Abstract:
INTRODUCTION
Characterization of Mycobacterium tuberculosis (MTB) isolates by DNA fingerprinting has contributed to tuberculosis (TB) control. The aim of this study was to determine the genetic diversity of MTB isolates from Tehran province in Iran.
METHODS
MTB isolates from 60 Iranian and 10 Afghan TB patients were fingerprinted by standard IS6110-restriction fragment length polymorphism (RFLP) analysis and spoligotyping.
RESULTS
The copy number of IS6110 ranged from 10-24 per isolate. The isolates were classified into 22 clusters showing ≥ 80% similarity by RFLP analysis. Fourteen multidrug-resistant (MDR) isolates were grouped into 4 IS6110-RFLP clusters, with 10 isolates [71% (95% CI: 45-89%)] in 1 cluster, suggesting a possible epidemiological linkage. Eighteen Iranian isolates showed ≥ 80% similarity with Afghan isolates. There were no strains with identical fingerprints. Spoligotyping of 70 isolates produced 23 distinct patterns. Sixty (85.7%) isolates were grouped into 13 clusters, while the remaining 10 isolates (14.2%) were not clustered. Ural (formerly Haarlem4) (n = 22, 31.4%) was the most common family followed by Central Asian strain (CAS) (n = 18, 25.7%) and T (n = 9, 12.8%) families. Only 1strain was characterized as having the Beijing genotype. Among 60 Iranian and 10 Afghan MTB isolates, 25% (95% CI: 16-37) and 70% (95% CI: 39-89) were categorized as Ural lineage, respectively.
CONCLUSIONS
A higher prevalence of Ural family MTB isolates among Afghan patients than among Iranian patients suggests the possible transmission of this lineage following the immigration of Afghans to Iran.
Keywords:
Mycobacterium tuberculosis; IS6110-RFLP; Spoligotyping.
INTRODUCTION
Tuberculosis (TB) is a global health problem that infects millions of people worldwide each year. According to the 2014 World Health Organization (WHO) global TB report, in 2013 there were 9.0 million new TB cases, which resulted in 1.5 million deaths11. World Health Organization (WHO). Global Tuberculosis report. WHO 2014.. Despite a promising decline in the incidence rate of TB in Iran from 36 cases per 100,000 people in 1990 to 17 cases per 100,000 people in 201022. World Health Organization (WHO). Global tuberculosis control. WHO 2011., TB continues to be a major public health concern in Iran. Risk factors such as drug addiction, human immunodeficiency virus (HIV) infection, drug-resistant Mycobacterium tuberculosis (MTB) strains, population ageing33. Alavi SM, Alavi L. Review on Epidemiology, Diagnosis, Occupational Hazards and Management of Pulmonary Tuberculosis in Elderly: A Guide for General Physicians Working in the Health Network Setting, Khuzestan, Iran. Jundishapur. J Microbiol 2013; 6: e6677., and sharing borders with 4 countries that have high rates of TB incidence, namely Afghanistan, Pakistan, Turkmenistan, and Iraq44. Mozafari M, Farnia P, Afraei M, Nezhad ZD, Masjedi MR, Velayati AA. Molecular diversity of Mycobacterium tuberculosis strains indifferent provinces of Iran. Iran J Microbiol 2013; 5:366-373.) (55. Haeili M, Darban Sarokhalil D, Fooladi AAI, Javadpour S, Hashemi A, Siavoshi F, et al. Spoligotyping and drug resistance patterns of Mycobacterium tuberculosis isolates from five provinces of Iran. Microbiology Open 2013; 2:988-996.,have contributed to a persistent problem of TB in Iran. According to a recent report, approximately 21 per 100,000 Iranians suffered from TB in 201366. World Health Organization (WHO). Global tuberculosis control. WHO 2013..
Tehran, the capital of Iran, is the 5th biggest city in the world. It has a population of more than 12 million people and an annual population growth rate of 1.4477. Statistical Centre of Iran. Selected findings of the 2011 national population and housing census 2011.. The population density (persons per km2) and average number of people per household in Tehran are 890 and 3.3, respectively77. Statistical Centre of Iran. Selected findings of the 2011 national population and housing census 2011.. Over the years, Tehran has experienced inward migration of people from elsewhere in Iran and from neighboring countries. More than 450,000 Afghans currently live in Tehran77. Statistical Centre of Iran. Selected findings of the 2011 national population and housing census 2011.. Significant developments in deoxyribonucleic acid (DNA) technology and molecular biology methods in the 21st century have greatly assisted the rapid diagnosis, immediate initiation of therapy, and identification of the source of infection of TB, which are crucial for stopping its further transmission. DNA fingerprinting based on IS6110 is an internationally standardized genotyping technique. IS6110 restriction fragment length polymorphism (RFLP) analysis relies on the analysis of IS6110 copy numbers and their locations within the genome of MTB. It is considered to be the most effective method for discriminating between MTB isolates among the molecular techniques that have been applied to the epidemiological study of TB88. Żaczek A, Brzostek A, Gorna A, Sajduda A, Wojtasik A, Dziadek J. Application of FLiP method for differentiation of Mycobacterium tuberculosis strains in comparison to commonly used methods, spoligotyping and MIRU-VNTR typing. Polish J Microbiol 2013; 62:73-76.. Even though it is time-consuming, labor-intensive, and requires successive DNA purification, and despite the difficulty of analyzing strains with a low copy number of IS6110 (< 6 copies), it is still the gold standard method for typing MTB strains55. Haeili M, Darban Sarokhalil D, Fooladi AAI, Javadpour S, Hashemi A, Siavoshi F, et al. Spoligotyping and drug resistance patterns of Mycobacterium tuberculosis isolates from five provinces of Iran. Microbiology Open 2013; 2:988-996.) (99. Supply P, Allix C, Lesjean S, Cardoso-Oelemann M, Rüsch-Gerdes S, Willery E, et al. Proposal for standardization of optimized mycobacterial interspersed repetitive unit-variable-number tandem repeat typing of Mycobacterium tuberculosis. J Clin Microbiol 2006; 44:4498-4510.. The spoligotyping method, which is a rapid polymerase chain reaction (PCR)-based method requiring small amounts of genomic DNA, has not been used as widely as RFLP, but it is suitable for differentiating MTB complex and for identifying the Beijing strains of MTB. It is also useful for investigating the evolutionary genetics of MTB when it is used in conjugation with other genotyping assays1010. Lari N, Rindi L, Sola C, Bonanni D, Rastogi N, Tortoli E, et al. Genetic diversity, determined on the basis of katG463 and gyrA95 polymorphisms, Spoligotyping, and IS6110typing, of Mycobacterium tuberculosis complex isolates from Italy. J Clin Microbiol 2005; 43:1617-1624.. Using RFLP together with spoligotyping in molecular epidemiological studies of MTB is recommended for the analysis of strains with a low copy number of IS6110. This study aimed to identify the transmission patterns of TB and the genetic diversity of MTB isolates from Tehran using the IS6110-RFLP and spoligotyping methods.
METHODS
Study population and data collection
In our descriptive study, 70 isolates of MTB were selected from a collection of culture-positive TB patients diagnosed between July 2013-January 2015. All isolates were supplied by the Regional TB reference laboratory and Baghiyatollah Hospital. The isolates were identified as MTB by standard biochemical tests including the niacin accumulation test, nitrate reduction test, and heat-labile catalase test. MTB isolates were subjected to drug susceptibility testing by the proportion method on Lowenstein-Jensen medium. Sixty (85.7%) patients were Iranian and the remaining 10 (14.2%) patients were Afghan.The molecular biology assays were carried out by technicians who did not have access to the results of phenotypic analyses.
Bacterial growth and chromosomal DNA isolation
Mycobacterium tuberculosis isolates were grown on Lowenstein-Jensen medium for 4 weeks. Chromosomal DNA was extracted by the method described by Van Soolingen et al.1111. Van Soolingen D, Hermans P, De Haas P, Soll D, Van Embden J. Occurrence and stability of insertion sequences in Mycobacterium tuberculosis complex strains: evaluation of an insertion sequence-dependent DNA polymorphism as a tool in the epidemiology of tuberculosis. J Clin Microbiol 1991; 29:2578-2586.. Briefly, 2-3 full loops of colonies were scraped, transferred to a micro tube that contained 400µl Tris-ethylenediaminetetraacetic acid (EDTA) buffer (pH 8), and inactivated by heating in a water bath at 80°C- for 30 min. After inactivation, 50µl of 10mg/mL lysozyme was added, vortexed, and incubated overnight at 37°C. Sodium dodecyl sulfate (SDS)/proteinase K (70µl of 10% [w/v] SDS and 5µl of 10mg/mL of proteinase K) were added, briefly vortexed, and incubated at 65°C for 10 min. One hundred microliters of 5M NaCl and N-cetyl-N,N,N-trimethylammonium bromide (CTAB)-NaCl solution (4.1g of NaCl and 10 g of CTAB per 100mL) was added and incubated at 65°C for 10 min. An equal volume of chloroform: isoamyl alcohol (24:1v/v) was added. After centrifugation for 5 min, 0.6 volumes of isopropanol was added to the supernatant liquid to precipitate the DNA. After incubation for 20 min at −20°C and centrifugation for 15 minat 12,000 × g and 4°C, the pellet was washed once with 70% (v/v) ethanol, and the air-dried pellet was dissolved in 20µL of Tris-EDTA buffer.
Preparation of DNA probe by PCR
The IS6110 probe was a PCR-amplified probe of 245 base pairs containing the sequence of the right arm of IS6110 from the M. tuberculosis H37Rv reference strain. Primers for the generation of the IS6110 probe were the oligonucleotides INS-1 (5ʹ-CGTGAGGGCATCGAGGTG GC-3ʹ) and INS-2 (5ʹ-GCGTAGGCGTCGGTGACAAA-3ʹ)1212. Van Embden J, Cave MD, Crawford JT, Dale J, Eisenach K, Gicquel B, et al. Strain identification of Mycobacterium tuberculosis by DNA fingerprinting: recommendations for a standardized methodology. J Clin Microbiol 1993; 31: 406-409.) (1313. Park Y-K, Bai G-H, Kim S-J. Restriction fragment length polymorphism analysis of Mycobacterium tuberculosis isolated from countries in the western pacific region. J Clin Microbiol 2000; 38: 91-197..
The PCR master mix was prepared with final reagent concentrations as follows; 5.0µL 10×Tth polymerase PCR buffer, 4.0µL dNTPs (2.5mM of each dNTP), 5.0µL of each primer (50µg/µL), 0.1µL Taqpolymerase (5U/µL), and 0.9µL water (Bioline, London, UK). Genomic DNA from the M. tuberculosis control strain H37Rv was added. Thermocycling conditions for PCR were 94°C for 2 min, followed by 35 cycles of 94°C for 1 min, 65°C for 1 min, and 72°C for 2 min. The final elongation step was 72°C for 5 min. The amplicon was detected using agarose gel electrophoresis with ethidium bromide staining. The PCR product was purified using the High Pure PCR Product Purification Kit according to the manufacturer's instructions (Roche, Basel, Switzerland). The eluted solution containing the purified PCR product was stored at -20°C.
Random labeling of DNA probe
The purified PCR product was labeled using the Gene Images(tm) random prime labeling protocol (Amersham Biosciences, Buckinghamshire, UK) and detected with a digoxigenin (DIG)-High Prime Kit (Roche) following the manufacturer's instructions. The labeled probe was stored in the dark at −20°C up to 6 months.
RFLP
DNA fingerprinting was performed by a standardized procedure as described by van Embden and colleagues1212. Van Embden J, Cave MD, Crawford JT, Dale J, Eisenach K, Gicquel B, et al. Strain identification of Mycobacterium tuberculosis by DNA fingerprinting: recommendations for a standardized methodology. J Clin Microbiol 1993; 31: 406-409.. Briefly, chromosomal DNA (2-3.5µg) was restricted with PvuII at 37 °C overnight. The restriction fragments were separated overnight by 1% (w/v) agarose gel electrophoresis in 1× Tris-borate-EDTA buffer. The fragments were transferred onto positively charged nylon membranes using the capillary method.
A probe of 245 base pairs in length was synthesized by PCR amplification using previously published primers and procedures, and was labeled using a DIG-High Prime Nucleic Acid Labeling and Detection Kit (Roche). Hybridization and detection were carried out according to a previously described method1414. Thierry D, Brisson-Noël A, Vincent-Levy-Frebault V, Nguyen S, Guesdon J, Gicquel B. Characterization of a Mycobacterium tuberculosis insertion sequence, IS6110, and its application in diagnosis. J Clin Microbiol 1990; 28: 2668-2673.. After hybridization, the insertion sequences were visualized colorimetrically using the DIG DNA Labeling and Detection Kit (Roche) according to the manufacturer's instructions.
Spoligotyping
Spoligotyping analysis of the MTB strains was performed essentially as described by Kamerbeek et al.1515. Kamerbeek J, Schouls L, Kolk A, Van Agterveld M, Van Soolingen D, Kuijper S, et al. Simultaneous detection and strain differentiation of Mycobacterium tuberculosis for diagnosis and epidemiology. J Clin Microbiol 1997; 35:907-914.. Briefly, genomic mycobacterial DNA was extracted from bacteria by a previously described method1111. Van Soolingen D, Hermans P, De Haas P, Soll D, Van Embden J. Occurrence and stability of insertion sequences in Mycobacterium tuberculosis complex strains: evaluation of an insertion sequence-dependent DNA polymorphism as a tool in the epidemiology of tuberculosis. J Clin Microbiol 1991; 29:2578-2586.. Oligonucleotides DRa and DRb were used as primers to amplify the whole DR region by PCR. The amplified biotinylated products were hybridized to a set of 43 immobilized oligonucleotides, each of which corresponded to one of the unique spacer DNA sequences within the DR locus. After hybridization, the membrane was washed, incubated with streptavidin-peroxidase conjugate, and finally developed by chemiluminescence (Amersham Biosciences). The DNA extract of M. tuberculosis H37Rv was used as a positive control.
Analysis of the results from RFLP and spoligotyping analyses
RFLP analysis was performed using an internationally standardized procedure as described by van Soolingen and colleagues1616. Van Soolingen D, Qian L, De Haas P, Douglas JT, Traore H, Portaels F, et al. Predominance of a single genotype of Mycobacterium tuberculosis in countries of east Asia. J Clin Microbiol 1995; 33:3234-3238.. The IS6110 fingerprint patterns were analyzed using GelCompar II software version 6.5 (Applied Maths, Sint-Martens-Latem, Belgium).
Spoligotyping results were compared in binary format with the results in the SITVIT2 database (Pasteur Institute of Guadeloupe) (available online at http://www.pasteur-guadeloupe.fr:8081/SITVITDemo). We defined a cluster as 2or more isolates from different patients with identical spoligotype patterns. Unique (nonclustered) spoligotypes were defined as those that did not cluster with any other sample in this study.
RESULTS
The RFLP patterns of 70 isolates obtained from 60 Iranian and 10 Afghan patients were determined. The age of the patients ranged from 16-84 years.Young patients (≤45 years old) represented 61% of the study population. Fourteen of the isolates were previously identified as multi-drug resistant (MDR) due to showing resistance to at least 2 first-line anti-TB drugs, including rifampicin and isoniazid.
IS6110 RFLP clustering
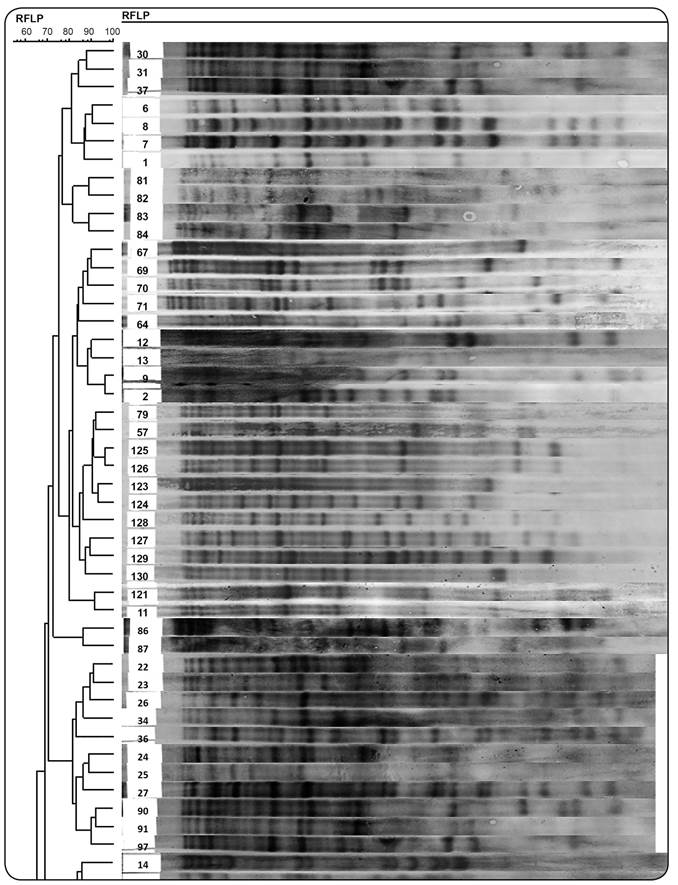
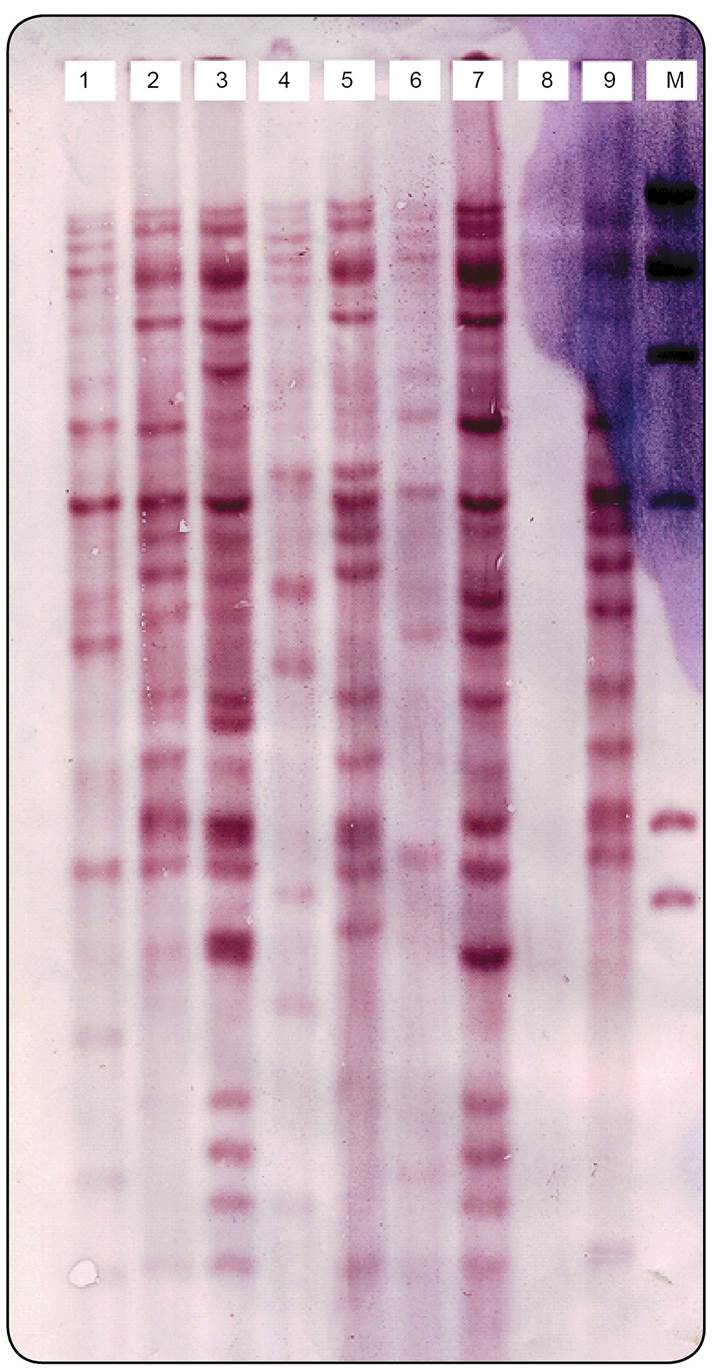
The copy number of IS6110 in each isolate was determined based on the number of bands hybridizing with the probe. The IS6110 copy number per isolate varied from 10-24 copies (Figure 1). According to band-based similarity coefficient of 80%, all fingerprinted isolates were grouped in to 22 clusters. The largest cluster contained 71% of the MDR isolates (n = 10). The remaining 4MDR isolates were distributed in 3 other clusters. All isolates from Afghan patients were grouped in 5 clusters and1 of these clusters contained only 4 Afghan isolates without any Iranian isolates. The remaining 6 Afghan isolates were clustered with those of Iranian patients. Figure 2 depicts part of an RFLP-based dendrogram of the studied isolates.
IS6110-based DNA fingerprints of Mycobacterium tuberculosis strains isolated from Tehran. M: marker; DNA: deoxyribonucleic acid.
A partial RFLP-based dendrogram of the analyzed Mycobacterium tuberculosis isolates, which were clustered based on the strains in each cluster having a band-based similarity coefficient of ≥ 80% as determined using GelCompar II software version 6.5 (Applied Maths, Sint-Martens-Latem, Belgium). RFLP: restriction fragment length polymorphism.
Spoligotyping
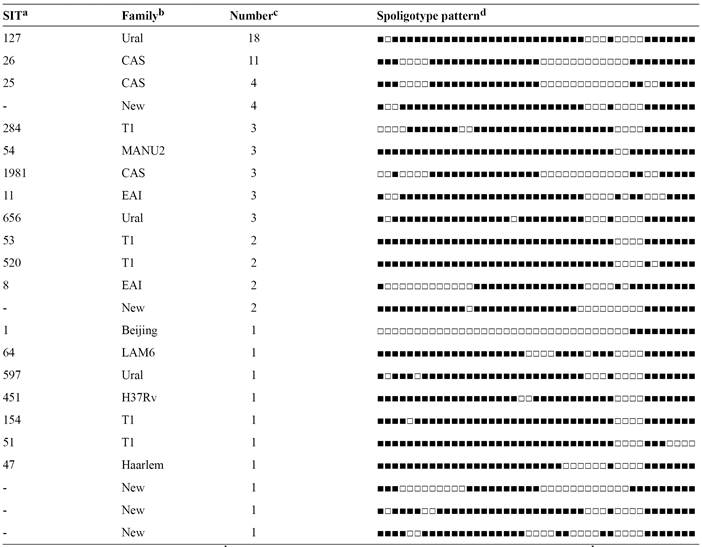
The spoligotyping of 70 isolates produced 23 different spoligo patterns (Table 1). Overall, 60 (85.7%) isolates were classified into 13 clusters, while the remaining 10 (14.2%) isolates did not form clusters due to having low similarity with the other isolates. The 2 largest clusters (Ural/Spoligotype International Type [SIT] 127 and Central Asian strains [CAS]/SIT26) contained 18 and 11 members respectively. Two clusters each contained 4 members, 5 clusters each had 3 members, and the remaining 4 clusters were each composed of 2 members. When considering all spoligotypes, the Ural (formerly Haarlem4) family was the most prevalent (n = 22, 31.4%), followed by CAS (n= 18, 25.7 %), T (n = 9, 12.8 %), East African Indian (EAI) (n = 5, 7%), MANU2 (n = 3, 4.2%), Latin American-Mediterranean (LAM) (n = 1, 1.4%), Beijing (n= 1, 1.4 %), Haarlem (n = 1, 1.4%), and H37Rv (n = 1, 1.4%). The obtained patterns from the remaining 9 isolates did not match spoligotyping patterns deposited in SITVIT2 database and were considered to represent orphan genotypes.Figure 3 illustrates a spoligotyping autoradiograph containing hybridization patterns of some of the studied samples.
Spoligotyping autoradiograph depicting hybridization patterns of amplified DNA samples obtained from Mycobacterium tuberculosis isolates from Tehran. DNA: deoxyribonucleic acid.
DISCUSSION
DNA typing of M. tuberculosis strains is based on the presence of polymorphisms that are generated by variability in both the copy numbers and chromosomal positions of IS6110 among clinical isolates of M. tuberculosis. Most members of M. tuberculosis complex have multiple copies of IS6110, but Mycobacterium bovis contains only 1 copy1717. Jagielski T, van Ingen J, Rastogi N, Dziadek J, Mazur PK, Bielecki J. Current methods in the molecular typing of Mycobacterium tuberculosis and other mycobacteria. BioMed Res Int 2014.. In this work, we did not find any isolate having less than 10 copies of IS6110 and copy numbers ranged from 10-24.
Alonso and colleagues reported that copy numbers of IS6110 ranged from 0-25 in their studied M. tuberculosis strains1818. Alonso H, Samper S, Martín C, Otal I. Mapping IS6110 in high-copy number Mycobacterium tuberculosis strains shows specific insertion points in the Beijing genotype. BMC genomics 2013; 14:422. A previous study from Iran showed that copy numbers of IS6110 in M. tuberculosis strains ranged from 5-8, and the mean was 111919. Doroudchi M, Kremer K, Basiri EA, Kadivar MR, Van Soolingen D, Ghaderi AA. IS6110-RFLP and spoligotyping of Mycobacterium tuberculosis isolates in Iran. Scand J Infect Dis 2000; 32:663-668..
The percentages of male and female patients in the study population were 51% and 48%, respectively. This is comparable to the proportions reported in a study from central Iran in 2013 (male = 48.5%, female = 51%)2020. Farazi A, Jabbariasl M, Tadayon K, Mossavar N, Keshavarz R, davood Hoseini S. Comparison of the genetic convergence between mycobacterium strains by three RFLP-based methods in central province of Iran. Iran J Med Sci 2014; 17:401..
The IS6110-RFLP fingerprinting analysis grouped 70 isolates into 22 clusters. Of 14 MDR isolates, 10 (71%) were grouped into 1 large cluster, signifying that these strains may have been epidemiologically linked, and their transmission rate was relatively higher than the other strains of MTB. Having similar fingerprints, which results in a higher clustering rate of isolates, has been attributed to recent transmission in the community2121. Roetzer A, Schuback S, Diel R, Gasau F, Ubben T, di Nauta A, et al. Evaluation of Mycobacterium tuberculosis typing methods in a 4-year study in Schleswig-Holstein, Northern Germany. J Clin Microbiol 2011; 49:4173-4178., whereas isolates that demonstrate unique fingerprints are likely to be associated with the reactivation of a latent infection2222. Kato-Maeda M, Small PM. Topic in Review: How molecular epidemiology has changed what we know about tuberculosis. West J Med 2000; 172:256.. Among the IS6110-RFLP clusters, 8 contained 5 or more isolates. The identification of clusters containing a large number of isolates indicates an over-representation of a group of closely related strains in a population. There was no cluster consisting of identical (100% similarity) fingerprints, meaning that each isolate had a unique fingerprinting result, which indicates that no laboratory cross-contamination occurred2323. Yang Z, Bates J, Eisenach K, Cave M. Secondary Typing of Mycobacterium tuberculosis Isolates with Matching IS6110 Fingerprints from Different Geographic Regions of the United States. J Clin Microbiol 2001; 39: 1691-1695..
Eighteen Iranian isolates were clustered with Afghan isolates due to sharing ≥ 80% similarity in their fingerprints. This indicates that there were transmissions of TB between members of the 2 ethnicities. While all 70 isolates showed unique fingerprinting patterns when analyzed using IS6110-RFLP, spoligotyping analysis grouped these 70 isolates into only 23 patterns. Thus, IS6110-RFLP is a more powerful method for discriminating M. tuberculosis isolates than the spoligotyping method. A study conducted in India reported results in line with our findings; they reported that IS6110-RFLP analysis can determine the minimum evolution in the M. tuberculosis genome and has more discriminatory power than spoligotyping does2424. Varma-Basil M, Kumar S, Arora J, Angrup A, Zozio T, Banavaliker JN, et al. Comparison of spoligotyping, mycobacterial interspersed repetitive units typing and IS6110-RFLP in a study of genotypic diversity of Mycobacterium tuberculosis in Delhi, North India. Mem I Oswaldo Cruz 2011; 106: 524-535..
Spoligotyping has been applied for genotyping of MTB isolates alongside RFLP, since it is suitable for distinguishing Beijing from non-Beijing family strains. In this study, 23 distinct patterns were obtained by spoligotyping analysis. Sixty isolates (85.7%) were grouped into 13 clusters containing 2-18 members. The largest cluster that formed based on the results of spoligotyping (Ural/SIT127), which contained 18 members, was further split into 11 IS6110-RFLP clusters. The second largest cluster (CAS/SIT26), which had 11 members, was split into 8 IS6110-RFLP clusters. Only 1 isolate was characterized as having a Beijing family genotype. Ural family (31.4%) was found to be the most frequent lineage followed by the CAS (25.7%) and T (12.8 %) families. The Ural lineage (formerly the H4 sub-lineage) has recently been excluded from the Haarlem family based on not harboring the mgtC545 (CGC→CAC) mutation2525. Abadia E, Zhang J, dos Vultos T, Ritacco V, Kremer K, Aktas E, et al. Resolving lineage assignation on Mycobacterium tuberculosis clinical isolates classified by spoligotyping with a new high-throughput 3R SNPs based method. Infect Genet Evol 2010; 10:1066-1074.. Our results are in agreement with those of a study performed by Haeili et al.55. Haeili M, Darban Sarokhalil D, Fooladi AAI, Javadpour S, Hashemi A, Siavoshi F, et al. Spoligotyping and drug resistance patterns of Mycobacterium tuberculosis isolates from five provinces of Iran. Microbiology Open 2013; 2:988-996., who reported that the Ural (34%), CAS (24%), and T (18.2%) families were the first, second, and third most predominant families, respectively, in MTB isolates obtained from 5 different provinces in Iran between 2010-2012. In another study performed by Kazemian et al.2626. Kazemian H, Haeili M, Kardan YJ, Rezaei F, Gizaw FS, Zahednamazi F, et al. Antimycobacterial activity of linezolid against multidrug-resistant and extensively drug-resistant strains of Mycobacterium tuberculosis in Iran. Int J Antimicrob Agents 2015; 45:668-670., who studied the genetic diversity of MDR and extensively drug-resistant MTB isolates from Iran, the Beijing (32.5%), Ural (22.5%), and CAS (10%) families were found to be the most common.
All Afghan samples (n = 10) belonged to 3 spoligotyping families including Ural (SITs 127 and 656, n = 7, 70%), T (SITs 284 and 520, n = 2, 20%), and EAI (SIT11, n = 1, 10%). Ural family genotypes were found in 70% (n = 7/10) of Afghan patients, whereas they constituted 25% (n = 15/60) of Iranian samples.
This study describes the molecular epidemiology of TB in Tehran province, which is inhabited by both native and immigrant populations. The higher prevalence of Ural family genotypes among Afghans suggests the possible transmission of this genotype following the immigration of Afghans to Iran mainly to the capital. However, our data cannot support making a confident judgment about this possibility due to some limitations of our study, as follows: a) the small size of our studied population (70 MTB isolates) does not represent all TB cases identified in Tehran during 2013-2015; and b) there were no sociodemographic or clinical data available to enable analysis of the linkage of RFLP/spoligotyping results with other epidemiological determinants. More population-based epidemiologic studies with a larger number of samples are required to provide a better understanding of TB transmission in Iran.
-
1World Health Organization (WHO). Global Tuberculosis report. WHO 2014.
-
2World Health Organization (WHO). Global tuberculosis control. WHO 2011.
-
3Alavi SM, Alavi L. Review on Epidemiology, Diagnosis, Occupational Hazards and Management of Pulmonary Tuberculosis in Elderly: A Guide for General Physicians Working in the Health Network Setting, Khuzestan, Iran. Jundishapur. J Microbiol 2013; 6: e6677.
-
4Mozafari M, Farnia P, Afraei M, Nezhad ZD, Masjedi MR, Velayati AA. Molecular diversity of Mycobacterium tuberculosis strains indifferent provinces of Iran. Iran J Microbiol 2013; 5:366-373.
-
5Haeili M, Darban Sarokhalil D, Fooladi AAI, Javadpour S, Hashemi A, Siavoshi F, et al. Spoligotyping and drug resistance patterns of Mycobacterium tuberculosis isolates from five provinces of Iran. Microbiology Open 2013; 2:988-996.
-
6World Health Organization (WHO). Global tuberculosis control. WHO 2013.
-
7Statistical Centre of Iran. Selected findings of the 2011 national population and housing census 2011.
-
8Żaczek A, Brzostek A, Gorna A, Sajduda A, Wojtasik A, Dziadek J. Application of FLiP method for differentiation of Mycobacterium tuberculosis strains in comparison to commonly used methods, spoligotyping and MIRU-VNTR typing. Polish J Microbiol 2013; 62:73-76.
-
9Supply P, Allix C, Lesjean S, Cardoso-Oelemann M, Rüsch-Gerdes S, Willery E, et al. Proposal for standardization of optimized mycobacterial interspersed repetitive unit-variable-number tandem repeat typing of Mycobacterium tuberculosis J Clin Microbiol 2006; 44:4498-4510.
-
10Lari N, Rindi L, Sola C, Bonanni D, Rastogi N, Tortoli E, et al. Genetic diversity, determined on the basis of katG463 and gyrA95 polymorphisms, Spoligotyping, and IS6110typing, of Mycobacterium tuberculosis complex isolates from Italy. J Clin Microbiol 2005; 43:1617-1624.
-
11Van Soolingen D, Hermans P, De Haas P, Soll D, Van Embden J. Occurrence and stability of insertion sequences in Mycobacterium tuberculosis complex strains: evaluation of an insertion sequence-dependent DNA polymorphism as a tool in the epidemiology of tuberculosis. J Clin Microbiol 1991; 29:2578-2586.
-
12Van Embden J, Cave MD, Crawford JT, Dale J, Eisenach K, Gicquel B, et al. Strain identification of Mycobacterium tuberculosis by DNA fingerprinting: recommendations for a standardized methodology. J Clin Microbiol 1993; 31: 406-409.
-
13Park Y-K, Bai G-H, Kim S-J. Restriction fragment length polymorphism analysis of Mycobacterium tuberculosis isolated from countries in the western pacific region. J Clin Microbiol 2000; 38: 91-197.
-
14Thierry D, Brisson-Noël A, Vincent-Levy-Frebault V, Nguyen S, Guesdon J, Gicquel B. Characterization of a Mycobacterium tuberculosis insertion sequence, IS6110, and its application in diagnosis. J Clin Microbiol 1990; 28: 2668-2673.
-
15Kamerbeek J, Schouls L, Kolk A, Van Agterveld M, Van Soolingen D, Kuijper S, et al. Simultaneous detection and strain differentiation of Mycobacterium tuberculosis for diagnosis and epidemiology. J Clin Microbiol 1997; 35:907-914.
-
16Van Soolingen D, Qian L, De Haas P, Douglas JT, Traore H, Portaels F, et al. Predominance of a single genotype of Mycobacterium tuberculosis in countries of east Asia. J Clin Microbiol 1995; 33:3234-3238.
-
17Jagielski T, van Ingen J, Rastogi N, Dziadek J, Mazur PK, Bielecki J. Current methods in the molecular typing of Mycobacterium tuberculosis and other mycobacteria. BioMed Res Int 2014.
-
18Alonso H, Samper S, Martín C, Otal I. Mapping IS6110 in high-copy number Mycobacterium tuberculosis strains shows specific insertion points in the Beijing genotype. BMC genomics 2013; 14:422
-
19Doroudchi M, Kremer K, Basiri EA, Kadivar MR, Van Soolingen D, Ghaderi AA. IS6110-RFLP and spoligotyping of Mycobacterium tuberculosis isolates in Iran. Scand J Infect Dis 2000; 32:663-668.
-
20Farazi A, Jabbariasl M, Tadayon K, Mossavar N, Keshavarz R, davood Hoseini S. Comparison of the genetic convergence between mycobacterium strains by three RFLP-based methods in central province of Iran. Iran J Med Sci 2014; 17:401.
-
21Roetzer A, Schuback S, Diel R, Gasau F, Ubben T, di Nauta A, et al. Evaluation of Mycobacterium tuberculosis typing methods in a 4-year study in Schleswig-Holstein, Northern Germany. J Clin Microbiol 2011; 49:4173-4178.
-
22Kato-Maeda M, Small PM. Topic in Review: How molecular epidemiology has changed what we know about tuberculosis. West J Med 2000; 172:256.
-
23Yang Z, Bates J, Eisenach K, Cave M. Secondary Typing of Mycobacterium tuberculosis Isolates with Matching IS6110 Fingerprints from Different Geographic Regions of the United States. J Clin Microbiol 2001; 39: 1691-1695.
-
24Varma-Basil M, Kumar S, Arora J, Angrup A, Zozio T, Banavaliker JN, et al. Comparison of spoligotyping, mycobacterial interspersed repetitive units typing and IS6110-RFLP in a study of genotypic diversity of Mycobacterium tuberculosis in Delhi, North India. Mem I Oswaldo Cruz 2011; 106: 524-535.
-
25Abadia E, Zhang J, dos Vultos T, Ritacco V, Kremer K, Aktas E, et al. Resolving lineage assignation on Mycobacterium tuberculosis clinical isolates classified by spoligotyping with a new high-throughput 3R SNPs based method. Infect Genet Evol 2010; 10:1066-1074.
-
26Kazemian H, Haeili M, Kardan YJ, Rezaei F, Gizaw FS, Zahednamazi F, et al. Antimycobacterial activity of linezolid against multidrug-resistant and extensively drug-resistant strains of Mycobacterium tuberculosis in Iran. Int J Antimicrob Agents 2015; 45:668-670.
-
This work was supported by a grant from Tehran University of Medical Sciences (Project number 24554).
Data availability
Data citations
Statistical Centre of Iran. Selected findings of the 2011 national population and housing census 2011.
Publication Dates
-
Publication in this collection
Mar-Apr 2016
History
-
Received
02 Jan 2016 -
Accepted
14 Mar 2016