ABSTRACT
Maize plants can establish beneficial associations with plant growth-promoting bacteria. However, few studies have been conducted on the characterization and inoculation of these bacteria in the Amazon region. This study aimed to characterize endophytic bacteria isolated from maize in the Amazon region and to assess their capacity to promote plant growth. Fifty-five bacterial isolates were obtained from maize grown in two types of ecosystems, i.e., a cerrado (savanna) and a forest area. The isolates were characterized by the presence of the nifH gene, their ability to synthesize indole-3-acetic acid (IAA) and solubilize calcium phosphate (CaHPO4), and 16S rRNA partial gene sequencing. Twenty-four bacteria contained the nifH gene, of which seven were isolated from maize plants cultivated in a cerrado area and seventeen from a forest area. Fourteen samples showed the capacity to synthesize IAA and only four solubilized calcium phosphate. The following genera were found among these isolates: Pseudomonas; Acinetobacter; Enterobacter; Pantoea; Burkholderia and Bacillus. In addition, eight isolates with plant growth-promoting capacity were selected for a glasshouse experiment involving the inoculation of two maize genotypes (a hybrid and a variety) grown in pots containing soil. Inoculation promoted the development of the maize plants but no significant interaction between maize cultivar and bacterial inoculation was found. A high diversity of endophytic bacteria is present in the Amazon region and these bacteria have potential to promote the development of maize plants.
Keywords:
Biological nitrogen fixation; Zea mays L.; 16S rRNA
RESUMO
Plantas de milho podem estabelecer associações benéficas com bactérias promotoras do crescimento vegetal. No entanto, poucos estudos de caracterização e inoculação com essas bactérias foram realizados na região Amazônica. O objetivo deste estudo foi caracterizar bactérias endofíticas isoladas de milho na região Amazônica e avaliar sua capacidade de promover o crescimento. Cinquenta e cinco isolados bacterianos foram obtidos de plantas de milho cultivadas em diferentes tipos de ecossistemas, cerrado (savana) e área de floresta. Estes foram caracterizados pela presença do gene nifH, capacidade de sintetizar ácido indol-3-acético (AIA), solubilizar fosfato de cálcio (CaHPO4) e sequenciamento parcial do gene 16S rRNA. Vinte e quatro bactérias apresentaram o gene nifH. Destas, sete são oriundas de plantas cultivadas em uma área de cerrado e 17 em área de floresta. Quatorze bactérias demonstraram a capacidade de solubilizar AIA e quatro de solubilizar fosfato de cálcio. Entre estas bactérias, foram encontrados os seguintes gêneros: Pseudomonas; Acinetobacter; Enterobacter; Pantoea; Burkholderia e Bacillus. Oito isolados com capacidade de promoção do crescimento vegetal foram selecionados para experimento em casa de vegetação com a inoculação em dois diferentes genótipos de milho (um híbrido e uma variedade) em vasos com solo. Quanto à promoção de crescimento vegetal, a inoculação promoveu o desenvolvimento de plantas de milho, mas não foi encontrada interação significativa entre a cultivar de milho e a inoculação bacteriana. Na região Amazônica há uma diversidade de bactérias endofíticas e estas possuem potencial para promover o desenvolvimento de plantas de milho.
Palavras-chave:
Fixação biológica de nitrogênio; Zea mays L.; 16S rRNA
INTRODUCTION
Some bacteria that inhabit the rhizosphere and/or colonize plant roots can contribute significantly to plant development. Such bacteria are known as plant growth-promoting bacteria (PGPB) (Kloepper and Schroth 1978Kloepper, J.W.; Schroth, M.N. 1978. Plant growth-promoting rhizobacteria on radishes. In: International Conference on Plant Pathogenic Bacteria, 41978Angers. Proceedings Angers: Station de Pathologie Vegetale et Phytobacteriologie, p.879-882.). The specific genera that inhabit the rhizosphere and roots that have been shown to promote plant growth include Arthrobacter, Azoarcus, Azospirillum, Bacillus, Burkholderia, Enterobacter, Gluconacetobacter, Herbaspirillum, Klebsiella, Paenibacillus, Pseudomonas and Serratia (Spaepen et al. 2009Spaepen, S.; Vanderleyden, J.; Okon, Y. 2009. Plant growth-promoting actions of rhizobacteria. Advances in Botanical Research, 51: 283-320.). The mechanisms by which these bacteria promote plant growth commonly include phosphate solubilization, production of growth regulators and biological nitrogen fixation (Gulati et al. 2009Gulati, A.; Vyas, P.; Rahi, P.; Kasana, R.C. 2009. Plant growth-promoting and rhizosphere-competent Acinetobacter rhizosphaerae strain BIHB 723 from the cold deserts of the Himalayas. Current Microbiology, 58: 371-377.; Dobbelaere et al. 2003Dobbelaere, S.; Vanderleyden, J.; Okon, Y. 2003. Plant growth-promoting effects of diazotrophs in the rhizosphere. Critical Reviews in Plant Science, 22: 107-149.; Döbereiner and Pedrosa 1987Döbereiner, J.; Pedrosa, F.O. 1987. Nitrogen-fixing bacteria in non-leguminous crop plants. Science Tech, Madison, USA, 155p.). Although many studies have been conducted on these plant growth-promoting microorganisms, broad applications of these microbes in agriculture are still in the early stages of development.
Maize is a staple crop grown worldwide for both human and livestock consumption. The plant has high nutritional requirements, especially nitrogen. Therefore, association of maize with nitrogen-fixing (diazotrophic) bacteria could be an alternative to supply this demand. In fact, evidence has demonstrated responses to inoculation in some situations. According to Okon and Labandera-Gonzales (1994Okon, Y.; Labandera-Gonzales, C.A. 1994. Agronomic applications of Azospirillum: an evaluation of 20 years worldwide field inoculation. Soil Biology and Biochemistry, 26: 1591-1601.), results from 20 years of research involving inoculation of grasses (including maize) with Azospirillum spp. showed a 30% increase in crop yield in 60 to 70% of the experiments. Nevertheless, there is considerable variation in the responses to inoculation because of the effects of both the plant genotype and the bacterial strain used (Garcia de Salomone and Döbereiner 1996Garcia de Salomone, I.E.; Döbereiner, J. 1996. Maize genotype effects on the response to Azospirillum inoculation. Biology and Fertility of Soils, 21: 193-196.; Garcia de Salomone et al. 1996Garcia de Salomone, I.E.; Döbereiner, J.; Urquiaga, S.; Boddey, R.M. 1996. Biological nitrogen fixation in Azospirillum strain-maize genotype associations as evaluated by 15N isotope dilution technique. Biology and Fertility of Soils, 23: 249-256. ).
In Brazil, inoculants containing strains of Azospirillum have recently begun be commercially used for inoculation of maize and wheat crops, with increases in grain yield of approximately 30% in some cases (Hungria et al. 2010Hungria, M.; Campo, R.J.; Souza, E.M.; Pedrosa, F.O. 2010. Inoculation with selected strains of Azospirillum brasilense and A. lipoferum improves yields of maize and wheat in Brazil. Plant and Soil, 331: 413-425.). However, despite the commercial launch of these inoculants, it is recognized that further research is required to characterize and select well-adapted bacteria for use as inoculants. A diverse range of PGPB genera have been isolated from maize plants, including Azospirillum (Döbereiner and Pedrosa 1987Döbereiner, J.; Pedrosa, F.O. 1987. Nitrogen-fixing bacteria in non-leguminous crop plants. Science Tech, Madison, USA, 155p.), Herbaspirillum (Baldani et al. 1986Baldani, J.I.; Baldani, V.L.D.; Seldin, L.; Döbereiner, J. 1986. Characterization of Herbaspirillum seropedicae nov. sp. a root associated nitrogen fixing bacterium. International Journal of Systematic Bacteriology, 36: 86-93.), Burkholderia (Perin et al. 2006Perin, L.; Martínez-Aguilar, L.; Paredes-Valdez, G.; Baldani, J.I.; Estrada-de-los-Santos, P.; Reis, V.M.; Caballero-Mellado, J. 2006. Burkholderia silvatlantica sp. nov., a diazotrophic bacterium associated with sugar cane and maize. International Journal of Systematic and Evolutionary Microbiology, 56: 1931-1937.) and others detected by studies using molecular techniques (Roesch et al. 2008Roesch, L.F.W.; Camargo, F.A.O.; Bento, F.M.; Triplett, E.W. 2008. Biodiversity of diazotrophic bacteria within the soil, root and stem of field-grown maize. Plant and Soil, 302: 91-104.).
In a recent study in the state of Roraima (located in a low-latitude Amazon region in northern Brazil), 55 bacterial isolates were obtained from four maize cultivars (two varieties and two hybrids) grown in different types of ecosystems: a cerrado (savanna) and a forest area. The present study aimed to characterize these isolates and evaluate their capacity to promote the growth of maize in a glasshouse.
MATERIALS AND METHODS
The isolates were obtained from two previous experiments conducted in experimental fields of the Embrapa Roraima: a cerrado (savanna) area in Boa Vista municipality (N 02º 15'00'' and W 60º 39'54''), and in a forest area in Cantá municipality (N 02º 39'48'' and W 60º 50' 15''), Roraima State, Brazil. Maize plants (hybrids BRS 1010 and BRS 3003 and varieties BRS 4157 and BR 106) were sampled at the flowering stage. The plants were separated into stems and roots, washed, superficially disinfected in sodium hypochlorite (2%), macerated and streaked onto Dyg's solid culture medium (Rodrigues Neto et al. 1986Rodrigues Neto, J.; Malavolta Júnior, V.A.; Victor, O. 1986. Meio simples para o isolamento e cultivo de Xanthomonas campestris pv. citri tipo B. Summa Phytopathologica, 12: 16.). A total of 537 isolates were obtained. These isolates were tested in the semi-solid culture medium BMGM (Estrada-de-los-Santos et al. 2001Estrada-de-los-Santos, P.; Bustilios-Cristales, R.; Caballero-Mellado, J. 2001. Burkholderia, a genus rich in plant-associated nitrogen fixers with wide environmental and geographic distribution. Applied and Environmental Microbiology, 67: 2790-2798.) without nitrogen to assess pellicle formation. Diazotrophic bacteria exhibit typical growth in a semi-solid medium without nitrogen, i.e., the formation of a pellicle near the surface of the medium. The pellicle formation is the first indication that they might be diazotrophic and this was used to select the isolates. Of the 537 isolates, 55 presented pellicle formation and were selected for characterization. These isolates were deposited in the Collection of Microorganisms of Embrapa Roraima.
The 55 isolates were grown in Dyg's liquid medium (Rodrigues Neto et al. 1986Rodrigues Neto, J.; Malavolta Júnior, V.A.; Victor, O. 1986. Meio simples para o isolamento e cultivo de Xanthomonas campestris pv. citri tipo B. Summa Phytopathologica, 12: 16.) for 48 h and then DNA was extracted from 1 mL of the cell suspension using the RBC Bioscience kit (cat. YGB300, Taiwan), following the instructions provided. The primers PolF and PolR were used to amplify the nifH gene according to the conditions described by Poly et al. (2001Poly, F.; Monrozier, L.J.; Bally, R. 2001. Improvement in the RFLP procedure for studying the diversity of nifH genes in communities of nitrogen fixers in soil. Research in Microbiology, 152: 95-103.). The subsequent tests were performed only for the 24 isolates that were positive for nifH gene amplification.
Indole-3-acetic acid (IAA) production was determined according to Sarwar and Kremer (1995Sarwar, M.; Kremer, R.J. 1995. Determination of bacterially derived auxins using a microplate method. Letters in Applied Microbiology, 20: 282-285.), with a few modifications. Isolates were grown in Dyg's liquid culture medium (48 h; 30 °C; 150 rpm in the dark) containing tryptophan (100 µg mL-1), with three replicates. After incubation, the cultures were centrifuged (5 min; 520 g) and 150 µl of each supernatant was treated with 100 µl of Salkowski's reagent (2% w/v of 0.5 M FeCl3 in 35% v/v of HClO4) in a polystyrene plate and incubated in the dark for 30 min at room temperature (27°C) to develop a pink color, which is indicative of IAA production. The diazotrophic Azospirillum brasilense strain (BR11001T = SP7), which is known to produce IAA (Radwan et al. 2002Radwan, T.; Mohamed, Z.K; Reis, V.M. 2002. Production of indole-3-acetic acid by different strains of Azospirillum and Herbaspirillum spp. Symbiosis, 32: 39-53.), was used as a positive control.
Calcium phosphate dibasic (CaHPO4) solubilization was assessed using solid GL medium (Sylvester-Bradley et al. 1982Sylvester-Bradley, R.; Askawa, N.; Latorraca, S.; Magalhães, F.M.M.; Oliveira, L.A.; Pereira, R.M. 1982. Levantamento quantitativo de microrganismos solubilizadores de fosfatos na rizosfera de gramíneas e leguminosas forrageiras na Amazônia. Acta Amazônica, 12: 12-22.). Inoculation of the culture medium with bacteria was performed in points over the culture medium with four replicates. The formation of a clear halo around the colonies (in contrast with opaque culture medium) was observed ten days after incubation at 28°C, indicating the capacity to solubilize phosphate.
Amplification of the 16S rRNA gene was performed for the 24 nifH-positive isolates using the Y1 (5' TGG CTCAGAACGAACGCTGGCGGC 3') (Young et al. 1991Young, J.P.W.; Downer, H.L.; Eardly, B.D. 1991. Phylogeny of the phototrophic Rhizobium strain BTAil by polymerase chain reaction-based sequencing of a 16S rRNA gene segment. Journal of Bacteriology, 173: 2271-2277.) and B3 primers (5' TACCTTGTTACGACTTCACCCCAGCT 3'). The PCR products were used directly for partial sequencing using the primer Y1. The sequencing was performed using a 3730xl DNA sequencer (Applied Biosystems). Good quality sequences (Phred 20) were compared with the GenBank database (www.ncbi.nlm.nih.gov) using the Basic Local Alignment Search Tool (BLAST). The isolate sequences and type strains were aligned using the ClustalW routine in MEGA 5.01 software (Tamura et al. 2007Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. 2007. MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Molecular Biololgy and Evolution, 24: 1596-1599.). A phylogenetic tree was constructed using the "neighbor-joining" method and Kimura's 2-parameter model (Kimura 1980Kimura, M. 1980. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. Journal of Molecular Evolution, 16: 111-120.) in MEGA 5.01, performing 1000 repetitions. The 16S rRNA gene sequences were deposited in the GenBank with accession numbers KC412136-KC412159.
Eight isolates (ERR 543, ERR 544, ERR 547, ERR 549, ERR 553, ERR 555, ERR 559 and ERR 566) were assessed for their ability to promote the growth of maize. These strains were selected based on the presence of plant growth-promotion features (nifH gene, IAA and phosphate solubilization) as well as the genotypic diversity of the 16S rRNA gene. The experiment was performed in a glasshouse. Soil (typical oxisol) was collected from a depth of 0-20 cm in an experimental cerrado area and was used for the experiment. Before starting the experiment, soil fertility analysis was performed (Embrapa 2009Embrapa. 2009. Manual de análises químicas de solos, plantas e fertilizantes. Brasília: Embrapa Informação Tecnológica, 627p.). The chemical and physical properties were as follows: pH of 6.4 (H2O); 1.39 cmolc dm-3 of Ca2+; 0.42 cmolc dm-3 of Mg2+; 0.06 cmolc dm-3 of K+; 0.01 cmolc dm-3 of Al3+; 1.91 cmolc dm-3 of H++Al3+; 76.50 mg dm-3 of P (Mehlich-1); 49.4% base saturation; 7.8 g dm-3 of organic matter; and clay, silt and sand contents of 4, 2 and 94%, respectively.
The pots contained approximately 1.5 dm-3 of soil and were fertilized with 32 mg of N as urea; 40 mg of P2O5 as simple superphosphate; 90 mg of K2O as KCl; 1.5 g of CaCl2.7H2O; 0.7 mg of MgSO4.7H2O and 5 ml of a micronutrient solution (g l-1) containing 0.8 H3BO3; 1.8 CuSO4.5H2O; 1.8 FeCl3.6H2O; 1.0 CoSO4.7H2O; 2.4 MnCl2.4H2O; 3.2 ZnSO4.7H2O and 0.02 Na2Mo4.2H2O. This represents the basic fertilizer applied under field conditions in Roraima state (approximately 50 kg ha-1 of N, 90 kg ha-1 of P2O5 and 120 kg ha-1 of K2O) (Valle et al. 2013Valle, I.C.A.; Alves, J.M.A.; Silva, L.S.; Uchôa, S.C.P.; Albuquerque, J.A.A.; Silva, D.C.O. 2013. Produção do milho híbrido 30F35HR cultivado na savana de Roraima em diferentes densidades de plantio. Agro@mbiente online, 7: 294-303.).
The treatments consisted of inoculation with eight bacterial isolates from the current study, a control without inoculation or nitrogen application; a control without inoculation but with mineral nitrogen added on the 21st day (32 mg pot-1 of N as urea, i.e., approximately 50 kg ha-1 of N, in addition to N applied at sowing) and a control inoculated with the Azospirillum brasilense strain (BR11001T = SP7) without nitrogen application. The isolates were grown in Dyg's liquid medium (48 h; 28 ºC; 150 rpm) and 0.1 mL of cell suspension (approximately 10-9 cells mL-1 evaluated by serial dilution on Dyg's solid medium) was applied to each seed.
The maize genotype variety BR 106 and hybrid BRS 1010 were tested in a 2 (maize cultivars) x 11 (N sources) factorial experiment with four completely randomized replications. Maize seeds were sterilized in 92% ethanol for 30 sec, 5% hydrogen peroxide for 5 min, washed 5 times with sterilized water, and sown (four seeds per pot). After emergence, only two plants were retained in each pot. The plants were harvested 40 days after emergence, prior to flowering when the nitrogen fixation rates are generally at a maximum in plants. Shoots were separated from roots and both parts were dried (65 °C for 5 days). The variables analyzed were shoot dry matter, root dry matter, total biomass dry matter and nitrogen content in the shoot (total-N) (Embrapa 2009Embrapa. 2009. Manual de análises químicas de solos, plantas e fertilizantes. Brasília: Embrapa Informação Tecnológica, 627p.). Data were submitted to analysis of variance using SISVAR software (Ferreira 2008Ferreira, D.F. 2008. SISVAR: a program for statistical analysis and teaching. Revista Symposium, 6: 36-41.) and the averages were contrasted using the Scott-Knott test at a 5 % probability level.
RESULTS
Out of the 55 isolates that produced pellicle formation in the nitrogen-free medium known to favor the growth of diazotrophics, 24 were positive for amplification of the nifH gene (Table 1). Seven of the positive isolates were obtained from maize plants collected in the cerrado and 17 were from the forest area (Table 1).
Origin, feature of plant-growth promotion and genotypic characterization of diazotrophic bacteria isolated from maize in the Amazon region, Roraima state, Brazil.
Among the 24 isolates positive for the nifH gene, 15 were also positive for IAA synthesis. Four were obtained from the cerrado area and 11 were obtained from the forest area (Table 1). Regarding the capacity to solubilize calcium phosphate, only 5 of the 24 isolates presented this feature (Table 1), all of which were obtained from maize cultivated in the cerrado area. Furthermore, only two isolates (ERR 543 and ERR 544) had the ability to produce IAA as well as to solubilize calcium phosphate (Table 1).
Partial 16S rRNA gene sequencing of the 24 isolates indicated the presence of the genera Pseudomonas (13 isolates), Burkholderia (4), Acinetobacter (2), Enterobacter (2), Pantoea (2), and Bacillus (1) (Table 1). Through phylogenetic analysis, it was possible to verify that 13 isolates belonging to the genus Pseudomonas were grouped with high similarity with Pseudomonas nitroreducens (Figure 1), 2 isolates (ERR 551 and ERR 544) were grouped with Acinetobacter soli, 2 were close to Enterobacter sp. (ERR 553 and ERR 565), 2 were close to Pantoea agglomerans (ERR 564 and ERR 566), 2 were close to B. gladioli (ERR 545 and ERR 546), 2 were close to B. cepacia (ERR 542 and ERR 543), and 1 of the genus Bacillus (ERR 547) was close to B. thuringiensis (Figure 1).
Neighbor joining phylogeny based on 16S rRNA gene sequences showing the relationships between diazotrophic maize (shown in bold) and other reference strains. Bootstrap values greater than 50% are indicated at the nodes. Bar: 5 substitution per 100 nucleotide positions.
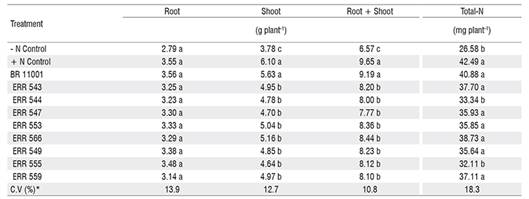
No significant interaction was observed between bacterial inoculation and maize genotype; therefore, only the average of the two cultivars is presented. The shoot and total dry matter of the inoculated and + Control nitrogen treatments were significantly different from those of the control without inoculation (- Control nitrogen), but similar responses were not observed for root dry matter (Table 2). In addition, the shoot and total dry matter of the plants inoculated with the new isolates were lower than in the + Control nitrogen treatment and the inoculation with Azospirillum brasilense (Table 2). Except for ERR 544 and ERR 555, all isolates showed a similar performance to that observed in the A. brasilense inoculation and + Control nitrogen treatment for total N, and these isolates presented values that were significantly higher than those in the - Control nitrogen treatment (Table 2).
DISCUSSION
Amplification of the nifH gene with the primers PolF and PolR was positive for 24 of the 55 isolates studied. The presence of the nifH gene is an indication of bacteria's ability to fix atmospheric nitrogen, although no nitrogenase activity was measured. Seventeen isolates were obtained from maize cultivated in a forest area and 7 from a cerrado area (Table 1). With respect to the plant growth-promotion features analyzed in this work, 17 isolates showed other direct mechanisms in addition to the presence of the nifH gene, i.e., IAA production or calcium phosphate solubilization (Table 1). IAA synthesis by microorganisms can stimulate root development, while the capacity to solubilize phosphate can nutrients more available to plants (Dobbelaere et al. 2003Dobbelaere, S.; Vanderleyden, J.; Okon, Y. 2003. Plant growth-promoting effects of diazotrophs in the rhizosphere. Critical Reviews in Plant Science, 22: 107-149.).
Similar to other studies, there was a high diversity of diazotrophic bacteria isolated from maize (Roesch et al. 2008Roesch, L.F.W.; Camargo, F.A.O.; Bento, F.M.; Triplett, E.W. 2008. Biodiversity of diazotrophic bacteria within the soil, root and stem of field-grown maize. Plant and Soil, 302: 91-104.; Montañez et al. 2012Montañez, A.; Blanco, A.R.; Barlocco, C.; Beracochea, M.; Sicardi, M. 2012. Characterization of cultivable putative endophytic plant growth promoting bacteria associated with maize cultivars (Zea mays L.) and their inoculation effects in vitro. Applied Soil Ecology, 58: 21-28.). Partial 16S rRNA sequencing revealed that the 24 bacterial isolates belonged to six different genera: Acinetobacter, Bacillus, Burkholderia, Enterobacter, Pantoea and Pseudomonas (Figure 1). All of these genera had previously been indicated as plant growth-promoting bacteria (Spaepen et al. 2009Spaepen, S.; Vanderleyden, J.; Okon, Y. 2009. Plant growth-promoting actions of rhizobacteria. Advances in Botanical Research, 51: 283-320.).
The isolates ERR 544 and ERR 551 were close to Acinetobacter soli, a species obtained from a forest soil (Kim et al. 2008Kim, D.; Baik, K.S.; Kim, M.S.; Park, S.C.; Kim, S.S.; Rhee, M.S.; Kwak, Y.S.; Seong, C.N. 2008. Acinetobacter soli sp. nov., isolated from forest soil. Journal of Microbiology, 46: 396-401.) but not a diazotrophic bacterium. However, within the genus Acinetobacter, A. oryzae has been described as diazotrophic in association with wild rice in China (Chaudhary et al. 2011Chaudhary, H.J.; Peng, G.; Hu, M.; He, Y.; Yang, L.; Luo, Y.; Tan, Z. 2011. Genetic Diversity of endophytic diazotrophs of the wild rice, Oryza alta and identification of the new diazotroph, Acinetobacter oryzae sp. nov. Microbial Ecology, 63: 813-821. ). In this study, we observed nifH gene amplification for both isolates (ERR 544 and ERR 551), indicating they may fix nitrogen. Moreover, Acinetobacter species have been found in the rhizosphere and roots and have shown the ability to promote plant growth through phosphate solubilization and/or the synthesis of auxins (Gulati et al. 2009Gulati, A.; Vyas, P.; Rahi, P.; Kasana, R.C. 2009. Plant growth-promoting and rhizosphere-competent Acinetobacter rhizosphaerae strain BIHB 723 from the cold deserts of the Himalayas. Current Microbiology, 58: 371-377.). According to our results, both isolates synthesized IAA, but only ERR 544 was able to solubilize phosphate (Table 1).
Four isolates belonged to the family Enterobacteriaceae according the 16S rRNA gene sequencing analysis and were close to the genera Enterobacter (ERR 553 and ERR 565) and Pantoea (ERR 564 and ERR 566) (Figure 1). Enterobacter and Pantoea are frequently found in the digestive tract of animals, and some of them are able to fix atmospheric nitrogen. They are also found in maize plants and are able to produce IAA (Montañez et al. 2012Montañez, A.; Blanco, A.R.; Barlocco, C.; Beracochea, M.; Sicardi, M. 2012. Characterization of cultivable putative endophytic plant growth promoting bacteria associated with maize cultivars (Zea mays L.) and their inoculation effects in vitro. Applied Soil Ecology, 58: 21-28.), similar to the isolates in the present study.
There were four isolates belonging to the genus Burkholderia, two of which were closely related to the Burkholderia cepacia complex (Figure 1), and three were able to solubilize calcium phosphates. Burkholderia spp. are known to promote the growth of grasses. They are able to fix nitrogen in free-living or endophytic form (Estrada-de-los-Santos et al. 2001Estrada-de-los-Santos, P.; Bustilios-Cristales, R.; Caballero-Mellado, J. 2001. Burkholderia, a genus rich in plant-associated nitrogen fixers with wide environmental and geographic distribution. Applied and Environmental Microbiology, 67: 2790-2798.), solubilize phosphates (Peix et al. 2001Peix, A.; Mateos, P.F.; Rodriguez-Barrueco, C.; Martinez-Molina, E.; Velazquez, E. 2001. Growth promotion of common bean (Pasheolus vulgaris L.) by a strain of Burkholderia cepacia under growth chamber conditions. Soil Biology and Biochemistry, 33: 1927-1935. ) and inhibit the growth of phytopathogens (Peix et al. 2001Peix, A.; Mateos, P.F.; Rodriguez-Barrueco, C.; Martinez-Molina, E.; Velazquez, E. 2001. Growth promotion of common bean (Pasheolus vulgaris L.) by a strain of Burkholderia cepacia under growth chamber conditions. Soil Biology and Biochemistry, 33: 1927-1935. ; Perin et al. 2006Perin, L.; Martínez-Aguilar, L.; Paredes-Valdez, G.; Baldani, J.I.; Estrada-de-los-Santos, P.; Reis, V.M.; Caballero-Mellado, J. 2006. Burkholderia silvatlantica sp. nov., a diazotrophic bacterium associated with sugar cane and maize. International Journal of Systematic and Evolutionary Microbiology, 56: 1931-1937.).
More than 50% of the isolates studied belonged to the genus Pseudomonas, which clearly indicates an abundance of isolates within this genus in association with maize in this Amazonian soil. Seven of them produced IAA but none were able to solubilize calcium phosphate (Table 1). In another study, Pseudomonas species were also found to be abundant in the maize rhizosphere, especially in sandy soils (Berge et al. 1991Berge, O.; Heulin, T.; Balandreau, J. 1991. Diversity of diazotroph populations in the rhizosphere of maize (Zea mays L.) growing on different French soils. Biology and Fertility of Soils, 11: 210-215.), but in a study in the South of Brazil, where the nifH gene was directly amplified from rhizospheric soil and maize plant tissue, Pseudomonas was one of the rarest genera (Roesch et al. 2008Roesch, L.F.W.; Camargo, F.A.O.; Bento, F.M.; Triplett, E.W. 2008. Biodiversity of diazotrophic bacteria within the soil, root and stem of field-grown maize. Plant and Soil, 302: 91-104.). Both areas where the maize was cultivated for the bacterial isolation in this study have sandy soil, but understanding the influence of soil characteristics on Pseudomonas populations requires further research.
The 13 Pseudomonas isolates were grouped together with P. nitroreducens (Figure 1), but because Pseudomonas species present high similarity in the 16S rRNA gene sequences (Tayeb et al. 2005Tayeb, L.A.; Ageron, E.; Grimont, F.; Grimont, P.A.D. 2005. Molecular phylogeny of the genus Pseudomonas based on rpoB sequences and application for the identification of isolates. Research in Microbiology, 156: 763-773.), other tests are necessary to confirm the taxonomic position of these isolates. Pseudomonas is one of the most diverse and ambiguous bacterial genera; it is found in several environments, such as sediments, plants, fungi, soil, water, the rhizosphere, and the phyllosphere of plants, oceans, animals, etc. Many Pseudomonas associated with plants promote growth and are able to produce phytohormones (Leveau and Gerards 2008Leveau, J.H.; Gerards, S. 2008. Discovery of a bacterial gene cluster for catabolism of the plant hormone indole 3-acetic acid. FEMS Microbiology Ecology, 65: 238-250.) and solubilize phosphates (Naik et al. 2008Naik, P.R.; Raman, G.; Narayanan, K.B.; Sakthivel, N. 2008. Assessment of genetic and functional diversity of phosphate solubilizing fluorescent pseudomonads isolated from rhizospheric soil. BMC Microbiology, 8: 230.). At least two species are diazotrophic: P. azotifigens and P. stutzeri (Hatayama et al. 2005Hatayama, K.; Kawai, S.; Shoun, H.; Ueda, Y.; Nakamura, A. 2005. Pseudomonas azotifigens sp. nov., a novel nitrogen-fixing bacterium isolated from a compost pile. International Journal of Systematic and Evolutionary Microbiology, 55: 1539-1544.), corroborating that the strains used in this study could be able to fix nitrogen.
The results obtained in the glasshouse trial showed that all isolates belonging to the six different genera were able to increase shoot dry matter compared with the control (Table 2). Some of the isolates increased the maize biomass and total-N by more than 30% compared with the control without inoculation (Table 2). Although the tested isolates have more than one mechanism for plant growth promotion, apparently biological nitrogen fixation was the most important because all of the isolates, except ERR 544, which belongs to the genus Acinetobacter, and ERR 555, which belongs to the genus Pseudomonas, contributed to significant N accumulation in shoots (Table 2).
In a study performed under field conditions in the state of Roraima with the inoculation of Herbaspirillum seropediacae in two maize cultivars (the hybrid BRS 1010 and the variety BSR 4157), a significant effect was observed in the grain yield of the hybrid due to inoculation, whereas a similar effect was not observed in the variety (Zilli et al. 2007Zilli, J.E.; Marson, L.C.; Reis, V.M.; Alves, G.C.; Baldani, L.D.; Cordeiro, A.C. 2007. Contribuição da bactéria diazotrófica Herbaspirillum seropedicae para o rendimento de grãos de arroz e milho em Roraima. Boa Vista: Embrapa Roraima, 20p. (Embrapa Roraima. Boletim de Pesquisa e Desenvolvimento, 6).). In this study, in contrast to the experiment conducted by Zilli et al. (2007) and other studies (Garcia de Salomone and Döbereiner 1996Garcia de Salomone, I.E.; Döbereiner, J. 1996. Maize genotype effects on the response to Azospirillum inoculation. Biology and Fertility of Soils, 21: 193-196.; Garcia de Salomone et al. 1996Garcia de Salomone, I.E.; Döbereiner, J.; Urquiaga, S.; Boddey, R.M. 1996. Biological nitrogen fixation in Azospirillum strain-maize genotype associations as evaluated by 15N isotope dilution technique. Biology and Fertility of Soils, 23: 249-256. ; Montañez et al. 2012Montañez, A.; Blanco, A.R.; Barlocco, C.; Beracochea, M.; Sicardi, M. 2012. Characterization of cultivable putative endophytic plant growth promoting bacteria associated with maize cultivars (Zea mays L.) and their inoculation effects in vitro. Applied Soil Ecology, 58: 21-28.), no influence of plant genotype was found in response to inoculation; the bacteria were the main factor promoting maize growth. Nevertheless, for these isolates, other genotypes must be tested to confirm this effect because it is known that the response to inoculation can result from the specificity of the plant-bacteria association, differences in the exudation of roots, and the efficiency of gas diffusion (Ladha et al. 1986Ladha, J.K.; Triol, A.G.; Daroy, M.L.G.; Caldo, G.; Ventura,W.; Watanabe, I. 1986. Plant associated N2-fixation (C2H2 reduction) by five rice varieties and relationship with plant growth characters as affected by straw incorporation. Soil Science and Plant Nutrition, 32: 91-106.).
The results of this study indicate a high diversity of Amazonian bacteria that are able to associate themselves with maize plants. These bacteria represent six distinct genera and are able to increase the growth of maize plants.
CONCLUSIONS
There is a high diversity of endophytic bacteria associated with maize plants in the Amazon region. These bacteria presented characteristics of plant growth-promoting rhizobacteria and the following genera were identified: Pseudomonas; Acinetobacter; Enterobacter; Pantoea; Burkholderia and Bacillus. The inoculation improved the growth of maize plants but no significant interaction between maize cultivar and bacterial inoculation was found.
ACKNOWLEDGEMENTS
We thank CNPq and Embrapa for the financial support in this study.
- Baldani, J.I.; Baldani, V.L.D.; Seldin, L.; Döbereiner, J. 1986. Characterization of Herbaspirillum seropedicae nov. sp. a root associated nitrogen fixing bacterium. International Journal of Systematic Bacteriology, 36: 86-93.
- Berge, O.; Heulin, T.; Balandreau, J. 1991. Diversity of diazotroph populations in the rhizosphere of maize (Zea mays L.) growing on different French soils. Biology and Fertility of Soils, 11: 210-215.
- Chaudhary, H.J.; Peng, G.; Hu, M.; He, Y.; Yang, L.; Luo, Y.; Tan, Z. 2011. Genetic Diversity of endophytic diazotrophs of the wild rice, Oryza alta and identification of the new diazotroph, Acinetobacter oryzae sp. nov. Microbial Ecology, 63: 813-821.
- Dobbelaere, S.; Vanderleyden, J.; Okon, Y. 2003. Plant growth-promoting effects of diazotrophs in the rhizosphere. Critical Reviews in Plant Science, 22: 107-149.
- Döbereiner, J.; Pedrosa, F.O. 1987. Nitrogen-fixing bacteria in non-leguminous crop plants. Science Tech, Madison, USA, 155p.
- Embrapa. 2009. Manual de análises químicas de solos, plantas e fertilizantes. Brasília: Embrapa Informação Tecnológica, 627p.
- Estrada-de-los-Santos, P.; Bustilios-Cristales, R.; Caballero-Mellado, J. 2001. Burkholderia, a genus rich in plant-associated nitrogen fixers with wide environmental and geographic distribution. Applied and Environmental Microbiology, 67: 2790-2798.
- Ferreira, D.F. 2008. SISVAR: a program for statistical analysis and teaching. Revista Symposium, 6: 36-41.
- Garcia de Salomone, I.E.; Döbereiner, J. 1996. Maize genotype effects on the response to Azospirillum inoculation. Biology and Fertility of Soils, 21: 193-196.
- Garcia de Salomone, I.E.; Döbereiner, J.; Urquiaga, S.; Boddey, R.M. 1996. Biological nitrogen fixation in Azospirillum strain-maize genotype associations as evaluated by 15N isotope dilution technique. Biology and Fertility of Soils, 23: 249-256.
- Gulati, A.; Vyas, P.; Rahi, P.; Kasana, R.C. 2009. Plant growth-promoting and rhizosphere-competent Acinetobacter rhizosphaerae strain BIHB 723 from the cold deserts of the Himalayas. Current Microbiology, 58: 371-377.
- Hatayama, K.; Kawai, S.; Shoun, H.; Ueda, Y.; Nakamura, A. 2005. Pseudomonas azotifigens sp. nov., a novel nitrogen-fixing bacterium isolated from a compost pile. International Journal of Systematic and Evolutionary Microbiology, 55: 1539-1544.
- Hungria, M.; Campo, R.J.; Souza, E.M.; Pedrosa, F.O. 2010. Inoculation with selected strains of Azospirillum brasilense and A. lipoferum improves yields of maize and wheat in Brazil. Plant and Soil, 331: 413-425.
- Kim, D.; Baik, K.S.; Kim, M.S.; Park, S.C.; Kim, S.S.; Rhee, M.S.; Kwak, Y.S.; Seong, C.N. 2008. Acinetobacter soli sp. nov., isolated from forest soil. Journal of Microbiology, 46: 396-401.
- Kimura, M. 1980. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. Journal of Molecular Evolution, 16: 111-120.
- Kloepper, J.W.; Schroth, M.N. 1978. Plant growth-promoting rhizobacteria on radishes. In: International Conference on Plant Pathogenic Bacteria, 41978Angers. Proceedings Angers: Station de Pathologie Vegetale et Phytobacteriologie, p.879-882.
- Ladha, J.K.; Triol, A.G.; Daroy, M.L.G.; Caldo, G.; Ventura,W.; Watanabe, I. 1986. Plant associated N2-fixation (C2H2 reduction) by five rice varieties and relationship with plant growth characters as affected by straw incorporation. Soil Science and Plant Nutrition, 32: 91-106.
- Leveau, J.H.; Gerards, S. 2008. Discovery of a bacterial gene cluster for catabolism of the plant hormone indole 3-acetic acid. FEMS Microbiology Ecology, 65: 238-250.
- Montañez, A.; Blanco, A.R.; Barlocco, C.; Beracochea, M.; Sicardi, M. 2012. Characterization of cultivable putative endophytic plant growth promoting bacteria associated with maize cultivars (Zea mays L.) and their inoculation effects in vitro. Applied Soil Ecology, 58: 21-28.
- Naik, P.R.; Raman, G.; Narayanan, K.B.; Sakthivel, N. 2008. Assessment of genetic and functional diversity of phosphate solubilizing fluorescent pseudomonads isolated from rhizospheric soil. BMC Microbiology, 8: 230.
- Okon, Y.; Labandera-Gonzales, C.A. 1994. Agronomic applications of Azospirillum: an evaluation of 20 years worldwide field inoculation. Soil Biology and Biochemistry, 26: 1591-1601.
- Peix, A.; Mateos, P.F.; Rodriguez-Barrueco, C.; Martinez-Molina, E.; Velazquez, E. 2001. Growth promotion of common bean (Pasheolus vulgaris L.) by a strain of Burkholderia cepacia under growth chamber conditions. Soil Biology and Biochemistry, 33: 1927-1935.
- Perin, L.; Martínez-Aguilar, L.; Paredes-Valdez, G.; Baldani, J.I.; Estrada-de-los-Santos, P.; Reis, V.M.; Caballero-Mellado, J. 2006. Burkholderia silvatlantica sp. nov., a diazotrophic bacterium associated with sugar cane and maize. International Journal of Systematic and Evolutionary Microbiology, 56: 1931-1937.
- Poly, F.; Monrozier, L.J.; Bally, R. 2001. Improvement in the RFLP procedure for studying the diversity of nifH genes in communities of nitrogen fixers in soil. Research in Microbiology, 152: 95-103.
- Radwan, T.; Mohamed, Z.K; Reis, V.M. 2002. Production of indole-3-acetic acid by different strains of Azospirillum and Herbaspirillum spp. Symbiosis, 32: 39-53.
- Rodrigues Neto, J.; Malavolta Júnior, V.A.; Victor, O. 1986. Meio simples para o isolamento e cultivo de Xanthomonas campestris pv. citri tipo B. Summa Phytopathologica, 12: 16.
- Roesch, L.F.W.; Camargo, F.A.O.; Bento, F.M.; Triplett, E.W. 2008. Biodiversity of diazotrophic bacteria within the soil, root and stem of field-grown maize. Plant and Soil, 302: 91-104.
- Sarwar, M.; Kremer, R.J. 1995. Determination of bacterially derived auxins using a microplate method. Letters in Applied Microbiology, 20: 282-285.
- Spaepen, S.; Vanderleyden, J.; Okon, Y. 2009. Plant growth-promoting actions of rhizobacteria. Advances in Botanical Research, 51: 283-320.
- Sylvester-Bradley, R.; Askawa, N.; Latorraca, S.; Magalhães, F.M.M.; Oliveira, L.A.; Pereira, R.M. 1982. Levantamento quantitativo de microrganismos solubilizadores de fosfatos na rizosfera de gramíneas e leguminosas forrageiras na Amazônia. Acta Amazônica, 12: 12-22.
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. 2007. MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Molecular Biololgy and Evolution, 24: 1596-1599.
- Tayeb, L.A.; Ageron, E.; Grimont, F.; Grimont, P.A.D. 2005. Molecular phylogeny of the genus Pseudomonas based on rpoB sequences and application for the identification of isolates. Research in Microbiology, 156: 763-773.
- Valle, I.C.A.; Alves, J.M.A.; Silva, L.S.; Uchôa, S.C.P.; Albuquerque, J.A.A.; Silva, D.C.O. 2013. Produção do milho híbrido 30F35HR cultivado na savana de Roraima em diferentes densidades de plantio. Agro@mbiente online, 7: 294-303.
- Young, J.P.W.; Downer, H.L.; Eardly, B.D. 1991. Phylogeny of the phototrophic Rhizobium strain BTAil by polymerase chain reaction-based sequencing of a 16S rRNA gene segment. Journal of Bacteriology, 173: 2271-2277.
- Zilli, J.E.; Marson, L.C.; Reis, V.M.; Alves, G.C.; Baldani, L.D.; Cordeiro, A.C. 2007. Contribuição da bactéria diazotrófica Herbaspirillum seropedicae para o rendimento de grãos de arroz e milho em Roraima. Boa Vista: Embrapa Roraima, 20p. (Embrapa Roraima. Boletim de Pesquisa e Desenvolvimento, 6).
Publication Dates
-
Publication in this collection
Apr-Jun 2016
History
-
Received
23 July 2015 -
Accepted
20 Oct 2015