Abstract
The enterococci are important nosocomial pathogens with a remarkable capacity of expressing resistance to several antimicrobial agents. Their ubiquitous nature and resistance to adverse environmental conditions take account for their ability to colonize different habitats and for their potential for easy spreading through the food chain. In the present study we evaluated the distribution of species and antimicrobial susceptibility among enterococcal isolates recovered from food obtained in retail stores in Rio de Janeiro, Brazil. The following species were identified among 167 isolates obtained from poultry meat and 127 from pasteurized milk: Enterococcus faecalis (62.6%), E. casseliflavus (17.3%), E. durans (6.5%), E. gallinarum (3.0%), E. gilvus (2.4%), E. faecium (2.0%), E. hirae (1.4%), and E. sulfureus (1.0%). The overall percentages of antimicrobial resistant isolates were: 31.2 % to tetracycline, 23.8% to erythromycin, 11.3% to streptomycin, 4.3% to chloramphenicol, 3.9% to gentamicin, 1.4% to norfloxacin, 1.1% to imipenem, 0.7% to ciprofloxacin, nitrofurantoin, and penicillin, and 0.4% to ampicillin. Intermediate resistance was detected in frequencies varying from 0.5% for linezolid to 58.2% for erythromycin. None of the isolates showed resistance to glycopeptides. High-level resistance to aminoglycosides was observed in 13.1% of the isolates. Multiresistance was observed in E. faecalis, E. casseliflavus, E. faecium, E. gallinarum, E. durans and E. gilvus.
Enterococcus; Antimicrobial resistance; enterococci from poultry; enterococci from pasteurized milk
ARTICLES
Antimicrobial resistance profiles of enterococci isolated from poultry meat and pasteurized milk in Rio de Janeiro, Brazil
Suely Aparecida Pimenta FracalanzzaI, + + Corresponding author: suely.fracalanzza@incqs.fiocruz.br ; Erica Miranda Damásio ScheideggerI ; Patricia Faria dos SantosI; Paola Cardarelli LeiteI; Lúcia Martins TeixeiraII
IDepartamento de Microbiologia, Instituto Nacional de Controle de Qualidade em Saúde-Fiocruz, Av. Brasil 4365, 21040-900 Rio de Janeiro, RJ, Brazil
IIInstituto de Microbiologia, Universidade Federal do Rio de Janeiro, Rio de Janeiro, RJ, Brazil
ABSTRACT
The enterococci are important nosocomial pathogens with a remarkable capacity of expressing resistance to several antimicrobial agents. Their ubiquitous nature and resistance to adverse environmental conditions take account for their ability to colonize different habitats and for their potential for easy spreading through the food chain. In the present study we evaluated the distribution of species and antimicrobial susceptibility among enterococcal isolates recovered from food obtained in retail stores in Rio de Janeiro, Brazil. The following species were identified among 167 isolates obtained from poultry meat and 127 from pasteurized milk: Enterococcus faecalis (62.6%), E. casseliflavus (17.3%), E. durans (6.5%), E. gallinarum (3.0%), E. gilvus (2.4%), E. faecium (2.0%), E. hirae (1.4%), and E. sulfureus (1.0%). The overall percentages of antimicrobial resistant isolates were: 31.2 % to tetracycline, 23.8% to erythromycin, 11.3% to streptomycin, 4.3% to chloramphenicol, 3.9% to gentamicin, 1.4% to norfloxacin, 1.1% to imipenem, 0.7% to ciprofloxacin, nitrofurantoin, and penicillin, and 0.4% to ampicillin. Intermediate resistance was detected in frequencies varying from 0.5% for linezolid to 58.2% for erythromycin. None of the isolates showed resistance to glycopeptides. High-level resistance to aminoglycosides was observed in 13.1% of the isolates. Multiresistance was observed in E. faecalis, E. casseliflavus, E. faecium, E. gallinarum, E. durans and E. gilvus.
Key words:Enterococcus - Antimicrobial resistance - enterococci from poultry - enterococci from pasteurized milk
The presence of antimicrobial resistant bacteria in foodstuffs of animal origin is becoming a matter of important concern as they may be transmitted to humans through the food supply. Therefore, protection of food supplies includes the microbiological quality and safety of commodities available for public consumption. While such concerns most frequently address pathogenic microorganisms, that present immediate risks to human health, there is a growing interest in commensal components of the microbiota associated with food (Giraffa et al. 1997, Giraffa 2002, Hayes et al. 2003, Mannu et al. 2003). Commensal bacteria contaminate food, water and the environment, and their dissemination can also be related to spreading of antimicrobial resistance determinants to pathogenic or to other commensal microorganisms, which may then infect or colonize human beings and animals (van den Bogaard & Stobbering 2000). In this context, members of genus Enterococcus are a matter of major attention, as they are ubiquitous bacteria widely distributed in a variety of habitats. They comprise a high proportion of saprophyte bacteria associated with the gastrointestinal tract of human beings and animals, and are usually found in large numbers in soil, water, food of vegetal origin, and especially of animal origin, such as beef, poultry and swine carcass (Franz et al. 1999, Giraffa 2002, Domig et al. 2003). Because of their abundance in faeces and also due to their ability of long last survival out of the enteric environment, the presence of enterococci is considered a indicator of fecal contamination and they are not considered "generally recognized as safe" (GRAS) (Domig et al. 2003, Mannu et al. 2003).
Although they are not considered as primary pathogens, enterococci rank among leading causes of nosocomial infections. In fact, some species of Enterococcus have currently a particular medical relevance, considering their role as cause of infections in predisposed humans and their remarkable capacity of acquiring resistance to several antimicrobials, leading to increasingly limited therapeutic options (Franz et al. 1999, Teixeira & Facklam 2003). In addition to the intrinsic resistance to several antimicrobials, the enterococci are characterized by their unique ability of exchanging genetic material (Leclercq 1997, Murray 1998), and to acquire resistance to many antimicrobial agents, including aminoglycosides, chloramphenicol, B-lactams, macrolides, quinolones, tetracycline, and, more recently, to the glycopeptides, represented by vancomycin and teicoplanin (Eaton & Gasson 2001, Teixeira & Facklam 2003, Huys et al. 2004). The occurrence of antimicrobial resistance among enterococci is not only restricted to the nosocomial setting, and therefore resistant strains carried on sources such as food may also act as potential reservoirs of antimicrobial resistance genes.
Monitoring the prevalence and antimicrobial resistance profiles of a bacterium with such a representativeness as the enterococci in different sources related to human and animals may contribute with insights on the possible exchange of strains and their antimicrobial resistance genes occurring among bacteria colonizing both animals and humans. Information on the prevalence of antimicrobial resistant enterococci in different food sources is, however, still limited in many regions. The purpose of this study was to determine the occurrence and distribution of the different species as well as the antimicrobial resistance profiles among enterococcal isolates recovered from foodstuffs of animal origin, represented by poultry and pasteurized milk, commercialized in Rio de Janeiro, RJ, Brazil.
MATERIALS AND METHODS
Bacterial strains and methods of isolation and identification - A total of 294 enterococcal isolates recovered from poultry meat (167; 56.8%) and from pasteurized milk (127; 43.2%) samples, obtained in retail stores located in Rio de Janeiro, Brazil, from October 2002 to October 2004 were examined. They were isolated from 50 food samples (25 poultry samples and 25 pasteurized milk samples) according to the methodology proposed by Andrews and June (1998). Briefly, poultry samples (25g) were placed in sterile plastic-bags containing 225 ml of buffered peptone (Oxoid Limited, Hampshire, UK) water, and then homogenized at 230 rpm for 2 min in a laboratory blender (Stomacher, Seward Limited, London, UK). Pasteurized milk samples (25 ml) were homogenized in 225 ml of buffered peptone water distributed in sterile flask. Aliquots of 1 ml of the homogenates were plated on Enterococcosel agar (Becton Dickinson Microbiology Systems, Sparks, MD, USA) plates. Up to ten typical colonies suggestive of Enterococcus were randomly selected from each primary isolation culture on Enterococosel agar for further identification.
The isolates were identified on the basis of cellular morphology following Gram stain, and results of biochemical testing, including catalase production, pyrrolidonylarylamidase activity, growth in 6.5% NaCl broth, hydrolysis of esculin in the presence of bile, pigment production, haemolytic activity, arginin hydrolysis, tolerance to tellurite, utilization of pyruvate, motility and acid production from the following carbohydrates: L-arabinose, mannitol, methyl-â-glucopyranoside (MGP), D-raffinose, sucrose, and sorbitol (Facklam & Collins 1989, Teixeira & Facklam 2003).
Antimicrobial susceptibility testing - Susceptibility to antimicrobials was evaluated for 282 isolates by the disk diffusion method, according to the Clinical and Laboratory Standard Institute (CLSI 2006a, b) guidelines. The following 14 antimicrobial agents were tested: ampicillin, chloramphenicol, ciprofloxacin, erythromycin, gentamicin, imipenem, linezolid, nitrofurantoin, norfloxacin, penicillin, streptomycin, teicoplanin, tetracycline, and vancomycin. Interpretation of susceptibility tests with imipenem were performed as recommended by Weinstein et al. (2004). Enterococcus faecalis ATCC 29212 and Staphylococcus aureus ATCC 25923 were used as quality control strains.
RESULTS
Isolation and identification of enterococcal strains - Two hundred and ninety four isolates characterized as Enterococcus were studied. Enterococcal isolates were recovered from all the poultry samples and from 18 (72%) of the pasteurized milk samples analyzed. Multiple isolates from a single sample where considered when two or more different species were detected or when isolates had differences in any of the phenotypic characteristics investigated.
Table I shows the distribution of the different enterococcal species identified among the isolates obtained from poultry and pasteurized milk samples. Overall, the species identified were: E. faecalis (62.6%), Enterococcus casseliflavus (17.3%), Enterococcus durans (6.5%), Enterococcus gallinarum (3.0%), Enterococcus gilvus (2.4%), Enterococcus faecium (2.0%), Enterococcus hirae (1.4%), and Enterococcus sulfureus (1.0%). Eleven (3.8%) isolates were not identified to the species level.
Antimicrobial susceptibility - The results of antimicrobial susceptibility testing obtained for 282 isolates are shown in Table II. Resistance to antimicrobials was found among the different Enterococcus species isolated from both foodstuff sources. The overall percentages of antimicrobial resistant isolates were: 31.2% to tetracycline, 24.1% to erythromycin, 11.3% to streptomycin, 4.3% to chloramphenicol, 3.9% to gentamicin, 1.4% to norfloxacin, 1.1% to imipenem, 0.7% to ciprofloxacin, nitrofurantoin, and penicillin, and 0.4% to ampicillin. None of the isolates showed full resistance to vancomycin, teicoplanin or linezolid. However, intermediate resistance to linezolid was detected in one (0.5%) E. faecalis isolate. Resistance to tetracycline was more frequently observed among E. faecium (80.0%), E. casseliflavus (39.2%), E. faecalis (32.6%), E. hirae (25.0%) E. gallinarum (22.2%), and E. gilvus (14.3%) isolates, while resistance to erythromycin was more frequent among E. faecium (60.0%), E. faecalis (30.2%), E. durans (15.8%), E. gilvus (14.3%), E. gallinarum (11.1%), and E. casseliflavus (9.8%). Resistance to imipenem was observed among 40.0% of the E. faecium isolates, and among 5.3% of those identified as E. durans. A high percentage (58.2%) of isolates presenting intermediate resistance to erythromycin was observed, distributed among the different species identified, except for E. sulfureus. Resistance to only one of the antimicrobials tested was observed in 26 (9.2%) isolates, including 20 recovered from meat poultry and 6 from pasteurized milk. Susceptibility to all antimicrobials tested was observed in 46 (16.3%) isolates, including 31 obtained from meat poultry and 15 from pasteurized milk.
The occurrence of high-level resistance to amino-glycosides (HLR-A) among the different species of enterococci identified according to their source is shown in Table III. HLR-A (gentamicin and/or streptomycin) was detected in 13.1% of the isolates (10.6% from poultry and 2.5% from pasteurized milk samples). High-level resistance to gentamicin (HLR-GE) was detected in 1.8% of the isolates. The frequency of HLR-GE was higher among isolates from poultry than among those recovered from pasteurized milk samples. Isolates presenting HLR-GE recovered from poultry were identified as E. casseliflavus and E. faecalis. High-level resistance to streptomycin (HLR-ST) was also more frequently detected among poultry isolates, involving E. faecalis and E. hirae. Isolates recovered from pasteurized milk samples and showing HLR-ST were identified as E. faecalis and E. hirae. Simultaneous high-level resistance to both gentamicin and streptomycin (HLR-GE/ST) was detected in 2.1% of the isolates, involving E. gallinarum and E. casseliflavus recovered from poultry, and E. durans isolated from pasteurized milk samples.
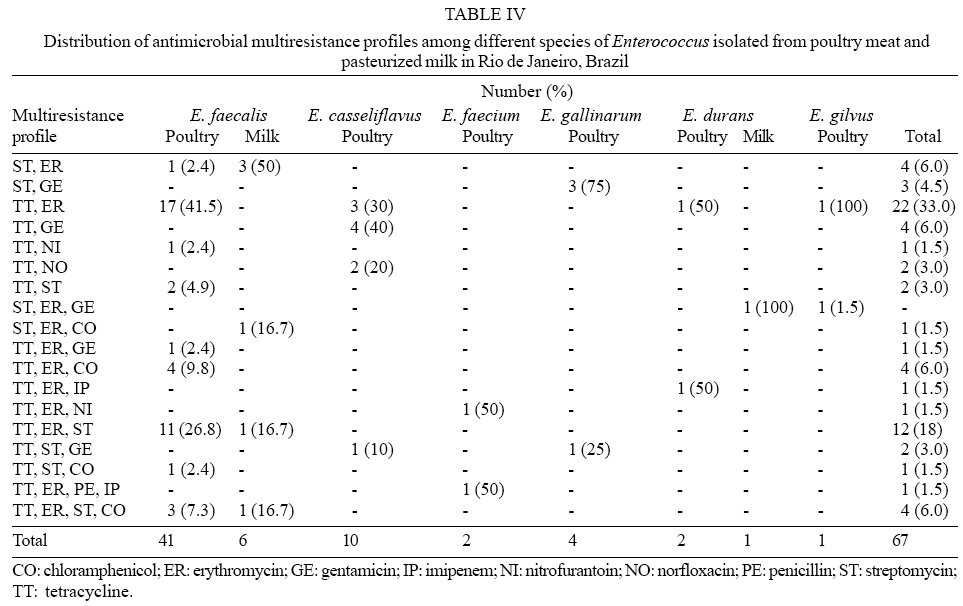
Among the 282 enterococcal isolates tested, 25.1% (21.3% from poultry and 2.5% from pasteurized milk) presented resistance to two or more of the antimicrobial agents tested. Sixty seven multiresistant isolates were found. E. faecalis was the species most frequently associated with multiresistance, corresponding to 61.2% of the multiresistant isolates from poultry, followed by E. casseliflavus (15.0%), E. gallinarum (6.0%), E faecium and E. durans (3.0%), and E. gilvus (1.5%). The species identified among multiple antimicrobial resistant isolates recovered from pasteurized milk samples were E. faecalis (9.0%) and E. durans (1.5%). A large proportion (41.5%) of the multiresistant E. faecalis isolates from poultry was simultaneously resistant to tetracycline and erythromycin, followed by 26.8% resistant to tetracycline, erythromycin and streptomycin. Among poultry isolates, 40% of those identified as E. casseliflavus were simultaneously resistant to tetracycline and gentamicin, while 75% of the E. gallinarum isolates were resistant to both streptomycin and gentamicin. Simultaneous resistance to streptomycin and to erythromycin was detected in 50% of the E. faecalis isolates from pasteurized milk (Table IV).
DISCUSSION
The enterococci have emerged as important opportunistic pathogens with a remarkable capacity of expressing resistance to several groups of antimicrobial agents, limiting the number of therapeutic options. They are associated with a variety of human infections, acquired mainly in the nosocomial setting, such as bacteremia, endocarditis, urinary tract infections and wound infections (Murray 1990, Teixeira & Facklam 2003, Andrade et al. 2006). On the other hand, species of the genus Enterococcus comprise a large proportion of the normal microbiota associated with the gastrointestinal tract of humans and animals (Giraffa 2002, Hayes et al. 2003). They can be found on almost everything that surrounds human beings, including foodstuff, mainly raw products of animal origin (meat and milk), and from sources associated with low hygienic conditions, where their presence is an indicator of fecal contamination (Mannu et al. 2003). The ubiquitous nature of the enterococci and resistance to adverse environmental conditions take account for their ability to colonize different habitats, and for their potential for easy spreading through the food chain. Therefore, the presence of antimicrobial resistant enterococci in foods has been a matter of growing concern. The present study reports the distribution of species and the antimicrobial resistance profiles among enterococcal isolates recovered from poultry meat and pasteurized milk samples obtained in retail stores in the city of Rio Janeiro, Brazil, in the period of October 2002 to October 2004.
Although differences in percentages were noticeable, E. faecalis was the predominant species recovered from both poultry (50.9%) and pasteurized milk (77.9%) samples, in accordance with previous studies (Franz et al. 1999, Gelsomino et al. 2001, Giraffa 2003). The second most frequent enterococcal species isolated from poultry was E. casseliflavus (26.3%), followed by E. gallinarum and E. gilvus (4.2% each), while E. durans (12.6%) was the second most frequent species recovered from milk, followed by E. casseliflavus (5.5%). The prevalence of E. faecium in the present study was low, even in poultry samples (3.0%), in contrast with other reports indicating E. faecium as the most frequent enterococcal species isolated from poultry (Hayes et al. 2003). Pigment-producing enterococcal species (E. casseliflavus, E. gilvus and E. sulfureus) accounted for 32.3% of the poultry isolates, and for 5.5% (E. casseliflavus only) of those recovered from milk. On the other hand, species with naturally occurring intrinsic low level resistance to vancomycin (E. casseliflavus and E. gallinarum) represented 30.5% of the isolates from poultry and 7.1% of the milk isolates.
The enterococci are intrinsically resistant to a large range of antimicrobials of therapeutic use, and they are also known by the ability to acquire and transfer resistance markers by a process mediated by genes present in plasmids and transposons that facilitate their dissemination (Murray 1990). Data on antibiotic resistance of enterococci associated with foodstuffs indicate that there is a strong epidemiological evidence of a link between the use of antibiotics in human medicine and animal husbandry and the emergence, spreading and persistence of resistant strains in animal products (van den Bogaard & Stobbering 2000). The major risk related to the resistance traits is that they are for the most part transferable to the enterococci and more virulent pathogens as well. In the present work enterococci showing resistance to ampicillin, chloramphenicol, erythromycin, gentamicin, imipenem, penicillin, nitrofurantoin, norfloxacin, streptomycin, and tetracycline were isolated from poultry meat as well as from milk. Resistance to tetracycline, erythromycin, and chloramphenicol was detected in percentages of 31.2%, 23.8%, and 4.3%, respectively. When intermediate results are added, the percentages of isolates with decreased susceptibility to these antimicrobials were 38.3%, 82.0%, and 5.7%, respectively. These percentages of resistance are significant, while lower or occasionally comparable to those previously reported for enterococci recovered from human sources in Brazil (Mondino et al. 2003, d'Azevedo et al. 2004 ) and other locations (Lopes et al. 2003). The observation of concomitant resistance to chloramphenicol, erythromycin, and tetracycline confirms the frequent resistance to multiple antibiotics among enterococcal isolates from humans sources in Brazil (Mondino et al. 2003, Vilela et al. 2006). Although a large proportion of isolates was susceptible to gentamicin and streptomycin, HLR-A was still detected in a significant percentage (12.8%) and comprising different species, including E. faecalis, E. casseliflavus, E. gallinarum, E. durans, and E. hirae. Interestingly, in the present study, HLR-ST to streptomycin was most frequently detectd among E. faecalis isolates, while HLR-GE to gentamicin was most frequently observed among E. casseliflavus. The occurrence of an isolate presenting intermediate results to linezolid is an important matter. Decreased susceptibility to linezolid, an antimicrobial agent that has been used in clinical practice for a relatively short period of time, has been sporadically detected in enterococci from human sources worldwide (Bersos et al. 2004). To our knowledge, however, it has not been found among isolates from other sources yet. In vitro susceptibility tests for linezolid can be performed by disk diffusion. However, non-susceptible isolates can produce indistinct zones of inhibition when tested by this method making interpretation of test results very difficult (Tenover et al. 2007). The isolate that was considered as intermediate to linezolid in the in the present work gave zones of inhibition varying from 20 to 23 mm in tests performed in different days and using disks obtained from different manufactures. Such variations around the cutoff value may lead to differences in interpretation. On the other hand presence of isolates with such important antimicrobial resistance markers in foodstuff is a cause of concern, considering the possibility of these microorganisms spread through the food chain. Therefore, we are planning to confirm such result by determining the minimum inibitory concentration for linezolid.
The results indicating absence of vancomycin-resistant enterococci (VRE) among isolates from poultry are consistent with previous observations in the United States (Butaye et al. 2001). In contrast, VRE were frequently isolated from raw meat products in European countries, probably as a result of selection of resistant populations, since the use of avoparcin, a glycopeptide applied as a growth promoter in animal food, was allowed in these countries earlier (Aaerestrup et al. 2000).
Multi-resistant enterococcal populations are commonly isolated from human beings, aquatic environments, sewage waters and can be introduced into human intestinal microflora by the consumption of food. The enterococcal resistance to multiple antibiotics with the propensity to acquire new traits continues to create therapeutic problems (Murray 1998). Their presence in food may represent a more serious risk of expanding antibiotic resistance through the food chain (Giraffa 2002). In this work, multiresistance to antimicrobials was particularly detected among E. faecalis, E. casseliflavus, and E. faecium recovered from poultry, and among E. gallinarum, E. faecalis and E. durans obtained from milk.
Our observations corroborate previous findings demonstrating that different species of Enterococcus can be frequently found in poultry as well as in pasteurized milk (Giraffa et al. 1997). Considering the rather high heat resistance of Enterococcus, it is probable that isolates would be detected in pasteurized milk. However, a lack of discipline during subsequent processing can lead to the secondary contamination of this product (Teuber et al. 1999). In addition, the presence of strains with clinically significant antimicrobial resistance markers indicate the need of effective control strategies to reduce contamination of foodstuff by these microorganisms, as they may represent potential reservoirs of virulence and resistance genes with access to human host adapted bacterial cells that can be disseminated through the food chain, and may represent significant cause of Public Health problems. Therefore, monitoring programs are essential to trace trends in the development and dissemination of resistance among isolates from different sources, especially for those consumers who are at highest risk, i.e. the elderly and the immunocompromised.
ACKNOWLEDGEMENTS
To Filomena Soares Pereira da Rocha and Jaqueline Martins Morais from Instituto de Microbiologia, Universidade Federal do Rio de Janeiro, for technical assistance.
Received 1 June 2007
Accepted 9 October 2007
Financial support: INCQS-Fiocruz, CNPq, Faperj, MCT/Pronex
- Aarestrup FM, Kruse H, Tast E Hammerum AM, Jensen LB 2000. Associations between the use of antimicrobial agents for growth promotion and the occurrence of resistance among Enterococcus faecium from broilers and pigs in Denmark, Finland, and Norway. Microb Drug Resist 6: 63-67.
- Andrade SS, Sader HS, Jones RN, Pereira AS, Pignatary ACC, Gales AC 2006. Increased resistance to first-line agents among bacterial pathogens isolated from urinary tract infections in Latin America: time for local guidelines? Mem Inst Oswaldo Cruz 101: 741-748.
- Andrews WH, June GA 1998. Food sampling and preparation of homogenates. In RL Merker (ed.), Food and drug administration bacteriological analytical manual, 8th ed., AOAC International, Gaithersburg, MD, p. 1-9.
- Bersos Z, Maniati M, Kontos F, Petinaki E, Maniatis AN 2004. First report of a linezolid-resistant vancomycin-resistant Enterococcus faecium strain in Greece. J Antimicrob Chemoth 53: 685-686.
- Butaye P, Devriese LA, Haesebrouck F 2001. Differences in antibiotic resistance patterns of Enterococcus faecalis and Enterococcus faecium strains isolated from farm and pet animals. Antimicrob Agents Chemother 45: 1374-1378.
- CLSI - Clinical and Laboratory Standard Institute 2006a. Performance standards for antimicrobial disk susceptibility tests. Approved Standard, 9th ed., CLSI document M2-A9. Clinical and Laboratory Standard Institute, Wayne, PA, p. 1-37.
- CLSI - Clinical and Laboratory Standard Institute 2006b. Performance standards for antimicrobial susceptibility testing 16th Informational Suplement, CLSI document M100-S16. Clinical and Laboratory Standard Institute, Wayne, PA, p. 1-173.
- d'Azevedo PA, Dias CAG, Lemos, SK, Bittencourt JAF, Teixeira LM 2004. Antimicrobial Susceptibility among Enterococcus Isolates from the City of Porto Alegre, RS, Brazil. Braz J Microbiol 35: 199-204.
- Domig KJ, Mayer HK, Kneifel W 2003. Methods used for the isolation, enumeration, characterisation and identification of Enterococcus spp.: 1. Media for isolation and enumeration. Int J Food Microbiol 88: 147-164.
- Eaton, TJ, Gasson MJ 2001. Molecular screening of Enterococcus virulence determinants and potential for genetic exchange between food and medical isolates. Appl Environ Microbiol 67: 1628-1635.
- Facklam RR, Collins MD 1989. Identification of Enterococcus species isolated from human infections by a conventional test scheme. J Clin Microbiol 27: 731-734.
- Franz CMAP, Holzapfel WH, Stiles ME 1999. Enterococci at the crossroads of safety? Int J Food Microbiol 47: 1-24.
- Gelsomino R, Vancanneyt M, Condon S, Swing J, Cogan TM 2001. Enterococcal diversity in the cheesemaking environment of an Irish Cheddar-type cheesemaking factory. Int J Food Microbiol 71: 177-178.
- Giraffa G 2002. Enterococci from foods. FEMS Microbiol Rev 26: 163-171.
- Giraffa G 2003. Funcionality of enterococci in dairy products. Int J Food Microbiol 88: 215-222.
- Giraffa G, Carminati D, Neviane E 1997. Enterococci isolated from dairy products: a review of risks and potential technological use. J Food Prot 60: 365-378.
- Hayes JR, Enghish LL, Carter PJ, Proescholt T, Lee KY, Wagner DD, White DG 2003. Prevalence and antimicrobial resistance of Enterococcus species isolated from retail meats. Appl Environ Microbiol 69: 7153-7160.
- Huys G, D'Haene K, Collard J-C, Swings J 2004. Prevalence and molecular characterization of tetracycline resistance in Enterococcus isolates from food. Appl Environ Microbiol 70: 1555-1562.
- Leclercq R 1997. Enterococci acquire new kinds of resistance. Clin Infect Dis 24: S80-S84.
- Lopes FS, Ribeiro T, Martins MP, Terreiro R, Crespo MTD 2003. Gentamicin resistance in dairy and clinical enterococcal isolates and in reference strains. J Antimicrob Chemother 52: 214-219.
- Mannu L, Paba A, Daga E, Comunian, R, Zannetti S, Duprè I, Sacchi LA 2003. Comparison of the incidence of virulence determinants and antibiotic resistance between Enterococcus faecium of animal and clinical origin. Int J Food Microbiol 88: 291-304.
- Mondino SSB, Castro ACD, Mondino PJJ, Carvalho MGS, Silva KMF, Teixeira LM 2003. Phenotypic and genotypic characterization of clinical and intestinal enterococci isolated from inpatients and outpatients in two Brazilian hospitals. Microbial Drug Resist 9: 167-174.
- Murray BE 1990. The life and times of the Enterococcus Clin Microbiol Rev 3: 46-65.
- Murray BE 1998. Diversity among multidrug-resistant enterococci. Emerg Infect Dis 4: 37-47.
- Teixeira LM, Facklam RR 2003. Enterococcus In PR Murray, EJ Baron, JH Jorgensen, MA Pfaller, RH Yolken (eds), Manual of Clinical Microbiology, 8th ed., American Society for Microbiology, Washington, DC, pp. 422-433.
- Tenover FC, Williams PP, Stocker S, Tompson A, Clarck LA, Limbago B, Carey RB, Poppe SM, Shinabarger D, McGowan Jr JE 2007. Accuracy of six antimicrobial susceptibility methods for testing linezolid against staphylococci and enterococci. J Clin Microbiol 45: 2917-2922.
- Teuber M, Miele L, Schwarz F 1999. Acquired antibiotic resistance in lactic acid bacteria from food. Antoine van Leeuwe-nhoek 76: 115-137.
- van den Bogaard AE, Stobbering EE 2000. Epidemiology of resistance to antibiotic. Links between animals and humans. Int J Antimicrob Agents 14: 327 -335.
- Vilela MA, Souza SL, Palazzo ICB, Ferreira JC, Morais Jr. M A, Darini ALC, Morais MMC 2006. Identification and molecular characterization of Van A-type vancomycin-resistant Enterococcus faecalis in Northeast of Brazil. Mem Inst Oswaldo Cruz 101: 716-719.
- Weinstein MP, Mirret S, Kannangara S, Monahan J, Harrel L J, Wilson A C, Reller B 2004. Multicenter evaluation of use of penicillin and ampicillin as surragates for in vitro testing of susceptibility of enterococci to imipenem. J Clin Microbiol 42: 3747-3751.
Publication Dates
-
Publication in this collection
31 Aug 2007 -
Date of issue
Nov 2007
History
-
Accepted
09 Oct 2007 -
Received
01 June 2007