identificadas em unidades hospitalares diferentes, sugerindo disseminação clonal no ambiente da instituição.
NOSOCOMIAL INFECTIONS
Extended-spectrum beta-lactamases among Enterobacteriaceae isolated in a public hospital in Brazil
Beta-lactamases de espectro estendido em Enterobacteriaceae isoladas de Hospital Público no Brasil
Milena DropaI; Livia C. BalsalobreI; Nilton LincopanII; Elsa M. MamizukaII; Thays MurakamiIII; Valéria C. CassettariIV; Fábio FrancoIV; Stella M. GuidaV; Angelica J. BalabakisV; Lilian F. PassadoreV; Silvia R. SantosV; Glavur R. MattéI; Maria H. MattéI
IPublic Health Laboratory, School of Public Health, University of São Paulo, Avenida Dr. Arnaldo 715, Cerqueira César, 01246-904 São Paulo, SP, Brazil
IILaboratory of Antimicrobial Resistance and Pharmaceutical Alternatives, Dept. Microbiology, Institute of Biomedical Sciences, University of São Paulo, São Paulo, Brazil. Av. Professor Lineu Prestes, 580, Bloco 17, Butantã, 05508-900 São Paulo, SP, Brazil
IIIDepartment of Scientific and Technological Policy, Institute of Geosciences, University of Campinas. Caixa Postal 6152, 13083-970 Campinas, São Paulo, Brazil
IVHospital Infection Control Committee, University Hospital, University of São Paulo, Av. Professor Lineu Prestes 2565, Butantã, 05508-000 São Paulo, SP, Brazil
VLaboratory of Clinical Microbiology, University Hospital, University of São Paulo, Av. Professor Lineu Prestes 2565, Butantã, 05508-000 São Paulo, SP, Brazil
Correspondence to Correspondence to: Milena Dropa Avenida Dr. Arnaldo 715, Cerqueira César 01246-904 São Paulo, SP, Brasil Phone: +55 11 30617753. Fax: +55 11 30833501 E-mail: milenadropa@usp.br
SUMMARY
Extended-spectrum β-lactamases (ESBL) in enterobacteria are recognized worldwide as a great hospital problem. In this study, 127 ESBL-producing Enterobacteriaceae isolated in one year from inpatients and outpatients at a public teaching hospital at São Paulo, Brazil, were submitted to analysis by PCR with specific primers for blaSHV, blaTEM and blaCTX-M genes. From the 127 isolates, 96 (75.6%) Klebsiella pneumoniae, 12 (9.3%) Escherichia coli, 8 (6.2%) Morganella morganii, 3 (2.3%) Proteus mirabilis, 2 (1.6%) Klebsiella oxytoca, 2 (1.6%) Providencia rettgeri, 2 (1.6%) Providencia stuartti, 1 (0.8%) Enterobacter aerogenes and 1 (0.8%) Enterobacter cloacae were identified as ESBL producers. BlaSHV, blaTEM and blaCTX-M were detected in 63%, 17.3% and 33.9% strains, respectively. Pulsed field gel eletrophoresis genotyping of K. pneumoniae revealed four main molecular patterns and 29 unrelated profiles. PCR results showed a high variety of ESBL groups among strains, in nine different species. The results suggest the spread of resistance genes among genetically different strains of ESBL-producing K. pneumoniae in some hospital wards, and also that some strongly related strains were identified in different hospital wards, suggesting clonal spread in the institutional environment.
Keywords: ESBL; Enterobacteriaceae; Klebsiella pneumoniae; Pulsed Field Gel Electrophoresis; Antimicrobial resistance.
RESUMO
Beta-lactamases de espectro estendido (ESBL) em enterobactérias são reconhecidas mundialmente como um grande problema hospitalar. Neste estudo, 127 Enterobacteriaceae produtoras de ESBL isoladas por um ano, de pacientes internados e ambulatoriais de um hospital público de ensino em São Paulo, Brasil, foram submetidas à análise pela PCR com iniciadores específicos para os genes blaSHV, blaTEM e blaCTX-M. Dos 127 isolados, 96 (75,6%) K. pneumoniae, 12 (9,3%) E. coli, 8 (6,2%) M. morganii, 3 (2,3%) Proteus mirabilis, 2 (1,6%) Klebsiella oxytoca, 2 (1,6%) Providencia rettgeri, 2 (1,6%) Providencia stuartti, 1 (0,8%) Enterobacter aerogenes e 1 (0,8%) Enterobacter cloacae foram identificados como produtores de ESBL. BlaSHV, blaTEM e blaCTX-M foram detectados em 63%, 17,3% e 33,9% das cepas, respectivamente. A genotipagem de K. pneumoniae por eletroforese em campo pulsado revelou quatro padrões moleculares principais e 29 perfis não relacionados. Os resultados da PCR demonstraram alta variedade de grupos de ESBL entre as cepas, em nove espécies diferentes. Os resultados sugerem a disseminação de genes de resistência entre cepas geneticamente diferentes de K. pneumoniae produtoras de ESBL em algumas unidades do hospital, e também que algumas cepas fortemente relacionadas foram identificadas em unidades hospitalares diferentes, sugerindo disseminação clonal no ambiente da instituição.
INTRODUCTION
Extended-spectrum beta-lactamases (ESBL) production by nosocomial pathogens are a major challenge for infection control committees in hospitals all over the world, once ESBL-producing strains and their encoding genes can stay permanently in hospitals, causing outbreaks and colonization (BRADFORD 2001, GUPTA et al. 2003, DENTON 2007). There are more than 300 ESBL described so far, most of them derived from groups TEM, SHV and CTX-M, with 150, 88 and 69 variants, respectively (see http://www.lahey.org/Studies).
In Latin America there is a high rate of ESBLs, although their distribution is still restricted to some genes and to specific locations (VILLEGAS et al. 2008). In SHV group, the variants SHV-2, SHV-5 and SHV-12 are the most common (PATERSON et al. 2003, VILLEGAS et al. 2008), although in Brazil there have already been reported the detection of SHV-4 (MENDES et al. 2004) and the characterization of SHV-27 (CORKILL et al. 2001). TEM variants have been described very recently in South America (PATERSON et al. 2003, VIGNOLI et al. 2005, VILLEGAS et al. 2008), and the most common are TEM-10, TEM-12 and TEM-26 (VILLEGAS et al. 2008). CTX-M ESBLs are endemic and widely distributed in our continent. They were reported in 1989 in Argentina (CTX-M-2) and, since then, several different enzymes from this group were reported in many countries, including Brazil, where CTX-M-8, -9 and -16 were described (VILLEGAS et al. 2008). Some recent Brazilian studies show the high prevalence of ESBL-carrying Enterobacteriaceae in healthcare centers and even in the community, especially in Klebsiella pneumoniae isolates (NOGUEIRA et al. 2006, MINARINI et al. 2007, MINARINI et al. 2008, SILVA DIAS et al. 2008).
The aim of this study was to identify groups of ESBL disseminated among enterobacteria isolated during one year in a teaching hospital, as well as to establish the genetic relationship among all ESBL-producing K. pneumoniae isolates.
MATERIAL AND METHODS
Setting: This study was carried out in a general public university hospital with 258 beds distributed among ten wards (Emergence Room - ER, Surgical Ward - SW, General Ward - AMW, Pediatric Ward - PW, Adult, Neonatal and Pediatric Intensive Care Units - A.ICU, N.ICU and P.ICU, Intermediate Care Unit - IMC, Neonatal Ward - NW and Maternity Ward - MW), with a mean turnover of 5500 patients-day/month.
Bacterial strains: During one year, 127 ESBL-producing Enterobacteriaceae strains were recovered from clinical samples of hospitalized patients. Isolates were responsible for community- or hospital-acquired infections, and for colonization.
Susceptibility test and ESBL production: Antimicrobial susceptibility tests, screening and confirmatory tests for ESBL production were carried out in Vitek System (BioMerieux, Hazlewood, MO). Double-disk synergy (Oxoid, Cambridge, UK) was also performed as a confirmatory test for all positive isolates (CLSI 2008).
Detection of bla genes: Total DNA extraction was carried out for all samples using the heat-shock technique (CHAPMAN et al. 2001). Isolates were tested by PCR for the β-lactamase groups SHV, TEM and CTX-M, with previously described oligonucleotides and reaction conditions (CAO et al. 2002). PCR products were analyzed by agarose gel electrophoresis.
Pulsed Field Gel Electrophoresis (PFGE): All K. pneumoniae isolates were submitted to DNA extraction (CDC 2004) and restriction endonuclease digestion with 10U XbaI. Separation of fragments was carried out at 6V/cm at 14 ºC for 20h. The linear pulse time was 3,51s to 30,82s. Band polymorphism generated by PFGE was analyzed and the molecular patterns were grouped primarily according to previously established criteria (TENOVER et al. 1995), and then according to Dice similarity coefficient and demonstrated graphically in dendrograms. PFGE patterns were also analyzed according to their distribution among hospital wards during the studied period, using the software Pajek as a tool (BATAGELJ and MRVAR 1996).
RESULTS
Bacterial strains: 127 ESBL-producing isolates were recovered from 86 patients among all hospital wards, except for the Maternity. Ninety-six (75.6%) strains were K. pneumoniae (ESBL-Kp), 12 (9.3%) Escherichia coli, 8 (6.2%) Morganella morganii, 3 (2.3%) Proteus mirabilis, 2 (1.6%) Klebsiella oxytoca, 2 (1.6%) Providencia rettgeri, 2 (1.6%) Providencia stuartti, 1 (0.8%) Enterobacter aerogenes and 1 (0.8%) Enterobacter cloacae. According to the hospital infection control committee, 29 (22.8%) isolates were responsible for hospital-acquired infections, 14 (11%) for community-acquired infections, and 84 (66.1%) for cases of colonization, from which 57 (67.9%) were recovered from routine samples and 27 (32.1%) were from surveillance cultures carried out by the committee.
Antimicrobial susceptibility tests and detection of ESBL production and genes: All strains were resistant to ampicillin and cephalotin, and susceptible to imipenem. Except for these results, the predominant antimicrobial susceptibility profiles for each PFGE pattern of K. pneumoniae strains are shown in Table 1, and for the other Enterobacteriaceae isolates are in Table 2. The combined discs method was able to detect the production of ESBL in all strains.
PCR: Genes blaSHV were detected in 80 (63%) isolates, blaCTX-M in 43 (33.9%) and blaTEM in 22 (17.3%). Among K. pneumoniae, blaSHV genes were detected in 71 (74%) strains. Twenty one (16.5%) strains produced some type of ESBL that could not be detected in PCR reactions, and thus were not classified as any of the groups studied. It was observed the detection of one to three groups of bla genes in each ESBL-producing isolate. All combinations of bla genes detected are shown in Tables 1 and 2.
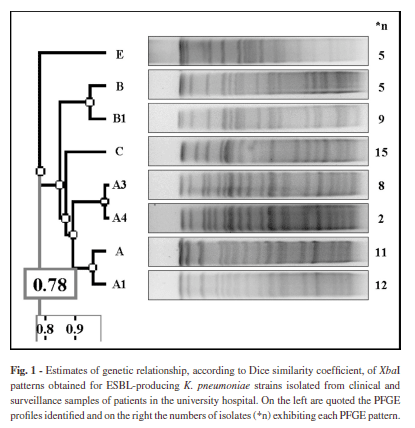
PFGE: Four main PFGE patterns (A, B, C and E) and four subtypes (A1, A3, A4 and B1) were obtained from 67 (69.8%) ESBL-Kp isolates (Fig. 1). The subtypes were classified according to previously established interpreting criteria (TENOVER et al. 1995). The other 29 (30.2%) strains presented unrelated profiles. Pattern A and subtypes A1, A3 and A4 were grouped and called only A*, and pattern B and subtype B1 were called only B*. Pattern A* was observed in 33 (34.4%) strains, pattern B* in 14 (14.6%) strains, pattern C in 15 (15.6%) strains, and pattern E in five (5.2%) strains. The distribution of the different PFGE patterns among the hospital wards during the studied period is shown in Fig. 2. From isolates showing genetically related profiles, 53 (79.1%) were responsible for colonizations. For this group detection of blaSHV genes was observed in 56 (83.6%) of 67 strains, blaTEM in five (7.5%) and blaCTX-M in seven (10.4%) isolates. In strains showing unrelated patterns (29), it was possible to observe detection of blaSHV genes in 22 (75.9%), blaTEM in 11 (37.9%) and blaCTX-M in 16 (55.2%) strains.
DISCUSSION
Production of ESBL has been frequently described worldwide, in several Enterobacteriaceae species, isolated from diverse clinical samples (QUINTEROS et al. 2003, CELENZA et al. 2006). In the present study, during one year it was detected one to three ESBL encoding genes (bla) in 127 isolates of nine Enterobacteriaceae species. It was not possible to establish the relationship among the several antimicrobial susceptibility patterns observed with the bla genes detected, once we carried out only a group identification. In regard to PFGE patterns, each of them presented a predominant susceptibility profile, although they had different bla genes combinations.
All the three groups of genes searched by PCR were detected, alone or in several combinations. Encoding genes for the SHV family were detected in 63% of the isolates, in six different species, from which 88.7% were K. pneumoniae. Most enzymes from this group are ESBL, except for SHV-1 and SHV-11, and according to the literature most K. pneumoniae strains posses the chromosomal gene blaSHV-1 (BABINI & LIVERMORE 2000, HOWARD et al. 2002, LIVERMORE 2008). Hence, the identification of blaSHV genes in the major part of ESBL-Kp strains in this study was already expected, although we cannot affirm those are ESBL-encoding genes, mostly in strains presenting more than one bla gene. On the other hand, blaSHV genes were also detected in other four species (E. coli, M. morganii, P. mirabilis and E. cloacae), what shows their spread potential among different strains. Group TEM was identified in 17.3% of isolates in five different species (K. pneumoniae, K. oxytoca, E.coli, M. morganii and P. mirabilis), generally in association with SHV and/or CTX-M genes. Therefore, it was not possible to presume if they were responsible for the ESBL phenotype or not, once they could be non-ESBL encoding genes associated to another gene expressing the extended-spectrum phenotype (LIVERMORE 2008). On the other hand, co-production of different ESBLs by a unique strain is a common finding in several studies (LI et al. 2003, PATERSON et al. 2003, VIGNOLI et al. 2005, MINARINI et al. 2007), so the carriage of two or three ESBL encoding genes by the TEM-producing strains of our study could also have happened. CTX-M ESBL was detected in 33.9% of isolates, in all species, except for P. rettgeri. This emerging group of enzymes is prevalent and endemic in South America (CELENZA et al. 2006, VILLEGAS et al. 2008) and, similarly to our work, some other studies in Argentina and Bolivia reported the finding of blaCTX-M genes in a great variety of Enterobacteriaceae species (QUINTEROS et al. 2003, CELENZA et al. 2006). ESBL-producing isolates in which none of the genes searched was detected could be carrying genes from other groups of extended-spectrum enzymes.
Klebsiella pneumoniae was the species with the highest number of isolates (75.6%) in this study. It is the most frequent ESBL-producing species, especially in Latin America, and its prevalence in Brazil is around 50%, while in the United States it is 5% and in Europe, 15 to 20% (SADER et al. 2001, GALES et al. 2002, NOGUEIRA et al. 2006, VILLEGAS et al. 2008). As a nosocomial pathogen, it can cause severe morbidity and mortality in hospitalized patients, and once it is part of intestinal flora, rectal colonization can be strongly associated to infection by ESBL- Kp (GUPTA et al. 2003). In addition to its intrinsic resistance to ampicillin due to the presence of chromosomal β-lactamase SHV-1 (BABINI & LIVERMORE 2000), it can also express resistance due to other β-lactamase classes like metalloenzymes and AmpC, and/or due to porine losses (MENDES et al. 2004, LINCOPAN et al. 2005, DENTON 2007). In Brazil, there are some reports of multi-drug resistant hospital-acquired K. pneumoniae (MENDES et al. 2004, LINCOPAN et al. 2005, PEIRANO et al. 2009). In the present study, four main PFGE patterns (A*, B*, C, and E) were obtained from 67 (69.8%) ESBL-Kp isolates. Figure 2 shows the distribution of all patterns among the hospital wards during the studied period, emphasizing that there was clearly four groups of genetically related strains and several isolated strains with genetic unrelated profiles. Patterns A* and B* were found mostly in the neonatal ward, neonatal ICU (B only) and pediatric ICU (A only), where a few unrelated patterns were identified (Fig. 2). These wards have patients with fragile host defenses, receiving excessive manipulation and a high antimicrobial intake, which could have been the predisposing factors for infections and spread of resistance genes (BRADFORD 2001, GUPTA et al. 2003). An outbreak took place in the Neonatal Ward during our study, and it is described elsewhere (CASSETARI et al. 2009). In Figure 2 we can see that pattern A* was isolated from June 2004 to January 2005, and pattern B* from December 2004 to February 2005. The "substitution" of pattern A* by B* in the neonatal ward was coincident with the infection control measures that were reinforced by the institution during the outbreak investigation. Although pattern C was identified in 15 strains, seven of them were isolated from a unique patient who was hospitalized in Neonatal ICU, Pediatric ICU and Pediatric Ward during all the study period (data not shown) (Fig. 2). Pattern E was identified in three strains from patients of Neonatal Ward and in two strains from patients of Pediatric ICU, between November and December of 2004. After this period, no more strains from this pattern were isolated until the end of the study period (July of 2005) (Fig. 2). K. pneumoniae isolated from samples of patients from Adult ICU, IMC, General and Surgical wards, showed only unrelated patterns (Fig. 2). According to another study, when isolates are genetically distinct, the group of resistant microorganisms can be originated due to excessive antimicrobial selective pressure, which favors resistant phenotypes within an unrelated group of strains, turning to be a problem of endemic resistance of different groups of organisms (TOSIN et al. 2003). Also, approximately 30% of those unrelated strains were isolated from patients who had been transferred from other hospitals or had previous hospital stays in our institution. In other reports, after genotyping K. pneumoniae isolates, it was detected several distinct patterns, suggesting horizontal transfer of resistance genes as a way of spread. Those strains contained blaCTX-M genes, which is exclusively located in mobile elements (LI et al. 2003, MINARINI et al. 2007). Similarly, in our study blaCTX-M genes were detected in a high number of isolates with unrelated molecular patterns (55.2%), and in a few strains of patterns A*, B*, C and E, what can suggest that in the environment of the studied institution there was also the spread of these resistance genes by conjugation. Some other observations in our results can reaffirm this kind of spread. TEM ESBLs were not detected in isolates from pattern A, but they were present in the subtype A1, which had one extra band in relation to A. The same occurred for subtypes A3 and A4, in which groups TEM and CTX-M were present only in the last one, what could have happened due to the presence of different plasmids among the isolates (LI et al. 2003). Despite the wide spread of ESBL encoding genes through plasmids, the high genetic variability found among Brazilian strains suggests that a continuous selection of resistant mutants occurs mostly due to the use of third-generation cephalosporins (SADER et al. 2001, GALES et al. 2002). Indiscriminate use of antimicrobials is strongly associated to multidrug resistance, colonization, hospital outbreaks and severe infections due to ESBL-producing microorganisms. This is considered a public health problem, and not only institutional, once solutions and control depend on local policies for restricted antimicrobial use (BRADFORD 2001, PATERSON et al. 2000).
According to the infection control committee of the studied hospital, ESBL-producing enterobacteria represent a great challenge for the control of infection and colonization in the institution, especially in Intensive Care Units. Our results suggest that the problem of dissemination of resistant clones and genes in hospital environment can occur due to contact with healthcare workers or to the patient's transferring between hospital units, once isolates presenting identical molecular patterns were observed in different wards, and some isolates showed patterns identical to the strain isolated from the hands of a nursing auxiliary in the institution (CASSETARI et al. 2009). Our data corroborate the importance of antibiotic use restriction and enhancement of prevention measures, mostly efficient hand-washing, contact isolation of colonized patients, appropriate cleansing and disinfection of materials and equipment, and monitoring of patients transference between hospital wards. Surveillance cultures are also of great value when appropriately applied (GUPTA et al. 2003).
In conclusion, several groups of bla genes were detected in clinical samples during one year in the studied institution, and further studies will be necessary to identify them precisely. Moreover, ESBL-Kp strains with unrelated molecular patterns carried bla genes located exclusively in mobile elements, suggesting horizontal gene transfer between isolates. On the other hand, similar and/or identical ESBL-Kp according to PFGE were disseminated among several hospital wards, suggesting spread in hospital environment.
ACKNOWLEDGEMENTS
This work was supported by a grant from the Collaborating Center of Sanitary Surveillance (CECOVISA - School of Public Health of University of São Paulo).
Received: 1 April 2009
Accepted: 18 June 2009
- 1. BABINI, G.S. & LIVERMORE, D.M. - Are SHV b-lactamases universal in Klebsiella pneumoniae?. Antimicrob. Agents Chemother., 44: 2230, 2000.
- 2. BATAGELJ, V. & MRVAR, A. - Pajek program for large network analysis. [online]. [cited 2008 Nov 30]. Available from: URL: http://vlado.fmf.uni-lj.si/pub/networks/pajek/, 1996.
- 3. BRADFORD, P.A. - Extended-spectrum β-lactamases in the 21st century: characterization, epidemiology, and detection of this important resistance threat. Clin. Microbiol. Rev., 14: 933-951, 2001.
- 4. CAO, V.; LAMBERT, T.; NHU, D.Q. et al. - Distribution of extended-spectrum β-lactamases in clinical isolates of Enterobacteriacea in Vietnam. Antimicrob. Agents Chemother., 46: 3739-3743, 2002.
- 5. CASSETTARI, V.C.; SILVEIRA, I.R.; DROPA, M. et al. - Risk factors for colonization of newborn infants during an outbreak of extended-spectrum β-lactamase-producing Klebsiella pneumoniae in an intermediate-risk neonatal unit. J. Hosp. Infect., 71: 340-347, 2009.
- 6. CELENZA, G.; PELLEGRINI, C.; CACCAMO, M. et al. - Spread of blaCTX-M-type and blaPER-2 β-lactamase genes in clinical isolates from Bolivian hospitals. J. Antimicrob. Chemother., 57: 975-978, 2006.
-
7CENTERS FOR DISEASE CONTROL AND PREVENTION CDC - PulseNet National molecular subtyping Network for foodborne and disease surveillance. One day standardized protocol for molecular subtyping of Escherichia coli O157:H7, non-typhoidal Salmonella serotypes, and Shigella sonnei by Pulsed Field Gel Eletrophoresis (PFGE). Atlanta, 2004.
- 8. CHAPMAN, P.A.; ELLIN, M.; ASHTON, R. & SHAFIQUE, W. - Comparison of culture, PCR and immunoassays for detecting Escherichia coli O157 following enrichment culture and immunomagnetic separation performed on naturally contaminated raw meat products. Int. J. Food Microbiol., 68: 11-20, 2001.
-
9CLINICAL AND LABORATORY STANDARDS INSTITUTE - Performance standards for antimicrobial disk tests Approved Standard. 18. ed. Wayne, CLSI, 2008. (CLSI document M100-S18 (M2)).
- 10. CORKILL, J.E.; CUEVAS, L.E.; GURGEL, R.Q.; GREENSILL, J. & HART, C.A. - SHV-27, a novel cefotaxime hydrolyzing b-lactamase, identified in Klebsiella pneumoniae isolates from a Brazilian hospital. J. Antimicrob. Chemother., 47: 463-465, 2001.
- 11. DENTON, M. - Enterobacteriaceae Int. J. Antimicrob. Agents, 29(suppl. 3): S9-S22, 2007.
- 12. GALES, A.C.; SADER, H.S. & JONES, R.N. - Urinary tract infection trends in Latin American hospitals: report from the SENTRY antimicrobial surveillance program (1997-2000). Diagn. Microbiol. infect. Dis., 44: 289-299, 2002.
- 13. GUPTA, A.; AMPOFO, K.; RUBENSTEIN, D. & SAIMAN, L. - Extended-spectrum β-lactamase-producing Klebsiella pneumoniae infections: a review of the literature. J. Perinatol., 23: 439-443, 2003 .
- 14. HOWARD, C.;. DAAL, A.; KELLY, G. et al. - Identification and minisequencing-based discrimination of SHV β-lactamases in nosocomial infection-associated Klebsiella pneumoniae in Brisbane, Australia. Antimicrob. Agents Chemother., 46: 659-664, 2002.
- 15. LI, C.R.; LI, Y. & ZHANG, P.A. - Dissemination and spread of CTX-M extended-spectrum β-lactamases among clinical isolates of Klebsiella pneumoniae in central China. Int. J. Antimicrob. Agents, 22: 521-525, 2003.
- 16. LINCOPAN, N.; McCULLOCH, J.A.; REINERT, C. et al. - First isolation of metallo-β-lactamase-producing multiresistant Klebsiella pneumoniae from a patient in Brazil. J. clin. Microbiol., 43: 516-519, 2005.
- 17. LIVERMORE, D.M. - Defining an extended-spectrum β-lactamase. Clin. Microbiol. Infect., 14(suppl. 1): 3-10, 2008.
- 18. MENDES, C.; KIFFER, C.; SEGURA, A. et al. -Klebsiella pneumoniae with multiple antimicrobial resistance. Braz. J. infect. Dis., 8: 109-111, 2004.
- 19. MINARINI, L.A.R.; GALES, A.C.; PALAZZO, I.C.V. & DARINI, A.L.C. - Prevalence of community-occurring extended-spectrum β-lactamase-producing Enterobacteriaceae in Brazil. Curr. Microbiol., 54: 335-341, 2007.
- 20. MINARINI, L.A.R.; CLÍMACO, E.C.; GUIMARÃES, D.B. et al. - Clonal transmission of ESBL-producing Klebsiella spp. at a university hospital in Brazil. Curr. Microbiol., 56: 587-591, 2008.
- 21. NOGUEIRA, K.S.; HIGUTI, I.H.; NASCIMENTO, A.J. et al. - Occurrence of extended-spectrum beta-lactamases in Enterobacteriaceae isolated from hospitalized patients in Curitiba, southern Brazil. Braz. J. infect. Dis., 10: 390-395, 2006.
- 22. PATERSON, D.L.; MULAZIMOGLU, L.; CASELLAS, J.M. et al. - Epidemiology of ciprofloxacin resistance and its relationship to extended-spectrum β-lactamase production in Klebsiella pneumoniae isolates causing bacteremia. Clin. infect. Dis., 30: 473-478, 2000.
- 23. PATERSON, D.L.; HUJER, K.M.; HUJER, A.M. et al - Extended-spectrum β-lactamases in Klebsiella pneumoniae bloodstream isolates from seven countries: dominance and widespread prevalence of SHV- and CTX-M-type β-lactamases. Antimicrob. Agents Chemother., 47: 3554-3560, 2003.
- 24. PEIRANO, G.; SEKI, L.M.; VAL PASSOS, V.L. et al - Carbapenem-hydrolysing beta-lactamase KPC-2 in Klebsiella pneumoniae isolated in Rio de Janeiro, Brazil. J. Antimicrob. Chemother., 63: 265-268, 2009.
- 25. QUINTEROS, M.; RADICE, M.; GARDELLA, N. et al - Extended-spectrum β-lactamases in Enterobacteriaceae in Buenos Aires, Argentina, Public Hospitals. Antimicrob. Agents Chemother., 47: 2864-2867, 2003.
- 26. SADER, H.S.; GALES, A.C.; PFALLER, M.A. et al - Pathogen frequency and resistance patterns in Brazilian hospitals: summary of results from three years of the SENTRY antimicrobial surveillance program. Braz. J. infect. Dis., 5: 200-214, 2001.
- 27. SILVA DIAS, R.C.; BORGES-NETO, A.A.; FERRAIUOLI, G.I.D. et al - Prevalence of AmpC and other β-lactamases in enterobacteria at a large urban university hospital in Brazil. Diagn. Microbiol. infect. Dis., 60: 79-87, 2008.
- 28. TENOVER, F.C.; ARBEIT, R.D.; GOERING, R.V. et al - Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel eletrophoresis: criteria for bacterial strain typing. J. clin. Microbiol., 33: 2233-2239, 1995.
- 29. TOSIN, I.; SILBERT, S. & SADER, H.S. - The use of molecular typing to evaluate the dissemination of antimicrobial resistance among Gram-negative rods in Brazilian Hospitals. Braz. J. infect. Dis., 7: 360-369, 2003.
- 30. VIGNOLI, R.; VARELA, G.; MOTA, M.I. et al - Enteropathogenic Escherichia coli strains carrying genes encoding the PER-2 and TEM-116 extended-spectrum β-lactamases isolated from children with diarrhea in Uruguay. J. clin. Microbiol., 43: 2940-2943, 2005.
- 31. VILLEGAS, M.V.; KATTAN, J.N.; QUINTEROS, M.G. & CASELLAS, J.M. - Prevalence of extended-spectrum β-lactamases in South America. Clin. Microbiol. Infect., 14: 154-158, 2008.
Publication Dates
-
Publication in this collection
26 Aug 2009 -
Date of issue
Aug 2009
History
-
Received
01 Apr 2009 -
Accepted
18 June 2009