Abstracts
The first dengue fever epidemic in the State of Rondônia (western region of Brazil) was recorded in 1997, without laboratory confirmation. Following this, there was an epidemic in Manaus, in the neighboring State of Amazon, in 1998, in which DENV-1 and DENV-2 viruses were isolated from patients. In the present paper, the serotype characterization of the dengue virus isolated from patients with clinically suspected dengue in Porto Velho, Rondônia, between 2001 and 2003 is described. One hundred and fifty blood samples were collected between the first and fifth days of symptoms. Seventy samples of virus isolates were subjected to dengue identification by means of RT-PCR using universal primers for the NS1 gene of DENV, which amplifies a 419 bp fragment. The amplicons obtained were subjected to enzymatic digestion to characterize the viral serotypes. All the samples analyzed were DENV-1. A nucleotide sequence randomly selected from one amplicon, which was also DENV-1, presented 98% similarity to sequences from Southeast Asia that were obtained from GenBank.
Dengue fever; RT-PCR; Enzymatic digestion
A primeira epidemia de febre do dengue no Estado de Rondônia, Região Ocidental do Brasil foi registrado em 1997, sem confirmação laboratorial. Em seguida, houve uma epidemia descrita em 1998, em Manaus, no vizinho Estado do Amazonas, onde os vírus DENV-1 e DENV-2 foram isolados de pacientes. No presente artigo, foi descrito a caracterização do sorotipo do vírus dengue isolado de pacientes com suspeitas clinicas de dengue em Porto Velho, Rondônia, entre 2001 a 2003. Foram coletadas 150 amostras de sangue, entre primeiro e quinto dia de sintomas. Setenta amostras de vírus isolados foram submetidas a identificação do dengue pela RT-PCR usando primers universais para gene da NS1 do DENV que amplifica um fragmento de 419pb. O amplicon obtidos foram submetidos a digestão enzimática para caracterização do sorotipo viral. Todas as amostras analisadas foram DENV-1. A seqüência nucleotídica de um dos amplicons aleatoriamente selecionada também DENV-1 demonstrou 98% similaridade com as seqüências do Sudeste Asiático obtidas no GenBank.
Febre do dengue; RT-PCR; Digestão enzimática
ARTICLES ARTIGOS
Characterization of dengue virus serotype 1 in epidemics in Porto Velho, Rondônia, in 2001-2003
Caracterização do sorotipo 1 do vírus da dengue de epidemias entre 2001-2003 em Porto Velho, Rondônia
Deusilene Souza Vieira; Eduardo Rezende Honda; Soraya Santos Pereira; Glauciane da Silva Bifano; Mauro Shugiro Tada; Weber Cheli Batista
Instituto de Pesquisa em Patologias Tropicais (IPEPATRO), Porto Velho, RO
Centro de Pesquisa em Medicina Tropical (CEPEM), Porto Velho, RO
Endereço para correspondência Endereço para correspondência: Dra. Deusilene Sousa Vieira. BR-364/Km 4,5, Caixa Postal 87 78900-970 Porto Velho, RO, Brasil. Telefax: 55 69 3225-3304 e-mail: deusylenebio@hotmail.com
ABSTRACT
The first dengue fever epidemic in the State of Rondônia (western region of Brazil) was recorded in 1997, without laboratory confirmation. Following this, there was an epidemic in Manaus, in the neighboring State of Amazon, in 1998, in which DENV-1 and DENV-2 viruses were isolated from patients. In the present paper, the serotype characterization of the dengue virus isolated from patients with clinically suspected dengue in Porto Velho, Rondônia, between 2001 and 2003 is described. One hundred and fifty blood samples were collected between the first and fifth days of symptoms. Seventy samples of virus isolates were subjected to dengue identification by means of RT-PCR using universal primers for the NS1 gene of DENV, which amplifies a 419 bp fragment. The amplicons obtained were subjected to enzymatic digestion to characterize the viral serotypes. All the samples analyzed were DENV-1. A nucleotide sequence randomly selected from one amplicon, which was also DENV-1, presented 98% similarity to sequences from Southeast Asia that were obtained from GenBank.
Key-words: Dengue fever. RT-PCR. Enzymatic digestion.
RESUMO
A primeira epidemia de febre do dengue no Estado de Rondônia, Região Ocidental do Brasil foi registrado em 1997, sem confirmação laboratorial. Em seguida, houve uma epidemia descrita em 1998, em Manaus, no vizinho Estado do Amazonas, onde os vírus DENV-1 e DENV-2 foram isolados de pacientes. No presente artigo, foi descrito a caracterização do sorotipo do vírus dengue isolado de pacientes com suspeitas clinicas de dengue em Porto Velho, Rondônia, entre 2001 a 2003. Foram coletadas 150 amostras de sangue, entre primeiro e quinto dia de sintomas. Setenta amostras de vírus isolados foram submetidas a identificação do dengue pela RT-PCR usando primers universais para gene da NS1 do DENV que amplifica um fragmento de 419pb. O amplicon obtidos foram submetidos a digestão enzimática para caracterização do sorotipo viral. Todas as amostras analisadas foram DENV-1. A seqüência nucleotídica de um dos amplicons aleatoriamente selecionada também DENV-1 demonstrou 98% similaridade com as seqüências do Sudeste Asiático obtidas no GenBank.
Palavras-chaves: Febre do dengue. RT-PCR. Digestão enzimática.
Dengue is at present the most prevalent virus-borne disease in the world, with particularly high incidence in tropical and subtropical regions6 9. The dengue virus belongs to the Flavivirus genus, and its genome contains a single-strand positive RNA molecule12 14. There are four dengue virus serotypes, which are designated DENV-1, DENV-2, DENV-3 and DENV-4 on the basis of their antigenic characteristics1 19 20. Dengue virus infections produce a range of clinical manifestations, going from asymptomatic and mild symptomatic infections known as dengue fever (DF), to severe disease that frequently has a lethal outcome and is known as dengue hemorrhagic fever (DHF) and dengue shock syndrome (DSS)7 13 16. The epidemiological situation of the Central, Southern and Caribbean areas of the Americas now resembles that of Southeast Asia, with over one million DF cases in 2002 and more than 30 countries affected. In the same year, DHF occurred in more than 20 Latin American countries with 17,000 cases including 225 fatalities8.
In Brazil, while suspected clinical DF cases had previously been recorded, the first dengue outbreak with laboratory confirmation occurred in 1981, in Boa Vista (northern Amazon area), with detection of DENV-1 and DENV-4 serotypes, probably derived from the neighboring Venezuelan endemic areas. However, widespread dissemination of DENV-1 only began after its first occurrence in Rio de Janeiro (southeastern Brazil), in 1986, when some urban areas in the Northeast were also affected18.
Rondônia State, in the western area of the Brazilian Amazon region, with 1,130,847 inhabitants11, has a strategic situation for the propagation of dengue viruses for a variety of reasons. It is situated in a tropical area and has an extensive border with Bolivia. In addition, the State of Rondônia is an important intersection point for river and roads transportation routes that connect with southern and central areas of Brazil and Bolivia. These have made it possible for new infectious agents to be imported, including different dengue serotypes. Over recent decades, the State of Rondônia has received hundreds of thousands of migrants from several regions of Brazil, and many of the state's inhabitants habitually travel back and forth to places where dengue outbreaks occur. Porto Velho, the state capital, with 373,917 inhabitants11 is in a particularly sensitive situation, principally because of the presence of a river port that receives most of the grain production from the central-western region of Brazil, for export to Europe and North America. The BR364 highway that connects Porto Velho to the central and southern states of Brazil has heavy traffic of trucks and buses, and the port area is known as an important stopover point for truck drivers and passengers coming or going to Manaus in the north or to Mato Grosso in the east. Finally, Porto Velho has a very poor sanitation system, which favors mosquito breeding in all outlying areas of the city.
The first dengue outbreak in Porto Velho occurred in 1999 but no serotyping of the dengue virus responsible was established. Our laboratory, which was created in June 2000, made it possible to start serological and molecular epidemiological studies. In the present communication, we present the molecular characterization of the dengue virus responsible for the Porto Velho outbreak that began at the end of 2001 and extended to the end of 200315.
MATERIALS AND METHODS
Study area and patient participation. From April 2001 to September 2003, 150 clinically suspected cases of dengue were followed at the Dr Hamilton Gondin Clinical Unit of the city of Porto Velho and at the Tropical Medicine Research Center of Rondônia (CEPEM), as shown in Table 1. Both of these clinical units are situated on the periphery of Porto Velho. Patients were invited to participate in the survey, and when they accepted the invitation, they signed an informed consent document for their participation that had been drawn up by the Ethics Committee of CEPEM. One blood sample was collected from each of the participants.
Sample collection and virus isolation. Blood samples of 5ml were collected in dry tubes from each patient and left at 37°C for approximately 30 minutes, for clot formation. One hundred microliters of serum was diluted tenfold in Leibowitz-15 tissue culture medium and inoculated into C6/36 (Aedes albopictus) cell monolayers in 24-well plates, one well for each patient. The excess serum was stored in a freezer at -70°C. The cells were cultivated in Leibowitz L-15 medium containing 10% fetal calf serum and an antibiotic-antimycotic solution (Invitrogen). Positive controls were prepared using C6/36 cells infected with dengue virus type 1 (Mochizuki strain), and the negative controls were C6/C36 cells that were kept uninfected. After inoculation, the microplates were incubated at 28°C for 7 days4 17. The viral infection was confirmed by the indirect immunofluorescence (IF) technique using dengue polyclonal specific antibodies that were kindly provided by Dr Vincent Deubel of the Pasteur Institute.
RNA Extraction. RNA was extracted from the infected C6/C36 cell culture supernatants by the TRIzol method5 with slight modifications3, as follows: 500µl of the samples were mixed with 750µl of TRIzol® and agitated vigorously in a vortex. After incubation of the mixture for five minutes at room temperature, 150l of chloroform was added and the mixture was agitated gently for 15 seconds and centrifuged at 12,000rpm for 15 minutes at 4°C. The RNA present in the upper aqueous phase was precipitated with 500µl of propanol alcohol and centrifuged at 14,000rpm for 10 minutes at 4°C. The propanol alcohol was discarded and the precipitated RNA was washed with 500µl of 75% ethanol and centrifuged at 14,000rpm for five minutes at 4°C. The excess ethanol was removed and the sediment was dried in a concentrator (Eppendorf, Germany). The RNA was suspended in 50µl of DEPC water and stored at -70°C.
Reverse transcription-polymerase chain reaction. For the reverse transcription, 5µl of the RNA extracts were added to a solution containing 3µg of the hexamer primer pd(N)6 (Invitrogen, USA) and 10mM of dNTPs (Invitrogen, USA). The sample was heated at 65°C for five minutes, and 4µl of 5X buffer, 2l of DTT 0.1M, 10U/l of RNase inhibitor and 200U/l of SuperScriptTM (Invitrogen, USA) were added to the solution, which had a final volume of 20µl. The mixture was incubated at 42°C for one hour and, finally, incubated at 70°C for 10 minutes in order to denature the reverse transcriptase.
For the polymerase chain reaction, 4µl of the cDNA were mixed with 50pmol of the AD3 and AD4 universal dengue primers that anneal to the NS1 gene of the virus9, and with 0.1mM of dNTPs, 1U of Taq DNA-polymerase (Invitrogen, USA), 5µl of 10x buffer and 42µl of distilled water. The cDNA was denatured at 94°C for five minutes, and the mixture was subjected to 35 thermal cycles of 94°C for one minute, 55°C for one minute and 72°C for one minute. The amplicons were detected after electrophoresis on 1.5% agarose gel stained with 1g/ml of ethidium bromide and viewed under UV light.
Assays with restriction enzyme. To characterize the dengue virus serotype 1, treatment with Kpn I was carried out, which promotes amplicon digestion in two fragments of 164bp and 256bp. Kpn I does not digest amplicons of dengue virus serotypes 2, 3 and 4. To characterize the dengue virus serotype 2, treatment with Hind III was carried out, which allows amplicon digestion in two fragments of 194 bp and 226 bp. Hind III does not digest amplicons of dengue virus serotypes 1, 3 or 43.
For digestion, the mixture included 8µl of PCR product containing amplicons, 10U/µl of the restriction enzyme and 1µl of buffer 10x (Invitrogen, USA). The mixture was incubated at 40°C for 3h. After incubation, the amplicon digestion products were electrophorized on 2.5% agarose gel stained with 1µg/ml of ethidium bromide and were then analyzed under UV light.
Cloning and sequencing of NS1 samples. For cloning and sequencing the NS1 gene of dengue virus type 1 from isolated samples, two pairs of primers were designed using the Edit Sequence of the DNA Star package. The first pair, named NS1-1, amplified one fragment of 498bp corresponding to the N terminal portion and a second pair named NS1-2 amplified one fragment of 558bp corresponding to the C terminal portion. For this experiment, three dengue virus amplicons derived from three different patients were cloned in pGEM® T easy vector, in accordance with the manufacturer's instructions. The sequencing of the primers is shown in Table 1.
The amplicons obtained by cloning with NS1-1 and NS1-2 pairs of primers were directly sequenced in an ABI PRISM 310 Genetic Analyzer (PE Biosynthesis, USA) using the Big Dye Terminator Cycle Sequencing kit (PE Biosynthesis, USA), in conformity with manufacturer's manual. The sequence obtained was subjected to a BLAST search on the NCBI home page, to ensure that the samples isolated were of the dengue virus.
RESULTS
Isolation and characterization of dengue virus. Out of the 150 C6/C36 cell cultures inoculated with sera from patients clinically suspected of dengue infection, 70 were positive for dengue virus by immunofluorescence and all of them gave positive RT/PCR results, with generation of the 419bp amplicon, as shown in Figure 1.
Restriction enzyme digestion of amplicons. The amplicons obtained from the 70 virus isolates were subjected to restriction enzyme digestion for characterization of dengue virus serotypes. All samples were characterized as having dengue virus type 1, as shown in Figure 2.
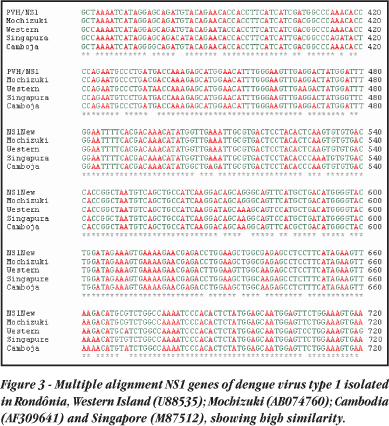
Nucleotide sequencing. One dengue NS1 amplicon from a virus isolated in Porto Velho was selected for nucleotide sequencing. The nucleotide sequence showed high homology with others from Southeast Asia and with previously sequencing, with 98% identification, as shown in Figure 3.
DISCUSSION
Dengue virus serotyping and characterization is important because it enables determination of the origin of a circulating virus and the potential impact of this pathogen on the population. RT-PCR, using universal dengue primers, followed by virus characterization by restriction enzymatic digestion is a quick and sensitive sequence of techniques that can be used for dengue epidemiological surveillance3 10.
In the present study, these techniques were used to analyze 150 serum samples from clinically suspected patients, which were collected during the epidemics that occurred in the city of Porto Velho from 2001 to 2003. Seventy samples inoculated in C6/36 cell cultures were shown to be positive for the infections, as detected by indirect immune fluorescence, and their corresponding cDNAs were amplified with the primers AD3 and AD4, which were developed by Henchal et al10, to produce amplified fragments of the NS1 gene with 419bp10.
All amplicons from the 70 positive samples were subjected to digestion by the enzymes Kpn I and Hind III for determination of viral serotypes 1 and 2, respectively. Two digestion fragments (256bp and 164bp) were observed for 60 samples corresponding to serotype 1 of the dengue virus. Ten samples were characterized as DENV-1 or DENV-2, and these were probably another circulating type.
The multiple alignment of sequences obtained between dengue type 1 isolated in Rondônia and dengue type 1 isolated in other places in the world (Genbank) and samples sequenced in Rio de Janeiro showed an identity of 98%. This degree of similarity suggests that the virus responsible for the Porto Velho dengue epidemics of 2001-2003 was derived from southern areas of the country and was brought in by people travelling overland. This is, for the moment, the entry route that should be subjected to greater epidemiological surveillance.
ACKNOWLEDGEMENTS
To Dr. Antônio Benedito and Sérgio De Paula (Molecular Virology Laboratory of the School of Medicine, University of Ribeirão Preto).
Recebido para publicação em 5/12/2006
Aceito em 2/4/2007
- 1. Chambers TJ, Hahn CS, Galler R, Rice CM. Flavivirus genome organization, expression and replication. Annual review of microbiology, 44: 649-688, 1990.
- 2. Chomczynski P, Sacchi N. Singer-step method of RNA isolation b acid guanidinium thiocyanate-phenol-chroform extraction. Analytical Biochemistry 162: 156-159, 1987.
- 3. De Paula SO, Pires Neto RJ, Correa JA, Assumpção SR, Costa ML, Lima DM, Fonseca BAL. The use of reverse transcription polymerase chain reaction (RT-PCR) for the rapid detection and identification of dengue virus in and endemic region. Transactions of the Royal Society of tropical Medicine and Hygiene 96: 266-269, 2002.
- 4. Figueiredo LTM. Uso de células de Aedes albopictus C6/36 na propagação e classificação de arbovírus das famílias Togaviridae, Flaviviridae, Bunyaviridae e Rhabdoviridae. Revista da Sociedade Brasileira em Medicina Tropical 23:13-18,1990.
- 5. Figueiredo LTM. Patogenia das infecções pelos vírus dengue. Virology 32:15-20, 1998.
- 6. Gubler DJ. Dengue and dengue hemorrhagic fever in the Americas. P.R. International journal of health sciences 6: 107-111, 1987a.
- 7. Gubler DJ. Dengue and dengue hemorrhagic fever: its history and resurgence as a global health problem. In: Gubler DJ, Kuno G (eds) Dengue and dengue hemorrhagic fever. CAB international, London United Kingdom, New York p.1-22, 1997.
- 8. Guzman MG, Kouri G. Dengue an dengue hemorragic fever in the Americas: lessons and challenges. Journal Clinical Virology 27:1-13, 2003.
- 9. Halstead SB. Pathogenesis of dengue: challenges to molecular biology. Science. 239: 476-481, 1988.
- 10. Henchal EA, Polo SI, Vorndam V, Yaemsiri C, Innis BI, Hoke CH. Sensitivity and specifity of a universal primer se for the rapid diagnosis of dengue virus infecions by polymerase chain reaction and nucleic acid hibridization. American Journal of Tropical Medicine and Hygiene 45:418-428, 1991.
-
11Ministério do Planejamento. Instituto Brasileiro de Geografia Estatística. Censo 2000.
- 12. Monath TP. Flaviviruses In: Fields BN, Knipe DM (eds) Virology. Raven New York, p.763-814, 1990.
- 13. Pinheiro FP, Corber SJ. Global situation of dengue and hemorrhagic fever, and its emergence in the Americas. World Health Organization 50:161-168, 1997.
- 14. Rice CM. Flaviviridae: the viruses and their replication. In: Fields BN, Knipe DM (eds) Fields virology, 3nd edition. Lippincott-Raven Publisher, Philadelphia, PA, p.931-959. 1996
- 15. Serviço Nacional de Atendimento Médico-SINAM/Agência de Vigilância Sanitária-AGEVISA/Secretária de Estado de Saúde de Rondônia-SESAU/RO. Casos notificados de dengue Estado de Rondônia, 2006.
- 16. Tauil PL. Urbanization and dengue ecology. Caderno de Saúde Pública. 17 (supl): 99-102, 2001.
- 17. Tesh RB. A method for the isolation and identification of dengue viruses, using mosquito cell culture. American Journal of Tropical Medicine and Hygiene 28: 1053-1059, 1979.
- 18. Teixeira MG, Costa MCN, Baffeto ML, Mota E. Dengue e febre hemorrágica do dengue no Brasil: que tipo de pesquisas a sua tendência, vigilância e experiências de controle indicam ser necessárias. Cadernos de Saúde Pública 21:1307-1315, 2005.
- 19. Vorndam V, Kuno G, Rosado N. A PCR-restriction enzyme technique for determining dengue virus subgroups within serotypes. Journal of Virology Methods 48: 237-244, 1994a.
- 20. Vorndam V, Nogueira RMR, Trent DW. Restriction enzyme analysis of American region dengue viruses. Archives of Virology 136: 191-196, 1994b.
Endereço para correspondência:
Publication Dates
-
Publication in this collection
17 July 2007 -
Date of issue
June 2007
History
-
Accepted
02 Apr 2007 -
Received
05 Dec 2006