Abstract
Rotavirus replication and virus assembly take place in electrodense spherical structures known as viroplasms whose main components are the viral proteins NSP2 and NSP5. The viroplasms are produced since early times after infection and seem to grow by stepwise addition of viral proteins and by fusion, however, the mechanism of viropIasms formation is unknown. In this study we found that the viroplasms surface colocalized with microtubules, and seem to be caged by a microtubule network. Moreover inhibition of microtubule assembly with nocodazole interfered with viroplasms growth in rotavirus infected cells. We searched for a physical link between viroplasms and microtubules by co-immunoprecipitation assays, and we found that the proteins NSP2 and NSP5 were co-immunoprecipitated with anti-tubulin in rotavirus infected cells and also when they were transiently co-expressed or individually expressed. These results indicate that a functional microtubule network is needed for viroplasm growth presumably due to the association of viroplasms with microtubules via NSP2 and NSP5.
rotavirus; viroplasms; microtubules; NSP2; NSP5
ARTICLES
Association of rotavirus viroplasms with microtubules through NSP2 and NSP5
Claudio Cabral-Romero; Luis Padilla-Noriega1 1 Corresponding author: lpadilla@servidor.unam.mx
Departamento de Biología Molecular, Instituto de Investigaciones Biomédicas, Universidad Nacional Autónoma de México, Apartado Postal 70-228, México, DF 04510, México
ABSTRACT
Rotavirus replication and virus assembly take place in electrodense spherical structures known as viroplasms whose main components are the viral proteins NSP2 and NSP5. The viroplasms are produced since early times after infection and seem to grow by stepwise addition of viral proteins and by fusion, however, the mechanism of viropIasms formation is unknown. In this study we found that the viroplasms surface colocalized with microtubules, and seem to be caged by a microtubule network. Moreover inhibition of microtubule assembly with nocodazole interfered with viroplasms growth in rotavirus infected cells. We searched for a physical link between viroplasms and microtubules by co-immunoprecipitation assays, and we found that the proteins NSP2 and NSP5 were co-immunoprecipitated with anti-tubulin in rotavirus infected cells and also when they were transiently co-expressed or individually expressed. These results indicate that a functional microtubule network is needed for viroplasm growth presumably due to the association of viroplasms with microtubules via NSP2 and NSP5.
Key words: rotavirus - viroplasms - microtubules - NSP2 - NSP5
Group A rotaviruses are the leading etiological agents of severe diarrheal disease in infants and young children worldwide (Parashar 2003). These viruses belong to the genus Rotavirus within the Reoviridae family and their genome consist of 11 double-stranded RNA segments that encode six structural proteins named VP1-VP4, VP6, and VP7, as well as six nonstructural proteins named NSP1 to NSP6 (Estes 2001).
Rotavirus replication takes place in the cytoplasm of infected cells in electrodense spherical structures known as "viroplasms" that can be found as early as 4 h after infection (Petrie et al. 1984). Viroplasms are composed of viral RNA and the proteins VP1, VP2, VP3, VP6, NSP2, and NSP5 (Petrie et al. 1982, 1984, Gallegos & Patton 1989), and while virion assembly occur in these structures (Petrie et al. 1982, Gallegos & Patton 1989) the mechanism of viroplasms formation is unknown. Recently, it was reported that the number of rotavirus viroplasms decrease with post-infection time (Eichwald et al. 2004), and while the diameter of single viroplasms increased with time, the total number of viroplasms per cell diminishes, suggesting that growth of these inclusion bodies occur by fusion and probably also by stepwise addition of viral components to the viroplasms surface (Eichwald et al. 2004).
In the viroplasms the viral mRNAs are replicated to produce genomic double stranded RNAs (dsRNA) which are simultaneously encapsidated into double layerded viral particles (DLPs), that contain the 11 dsRNA at the core of the particle surrounded by the inner and intermediate protein layers of VP2 and VP6, respectively. The third layer formed by VP4 and VP7 is acquired by budding of the DLP particles into the endoplasmic reticulum (ER), a process that requires assistance of the viral non-structural protein 4 (NSP4), whereby a transient envelope is acquired, that is finally lost within the ER along with NSP4. The fundamental role of NSP4 in this step is evident by the reduction in size of viroplasms that is observed when the NSP4 gene is silenced though RNA interference (Lopez et al. 2005).
Rotavirus infection changes the intracellular distribution of microtubule associated protein 2 (MAP2) and subviral particles normally located in viroplasms have been co-purified with MAP2 (Weclewicz et al. 1993), suggesting a possible relationship of these particles, or the viroplasms where they are assembled, with microtubules.
In addition, other members of the Reoviridae family replicate in association with the cytoskeleton. The prototype of the Reoviridae family is the genus Reovirus that requires an intact microtubule network for virus assembly (Dales 1963, Parker et al. 2002). Likewise, members of the genus Orbivirus within the Reoviridae form viral inclusion bodies associated to intermediate filaments (Eaton et al. 1987). However, the role of the cytoskeleton in rotavirus viroplasms formation is unknown.
NSP2 is a 34 KDa protein with an octameric array that binds to RNA in a non-specific manner, is the most abundant protein of viroplasms (Petrie et al. 1984) and is essential for viroplasms formation (Ramig & Petrie 1984). NSP5 is a fosfo-glycoprotein that presents different isoforms that differ in the phosphorylation level with molecular masses from 28 to 32-34 KDa (Afrikanova et al. 1996, Blackhall et al. 1997, Poncet et al. 1997). NSP2 and NSP5 are located in the viroplasms of rotavirus infected cells, interact with each other and also with other viral proteins, NSP2 with the viral RNA polymerase VP1, and NSP5 with VP2 and NSP6 (Aponte et al. 1996, Berois et al. 2003, Torres-Vega et al. 2000). The co-expression of NSP2 and NSP5 in cells, in the abscence of other viral proteins, results in viroplasm like-structures, VLS (Fabretti et al. 1999), however when expressed individually they produce a diffuse distribution, hence these proteins are considered the minimum components of the viroplasms.
In this report, we present evidence of the association of rotavirus viroplasms with microtubules, possibly through NSP2 and NSP5, as well as data that support the requirement of an intact microtubule network for viro-plasms growth. This is the first cellular element described to be involved in viroplasm formation.
MATERIALS AND METHODS
Cells and viruses - The African green monkey kidney epithelial cell lines MA104 and BSC1 were cultured in modified Eagles medium (MEM) suplemented with 10% fetal calf serum (Invitrogen, Carlsbad, CA). The simian rotavirus strain SA11 was propagated in MA104 cells as described by Estes et al. (1979).
Plasmid constructs - Rotavirus SA11 genes 8 and 11 that code for NSP2 and NSP5, respectively, were amplified from genomic dsRNA with the Thermoscript reverse transcription-polymerase chain reaction (RT-PCR) system (Invitrogen) and cloned into the NcoI-BamHI sites of the plasmid p.VOTE1 (Ward et al. 1995). The following specific primers were used for the amplification of the NSP2 gene (GeneBank accession no. J02353): 5'-GTTATCC ATGGCTGAGCTAGC-3' and 5'-GTTGGATCCTTAAA TTCCTATTTGAGAA-3'. The primers used for amplification of the NSP5 gene (GeneBank accession no. M28347) were 5'-GTATACCATGGCTCTCAGTATT GACGTG-3' and 5'-GTAGGATCCTTACAAATCTTCAATCAA-3'. The NSP2 gene from SA11 was also cloned into the NcoI-BamHI sites of the plasmid pET28b (Novagen, Madison, WI) to obtain the construct pbac-NSP2 for NSP2 expression in bacteria. The following primers were used for amplification of the NSP2 gene: 5' -GTTATCCATGGCTGAG CTAGC- 3' and 5' -GTCTACTCGAGAATTCCTATTT GAGA- 3'. In this construct a six histidines tail was added in the carboxy terminus to facilitate its purification by affinity chromatography. The inserts in all these plasmids were sequenced in both strands.
NSP2-his 6 purification - NSP2 expression in Escherichia coli BL21-DE3 transformed with pbacNSP2 was induced with 1 mM of IPTG for 5 h at 37°C. Bacteria were disrupted in lysis buffer (PBS plus protease inhibitor cocktail EDTA-free, Roche, Mannheim, Germany), and NSP2 was solubilized with 8 M urea and purified using Ni-nitrilo-acetic acid (Ni-NTA) columns (QIAGEN, Germany), as indicated by the provider.
Transient expression of rotavirus proteins using the vaccinia T7 RNA polymerase hybrid system - Rotavirus proteins NSP2 and NSP5 were expressed in a transient vaccinia virus expression system (Fuerst et al. 1986) by infection with the vaccinia virus strain vTF7-3 for expression of the T7 RNA polymerase, and co-transfection with plasmids pNSP2 and/or pNSP5 for transcription of the rotavirus genes from a T7 RNA polymerase promoter. 5 ´ 105 BSC1 cells per well were seeded in six well cell culture clusters, and incubated 12-24 h. The cells were then infected at an MOI of 10 with vTF7-3, and 1 h post-infection the cells were transfected with 2.5 ml of Lipofectamine-2000 reagent (Invitrogen) in a transfection mixture prepared according to provider, that contained 100 ng of the plasmid DNA. Eighteen h after infection the cells were washed with 1 ml of PBS per well, scraped, and aliquoted in two microcentrifuge tubes with 500 ml each one. The cells were centrifuged and the pellets were resuspended in 20 ml of lysis buffer (50 mM Tris-HCl pH 7.5 plus protease inhibitor for mammalian cells, Sigma Cat. P4083), and disrupted in a cup sonicator for 1 min on ice. The disrupted cells were mixed with 20 ml of Laemmli sample buffer 2 ´, boiled for 3 min and run in a 12% SDS-PAGE (Laemmli 1970) for analysis by Western blot.
Production of anti-NSP2 serum - The purified NSP2 was administrated intramuscularly to guinea pigs in four doses of 100 mg of recombinant protein plus Freund's adjuvant (Sigma-Aldrich, San Louis, MS), complete for the first dose and incomplete for the subsequent ones. Two weeks after the last dose the guinea pigs were bled, and the serum was separated.
Western blots - One half well of the cells from 6 well plates were resolved in a 12% SDS-PAGE (Laemmli 1970). The proteins were electrotransferred onto nitrocellulose membranes (Biorad, California) in a transfer buffer containing methanol (25 mM Tris, 192 mM glycine, 20% methanol and 0.1% SDS). Membranes were blocked with 5% skim milk in 5% tween PBS for 1 h at room temperature (RT) and incubated overnight at 4°C with the following antibodies: anti-NSP2 guinea-pig serum diluted 1:1000, or anti-b tubulin monoclonal antibody from ascites fluid diluted 1:1000 (anti-b tubulin D-10: sc-5274, Santa Cruz, CA), or anti-NSP5 rabbit serum diluted 1:1500 (kindly provided by C Arias, Institute of Biotechnology, National Autonomous University of Mexico). After three washes with 0.1% tween in PBS, peroxidase labeled-protein A (Kirkegaard and Pery Laboratories Inc., Maryland) was added as a secondary antibody and incubated for 30 min at RT. Membranes were washed with 0.1% tween in PBS and developed using a comercial substrate according to the instructions of the provider (SuperSignal West Femto Stable, Pierce, Rockford, IL).
Immunofluorescence microscopy - 4 ´ 104 cells per well were seeded in 16 well chambered slides (Nalge Nunc, Rochester, NY), incubated 12-24 h and infected with vaccinia virus strain vTF7-3 (Fuerst et al. 1986) at a MOI of 0.1. In some experiments nocodazole (15 mg/ml) was used to interfere with microtubule function at different times post-infection. 24 h after infection the cells were washed three times with cold PBS, fixed and permeabilized with cold 2% paraphormaldehyde (PFA), 0.1% triton X-100 for 30 min at 4°C. The PFA was eliminated, washing three times with cold PBS and incubated for 1 h at 4°C with an anti-NSP5 serum diluted 1:400 in cold PBS, anti-b tubulin monoclonal antibodies diluted 1:30 (anti-b tubulin D-10: sc-5274, Santa Cruz) or an anti-NSP2 guinea-pig serum diluted 1:100 in cold PBS. After three washes with cold PBS, the cells were incubated for 1 h at 4°C with FITC-conjugated goat anti-guinea-pig antibodies (Zymed, South San Francisco, CA), FITC-conjugated goat anti-mouse antibodies (Zymed) or Texas Red dye-conjugated goat anti-rabbit antibodies (Jackson ImmunoResearch, West Grove, PA). Cells were washed three times with cold PBS, nuclei stained with DAPI (Chemicon, Temecula, CA) and mounted with DAKO mounting medium (Dako-Corporation, Carpinteria, CA). Samples were analyzed by conventional immunofluorescence microscopy, using an Olympus IX70 microscope and confocal microscopy with a Zeiss LSM Pascal confocal microscope.
Co-immunoprecipitations - Cell lysates were prepared from one well (of 6 wells plates) in lysis EBC buffer (50 mM, pH 8 Tris- HCl, 120 mM NaCl, 0.5% NP-40 and protease inhibitor-SIGMA P-8340). Cellular debris and insoluble proteins were eliminated by centrifugation for 15 min at 14,000 g. Soluble proteins were incubated with anti-b tubulin monoclonal antibodies (anti-b tubulin D-10: sc-5274, Santa Cruz) plus protease inhibitor (Sigma-Aldrich) for 1 h at 4°C with agitation. Protein A-Sepharose (Sigma-Aldrich) was added and incubated for 30 min more at 4°C with agitation. After three washes with buffer NETN (20 mM pH 8 Tris, 1 mM EDTA, 100 mM NaCl and 0.5% NP40), samples were resolved in a 12% SDS-PAGE (Laemmli 1970). Western blot assays were used to detect NSP2 or NSP5 in the immunoprecipitated samples.
RESULTS
Rotavirus viroplasms are caged by microtubules - Aiming to determine if the microtubules have a role in viroplasms formation, we determined if they colocalize with viroplasms in BSC1 cells infected with rotavirus SA11. Eighteen h post-infection (PI) the cells were fixed and viroplasms and microtubules were visualized by immunofluorescence and confocal microscopy. We observed that the surface of viroplasms co-localized with microtubules when viroplasms were detected with specific anti-NSP5 or anti-NSP2 sera (Fig. 1). The internal part of viroplasms does not co-localize with microtubules, but they are rather "caged" by the microtubule network. This result suggests a possible association of rotavirus viroplasms with microtubules.
Determination of the size and number of rotavirus viroplasms throughout infection - We determined the number of viroplasms in BSC1 cells infected with rotavirus SA11 at different times after infection. At 4, 12 or 24 h PI the cells were fixed and exposed to the serum anti-NSP2 or anti-NSP5 and analyzed by immunofluorescence microscopy. As previously observed by others (Petrie et al. 1984) we observed that both, NSP2 and NSP5 were localized in viroplasms since 4 h PI (Fig. 2), a time when viroplasms looked like punctual structures that were on average 30 per cell with a diameter of 0.5 m. At 12 h PI the number of viroplasms decreased to an average of 20 per cell with a diameter of 1.5-2 m, and at 24 h PI the number of viroplasms further decreased to 15 per cell with a diameter of 3-4 m. These result confirmed that the number of viroplasms decreased throughout infection, while size continuously increases, as previously shown by Eichwald et al. (2004).
Viroplasms formation is stopped by nocodazole - We decided to use nocodazole, an inhibitor of tubulin assembly into microtubules, to determine if these structures are necessary for viroplasms formation. The drug was added (15 mg/ml final concentration) to rotavirus infected cells at different times after infection and at the end of the incubation period of 18 h after infection the viroplasms were visualized by immunofluorescence microscopy using anti-NSP5 serum. In the absence of nocodazole, typical morphology of rotavirus viroplasms was observed with a diameter of 3-4 m (Fig. 3), but if nocodazole was added since 4 h after infection, just small punctuate structures were observed with a diameter of 0.5 m, while identical results were obtained if drug was added since 1 h PI (data not shown). Viroplasms of intermediate size were visualized with a diameter of 1.5-2 m when drug was added at 8 h after infection, and they were full-size when drug was added at the end of the infection, 1 h before fixing cells (Fig. 3C). The number of viroplasms as well as their morphology was similar to those observed in Fig. 2 according to the number of hours of incubation in the absence of nocodazole, indicating that this drug effectively stops viroplasm growth impeding any further fusion among them, in a process that is accompanied by a reduction in number. As a control we confirmed that 5 h of treatment with nocodazole disrupt the microtubules (Fig. 3E, F). These results show that viroplasms need an intact microtubule network for their growth.
Viroplasms are associated to microtubules in rota-virus infected cells - Based on the close interaction and co-localization of the surface of viroplasms with the microtubule network, and since NSP2 and NSP5 are considered as the elemental constituents of viroplasms, we hypothesized that either or both of these proteins could be the physical link between the viroplasms and microtubules. We performed co-immunoprecipitation assays with anti-tubulin in lysates of rotavirus infected cells obtained 24 h PI, and determined the presence of the most abundant protein of viroplasms, NSP2, by Western blot with a serum anti-NSP2. We found that NSP2 was co-immunoprecipitated with anti-tubulin (Fig. 4, lane 1), suggesting that viroplasms are associated to microtubules at least late after infection.
VLS are also associated to microtubules - Since viro-plasms are composed of many viral proteins and RNAs, and any of these could be responsible of binding to microtubules, we decided to produce VLS by co-expression in a vaccinia system of NSP2 and NSP5, the elementary components of viroplasms, aiming to determine if either or both are the physical link between viroplasms and microtubules. We confirmed that NSP2 and NSP5 are co-expressed in cells transfected with the plasmids pNSP2 and pNSP5 in the absence of any other viral protein (Fig. 5). When NSP2 and NSP5 are co-expressed viroplasm-like structures are formed (Fig. 6) as previously shown by others (Fabretti et al. 1999). We found that both NSP2 and NSP5 were co-immunoprecipitated with anti-tubulin in a lysate of cells co-expressing both viral proteins, as can be seen in Fig. 7 (lane 2), but they were not immuno-repcipitated with anti-actin, as a control (Fig. 7, lane 1). These results suggest that VLS associate to microtubules through either or both NSP2 and NSP5 while other viral proteins or RNAs that constitute the viroplasms do not seem to be essential for binding of viroplasms to microtubules.
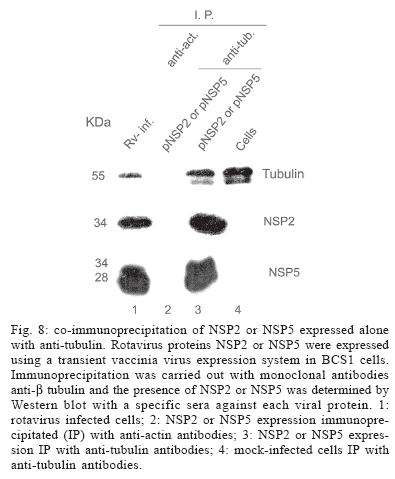
NSP2 and NSP5 expressed alone bind independently to microtubules - In order to analyze if NSP2 or NSP5 can independently bind to microtubules in the absence of VLS, we expressed NSP2 and NSP5 individually in BSC1 cells. We performed immunoprecipitation with anti-tubulin and determined the presence of the viral proteins in immunoprecipitates by Western blot. As shown in Fig. 8 NSP2 was present in the tubulin immunoprecipitates of cells expressing NSP2, and not in lysates immunoprecipitated with anti-actin as a control (Fig. 8, lane 2). Similarly, NSP5 was co-immunoprecipitated with anti-tubulin in cells expressing NSP5 (Fig. 8), and was not detected in the lysates immunorecipitated with anti-actin. These results suggest that both NSP2 and NSP5 can bind to microtubules independently of each other in conditions where VLS are not formed.
NSP2 and NSP5 differ in relative efficiency to bind to microtubules throughout infection - To study the dynamics of the association of NSP2 and NSP5 with microtubules during rotavirus infection, we obtained lysates of cells infected with rotavirus at 4, 12, and 24 h PI, and determined the interaction of tubulin with NSP2 and NSP5 by co-immunoprecipitation assays. Interestingly, NSP2 could be detected immunoprecipitating with anti-tubulin at the earliest time assayed 4 h, while NSP5 could not be detected at 4 h but was found associated to microtubules at 12 h PI (Fig. 9A), when the level of NSP2 bound to microtubules was maximal. 24 h PI the quantity of NSP2 and NSP5 bound to microtubules decreased (Fig. 9A, lane 5), being more evident for NSP5. However, when we repeated the assay with twice as much cells as in Fig. 9A, we could detect a weak band of NSP5 co-immunoprecipitating with anti-tubulin (Fig. 9B). From these results it is clear that the ratio of NSP5 to NSP2 associated to microtubules is different at different times PI, whereby NSP5 seems to be less effectively associated to microtubules at early than late times PI. These results suggest that NSP2 binds and remains associated to microtubules, while NSP5 has a variable efficiency of association to microtubules throughout infection.
DISCUSSION
The prototype of the Reoviridae family, reovirus needs an intact microtubule network to form their inclusion bodies (Dales 1963, Parker et al. 2002), and other members like bluetongue viruses form their inclusion bodies associated to intermediate filaments (Eaton et al. 1987). On the other hand, the viroplasms of rotavirus are formed by numerous viral proteins and RNAs, however NSP2 and NSP5 seem to be their elementary components, since these proteins are sufficient to form VLS when co-expressed in the absence of other viral components (Fabretti et al. 1999). In this study we found that NSP5, that together with NSP2 is essential for viroplasms formation, co-localize in the periphery of the viroplasms with microtubules, as determined by immunofluosrecence microscopy, hence the viroplasms seem to be caged by microtubule networks. This finding suggests that microtubules interact with the surface of viroplasms, hence we decided to study if microtubules are necessary for viroplasms formation.
By using nocodazole to inhibit the assembly of microtubules we found that the size of viroplasms was inversely proportional to the length of nocodazole exposure, while its number was directly proportional to nocodazole exposure, i. e. viroplasms cease to grow under the effect of nocodazole, in a process that is normally associated with a reduction in the number of viroplasms. Accordingly we observed numerous small punctuate viroplasms when nocodazole was added since early times after infection, suggesting that nocodazole does not interfere with the earliest events of viroplasms formation or "sowing", but only with their growth. The growth of viroplasms has been proposed to depend on addition on their components to the surface of viroplasms as well as on their fusion (Eichwald et al. 2004). The finding that viroplasms cease to grow completely in the presence of nocodazole strongly suggest that this drug prevents the fusion of viroplasmas as well as the associated reduction in their number, which clearly do not occur in the presence of this drug, while it is less clear if the proposed superficial growth of viroplasms is also affected by nocodazole. Since nocodazole binds to free tubulin and not to the polymer, affecting the assembly of tubulin subunits, we hypothesize that viroplasm formation depends of the constant addition of free tubulin to the polymer.
Since our data demonstrated the dependence of viroplasms growth on the dynamics of tubulin polymerization, we aimed to determine if viroplasms are bound to microtubules in rotavirus infected cells, and found that indeed NSP2, the main component of viroplasms, is co-immunoprecipitated with anti-tubulin, suggesting that in rotavirus infected cells viroplasms are bound to microtubules. From this result it is not clear what components of the viroplasms are responsible of their interaction with microtubules.
Since the elementary constituents of viroplasms are NSP2 and NSP5 as evidence by their ability to form VLS, we co-expressed these proteins and determined their ability to bind to microtubules as VLS complexes as well as when they were individually expressed. We found that as expected both proteins were co-immunoprecipitated with anti-tubulin when they were co-expressed, but surprisingly they also were co-immunoprecipitated with anti-tubulin when they were expressed individually. These data suggest that either or both of these proteins are responsible of the association of viroplasms to microtubules. We can not discard however, that other viral proteins participate in binding of rotavirus viroplasms to microtubules.
Other rotavirus proteins are bound to microtubules throughout infection, like NSP4 and VP4 (Xu et al. 2000, Nejmeddine et al. 2000); but they are not located in viro-plasms, since VP4 apparently assembles on preformed double layered viral particles at the final steps of morphogenesis, while NSP4 seem to be incorporated in DLPs that bud to the ER and interestingly it also affects viroplasms growth as demonstrated by the failure of viroplasms to grow upon transfection with NSP4-specific interfering RNA (RNAi) (Lopez et al. 2005). Since NSP4 is involved in budding of DLPs to the ER, and the interference with its function affects viroplasms growth, it can be inferred that growth of viroplasms need the clearance of DLPs formed at their surface by budding to the ER. We presume that any further viroplasms growth requires addition of the "building blocks" NSP2 and NSP5 to the surface of preformed viroplasms or viroplasms fusion and both processes are apparently dependent on continuous microtubule assembly.
Association of other viral proteins with microtubules have been demonstrated by co-immunoprecipitation assays, like Ebola virus matrix protein VP40 (Ruthel et al. 2005). Adenovirus binds to microtubules in a process mediated by dynein (Kelkar et al. 2004). We do not know however if the association of NSP2 and NSP5 with microtubules is directly to tubulin or mediated by other protein, like microtubule associated proteins (MAPs).
When the association of NSP2 and NSP5 with microtubules was analyzed at different times after infection, we found that NSP2 binds to microtubules as early as 4 h after infection, and remains associated throughout infection with little difference in the amount of protein detected. In contrast, NSP5 seem to have variable efficiency of association to microtubules, barely at 4 h but strongly at 12 h after infection. It is possible that NSP2 may bind first and than NSP5 in a sequential process in which NSP2 probably has an active role to recruit NSP5 and other viral proteins that form the viroplasms. Previous reports showed that a rotavirus mutant with a temperature sensitive lesion in the NSP2 gene does not form viroplasms (Raming & Petrie 1984), demonstrating the essential role of NSP2 in viroplasms formation.
The highest level of NSP2 and NSP5 bound to microtubules was reached at 12 h after infection that coincide with the maximal plus- and minus-strand RNA synthesis (Stacy-Phipps & Patton 1987), and infectious virus yield (McCrae & Faulkner-Valle 1981). Both NSP2 and NSP5 are involved in rotavirus replication, specifically in the packaging of the 11 double stranded RNA segments into the subviral particle (Kattoura et al. 1992, Taraporewala & Patton 2001, Vende et al. 2002). These data are consistent with the maximum level of these proteins bound to microtubules is at 12 h after infection. Both proteins NSP2 and NSP5 remain at viroplasms 24 h after infection, although the quantity of these proteins bound to microtubules decreased late after infection, and this is more evident for NSP5. The delay in the association of NSP5 to microtubules at early times PI combined with the intensity of its association at 12 h PI may reflect the dependence of the process on modifications like phosphorilation of this protein. In addition, it seems that the synthesis of NSP2 and NSP5 decreased at 24 h after infection, explaining the decrease in the amount of these proteins bound to microtubules at a time when both NSP2 and NSP5 are observed mostly at viroplasms and not diffuse through the cytoplasm as determined by immunofluosrecence.
In summary, our results suggest that rotavirus viro-plasms are associated to microtubules and an intact microtubule network is necessary for viroplasms formation. This is the first cellular element described to be involved in viroplasms formation.
ACKNOWLEDGEMENTS
To Simón Guzmán-León and Carolina Piña-Vázquez for excellent technical assistance. To Georgina Díaz and Gerardo Arrelín from the Instituto de Investigaciones Biomédicas, Universidad Nacional Autónoma de México, and Sergio Maldonado for their help to produce the hiperimmune anti-NSP2 serum. To Dr Carlos Arias for the generous gift of the anti-NSP5 serum, and Dr Bernard Moss form the National Institute of Health, Bethesda, MA, for providing the vaccinia vTF7-3 and the plasmid pVOTE1. To Drs Luis Vaca-Domínguez and Félix Recillas-Targa for critical advice throughut the PhD studies of CCR.
Received 10 February 2006
Accepted 10 April 2006
Finantial partial support: DGAPA-UNAM, grant IN222402, Consejo Nacional de Ciencia y Tecnología, México, grant 33371-N
This is part of the PhD manuscript of first author.
- Afrikanova I, Miozzo MC, Giambiagi S, Burrone OR 1996. Phosphorylation generates different forms of rotavirus NSP5. J Gen Virol 77: 2059-2065.
- Aponte C, Poncet D, Cohen J 1996. Recovery and characterization of the replicase complex in rotavirus-infected cells by using a monoclonal antibody against NSP2. J Virol 70: 985-991.
- Berois M, Sapin C, Erk I, Poncet D, Cohen J, 2003. Rotavirus nonstructural protein NSP5 interacts with major core protein VP2. J Virol 77: 1757-1763.
- Blackhall J, Fuentes A, Hansen K, Magnusson G 1997. Serine protein kinase activity associated with rotavirus phosphoprotein NSP5. J Virol 71: 138-144.
- Dales S 1963. Association between the spindle apparatus and reovirus. Proc Natl Acad Sci USA 50: 268-275.
- Eaton BT, Hyatt AD, White JR 1987. Association of bluetongue virus with the cytoskeleton. Virology 157: 107-116.
- Eichwald C, Rodriguez JF, Burrone OR 2004. Characterization of rotavirus NSP2/NSP5 interactions and the dynamics of viroplasm formation. J Gen Virol 85: 625-634.
- Estes, MK, Garham DY, Gerba CP, Smith EM 1979. Simian rotavirus SA11 replication in cell cultures. J Virol 31: 810-815.
- Estes MK 2001. Rotaviruses and their replication. In DN Knipe, PM Howley (eds), Virology, Lippincott Williams & Wilkings, Philadelphia, p. 1747-1785.
- Fabbretti E, Afrikanova I, Vascotto F, Burrone OR 1999. Two non-structural rotavirus proteins NSP2 and NSP5 form viroplasms like-structures in vivo. J Gen Virol 80: 333-339.
- Fuerst TR, Niles EG, Studier FW; Moss B 1986. Eukaryotic transient-expression system based on recombinant vaccinia virus that sinthesizes bacteriophage T7 RNA polymerase. Proc Natl Acad Sci USA 83: 8122-8126.
- Gallegos CO, Patton JT 1989. Characterization of rotavirus replication intermediates: a model for the assambly of single-shelled particles. Virology 172: 616-627.
- Kattoura MD, Clapp LL, Patton JT 1992. The rotavirus nonstructural protein NS35, posseses RNA-binding activity in vitro and in vivo. Virology 192: 698-708.
- Kelkar SA, Pfister KK, Crystal RG, Leopold PL 2004. Cytoplasmic dynein mediates adenovirus binding to microtubules. J Virol 78: 10122-10132.
- Laemmli UK 1970. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680-685.
- Lopez T, Camacho M, Zayas M, Najera R, Sanchez R, Arias CF, Lopez S 2005. Silencing the morphogenesis of rota-virus. J Virol 79:184-192.
- McCrae MA and Faulkner-Valle GP 1981. Molecular biology of rotavirus. I. Characterization of basic growth parameters and pattern of macromolecular synthesis. J Virol 39: 490-496.
- Nejmeddine M, Trugnan G, Spain C, Kohli E, Svensson L, Lopez S, Cohen J. 2000. Rotavirus spike protein VP4 is present at the plasm membrane and is associated with microtubules in infected cells. J Virol 74: 3313-3320.
- Parashar UD, Hummelman EG, Bresee JS, Miller MA, Glass RI 2003. Global illness and deaths caused by rotavirus disease in children. Emerg Infect Dis 9: 565-572.
- Parker JS, Broering TJ, Kim J, Higgins DE, Nibert ML 2002. Reovirus core protein m2 determines the filamentous morphology of viral inclusion bodies by interacting with and stabilizing microtubules. J Virol 76: 4483-4496.
- Petrie BL, Graham DY, Hanssen H, Estes MK 1982. Localization of rotavirus antigens in infected cells by ultrastructural immunocytochemistry. J Gen Virol 63: 457-467.
- Petrie BL, Greenberg HB, Graham DY, Estes MK 1984. Ultrastructural localization of rotavirus antigens using colloidal gold. Virus Res 1: 132-152.
- Poncet D, Lindenbaum P, Lharidon R, Cohen J 1997. In vivo and in vitro phosphorylation of rotavirus NSP5 correlates with its localization in viroplasms. J Virol 71: 34-41.
- Ramig RF, Petrie BL 1984. Characterization of temperature-sensitive mutants of simian rotavirus SA11: protein synthesis and morphogenesis. J Virol 49: 665-673.
- Ruthel G, Demmin GL, Kallstrom G, Javid MP, Badie SS, Will AB, Nelle T, Schokman R, Nguyen TL, Carra JH, Bavari S, Aman MJ 2005. Association of ebola virus matrix protein VP40 with microtubules. J Virol 79: 4709-4719.
- Stacy-Phipps S, Patton JT 1987. Synthesis of plus- and minus-strand RNA in rotavirus-infected cells. J Virol 61: 3479-3484.
- Taraporewala ZF and Patton JT 2001. Identification and characterization of the helix-desestabilizing activity of rotavirus nonstructural protein NSP2. J Virol 75: 4519-4527.
- Torres-Vega MA, Gonzalez RA, Duarte M, Poncet D, Lopez S, Arias CF 2000. The C-terminal domain of rotavirus NSP5 is essential for its multimerization, hyperphosphorylation and interaction with NSP6. J Gen Virol 81: 821-830.
- Vende P, Taraporewala ZF, Patton JT 2002. RNA-binding activity of the rotavirus phosphoprotein NSP5 includes affinity for double-stranded RNA. J Virol 76: 5291-5299.
- Weclewicz K, Svensson L, Billger M, Holmberg K, Wallin M, Kristensson K 1993. Microtubule associated protein 2 appears in axons of cultured dorsal root ganglia and spinal cord neurons after rotavirus infection. J Neurosci Res 36: 173-182.
- Xu A, Bellamy AR, Taylor JA 2000. Immobilization of the early secretory pathway by a virus glycoprotein that binds ro microtubules. EMBO J 19: 6465-6474.
Publication Dates
-
Publication in this collection
25 Oct 2006 -
Date of issue
Sept 2006
History
-
Accepted
10 Apr 2006 -
Received
10 Feb 2006