Abstract
We show for the first time that the ventral diverticulum of the mosquito gut (impermeable sugar storage organ) harbors microorganisms. The gut diverticulum from newly emerged and non-fed Aedes aegypti was dissected under aseptic conditions, homogenized and plated on BHI medium. Microbial isolates were identified by sequencing of 16S rDNA for bacteria and 28S rDNA for yeast. A direct DNA extraction from Ae. aegypti gut diverticulum was also performed. The bacterial isolates were: Bacillus sp., Bacillus subtilis and Serratia sp. The latter was the predominant bacteria found in our isolations. The yeast species identified was Pichia caribbica.
Aedes aegypti; gut diverticulum; microbiota; Serratia sp
ARTICLES
First isolation of microorganisms from the gut diverticulum of Aedes aegypti (Diptera: Culicidae): new perspectives for an insect-bacteria association
Desiely Silva GusmãoI; Adão Valmir SantosI; Danyelle Cristine MariniII; Érica de Souza RussoI; Anelise Maria Dias PeixotoI; Maurício Bacci JúniorIII; Marília Amorim Berbert-MolinaI; Francisco José Alves LemosI, + + Corresponding author: franze@uenf.br
ILaboratório de Biotecnologia, Universidade Estadual do Norte Fluminense Darcy Ribeiro, Av. Alberto Lamego 2000, 28013-602 Campos dos Goytacazes, RJ, Brasil
IIFaculdades Integradas Maria Imaculada, Mogi-Guaçu, SP, Brasil
IIILaboratório de Evolução Molecular, Universidade Estadual Paulista, Rio Claro, SP, Brasil
ABSTRACT
We show for the first time that the ventral diverticulum of the mosquito gut (impermeable sugar storage organ) harbors microorganisms. The gut diverticulum from newly emerged and non-fed Aedes aegypti was dissected under aseptic conditions, homogenized and plated on BHI medium. Microbial isolates were identified by sequencing of 16S rDNA for bacteria and 28S rDNA for yeast. A direct DNA extraction from Ae. aegypti gut diverticulum was also performed. The bacterial isolates were: Bacillus sp., Bacillus subtilis and Serratia sp. The latter was the predominant bacteria found in our isolations. The yeast species identified was Pichia caribbica.
Key words:Aedes aegypti - gut diverticulum - microbiota - Serratia sp.
Aedes aegypti (Diptera: Culicidae, Aedini) is the main urban vector for the human diseases yellow fever and dengue fever (Nasci & Miller 1996). The vector control, mainly by insecticide application and elimination of oviposition sites, has been used as the best solution to decrease the diseases incidence. Explore new strategies for blocking the insect transmitted diseases such as dengue fever is urgent in Tropical countries (Sperança & Capurro 2007). Microorganisms associated with the insect may have an important role for human infectious diseases epidemiology. Once a key role bacterium is discovered, it may be modified in order to affect the pathogen development and, consequently, the disease transmission (Azambuja et al. 2005, Riehle & Jacobs-Lorena 2005).
The alimentary canal of the mosquito is composed by the foregut, midgut and hindgut. The foregut is involved primarily with ingestion, conduction and storage of food (Romoser 1996). In the alimentary canal of the mosquito three diverticula arise near the posterior end of the esophagus: two from the dorso-lateral and one from the ventral wall of the gut, all surrounded by a thin impermeable cuticle. The ventral diverticulum (VD), or crop, is large and may extend into the abdomen (Dapples & Lea 1974). Usually all three diverticula are filled with air bubbles and are used as food reservoirs (Thompson 1905, Consoli & Lourenço-de-Oliveira 1994). The sugar meal, such as floral nectar, is stored in the diverticula and passes slowly to the midgut, where it is digested (Thompson 1905).
Microorganisms play important and often essential roles in the growth and development of many insect species. Despite the importance of these microbial associations with insects there are relatively few studies that elucidate their components and their roles in the interactions. Insects that rely on nutritionally poor diets tend to possess bacterial endosymbionts. Aphids, for example, which subsist solely on plant sap, harbor Buchnera spp., believed to provide amino acids and vitamins to their hosts (Douglas 1989). Blood is known to be severely deficient in essential B vitamins and some amino acids. Bloodsucking arthropods such as ticks, lice, bedbugs, reduviid bugs, and tsetse flies usually harbor symbiotic microorganisms (Buchner 1965). Symbiotic relationships have already been reported for blood feeding insects and some are very well established, as in the case of Rhodnius prolixus that maintains an association with the actinomycete bacteria Rhodococcus rhodnii (Dasch et al. 1984); and tsetse flies genus Glossina, which harbor three distinct symbiotic microorganisms: Wigglesworthia glossinidia, Sodalis glossinidius (commensal), and the parasitic microbe Wolbachia pipientis (Aksoy et al. 1997, Chen et al. 1999).
Many key questions about bacteria within the mosquito's midgut remain largely unanswered, and obligate bacteria have not been identified in mosquito's digestive canal to date. Some works have already reported the isolation of microorganisms from the midgut of Culex quinquefasciatus, Anopheles dirus, Anopheles albimanus, and Ae. aegypti (Pumpuni et al. 1996, Khampang et al. 1999, Luxananil et al. 2001, Pidiyar et al. 2002, 2004, Gonzalez-Ceron et al. 2003). The microbiota of Anopheles gambiae and Anopheles funestus was also investigated by 16S rDNA sequence by Lindh et al. (2005). The objective of this work was to investigate if Ae. aegypti VD harbored microorganisms.
MATERIALS AND METHODS
Mosquitoes - Insects were obtained from colonies of Ae. aegypti (Rockfeller strain), maintained in the insectary of the Laboratory of Biotechnology (UENF, Campos dos Goytacazes, Brazil). Mosquitoes were reared at 27ºC and provided with sterile 10% sucrose. A glass container filled with sterile distilled water was kept inside each cage to maintain the humidity. Larvae were fed on a sterile minced commercial mouse food. Pupae were rinsed and transferred to sterile distilled water and maintained in separate cages in aseptic conditions until adult emergence. All mosquitoes were newly emerged non-fed females (no blood or sucrose feeding), and were dissected on the same day of emergence.
Isolation of microorganisms - In order to surface sterilize, mosquitoes were rinsed serially, for 1 min, in the following solutions: sodium hypochlorite (1%), sterile phosphate-buffered saline (PBS) (81 mM Na2HPO4, 19 mM NaH2PO4, 150 mM NaCl, pH 7.4) and ethanol (70%). Finally, the insects were rinsed three times in PBS/1 min. Aliquots of 100 µl from the last PBS washes were plated in brain heart infusion (BHI) agar as control groups of the surface sterilization process. The sterilizations and dissections were performed in a laminar flow hood. Mosquitoes were dissected under a microscope, in a double cavity glass slide containing sterile PBS. The VD was carefully separated from the midgut, rinsed in sterile PBS, and transferred into a 1.5 ml tube, containing 200 µl of BHI. This procedure was repeated until five crops were obtained. The tubes were mixed thoroughly with a pestle, and an aliquot of 100 µl was transferred to a 50 ml Erlenmeyer containing 20 ml of BHI medium and 100 µl glucose. An aliquot of 100 µl was also transferred to Petri dishes containing BHI agar. The Erlenmeyers were incubated at 28ºC for 24-48 h, under agitation (80 rpm). After incubation the cultures were serially diluted (10-1 through 10-7) and an aliquot of 100 µl of each one was transferred to Petri dishes containing BHI agar. Plates were incubated at 28ºC for 24-48 h. Bacterial isolates were maintained at -70ºC in a solution of glycerol (15%) for further identification. We carried out a total of seven isolation assays, from different groups of Ae. aegypti, at different times. At the end, a total of 35 crops were sampled.
Identification of microorganisms - Microorganisms were first screened based on colony characteristics, morphology of isolates, Gram staining and motility by the hanging drop technique.
DNA extraction from bacterial isolates and amplification of rDNA 16S - The DNA extraction was adapted from Ausubel et al. (1992). The rDNA 16S was amplified using the following universal primers: 27f (Lane et al. 1985) and 1492r (Delong 1992). Polymerase chain reaction (PCR) was performed with Ready-To-Go kit (Amersham Pharmacia Biotech), template DNA solution (2 µl/100 hg), 27f primer (1 µl/6 rmol), 1492r primer (1 µl/6 rmol), 25 mM MgCl2 (1.5 µl), and 17.5 µl of ultra pure water. Cycling parameters for the PCR included an initial denaturation step at 95ºC/5 min, followed by 35 cycles of a denaturation step at 95ºC/1 min, a primer annealing step at 50ºC/1 min, an extension step at 72ºC/3 min, and a final step at 72ºC/4 min. The 16S rDNA amplification generated a product of approximately 1500 bp.
Amplification of a section rDNA 28S - Yeast DNA was extracted and purified as described above. The divergent D1/D2 domain (nucleotides 63-642 for Saccharomyces cerevisiae) at the 5'end of the large-subunit rRNA gene was symmetrically amplified with primers NL1 and NL4 (O'Donnell 1993). Each PCR was performed with the Ready-To-Go kit (Amersham Pharmacia Biotech) adding solution containing DNA (1 µl), NL-1 primer (1.6 µl/6 rmol), NL-4 primer (1.1 µl/6 rmol), 25mM MgCl2 (1.5 µl), and 17.8 µl of ultra pure water.
DNA extraction from Ae. aegypti gut diverticulum - Ae. aegypti females were dissected and the VD were carefully separated from the midgut under aseptic conditions. The DNA extraction was performed with the Wizardâ genomic DNA purification kit (Promega TM050).
Cloning - After purified, fragments were cloned using the Kit pGem-T Easy Vector Systems (Promega, USA) in Escherichia coli DH5a.
Sequencing reaction - The sequencing reaction was prepared as follows: Big Dye Terminator Kit (Applied Biosystems, Foster City, CA) (2 µl), "Save Money" buffer (200 mM Tris HCl, pH 9.0, 5 mM MgCl) (2 µl), forward T7 or reverse SP6 promoters (1 µl/5.0 rmoles), DNA (2 µl /100 hg) and ultra pure water (2 µl). PCR conditions were 96ºC/10 min, 50ºC/5 min, and 60ºC/4 min, 40 cycles. The sequencing of the 16S rRNA gene from the bacterial clones was done using a 27f primer (5'-end; 500 nucleotide region) and from the yeast it was done using the NL1 primer. Sequencing was carried out on an ABI Prism 377 DNA sequencer (Applied Biosystems).
Sequences alignment - The sequences obtained were initially compared to the ones deposited at the GenBank, using BLAST (http://www.ncbi.nlm.nih.gov/BLAST). The generated sequences and their homologs, which were retrieved from the GenBank, were aligned using the CLUSTAL W 1.4 program (Thompson et al. 1994). Phylogenetic trees were obtained by "neighbor-joining" (PAUP program, Swofford 2000).
Light microscopy - Samples of the VD were fixed in 2.5% glutaraldehyde, 0.1 M sodium cacodylate, pH 7.2, for 12 h at room temperature. Tissues were dehydrated in acetone, embedded in Epon and sections of 0.6 µm were cut with a diamond knife. Samples were stained by to-luidine blue and examined under a Zeiss light microscope.
Measurement of luminal gut pH - Gut pH was measured in vivo by allowing adult mosquitoes to ingest pH indicator dye dissolved in 10% sucrose solution, pH 6.5 (adjusted with 0.2 M NaOH) and observing the resulting color changes in their diverticulum. The indicator dye used was 0.5% bromocresol purple [transition interval: pH 6.8 (purple) to pH 5.2 (yellow)]. Mosquitoes were dissected immediately after ingestion of indicator and 12 and 24 h later in order to observe the color changing in VD content. As the pH of the initial solution was 6.5 (dark red), we would expect the color to change gradually to yellow at lower pH.
Medium acidification by Serratia sp. - A pH assay in test tubes and Petri dishes was also carried out in order to confirm that Serratia sp. isolated from Ae. aegypti VD had the ability to acidify the culture medium. Cultures of Serratia sp. were transferred to test tubes containing 2 ml of BHI with 2.0% glucose and 0.5% bromocresol purple (pH 6.5). Tubes were incubated at 28ºC and observed after 12 and 24 h to see the color changes. The same test was done in Petri dishes containing BHI agar with 2.0% glucose and 0.5% bromocresol purple (pH 6.5).
RESULTS
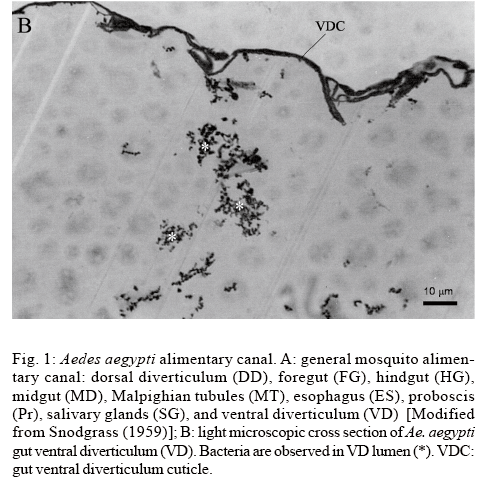
We successfully isolated microorganisms from all tested samples. In the preliminary analyses, we observed the existence of Gram-positive diplococci and Gram positive and Gram-negative rods in the VD lumen of Ae. aegypti. A diagrammatic drawing of the mosquito alimentary canal showing the position of the VD and midgut is seen in Fig. 1A. In Fig. 1B, microorganisms are observed within the lumen of the VD.
A total of 21 isolates of microorganisms were obtained from VD cultures from all the seven assays. The Gram-negative and Gram-positive rod-shape bacteria corresponded to nine isolates (42.8%) and five isolates (23.8%), respectively. The cocci corresponded to five isolates (23.8%), and yeasts to two isolates (9.5%). DNA extraction from isolates allowed us to identify Serratia sp., Bacillus sp., Bacillus subtilis and Pichia caribbica. We were also able to obtain nine yeast clones belonging to the Saccharomicetaceae family, and two were identified as P. caribbica. Four bacterial clones of the family Enterobacteriaceae identified as Serratia sp. were also obtained from the DNA extracted directly from the VD (Table).
The 16S rDNA sequences obtained from bacterial isolates and also from the extraction of the VD, allowed us to identify Serratia sp. as the predominant bacteria in Ae. aegypti VD (6 out of 7 isolations). Gene sequence similarity studies are shown in Table. The sequences obtained in this work were deposited in GenBank database (accession numbers are shown in Table).
The use of pH indicators allowed the direct and non-invasive measurement of the mosquito alimentary canal. The experiments revealed that the pH of the VD content decreased after 24 h of ingestion of the indicator. As seen in Fig. 2A-C, the VD content of the insect dissected right after feeding had a pH of 6.5 (dark red). The color of the VD content changed to bright red after 12 h and turned to yellow in 24 h (pH < 5.2).
The test tubes assays indicated that Serratia sp. isolated from Ae. aegypti VD had the ability to acidify the culture medium. After 24 h of incubation the pH changed from 6.5 (control tube) to 5.0 (Fig. 2E). Serratia sp. growing on BHI agar also acidified the medium, as observed by the yellow spots surrounding bacterial streaks (Fig. 2D).
DISCUSSION
Some works have already reported the presence of bacteria in the midgut of mosquitoes (DeMaio et al. 1996, Pumpuni et al. 1996, Luxananil et al. 2001, Zayed & Bream 2004), but have not described any role for these microorganisms in the insect physiology. We have observed, under scanning electron microscopy, a dramatically high number of small rodlike bacteria embedded in the food bolus 48 h after blood ingestion (DS Gusmão 2006, unpublished observations). DeMaio et al. (1996), Pumpuni et al. (1996) and Zayed and Bream (2004) also observed the increase in bacterial density in mosquito midgut after blood feeding. DeMaio et al. (1996) and Pumpuni et al. (1996) reported a frequent presence of Gram-negative rods belonging to Enterobacteriaceae in adults of Culex as well as Anopheles species. In laboratory-reared and wild-caught sandflies all predominant species identified by Dillon et al. (1996) belonged to Enterobacteriaceae. The genus Serratia has a wide host range and has been isolated from various insect groups (Krieg 1987). Serratia sp. is member of the Enterobacteriaceae family (facultative anaerobic, Gram-negative, cytochrome oxidase negative and catalase positive). In general, Serratia marcescens is reported most frequently as a pathogen of insectary-reared insects (Krieg 1987, Grimont & Grimont 1992). S. marcescens was identified in intestinal tube of Lutzomyia longipalpis, and was more abundant in sandflies fed on blood and sugar than the ones fed only on blood (Oliveira et al. 2001). Iverson et al. (1984) reported the presence of Serratia liquefaciens and S. marcescens in all sugar beet root maggot (Diptera: Oititidae) developmental stages, suggesting a symbiosis and a nutritional interdependence between these bacteria and the insect. Gonzalez-Ceron et al. (2003) studied the microorganisms present in the midguts of An. albimanus and identified several Serratia and Enterobacter species including S. marcescens, E. cloacae, and E. amnigenus. In Ae. triseriatus, Culex pipiens and Psorophora columbiae mosquitoes, S. marcescens was within the species most frequently isolated (DeMaio et al. 1996). S. marcescens with ability to lyse erythrocytes was also isolated from the gut larvae of the blood feeding insect R. prolixus (Azambuja et al. 2004).
Moll et al. (2001) describe an effective mechanism to eliminate gut microorganisms during mosquito metamorphosis and adult emergence. Bacteria found in recently emerged non-fed adults are present from the larval and pupal stage and, therefore, have some adaptations to overcome this mechanism.
According to Solé et al. (1997, 2000), the addition of glucose or other sugar to cell suspensions of S. marcescens resulted in the acidification of the medium. Their results suggest that the acidification of the medium could be due to the presence of organic acids resulting from bacterial metabolism. These results support our findings, indicating that the bacteria present in the VD may have a role in the sugar metabolism process. The acid pH we observed in Ae. aegypti VD may be a result of the sucrose oxidation by Serratia spp., as seen on the test tubes and Petri dishes assays (Fig. 2D, E).
Recently, Favia et al. (2007) reported that the main bacteria species associated with adults and larvae of the mosquito Anopheles stephensi are from the genus Asaia, that is related to acetic acid bacteria. They found Asaia sp. in various An. stephensi organs and stages but did not specify if they found it associated with the VD. We suppose that Asaia may also be present in An. stephensi VD once adult mosquitoes, as hypothesized by the authors, could obtain these bacteria from the nectar of tropical flowers, which are Asaia's natural habitat. Therefore, Serratia and probably Asaia could be contributing to acidify the VD lumen when metabolizing its sugar content.
There are no reports about the VD pH in Ae. aegypti to date. Gontijo et al. (1998) measured the pH in Lu. longipalpis esophageal diverticulum (crop) with bromocresol purple, the same indicator dye used in our study. They obtained a pH between 6.0 and 6.5, higher than our results.
Our findings may open new windows for understanding Aedes-bacteria interaction. As the VD has no direct connection with the midgut (Fig. 2B), the presence of bacteria inside this compartment may imply in a more specific insect-microorganism interaction than previously described. Unlike other insects, Ae. aegypti may not have a special structure for harboring friendly bacteria. Thus, the VD would provide a good environment for these bacteria that would be protected from the direct flux of food (blood) in the gut. Bacteria would remain in small numbers in the VD and would be released to the gut when convenient. As Enterobacteriaceae have the ability to ferment sugar (Ewing 1986), we could also hypothesize that one possible role for Serratia sp. would be to metabolize the sugar contained in the diverticulum, as seen in our in vitro assays, releasing important molecules to adult physiology. Once Serratia sp. was found in 85% of our assays and the bacteria embedded in the food bolus also seemed to be Serratia, we could also hypothesize that there is a more complex bacterial interaction during the digestion process, than simply a fortuitous "passing through" the alimentary canal.
ACKNOWLEDGEMENTS
To Center for the Study of Social Insects, UNESP, Rio Claro, São Paulo, Brasil, for performing the sequence reactions.
Received 12 June 2007
Accepted 23 November 2007
Financial support: CNPq, FAPERJ.
- Aksoy S, Chen X, Hypsa V 1997. Phylogeny and potential transmission routes of midgut-associated endosymbionts of tsetse (Diptera: Glossinidae). Insect Mol Biol 6: 83-190.
- Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K 1998. Current protocols in molecular biology, Vol. 1, John Wiley & Sons, New York.
- Azambuja P, Feder D, Garcia ES 2004. Isolation of Serratia marcescens in the midgut of Rhodnius prolixus: impact on the establishmet of the parasite Trypanosoma cruzi in the vector. Exp Parasitol 107: 89-96.
- Azambuja P, Garcia ES, Ratcliffe NA 2005. Gut microbiota and parasite transmision by insect vectors. Trends Parasitol 21: 568-572.
- Buchner P 1965. Endosymbionts of animals with plant microorganisms John Wiley & Sons, New York, 909 pp.
- Chen X, Li S, Aksoy S 1999. Concordant evolution of a symbiont with its host insect species: molecular phylogeny of genus Glossina and its bacteriome-associated endosymbiont, Wigglesworthia glossinidia J Mol Evol 48: 49-58.
- Consoli RAGB, Lourenço-de-Oliveira R 1994. Principais Mosquitos de Importância Sanitária no Brasil, Fiocruz, Rio de Janeiro, 228 pp.
- Dapples CG, Lea AO 1974. Inner surface morphology of the alimentary canal in Aedes aegypti (L.) (Diptera: Culicidae). Int J Insect Morphol Embryol 3: 433-442.
- Dasch GA, Weiss E, Chang K 1984. Endosymbionts of insects. In NR Krieg, JG Holt, Bergey's Manual of Systematic Bacteriology, Vol.1, Williams and Wilkins, Baltimore, MD, p. 811-833.
- Delong EF 1992. Archea in coastal marine environments. Proc Natl Acad Sci USA 89: 5685-5689.
- DeMaio J, Pumpuni CB, Kent M, Beier JC 1996. The midgut bacterial flora of wild Aedes triseriatus, Culex pipiens and Psorophora columbiae mosquitoes. Am J Trop Med Hyg 54: 219-223.
- Dillon RJ, El Kordy E, Shehata M, Lane RP 1996. The prevalence of a microbiota in the digestive tract of Phlebotomus papatasi Ann Trop Med Parasitol 90: 699-673.
- Douglas AE 1989. Mycetocyte symbiosis in insects. Biol Rev Camb Philos Soc 64: 409-434.
- Ewing WH 1986. Edwards and Ewing's Identification of Enterobacteriaceae, 4th ed., Elsevier, New York, 536 pp.
- Favia G, Ricci I, Damiani C, Raddadi N, Crotti E, Marzorati M, Rizzi A, Urso R, Brusetti L, Borin S, Mora D, Scuppa P, Pasqualini L, Clementi E, Genchi M, Corona S, Negri I, Grandi G, Alma A, Kramer L, Esposito F, Bandi C, Sacchi L, Daffonchio D 2007. Bacteria of the genus Asaia stably associate with Anopheles stephensi, an Asian malarial mosquito vector. Proc Natl Acad Sci USA 104: 9047-9051.
- Gontijo NF, Almeida-Silva S, Costa FF, Mares-Guia ML, Williams P, Melo MN 1998. Lutzomyia longipalpis: pH in the gut, digestive glycosidases, and some speculations upon Leishmania development. Exp Parasitol 90: 212-219.
- Gonzalez-Ceron L, Santillan F, Rodriguez MH, Mendez D, Hernandez-Avila JE 2003. Bacteria in midguts of field-collected Anopheles albimanus block Plasmodium vivax sporogonic development. J Med Entomol 40: 371-374.
- Grimont F, Grimont PAD 1992. The genus Serratia In A Balows, HG Trüper, M Dworkin, W Harder, KH Schleifer (eds), The Prokaryotes. A Handbook on the Biology of Bacteria: Ecophysiology. Isolations, Identification, Applications, Vol. 1, Springer, New York, NY, USA, p. 2822-2248.
- Iverson KAL, Bromel MC, Anderson AW, Freeman TP 1984. Bacterial symbionts in the sugar beet root maggot, Tetanops myopaeformis (von Roder). Appl Environ Microbiol 47: 22-27.
- Khampang P, Chungjatupornchai W, Luxananil P, Panyim S 1999. Efficient expression of mosquito-larvicidal proteins in a Gram-negative bacterium capable of recolonization in the guts of Anopheles dirus larva. Appl Microbiol Biotechnol 51: 79-84.
- Krieg A 1987. Diseases caused by bacteria and other prokaryotes. In JR Fuxa, Y Tanada, Epizootiology of insect diseases ,John Wiley & Sons, New York, p. 323-355.
- Lane DJ, Pace B, Olsen GJ, Stahl DA, Sogin ML, Pace NR 1985. Rapid determination of 16S ribosomal sequences for phylogenetic analyses. Proc Natl Acad Sci USA 82: 6955-6959.
- Lindh JM, Terenius O, Faye I 2005. 16S rRNA gene-based identification of midgut bacteria from field-caught Anopheles gambiae sensu lato and A. funestus mosquitoes reveals new species related to known insect symbionts. Appl Environ Microbiol 71: 7217-7223.
- Luxananil P, Atomi H, Panyim S, Imanaka T 2001. Isolation of bacterial strains colonizable in mosquito larval guts as novel host cells for mosquito control. J Biosci Bioeng 92: 342-345.
- Moll RM, Romoser WS, Modrzakowski MC, Moncayo AC, Lerdthusnee K 2001. Meconial peritrophic membranes and the fate of midgut bacteria during mosquito (Diptera: Culicidae) metamorphosis. J Med Entomol 38: 29-32.
- Nasci RS, Miller BR 1996. Culicine mosquitoes and the agents they transmit. In BJ Beaty, WC Marquardt (eds), The Biology of Disease Vectors, 1st ed., Colorado University Press, Niwot, CO, p. 85-97.
- O'Donnell K 1993. Fusarium and its near relatives. In DR Reynolds, JW Taylor (eds), The fungal holomorph: Mitotic and pleomorphic speciation in fungal systematics, CAB International, Wallingford, p. 225-233.
- Oliveira SMP, Morais BA, Gonçalves CA, Giordano-Dias CM, Vilela ML, Brazil RP, Almeida, JM, Asensi MD, Mello RP 2001. Microbiota do trato digestivo de fêmeas de Lutzomyia longipalpis (Lutz & Neiva, 1912) (Diptera: Psychodidae) provenientes de colônias alimentadas com sangue e com sangue e sacarose. Cad Saude Publica 17: 229-232.
- Pidiyar VJ, Jangid K, Patole MS, Shouche Y 2004. Studies on cultured and uncultured microbiota of wild Culex quinquefasciatus mosquito midgut based on 16S ribosomal RNA gene analysis. Am J Trop Med Hyg 70: 597-603.
- Pidiyar VJ, Kaznowski A, Badri NN, Patole MS, Shouche YS 2002. Aeromonas culicicola sp. nov., from the midgut of Culex quinquefasciatus Int J Syst Evol Microbiol 52: 1723-1728.
- Pumpuni CB, Demaio J, Kent M, Davis JR, Beier JC 1996. Bacterial population dynamics in three anopheline species: the impact on Plasmodium sporogonic development. Am J Trop Med Hyg 54: 214-218.
- Riehle MA, Jacobs-Lorena M 2005. Using bacteria to express and display anti-parasite molecules in mosquitoes: current and future strategies. Insect Biochem Mol Biol 35: 699-707.
- Romoser WS 1996. The vector alimentary system. In BJ Beaty, WC Marquardt (eds), The Biology of Disease Vectors, Colorado University Press, Niwot, CO, p. 298-317.
- Snodgrass RE 1959. The anatomical life of the mosquito. Smithsonian Miscellaneous Collections 139: 1-87.
- Solé M, Francia A, Rius N, Loren JG 1997. The role of pH in the "glucose effect" on prodigiosin production by non-proliferating cells of Serratia marcescens Lett Appl Microbiol 25: 81-84.
- Solé M, Rius N, Loren JG 2000. Rapid extracellular acidification induced by glucose metabolism in non-proliferating cells of Serratia marcescens Internatl Microbiol 3: 39-43.
- Sperança MA, Capurro ML 2007. Perspectives in the control of infectious diseases by transgenic mosquitoes in the post-genomic era - A Review. Mem Inst Oswaldo Cruz 102: 425-433.
- Swofford DL 2000. PAUP: Phylogenetic Analysis Using Parsimony, version 4.0b4a. Smithsonian Institution and Sinauer Associates, Sunderland, MA.
- Thompson JD, Higgins DG, Gibson TJ 1994. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22: 388-392.
- Thompson MT 1905. Alimentary canal of the mosquito. Proc Boston Soc Nat Hist 32: 145-202.
- Vaughan-Martini A, Kurtzman CP, Meyer SA, O'Neill EB 2005. Two new species in the Pichia guilliermondii clade: Pichia caribbica sp. nov. the ascosporic state of Candida fermentati, and Candida carpophila comb. nov. FEMS Yeast Res 5: 463-469.
- Zayed ME, Bream AS 2004. Biodiversity of the microbial flora associated with two strains of Culex pipiens (Diptera: Culicidae). Commun Agric Appl Biol Sci 69: 229-34.
Publication Dates
-
Publication in this collection
18 Jan 2008 -
Date of issue
Dec 2007
History
-
Received
12 June 2007 -
Accepted
23 Nov 2007