Abstract
The role of intracellular free polyamine (putrescine and spermidine) pools in multiple resistance to aminoglycoside antibiotics was investigated among in vitro selected kanamycin-resistant Escherichia coli J53 mutants expressing diminished oligopeptide-binding protein (OppA) levels and/or defective ornithine decarboxylase (ODC) activity. The results suggest that diminished OppA content, but not defective ODC activity expression, increased the relative concentration of free spermidine as compared to the wild type strain. Moreover, by adding exogenous polyamines or polyamine synthesis inhibitors to cultures with different mutant strains, a direct relationship between the intracellular OppA levels and resistance to kanamycin was revealed. Collectively these results further suggest a complex relation among OppA expression, aminoglycoside resistance and polyamine metabolism.
aminoglycoside resistance; polyamines; oligopeptide binding protein A; Escherichia coli; oligopeptide permease
DRUG RESISTANCE
Intracellular polyamine pools, oligopeptide-binding protein A expression, and resistance to aminoglycosides in Escherichia coli
Maria BR AcostaI; Rita C Café FerreiraI; Luís CS FerreiraI; Sérgio Olavo P CostaII,1 1 Corresponding author. E-mail: sopdcost@usp.br
IDepartamento de Microbiologia, Instituto de Ciências Biomédicas, Universidade de São Paulo, Av. Prof. Lineu Prestes 1374, Cidade Universitária, 05508-000 São Paulo, SP, Brasil
IIPós-Graduação em Saúde Coletiva, Universidade Católica de Santos, Santos, SP, Brasil
ABSTRACT
The role of intracellular free polyamine (putrescine and spermidine) pools in multiple resistance to aminoglycoside antibiotics was investigated among in vitro selected kanamycin-resistant Escherichia coli J53 mutants expressing diminished oligopeptide-binding protein (OppA) levels and/or defective ornithine decarboxylase (ODC) activity. The results suggest that diminished OppA content, but not defective ODC activity expression, increased the relative concentration of free spermidine as compared to the wild type strain. Moreover, by adding exogenous polyamines or polyamine synthesis inhibitors to cultures with different mutant strains, a direct relationship between the intracellular OppA levels and resistance to kanamycin was revealed. Collectively these results further suggest a complex relation among OppA expression, aminoglycoside resistance and polyamine metabolism.
Key words: aminoglycoside resistance - polyamines - oligopeptide binding protein A - Escherichia coli - oligopeptide permease
Bacterial resistance to aminoglycosides involving multiple mechanisms varying from mutational modification of ribosomal targets, production of inactivating enzymes, and reduced drug uptake due to cell envelope impermeability, poses a threat to the clinical use of these antibiotics (Davies & Wright 1997). Although production of aminoglycoside modifying enzymes is usually regarded as the most relevant trait in clinically relevant gram-negative bacterial species, resistance ascribed to reduced aminoglicoside uptake represents a significant cause of concern especially among bacterial species involved in nosocomial infections (Bryan et al. 1976, Price et al. 1981).
The uptake of aminoglycosides in gram-negative bacteria is a rather complex process and remains ambiguous in several aspects in regards of transport of the antibiotics through the cell envelope. An initial electrostatic binding to negatively charged outer membrane components is followed by an energy-dependent import phase that is modulated according to the external antibiotic concentration. This process seems to involve various pathways requiring energy from both the membrane electrical potential and ATP-hydrolysis (Taber et al. 1987).
Aminoglycoside resistance due to reduced cell envelope permeability has een attributed to a reduced proton electrochemical gradient across the membrane, a consequence of a defective respiratory chain component (Damper & Esptein 1981, Muir et al. 1985). Under laboratory conditions, such aminoglycoside resistant Escherichia coli K12 mutants express reduced growth rates and restricted ability to ferment alternative substrates (Muir et al. 1985). Therefore, bacterial strains expressing such resistance mechanisms would have impaired ability to compete with antibiotic sensitive strains, thus limiting the threat of dissemination in both nosocomial and community environments.
The observation that some in vitro selected amino-glycoside resistant E. coli mutants expressed reduced oligopeptide-binding protein A (OppA) levels suggests that the oligopeptide (Opp) transport system could also play a role in the active transport of aminoglycoside (Kashiwagi et al. 1992, 1998). Moreover, the observation that intracellular polyamine pools regulate OppA expression indicated that polyamines could indirectly affect the cell envelope permeability to aminoglycoside antibiotics (Kashiwagi et al. 1998). Indeed the high incidence of reduced OppA expression and defective production of enzymes involved in polyamine synthesis, detected both among in vitro selected E. coli K12 mutants as well as among pathogenic strains isolated from clinical settings, further emphasizes the relationship between aminogly-coside resistance, OppA and polyamine metabolism (Rodriguez & Costa 1999, Acosta et al. 2000).
OppA is the soluble periplasmic protein of the Opp transport system engaged in the active uptake of peptides up to five residues found both among gram-negative and gram-positive bacteria (Hiles & Higgins 1986, Monnet 2003). Mutations affecting OppA expression are pleotropic and, depending upon the bacterial species, may affect adhesion to host cell proteins and tissues, development of the competence state, cell-cell communication, and sporulation (Monnet 2003).
As previously reported by our group, plating of E. coli J53 strains on media containing kanamycin resulted in the high frequency selection of mutants expressing multiple resistance to aminoglycosides (Acosta et al. 2000, Monteiro et al. 2003). Some mutants had colonies smaller than those produced by the parental strain and did not grow in succinate minimal plates, indicating the expression of impaired respiratory chain component, whereas others with no detectable growth impairment were shown to express reduced OppA levels and/or defective activity of ornithine/arginine decarboxylases (Acosta et al. 2000, Monteiro et al. 2003).
In the present work we further investigate the possible links between aminoglycoside resistance, expression of OppA and polyamine metabolism using a set of E. coli K12 mutants selected under in vitro conditions for multiple resistance to aminoglycosides. Detection of free putrescine and spermidine pools in resistant mutants expressing altered OppA or ODC levels, as well as the effects of exogenous polyamines and inhibitors of polyamine synthesis on the sensitivity to aminoglycosides and OppA expression, confirm the complex regulation of the bacterial resistance to these antibiotics.
MATERIALS AND METHODS
Bacterial strain and culture conditions - The parental E. coli J53 (met pro thi) (Clowes & Rowley 1954) was grown in modified Luria broth (L2) (2% tryptone, 2% yeast extract, 1% NaCl, 1.5% agar) or MM medium (Davis & Mingioli 1950) containing 10 µg/ml of each methionine and proline and 1 µg/ml of thiamine. All cultures were obtained under aerated conditions in an orbital shaker (120 rpm) at 37 °C. Aminoglycoside-resistant mutants were selected in L2 medium containing kanamycin at a final concentration of 20 µg/ml, as previously reported (Acosta et al. 2000). The m1 mutant was previously isolated and described by Kashiwagi et al. (1998). The R26, R60, and R69 mutants were selected on L2 plates. All selected resistant mutants displayed growth rates similar to the parental strain and were able to grow on succinate as the sole carbon source, an indication of functional aerobic metabolism (Monteiro et al. 2003). The polyamine synthesis inhibitors cyclohexilamine (CHA) and methylglyoxal bis(guanyl-hydrazone) (MGBG) were added to the MM medium before inoculation with the different strains at a final concentration of 100 mg/ml. Cell growth in liquid medium was monitored by measuring the absorbance at 600 nm in a Hitachi (model U-2000) spectrophotometer. All reagents and antibiotics used in this study were purchased from Sigma (Sigma-Aldrich Co., St Louis).
Determination of the free polyamine pools - Cultures were prepared with overnight grown inocula diluted 1:100 dilution in MM medium at 37 °C and bacteria harvested at an optical density of 0.6 at A600nm. Free polyamines were extracted after suspension of the bacterial cells in 5% perchloric acid followed by neutralization with 0.5 N NaOH, as previously described (Smith & Davies 1985). The polyamine dansylation was carried out with dansyl chloride in saturated sodium carbonate for 18 h at room temperature in the dark (Smith & Davies 1985). Dansylated polyamines were kept at-20 °C until analysis in a Shimadzu high-performance chromatography system (Shim-Pack CLC ODS model CMB-10 A), according to previously described procedures (Smith & Davies 1985). Dansyl polyamine fluorescence was measured with an excitation wavelength of 365 nm and an emission wavelength of 510 nm. Quantification of free putrescine and polyamine levels were based on integration of eluant peak areas at respective retention times (18 min for putrescine and 23.5 min for spermidine). Quantification of cellular free polyamine levels were determined with purified putrescine and spermidine (2.5 pmol to 50 pmol) subject to the same purification conditions. Total protein contents were determined according to the method of Lowry et al. (1951) using bovine serum albumin (5 mg/ml stock solution) as standard.
Detection of ODC activity - ODC activity was evaluated after growth of isolated colonies in Möeller decarboxylase broth supplemented with 1% ornithine and layered with mineral oil, as previously described (Davis et al. 1992). After overnight incubation at 37 °C, a brown/purple color was indicative of positive enzymatic activity, whereas the yellow color, the unchanged medium color, indicated lack of enzyme activity.
MIC determination - MICs of kanamycin, neomycin, streptomycin, gentamicin, and tobramycin were determined in L2 agar plates containing two-fold serial dilution of each tested aminogly-coside and seeded with approximately 104 CFU. Results were read after overnight incubation at 37 °C. Resistance was defined as a MIC value of at least four-fold higher than the value determined for the parental strain under the same assay conditions.
Isolation of periplasmic proteins - Periplasmic protein fractions were obtained after chlorophorm extraction, as previously described (Ames et al. 1984). Overnight culture aliquots were submitted to gentle chlorophorm treatment followed by a concentration step with acetone.
SDS-PAGE and immunoblotting - Proteins were sorted in 9% polyacrylamide gels (C = 5%) set at 100 V for 2 h under denaturating conditions. Electroblotting to nitrocellulose membranes was performed in a model EPS 500/400 Semi-dry Electro-blotter Unit (Pharmacia, Upsalla) at an electrical current of 0.8 mA/cm2 of gel. The membranes were blocked with phosphate-buffered saline (PBS) containing 1% bovine serum albumin and 0.05% Tween 20, according to standard procedures (Sambrook et al. 1989). Monospecific polyclonal rabbit anti-OppA serum (kindly supplied by Dr K Igarashi, Chiba University, Japan) was used at a final dilution of 1:2,000 in PBS-0.5% Tween-20 and incubated for 1 h at room temperature. After washing, the membranes were incubated with goat anti-rabbit IgG horseradish peroxidase conjugate (Sigma) for 1 h. Reactive protein bands were visualized with 3,3-diami-nobenzidine and hydrogen peroxide. Periplasmic extracts were also analyzed by SDS-PAGE followed by silver staining of gels, according to a previously described procedure (Morrisey 1981). All samples analyzed were also electrophoresed and stained with silver to confirm that equal loads were applied to each slot. Band intensities and quantification of OppA expression in immunoblots were determined densitometrically in the Eagle Eye gel documentation system (Stratagene, La Jolla).
RESULTS
Free putrescine and spermidine pools in aminogly-coside resistant E. coli mutants - The free polyamine pools of a set of four aminoglycoside resistant mutants were analyzed during exponential growth in minimal medium. The m1 and R60 mutants were defective in OppA expression and had no detectable ODC activity; R69 mutant expressed reduced OppA levels but normal ODC activity, while the R26 mutant had normal OppA levels but no ODC activity. Densitometric analysis of the residual OppA expression in periplasmic fractions of the resistant strains, carried out in immunoblots developed with OppA-specific serum, confirmed that the m1 mutant expressed approximately 20-30% of OppA content detected in the parental strain, as previously reported (Kashiwagi et al. 1998). The OppA expression measured in the other three aminoglycoside resistant mutants ranged from 5-10% in the R60 and R69 mutants to approximately 95-100% in the R26 mutant. All selected mutants expressed multiple resistance to aminoglycosides and MIC values at least eight times higher than those expressed by the parental strain. The J53 strains had MIC values of 8 µg/ml for kanamycin, neomycin and streptomycin and 2 mg/ml for gentamicin and tobramycin, whereas the resistant mutants could either grow at antibiotic concentrations above 128 µg/ml (kanamycin, neomycin, and streptomycin) or express a MIC value of 32 µg/ml (gentamicin and tobramycin), except for the R26 mutant which had MIC values of 128 µg/ml for streptomycin and 16 µg/ml for both gentamicin and tobramycin.
Determination of the total free putrescine and spermidine pools of the aminoglycoside resistant mutants, as well as the parental strain, revealed that the m1 mutant had approximately a three-fold reduction of total free polyamine content, mainly represented by a drastic reduction of free putrescine, with regard to the parental strain (Table). In contrast, the R60 mutant expressed an increase in the total free polyamine content as compared to the J53 strain, represented by an approximate ten-fold increase in spermidine levels and a two-fold increase in the soluble putrescine levels (Table). The remaining two mutants expressed contrasting results with regard to spermidine (higher in R69 and lower in R26) while the putrescine pools were reduced by approximately half (R69 mutant) or slightly reduced (approximately 80%) as compared with the free polyamine pools of the parental J53 strain (Table). The spermidine/putrescine (S/P) ratio of all resistant mutants defective in OppA expression displayed roughly a five-fold increase (S/P ratios ranging from 0.66 to 0.69) as compared to the parental strain (S/P ratio of 0.12). The enhanced S/P ratio observed in the OppA deficient strains were mainly attributed to higher relative spermidine levels which were increased from approximately 10% in the parental strain to 40% of the total free polyamine pool (Table). In contrast, the R26 mutant expressing normal OppA levels but defective in ODC activity had a S/P ratio of 0.1.
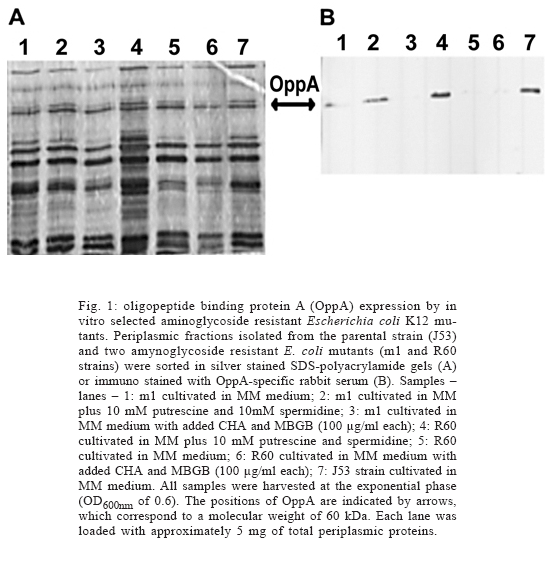
Effect of exogenous polyamines and polyamine synthesis inhibitors on OppA expression and aminogly-coside resistance - Addition of polyamines to the growth medium enhanced OppA levels detected in periplasmic extract of m1 and R60 mutants, while the opposite effect was observed in cultures prepared with polyamine synthesis inhibitors (Fig. 1). Moreover, addition of free polyamines to the growth medium enhanced sensitivity to kanamycin in both mutants, while polyamine synthesis inhibitors enhanced resistance (Fig. 2). The MIC values for kanamycin, as well as for other aminoglycosides, expressed by the two antibiotic resistant mutants were reduced or increased by at least two-fold in MM plates containing exogenous polyamines or inhibitors of poly-amine synthesis, respectively.
DISCUSSION
Polyamines, such as putrescine and spermidine, are important nitrogen-containing polycationic compounds required by all living cells for both protein and nucleic acids synthesis (Tabor & Tabor 1985). Under certain physiological conditions polyamines contribute to the structural stabilization of acidic polymeric molecules such as RNA and DNA, but a significant fraction of the cellular pool is found in free form and can interact with untrans-lated specific mRNA regions post-transcriptionally controlling the expression of several genes, a controlling mechanism recently named as the polyamine modulon (Igarashi & Kashiwagi 2000, Yoshida et al. 2004). In the present work we investigated a possible link between polyamines and multiple resistance to aminoglycosides by measuring the free polyamine pools of E. coli K12 mutants selected for multiple resistance to aminolyco-sides. The total free polyamine pools of four resistant mutants revealed that relationship between free polyamines and aminoglycoside resistance is complex and ambigous. One mutant (m1) expressed reduced polyamine levels, another (R60) expressed enhanced levels, while the remaining two mutants (R69 and R26) expressed levels similar to the J53 strain, although all expressed similar resistance levels to multiple aminoglycoside antibiotics.
The finding that the spermidine/putrescine ratio of three resistant mutants with reduced OppA levels was higher than that detected in the parental strain was intriguing. In these mutants the relative concentration of spermidine was at least five-fold higher than the levels detected in the parental strain suggesting that polyamine-mediate OppA control may rely on the balance of the two polyamine forms rather than the absolute cellular concentration of one polyamine species. The observations concerning ODC activity and aminoglycoside resistance were unclear. The three defective ODC mutants expressed quite distinct free polyamine levels, and no direct correlation could be drawn based on ODC activity, OppA expression and aminoglycoside resistance. ODC knockout mutants were reported to express reduced total polyamine levels (Igarashi et al. 1986). The unknown nature of the genetic defects causing the defective ODC expression in the tested mutants prevents the establishment of any conclusion regarding a possible link between ODC activity and aminoglycoside resistance. Future experiments based on polyamine synthesis knockout mutants should address the role of ODC, as well as other enzymes involved in polyamine metabolism, and aminoglycoside resistance.
Polyamine auxotrophic E. coli mutants have previously been shown to be resistant to streptomycin when deprived of putrescine (Goldemberg & Algranati 1981). The resistance phenotype was proposed to reflect an alteration of aminoglycoside ribosomal targets. In such mutants, the resistance phenotype associated with polyamine deprivation was restricted to a few antibiotics, such as streptomycin, as opposed to the resistant mutants affected on aminoglycoside uptake, which are resistant to most, if not all, tested aminoglycosides. The data presented in this study further indicate that inbalances in polyamine metabolism may indirectly affect aminoglyco-side resistance via control of OppA expression (Igarashi et al. 1997, Kashiwagi et al. 1998, Acosta et al. 2000). Results based on the m1 and R60 demonstrated that addition of polyamine synthesis inhibitors reduced OppA expression and increased resistance to aminoglycosides. On the other hand, incorporation of exogenous polyamines to growth media resulted in opposite effects enhancing both OppA expression and sensitivity to aminoglycosides.
A putative common uptake system for polyamines and aminoglycosides was initially proposed as an explanation of the streptomycin suppressive effect on the uptake of spermidine and putrescine (Höltje 1978). The cloning of polyamine transport encoding genes did not confirm the existence of a common uptake system capable of handling aminoglycosides and polyamines (Kashiwagi et al. 1990), but later reports clearly demonstrated that polyamines have a role in the uptake of peptides via activation of OppA expression (Igarashi et al. 1997, Kashiwagi et al. 1998). Our data represent additional evidences that polyamines can indeed affect OppA expression and, thus, indirectly regulate sensitivity to aminoglycosides. Moreover, the enhanced frequency of aminoglycoside resistant strains expressing low OppA levels among E. coli strains isolated from hospital infections indicate that findings similar to those reported for the E. coli J53 may also be valid to clinically relevant strains (Acosta et al. 2000)
Collectively, these evidences support the hypothesis that polyamines play a role in the multiple resistance to aminoglycosides in E. coli K12 via modulation of OppA expression, thus affecting the active transport of these antibiotics through the oligopeptide uptake system. Future experiments based on precise knockout mutants affected in OppA expression and genes encoding polyamine biosynthetic pathways as an attempt to further explain the link between polyamines, OppA expression and multiple resistance to aminoglycosides are clearly relevant.
Received 2 June 2005
Accepted 18 October 2005
Financial support: Fapesp, CNPq
- Acosta MBR, Ferreira RCC, Padilla G, Ferreira LCS, Costa SOP 2000. Altered expression of oligopeptide-binding protein (OppA) and aminoglycoside resistance in laboratory and clinical Escherichia coli strains. J Med Microbiol 49: 409-413.
- Ames GFL, Prody C, Kustu S 1984. Simple, rapid and quantitative release of periplasmic proteins by chlorophorm. J Bacteriol 160: 1181-1183.
- Bryan LE, Haraphogse R, Van den Elzen HM 1976. Gentamicin resistanse in clinical isolates of Pseudomonas aeruginosa associated with diminished gentamicin accumulatio and no detectable enzymatic modifiation. J Antibiot 29: 743-753.
- Clowes RC, Hayes W 1968. Experiments in Microbial Genetics, Blackwell Scientific Publications, Oxford.
- Damper PD, Epstein W 1981. Role of the membrane potential in bacterial resistance to aminoglycoside antibiotics. Antimicrob Agents Chemother 20: 803-806.
- Davies J Wright GD 1997. Bacterial resistance to aminoglycoside antibiotics. Trends Microbiol 5: 234-240.
- Davis BD, Mingioli ES 1950. Mutants of Escherichia coli requiring methionine or vitamin B12. J Bacteriol 60: 17-28.
- Davis BD, Morris DR, Coffino P 1992. Sequestered end products and enzyme regulation: the case of ornithine decarboxylase. Microbiol Rev 56: 280-290.
- Höltje JV 1978. Streptomycin uptake via an inducible polyamine transport system in Escherichia coli Eur J Biochem 86: 345-351.
- Goldemberg SH, Algranati ID 1981. Polyamine requirement for streptomycin action on protein synthesis in bacteria. Eur J Biochem 117: 251-255.
- Hiles ID, Higgins CF 1986. Peptide uptake by Salmonella typhimurium The periplasmatic oligopeptide-binding protein. Eur J Biochem 158: 561-567.
- Igarashi K, Kashiwagi K 2000. Polyamines: mysterious modulators of cellular functions. Biochem Biophys Res Comm 271: 559-564.
- Igarashi K, Kashiwagi K, Hamasaki H, Miura A, Kakegawa T, Hirose S, Matsuzaki S 1986. Formation of compensatory polyamine by Escherichia coli polyamine-requiring mutants during growth in the absence of polyamines. J Bacteriol 166: 128-134.
- Igarashi K, Saisho T, Yuguchi M, Kashiwagi K 1997. Molecular mechanism of polyamine stimulation of the synthesis of oligopeptide-binding protein. J Biol Chem 272: 4058-4064.
- Kashiwagi K, Hosokawa N, Furuchi T, Kobayashi H, Sasakawa C, Yoshikawa M, Igarashi K 1990. Isolation of polyamine transport-deficient mutants of Eschericha coli and cloning of the genes for polyamine transport proteins. J Biol Chem 265: 20893-30897.
- Kashiwagi K, Miyaji A, Ideka S, Tobe T, Sasakawa C, Igarashi K 1992. Increase of sensitivity to aminoglycoside antibiotics by polyamine-induced protein (oligopeptide-binding protein) in Escherichia coli J Bacteriol 174: 4331-4337.
- Kashiwagi K, Tsuhako MH, Sakata K, Saisho T, Igarashi A, Costa SOP, Igarashi K 1998. Relationship between spontaneous aminoglycoside resistance in Escherichia coli and a decrease in oligopeptide binding protein. J Bacteriol 180: 5484-5488.
- Lowry OH, Rosenbrough NJ, Farr AL, Randall RJ 1951. Protein measurement with the Folin phenol reagent. J Biol Chem 193: 265-275.
- Monnet V 2003. Bacterial oligopeptide-binding proteins. Cell Mol Life Sci 60: 2100-2114.
- Monteiro ACM, Ferreira RCC, Padilla G, Costa, SOP, Ferreira LCS 2003. Environmental and genetic factors affecting mutability to aminoglycoside antibiotics among Escherichia coli K12 strains. Genet Mol Biol 26: 221-227.
- Morrissey JM 1981. Silver stain for protein in polyacrylamide gels: a modified procedure with enhanced uniform sensitivity. Anal Biochem 117: 307-310.
- Muir ME, Ballesteros M, Wallace BJ 1985. Respiration rate, growth rate and the accumulation of streptomycin in Escherichia coli J Gen Microbiol 131: 2573-2579.
- Price KE, Kresel PA, Farchione LA, Siskin SB, Karpow SA 1981. Epidemiological studies of aminoglycoside resistance in the U.S.A. J Antimicrob Chemother 8: 89-105.
- Rodriguez MB, Costa SOP 1999. Spontaneous Escherichia coli kanamycin-resistant mutant shows altered periplasmic oligopeptide permease protein, OppA, and impermeability to aminoglycosides. Rev Microbiol 30: 153-156.
- Sambrook J, Fritsch EF, Maniatis T 1989. Molecular Cloning: A Laboratory Manual, 2nd ed., Cold Spring Harbor Laboratory, New York.
- Smith MA, Davis PJ 1985. Separation and quantification of polyamines in plant tissue by high performance liquid chromatography of their dansyl derivates. Plant Physiol 78: 89-91.
- Taber HW, Mueller JP, Miller PF, Arrow AS 1987. Bacterial uptake of aminoglycoside antibiotics. Microbiol Rev 51: 439-457.
- Tabor CW, Tabor H 1985. Polyamines in microorganisms. Microbiol Rev 49: 81-99.
- Yoshida M, Kashiwagi K, Shigemasa A, Taniguchi S, Yamamoto K, Makinoshima H, Ishihama A, Igarashi K 2004. A unifying model for the role of polyamines in bacterial cell growth, the polyamine modulon. J Biol Chem 279: 46008-46013.
Publication Dates
-
Publication in this collection
09 Jan 2006 -
Date of issue
Nov 2005
History
-
Received
02 June 2005 -
Accepted
18 Oct 2005