Abstract
For many years, prokaryotic cells were distinguished from eukaryotic cells based on the simplicity of their cytoplasm, in which the presence of organelles and cytoskeletal structures had not been discovered. Based on current knowledge, this review describes the complex components of the prokaryotic cell cytoskeleton, including (i) tubulin homologues composed of FtsZ, BtuA, BtuB and several associated proteins, which play a fundamental role in cell division, (ii) actin-like homologues, such as MreB and Mb1, which are involved in controlling cell width and cell length, and (iii) intermediate filament homologues, including crescentin and CfpA, which localise on the concave side of a bacterium and along its inner curvature and associate with its membrane. Some prokaryotes exhibit specialised membrane-bound organelles in the cytoplasm, such as magnetosomes and acidocalcisomes, as well as protein complexes, such as carboxysomes. This review also examines recent data on the presence of nanotubes, which are structures that are well characterised in mammalian cells that allow direct contact and communication between cells.
prokaryotic cells; eukaryotic cells; prokaryotic cytoskeleton homologues; carboxysomes; acidocalcisomes; nanotubes
REVIEW
Prokaryotic cells: structural organisation of the cytoskeleton and organelles
Wanderley de SouzaI, II, III, + + Corresponding author: wsouza@biol.ufrj.br
ILaboratório de Ultraestrutura Celular Hertha Meyer, Instituto de Biofísica Carlos Chagas Filho, Centro de Ciências da Saúde, Universidade Federal do Rio de Janeiro, Av. Brigadeiro Trompowsky s/n Bloco G, 21941-900 Rio de Janeiro, RJ, Brasil
IIDiretoria de Programas, Instituto Nacional de Ciência e Tecnologia de Biologia Estrutural e Bioimagem, Rio de Janeiro, RJ, Brasil
IIIInstituto Nacional de Metrologia, Qualidade e Tecnologia, Rio de Janeiro, RJ, Brasil
ABSTRACT
For many years, prokaryotic cells were distinguished from eukaryotic cells based on the simplicity of their cytoplasm, in which the presence of organelles and cytoskeletal structures had not been discovered. Based on current knowledge, this review describes the complex components of the prokaryotic cell cytoskeleton, including (i) tubulin homologues composed of FtsZ, BtuA, BtuB and several associated proteins, which play a fundamental role in cell division, (ii) actin-like homologues, such as MreB and Mb1, which are involved in controlling cell width and cell length, and (iii) intermediate filament homologues, including crescentin and CfpA, which localise on the concave side of a bacterium and along its inner curvature and associate with its membrane. Some prokaryotes exhibit specialised membrane-bound organelles in the cytoplasm, such as magnetosomes and acidocalcisomes, as well as protein complexes, such as carboxysomes. This review also examines recent data on the presence of nanotubes, which are structures that are well characterised in mammalian cells that allow direct contact and communication between cells.
Key words: prokaryotic cells - eukaryotic cells - prokaryotic cytoskeleton homologues - carboxysomes - acidocalcisomes - nanotubes
Early morphological studies using light microscopy and even the initial observations made with transmission electron microscopes indicated major differences in the organisation of prokaryotic and eukaryotic cells. While eukaryotic cells were characterised as exhibiting internal organisation associated with various membrane-bound organelles, bacteria displayed a homogeneous internal organisation. Although this idea predominated for many years, detailed analysis of the structural organisation of prokaryotic cells, especially using modern structural approaches in association with genomic and proteomic analyses, has indicated that this view is incorrect. Indeed, the available information now points to the existence of an assemblage of complex proteinaceous structures localised in well-defined regions of the prokaryotic cell, as well as the presence of several membrane-bound organelles, thus indicating that there are subcompartments in these cells. The existence of a complex cytoskeleton formed by proteins resembling those involved in the assembly of microtubules, microfilaments and intermediate filaments further indicates that some structural features that are usually associated with fundamental physiological processes, such as cell shape maintenance, cell division and cell locomotion, have been maintained throughout the evolutionary process. In addition, it has been shown that complex processes, such as protein glycosylation (Benz & Schmidt 2002) and cell-cell adhesion, that were previously recognised only in eukaryotic cells are also common in bacteria that are committed to each other and develop into multicellular structures (Keim et al. 2004). These points will be discussed in greater detail later. In this brief review, I will discuss only the structures and organelles that have been better characterised or identified in prokaryotic cells over the last 10 years.
Methodological considerations - A detailed overview of all the methodologies that have played an important role in the identification and characterisation of the structures mentioned herein is beyond the scope of this review. However, for the sake of clarity, I will briefly mention a few approaches that have been highly relevant in this field.
First, I will comment on the advances in light and electron microscopy. In the case of light microscopy, it is important to mention the development of techniques that allow the insertion of portions of genes encoding fluorescent proteins into the gene encoding a specific protein, allowing its precise localisation using fluorescence microscopy. This approach has become more powerful with the recent development of high-resolution fluorescence microscopes. A resolution as high as 20 nm is achievable via stimulated emission depletion microscopy. For instance, using this approach, new information was obtained regarding the localisation of the tubulin-like protein FtsZ in Bacillus subtilis (Jennings et al. 2011). With respect to transmission electron microscopy, it is well known that "fragile structures", especially those formed through the assembly of protein subunits, may disassemble or even be fragmented during conventional processing of biological samples for this technique. The best way to preserve these structures is through the use of physical fixation, especially via high pressure freezing with liquid nitrogen under conditions in which ice crystals are not formed (review in Heuser 2011). Following this initial fixation step, the samples can be processed for plastic embedding according to the freeze-substitution method in which the temperature is gradually increased from -196ºC to -20ºC, then infiltrated with resins, such as Unicryl, Lowicryl or LR Gold, that are then polymerised at the same temperature. Subsequently, thin sections are obtained through conventional procedures and analysed using an electron microscope (review in Hurbain & Sachse 2011). Another approach is to obtain thin frozen sections following quick freezing, which are then cryo-transferred to an electron microscope and observed while still in a frozen state. Alternatively, if samples are sufficiently thin, it is possible to freeze an entire bacterium and observe it at low temperatures with an electron microscope. More information can always be obtained if images from a sample are acquired at different angles (for instance, from -60º to +60º, at intervals of 1º). Using this approach, known as electron tomography, tomograms are acquired and a 3D reconstruction of a structure can be obtained, offering novel views of biological structures (review in Hoenger & Bouchet-Marquis 2011).
Second, it is important to mention the advances in sequencing techniques that allow complete sequencing of the genome of a bacterium in a few hours. Additionally, through proteomic analysis, the complete set up of a protein can be obtained. With the use of bioinformatics tools, it is then possible to apply genomic and proteomic data in comparisons with data previously deposited in gene and protein databanks and check for potential homologies between genes and proteins identified in bacteria with those previously characterised in other cell types (reviews in Hu et al. 2011, Sabidó et al. 2011, Terfve & Saez-Rodriguez 2012).
The prokaryote cytoskeleton - Cytoskeletal proteins exist in all eukaryotic cells and perform functions varying from maintaining cell shape to participation in intracellular movements, cell division and locomotion of the cell, among others. Cytoskeletal structures are generally formed by a major protein that is able to interact with other proteins, commonly designated cytoskeleton-associated proteins. This binding to cytoskeleton-associated proteins permits functions such as interaction with other molecules, control of stability and induction of cycles of polymerisation and depolymerisation. Three major classes of cytoskeletal structures are generally considered: microtubules, microfilaments and intermediate filaments. This area of inquiry has recently been the subject of intense investigation. Here, I will note the most relevant data from a cell biology perspective in prokaryotic cells.
Tubulin-like homologues - The processes involved in the division of bacteria, especially Escherichia coli, have been the subject of intense investigation for many years. Observations using light microscopy and electron microscopy have established that during division, a bacterial cell increases in size, assumes a long rod-like shape and the inner membrane then invaginates. Inner membrane invagination is followed by invagination of the outer membrane and the cell wall, forming a septum that divides the cell in half. Genetic studies have indicated the existence of genes controlling the septation process. These genes were further analysed using temperature-sensitive mutants. Under certain temperatures, septa were not formed, generating filamentous cells. These genes were designated fts genes, for "filamentous temperature-sensitive mutants". One of these proteins, known as FtsZ, was further characterised using an immunocytochemical approach in association with transmission electron microscopy. It was then shown that gold particles indicating the presence of FtsZ displayed a diffuse distribution in most bacteria, but concentrated in the form of a ring localised mainly in the centre of cells that were about to divide. As division progressed, the ring was observed to constrict and labelling concentrated at the tip of the invaginating membrane (Fig. 1, diagrammatical view). This structure was designated the Z-ring (Bi & Lutkenhaus 1991). Subsequently, these studies were extended using immunofluorescence microscopy, in which the resolution is lower, but examination of a large number of cells is possible. It was then shown that the FtsZ-ring was assembled in all cells very early in the cell cycle and that at non-permissive temperatures, at which long filamentous bacteria are formed, multiple Z-rings were localised at precise intervals along the cell (Fig. 2) (Levin & Losick 1996). Another significant advance in this field has come from studies using an FtsZ-GFP chimera along with deconvolution during acquisition of immunofluorescence images, thus allowing more precise localisation of the protein (Ma et al. 1996). Using this approach, the ring structure was clearly observable in a three-dimensional view and it was shown that the Z-ring is assembled very early in the bacterial life cycle. Small differences in the localisation of the Z-ring in different species were noted. For instance, in E. coli, the ring was not observed at the end of cells, whereas in B. subtilis, small spots remained at separation sites after a new Z-ring had formed (Levin & Losick 1996).
After these initial studies, several groups sought to detect FtsZ in different species. It is now known that FtsZ is found in both gram-positive and gram-negative bacteria, as well as in some species of archaea. Genomic studies have shown that two Chlamydia species do not express FtsZ (Stephens et al. 1998, Kalman et al. 1999). Both of these species are obligate parasites that live in membrane-bound cytoplasmic vacuoles. It has been suggested that they use the host cell machinery to carry out their division. Two free-living prokaryote species have also been found not to express FtsZ. One is Aeropyrum pernix, an archaeon that lives at 90ºC in oceanic thermal vents (Sako et al. 1996) and the other is Ureaplasma uralyticum, a mycoplasma that lives in a host, but can be cultivated in axenic medium.
An ftsZ gene has been detected in the plant species Arabidopsis thaliana. Although this gene is encoded in the nuclear genome, the protein is synthesised in the cytoplasm and transported to the chloroplast due to the presence of a signal sequence (Osteryoung & Vierling 1995). Subsequently, the same group demonstrated the existence of two genes for this protein in Arabidopsis: FtsZ1 is transported to the chloroplast, while FtsZ2 remains on the cytoplasmic side of the chloroplast (Osteryoung et al. 1998). Antisense experiments showed that both genes are essential for chloroplast division. FtsZ has not been found in the mitochondria of most species examined to date, though this organelle has a prokaryotic origin. This protein was likely replaced with dynamin in mitochondria (review in Erickson 2000). However, FtsZ has been detected in the golden-brown alga Mallomonas splendens, localised in patches on the mitochondrial membrane (Beech et al. 2000).
Initial sequential analysis showed that FtsZ contains a short segment of amino acids that is virtually identical to the tubulin signature motif found in all alpha, beta and gamma tubulins (de Boer et al. 1992, Raychaudhuri & Park 1992, Mukherjee et al. 1993). Further analysis identified a dozen additional completely conserved residues. Identification of the molecular structure of the FtsZ protein showed that the folds of tubulin and FtsZ are identical, except for small segments at the amino and carboxyl termini (Lowe & Amos 1998, Nogales et al. 1998). Subsequently, it was shown that in vitro FtsZ assembles into protofilaments and rods (Bramhill & Thompson 1994, Mukherjee & Lutkenhaus 1994). The protofilaments consist of a head-to-tail linear polymer of FtsZ (Mukherjee & Lutkenhaus 1994, Erickson et al. 1996, Lowe & Amos 1999, Oliva et al. 2004, review in Margolin 2005). Structural analysis of FtsZ polymers shows that they are similar to tubulin polymers (Erickson et al. 1996). However, unlike microtubules, FstZ protofilaments do not assemble into higher-order tubules (Fig. 3). Additionally, FstZ protofilaments do not exhibit an FstZ-GTP cap. The structure of filaments formed from the tubulin/FtsZ-like protein TubZ of Bacillus thuringiensis has been determined by combining crystallography and electron microscopy analyses. This combined analysis showed a double helical superstructure, which is unusual for tubulin (Aylett et al. 2010).
How FtsZ functions remains unclear - It has been suggested that FtsZ participates in a minimal type of division machinery containing approximately 15,000 molecules of the protein per cell (Dai & Luthkenhaus 1992) that form protofilaments encircling the bacterium approximately 30 times. The protofilament would be attached to the membrane either via proteins or through direct association with lipids. Two new proteins designated FtsK and ZipA could participate in this process. ZipA is an integral membrane protein that is essential for E. coli division (Hale & de Boer 1997). It contains an amino-terminal transmembrane anchor and a cytoplasmic domain consisting of three parts: a 60 amino acid charged domain, a 100 amino acid domain rich in proline and asparagine and a 144 amino acid globular carboxy-terminal domain. FtsZ can assemble into the Z-ring without ZipA, but recruitment of ZipA to the Z-ring requires FtsZ (Hale & de Boer 1999, Liu et al. 1999). Some constriction may occur, most likely due to conformational changes taking place in the protofilament, which may exhibit a straight conformation once it is attached to the membrane and then adopt a curved conformation upon dissociation from the membrane (Erickson 1997). The FtsZ ring consists of many bundles of protofilaments that continuously extend and shrink due to the addition of GTP-FtsZ and the release of GDP-FtsZ (review in Graumann 2004). In the presence of GTP, protofilaments are usually straightened, whereas in the presence of GDP, protofilaments favour a curved conformation.
It is important to note that FtsZ serves as a target for several auxiliary proteins. For instance, FtsA and ZipA appear to stabilise the Z-ring and ZipA is also involved in the crosslinking of FtsZ protofilaments, whereas SulA, EzrA and MinC are inhibitors of FstZ polymerisation. All of these proteins interact at the level of division site placement. They appear to be recruited in a linear order, with FtsA and ZipA being required for all the other proteins to be recruited (reviews in Margolin 2005, Moller-Jensen & Lowe 2005).
In addition to FtsZ, other tubulin homologues have been found in some prokaryotes. BtubA and BtubB were found in Prosthecobacter dejongeii (Schlieper et al. 2005, review in Shih & Rothfield 2006, Lowe & Amos 2009). The sequence homology between the bacterial proteins and tubulin varies considerably: the homology is 17% for FstZ and 35% for BtubA/B relative to tubulin (Schlieper et al. 2005, review in Graumann 2007). However, sequencing studies have shown that FstZ and GtubA/B do not correspond to either alpha or beta tubulin, but contain mosaic sequences with intertwining features from both types of tubulin proteins (Martin-Galiano et al. 2011). Isolated BtubA and BtuB proteins co-assemble in a 1:1 ratio into protofilament polymers that hydrolyse GTP (Schlieper et al. 2005). The genes encoding these proteins were most likely transferred from a eukaryotic cell via horizontal gene transfer, diverging shortly after tubulin gene duplication (Schlieper et al. 2005, Martin-Galiano et al. 2011).
Because FstZ is involved in cell division, it has become a target for the development of new drugs able to inhibit prokaryotic proliferation. Special emphasis has been placed on the development of drugs that do not interfere with eukaryotic tubulin. Indeed, it has been shown that viriditoxin, a natural compound isolated from Aspergilus viridinutans, inhibits FstZ polymerisation at concentrations in the µg/mL range (Wang et al. 2003). Several other drugs, such as SRI-3072, a taxane derivative, sanguinarine, a 3-(2-indolyl) piperidine derivative, and a carboxybiphenyllindole derivative, have been shown to interact with FstZ (review in Volmer 2006).
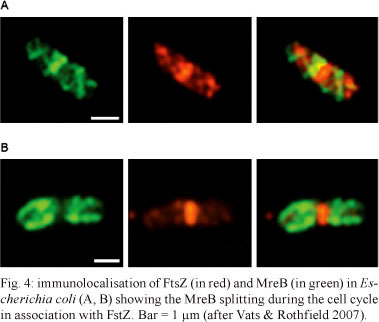
Actin-like homologues - Since the late 1980s, it has been known that the gene mreB (from murein cluster e) is involved in establishing and maintaining the shape of prokaryotic cells (reviewed in Graumann 2004). A lack of mreB in E. coli causes cell death. Other MreB-like proteins have been identified in B. subtilis, namely Mb1 and MreBH. Depletion of these proteins leads to bending and distortion of the cell's shape, leading to the appearance of banana-shaped cells (Abhayawardhane & Stewart 1995, Jones et al. 2001, Souto & Graumann 2003). Genes encoding both proteins have been found in almost all rod-shaped species and are absent from species that grow as cocci (Daniel & Errington 2003). It has been shown that MreB exists during most of the cell cycle and it splits into two equivalent structures (Fig. 4) in a process that is triggered via membrane association with the tubulin-like protein FstZ (Vats & Rothfield 2007). Further studies have shown that these proteins are involved in the maintenance of cell shape and viability. Immunofluorescence microscopy of cells transfected with GFP-tagged MreB and Mb1 showed that they form filamentous structures in a helically arranged pattern localised below the bacterial membrane. Mb1 filaments extend from pole to pole, while MreB filaments were not detected at the poles (Jones et al. 2001). These observations led to the suggestion that MreB is involved in the control of cell width, while Mb1 controls cell length.
A breakthrough in this area of inquiry came from work showing that MreB is a member of the actin family and that it forms actin-like double filaments that run in parallel and with the same linear orientation, in contrast to what is observed for actin, which adopts a helical organisation (Van den Ent et al. 2001). Furthermore, unlike actin, which favours ATP over GTP, MreE can use both ATP and GTP equally well for the formation of filaments. MreE filaments are more rigid than actin filaments and they tend to assemble into bundles. In contrast, actin filaments are assembled as single filaments (Esue et al. 2006). The spacing between MreB proteins along protofilaments is 5.1 Å, while the spacing between subunits of eukaryotic actin is 55 Å (Fig. 5). These proteins change their pattern of localisation according to the cell cycle stage (Carballido-Lopez & Errington 2003, Figge et al. 2004). Bundles of filaments move continuously and in parallel along helical tracks (Defeu-Soufo & Graumann 2004). It has also been suggested that these proteins are involved in the organisation of the cell wall, leading to the cells acquiring their typical rod-like shape (Daniel & Errington 2003). Other data also demonstrate the involvement of actin-like proteins in the segregation of plasmids and whole chromosomes (Van den Ent et al. 2002).
ParM belongs to another class of actin homologue that differs with respect to its sequence and polymerisation dynamics from the MreB class. While the MreB class is always encoded in the bacterial chromosome and regulates a wide array of cellular functions, the ParM class always interacts with extrachromosomal plasmids and appears to be dedicated to the segregation of plasmids, functioning as a motor. As filaments grow, they push plasmids to the cell poles prior to cell division (Moller-Jensen et al. 2003, review in Gitai 2005). Two other Par proteins, ParA and ParB, have been characterised in Caulobacter crescentus. Par A forms linear polymers in vitro. Par B is a centromere-binding protein that binds to and destabilises the Par A structure, causing centromere movement to opposite poles (Ptacin et al. 2010). This observation suggests that prokaryotic cells use mechanisms resembling well-characterised mechanisms involved in the mitotic machinery operating in eukaryotic cells.
No other cytoskeletal motor proteins, such as dynein, kinesin or myosin, have been identified in prokaryotes thus far.
Intermediate-like filament protein homologues - Studies using the bacterium C. crescentus, which presents a vibrio shape with a crescent form, show that this shape is due to the presence of a protein known as CreS-protein, or crescentin. The corresponding gene was identified and it was found that when it was deleted, cells still grew but lost their typical shape, resulting in straight cells (Ausmees et al. 2003). Immunofluorescence microscopy showed that the protein localises in a filamentous form on the concave side of cells and along the inner cell curvature over the entire length of the helical cells. The available data indicate that the CreS protein induces cell bending and creates asymmetry along the length of a cell. This protein has also been found in other species, including Helicobacter pylori. Further analysis of this protein showed that it exhibits a high content of coiled-coils and strutters, which are positions where the coiled-coils are disrupted. This organisation resembles that found in proteins such as keratin, which forms intermediate filaments. Indeed, isolated CreS protein assembles into 10 nm thick filaments in vitro without requiring cofactors or energy. It was found that crescentin shares approximately 25% sequence identity and 40% similarity with eukaryotic intermediate filament proteins (Ausmees et al. 2003, Ausmees 2006).
A second type of intermediate-like filament has been characterised in several spirochete genera, including Treponema, Spirochaeta, Leptonema and Diplocalyx. It appears as a filamentous ribbon-like structure composed of two-six filaments, which are severed during cell division such that half of them remain in each daughter cell. The width of the filaments is approximately 6 nm and the space between them is approximately 10.4 nm. Based on the thickness of these filaments, they are very different from typical intermediate filaments of mammalian cells, which exhibit a thickness of approximately 10 nm. The protein that forms these filaments is encoded by a gene known as cfpA and the corresponding protein is CfpA, which is a unique peptide of 82 kDa (Masuda & Kawata 1989, You et al. 1996, Izard et al. 1999). There is no sequence similarity between CfpA and other known proteins and it does not contain any predicted extended coiled-coil domains (Izard et al. 1999). Electron microscope tomography revealed that the filaments are part of a complex in which one set of components anchors the filaments to the membrane while another set establishes bridges between the filaments and cytoplasmic components. In addition, the filaments do not contact each other (Izard et al. 2004, Izard 2006).
Although this has not yet been studied in detail, other proteins have also been implicated in the assembly of intermediate filament-like structures presenting typical coiled-coil domains. These structures include the fibrillar structures found in Spiroplasma melliferum formed by a 55-59 kDa protein. The assembled filaments are located beneath the membrane and are arranged in parallel ribbons extending along the length of the spiral-shaped organism and their expression is dependent upon the cfpA gene (Townsend et al. 1980, Trachtenberg 1998, Izard et al. 1999, Kurner et al. 2005). Another filamentous structure is related to the protein AgIZ. The C-terminal region of this protein includes heptad repeats, which are a characteristic of the rod region of coiled-coils of proteins, such as the myosin heavy chain observed in Myxococcus xanthus (Yang et al. 2004). Expression of this protein in E. coli results in the formation of a parallel array of filaments. Finally, another type of structure has been described based on cytoskeletal-like structures that have been observed in both intact and detergent-extracted Mycoplasma pneumoniae (Regula et al. 2001, Hegermann et al. 2002).
Protein glycosylation - For many years, protein glycosylation was considered to be a characteristic feature of eukaryotic cells, which possess an elaborate system of cisternae forming the endoplasmic reticulum and Golgi complex systems. One of the first indications that protein glycosylation also takes place in prokaryotic cells came from work on Halobacterium halobium and Clostridia, in which the existence of an S-layer glycoprotein accounting for approximately 50% of the cell envelope proteins was demonstrated. Subsequently, other glycosylated proteins have been found in structures such as flagella and pili. Glycoproteins appear to be modified primarily by short-chain carbohydrate moieties (mono to trisaccharide substituents). The consensus sequence for N-glycosylation (Ser/Thr-Xaa-Asn) has been identified in Mycoplasma tuberculosis, while a sequence motif for O-glycosylation has not been identified. There are several excellent reviews on this subject (Schaffer et al. 2001, Benz & Schmidt 2002).
Protein complexes - At present there, is a great deal of evidence of the presence of protein complexes localised in the cytoplasm. These complexes usually consist of several proteins that associate with each other, forming a structure that can be recognised in thin sections using electron microscopy in many cases. If the complex is too small, it is not directly recognised, but can be isolated through differential centrifugation and then subjected to structural analysis. The biological way to form a protein complex is to combine proteins involved in a well-defined metabolic pathway. Therefore, we can consider protein complexes as defining a certain subcompartment of the cell. Because these complexes are not surrounded by a typical unit membrane, I prefer to refer to them as protein complexes rather than protein organelles, as described by others. One good example of a protein complex in a eukaryotic cell is the proteasome, which is a structure involved in the degradation of proteins.
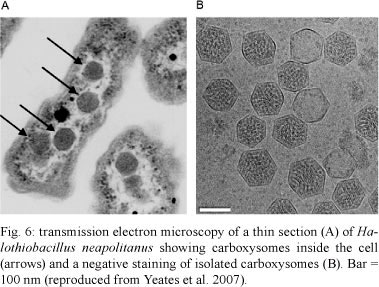
In the case of prokaryotic cells, cytoplasmic structures initially referred to as inclusion bodies or polyhedral bodies have been observed since 1956 in the cyanobacterium Phormidium uncinatum (Drews & Niklowitz 1956). Interest in this field increased when these bodies were identified and isolated from the chemoautotrophic bacterium Halobacterium neapolitanus and were shown to be composed of a surrounding shell and a matrix containing the enzyme ribulose-1-5-biphosphate carboxylase, also abbreviated as RuBisCO. They were then designated as carboxysomes (Shively et al. 1973). Carboxysomes are considered to exist in all cyanobacteria and in a limited number of sulphur and nitrifying bacteria (reviews in Bobik 2006, Yeates et al. 2007, 2008). In addition to RuBisCO, a set consisting of seven-11 other proteins is found in carboxysomes.
Carboxysomes play a central role in the final stage of the carbon-concentrating mechanism through which autotrophic microorganisms accumulate inorganic carbon to enhance CO2 fixation. The process also involves the conversion of bicarbonate to CO2 (Fig. 6).
Carboxysomes exhibit an icosahedral structure with nearly flat triangular faces (Schmid et al. 2006) and usually show a cross-sectional diameter of approximately 100-150 nm. Therefore, they are approximately 1,000 times larger than a ribosome. Their shell measures approximately 3-4 nm and is composed of several proteins designated as CsoS1. It is important to note that homologues of the carboxysome shell proteins have been reported in Salmonella (Chen et al. 1994).
Organelles - Evidence now exists supporting the presence of membrane-bound organelles in prokaryotic cells. Here, we will describe the organelles that are better characterised, such as the magnetosomes, lipid bodies and acidocalcisomes.
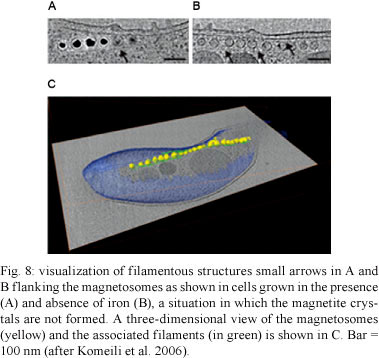
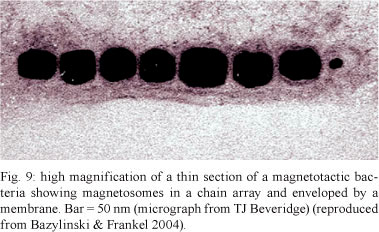
Magnetosomes are found in magnetotactic bacteria, which are mostly aquatic prokaryotes that swim along geomagnetic field lines (Blakemore 1975). These bacteria are able to synthesise a magnetic mineral crystal whose morphology is species specific. These bacteria mineralise either iron oxide magnetosomes, which contain crystals of magnetite (Fe3O4) (Frankel et al. 1979), or iron sulphide magnetosomes (Farina et al. 1990, Heywood et al. 1990, Mann et al. 1990), which contain crystals of greigite (cubic Fe3S4), mackinawite (tetragonal FeS) and cubic FeS (Posfai et al. 1998). Other iron sulphide minerals have also been identified (review by Bazylinski & Frankel 2004). Recent studies point to the existence of a novel group of sulphate-reducing bacteria that present two different magnetosome gene clusters, suggesting that one may be responsible for greigite biomineralisation and the other for magnetite (Lefévre et al. 2011). In most cases, the magnetosomes are arranged in one or more chains, which is a process in which filaments of actin-like proteins seem to be involved (Figs 7-9). Each magnetosome is 35-120 nm long and displays roughly cuboidal, elongated prismatic or tooth, bullet or arrowhead-shaped crystals. The magnetosome is surrounded by a 3-4 nm thick lipid bilayer containing phospholipids, fatty acids and some proteins (Gorby et al. 1988). A 16 kDa protein with GTPase activity was detected in the magnetosome membrane of Magnetospirillum magneticum (Okamura et al. 2001). Proteins involved in the transport of iron have also been found. Freeze-fracture studies clearly showed that the magnetosome membrane displayed intramembranous particles on both fracture faces. Indeed, a large number of proteins have been identified in the membrane (review by Bazylinski & Frankel 2004).
The biogenesis of magnetosomes seems to involve several steps including (i) magnetosome vesicle formation, most likely derived from the bacterial membrane, (ii) arrangement of the vesicles into chains, a process involving cytoskeletal components (Komeili et al. 2006), (iii) iron uptake by the bacterium, (iv) iron transport into the magnetosome and (v) controlled biomineralisation within the organelle (reviewed by Frankel & Bazylinski 2006). Genomic analysis indicates that approximately 28 conserved genes are associated with the biogenesis and function of magnetosomes. Approximately 20 proteins are also involved in these processes. The assembly of the magnetosome chain involves an acidic protein that links the magnetosomes to an actin-like cytoskeleton (reviewed by Schuller 2008).
Acidocalcisomes are acidic calcium storage organelles that have been described and characterised in several parasitic protozoa (Seufferheld et al. 2003, reviewed by Docampo et al. 2005). Based on morphological, biochemical and physiological data, it was shown that they correspond to structures known as volutin granules that have been found in several cell types, including prokaryotic cells. Acidocalcisomes were characterised in Agrobacterium tumefaciens (Fig. 10) based on the following experiments: (i) electron microscopy of thin sections showed the presence of membrane-bound organelles with an electron-dense content, (ii) X-ray microanalysis showed the presence of phosphorous, magnesium, potassium and calcium in the electron-dense content of the organelle, (iii) pyrophosphatase was localised to the organelle using immunofluorescence microscopy and (iv) it was indicated that the organelle was acidic based on staining with the LysoSensor blue DND-167 dye (Seufferheld et al. 2003).
Other organelles include (i) a large number of vesicles in photosynthetically grown cells of several purple bacteria, known as chromatophores (Cohen-Bazire & Kunisawa 1963), (ii) sac-like thylakoids in cyanobacteria (Jensen 1993), (iii) inclusions containing poly-β-hydroxybutyrate, the sulphur globules of Thiorhodaceae and the gas vacuoles of some aquatic bacteria, (iv) the pirellulosome found in Planctomycetales and (v) the anammoxosome of Candidatus species (Lindsay et al. 2001).
Additionally, many prokaryotic cells are able to accumulate large amounts of lipids into cytoplasmic lipid bodies. In some species, triacylglycerols predominate, as in the case of Mycobacterium sp. In others, such as Acinetobacter and Micrococcus, the accumulation of wax esters predominates. Polyhydroxyalkanoates are also found in Bacillus megaterium (reviewed by Waltermann & Steinbuchel 2005). Initial electron microscopy studies suggested that the lipid inclusions were surrounded by a membrane or envelope that stabilises highly hydrophobic content in the aqueous cytoplasmic environment (review by Alvarez & Steinbuchel 2002). However, it is currently thought that lipid inclusions are surrounded by a thin phospholipid monolayer boundary containing associated proteins (reviewed by Waltermann & Steinbuchel 2005, Murphy 2011).
Cytonemes and tunnelling nanotubes - While analysing the imaginal discs of Drosophila, Ramirez-Weber and Kornberg (2000) observed the presence of filopodia-like structures up to 100 µm in length rich in actin connecting different cells. They designated these structures "cytonemes" after the Latin term for "cell thread". Since then, similar structures have been observed in many cell types (reviewed by Sherer & Mothes 2008). This is clearly a very thin (approximately 300 nm in diameter), highly dynamic and fragile structure. It is more easily found in cell cultures in which the cell surface has been labelled with a fluorescent dye or stained to detect actin filaments and observed using immunofluorescence microscopy. Observation of cytonemes using electron microscopy requires careful processing to avoid rupturing their very thin and delicate structure.
Studies have investigated the functional roles played by the cytonemes/tunnelling nanotubes. It is clear that these structures establish close contact between cells and that the transport of molecules and even small organelles, such as endocytic and secretory vesicles as well as prions, takes place through these structures (reviewed by Gerdes & Carvalho 2008, Davis & Sowinski 2008, Sherer & Mothes 2008, Gousset et al. 2009). Some bacteria and viruses have been shown to use this type of structure as a type of cable following its binding to their extracellular surface (Onfelt et al. 2006, Sherer et al. 2007).
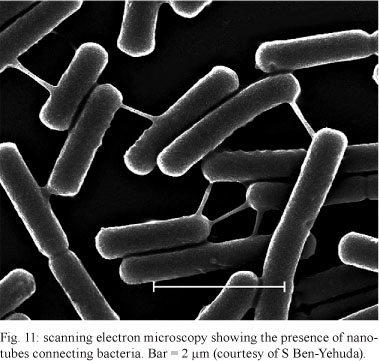
More recently, nanotube-like structures were also reported in the bacteria B. subtilis, E. coli and Staphylococcus aureus (Dubey & Ben-Yehuda 2011). These authors showed that GFP molecules are transferred from one cell to the other via nanotubes. In addition, bacteria use nanotubes for the transfer of molecules involved in resistance to antibiotics. Electron microscopy revealed the presence of nanotubes with a width varying from 30-130 nm connecting bacteria. Nanotubes can reach a length of 1 µm (Fig. 11).
Perspectives - Significant advances in the fields covered in this review are expected over the next few years, mainly due to increased use of high-resolution microscopy techniques, proteomics and genomics. For instance, we expect that proteomics analyses of isolated filament complexes will reveal new proteins associated with the major ones characterised to date. Previous experiences regarding the cytoskeletal structures of mammalian cells have shown that for each major protein involved, there are many others that play a role in the control of the assembly and disassembly of the filaments. The use of high-resolution electron microscopy, especially cryo-electron tomography, will reveal new details about the association of the filaments with the plasma membrane and other structures. This information is important with respect to analysing the functionality of the various structures discussed.
ACKNOWLEDGEMENTS
To several colleagues and students, who made the suggestion to write this review, to Drs Marlene Benchimol, Narcisa L Cunha e Silva, Thais Souto-Padrón and Marcos Farina, who critically reviewed the paper and made suggestions to improve it, to several colleagues and editors, which authorized the use of previously published figures, and to David Stracker, who revised the paper.
Received 23 August 2011
Accepted 23 February 2012
Financial support: CNPq, FINEP, CAPES, DECIT-MS, FAPERJ
- Abhayawardhane Y, Stewart GC 1995. Bacillus subtilis possesses a second determinant with extensive sequence similarity to the Escherichia coli merB morphogene. J Bacteriol 177: 765-773.
- Alvarez HM, Steinbuchel A 2002. Triacylglycerols in prokaryotic microorganisms. App Microbiol Biotechnol 60: 367-376.
- Ausmees N 2006. Intermediate filament-like cytoskeleton of Caulobacter crescentus J Mol Microbiol Biotechnol 11: 152-158.
- Ausmees N, Kuhn JR, Jacobs-Wagner C 2003. The bacterial cytoskeleton: an intermediate filament-like function in cell shape. Cell 115: 705-713.
- Aylett CHS, Wang Q, Michie KA, Amos LA Lowe J 2010. Filament structure of bacterial tubulin homologue TubZ. Proc Natl Acad Sci USA 107: 19766-19771.
- Bazylinski DA, Frankel RB 2004. Magnetosome formation in prokaryotes. Nature Rev Microbiol 2: 217-230.
- Beech PL, Nheu T, Schultz T, Herert S, Lithgow T, Gilson PR, McFadden GI 2000. Mitochondrial ftsZ in a chromophyte alga. Science 287: 1276-1279.
- Benz I, Schmidt A 2002. Never say never again: protein glycosylation in pathogenic bacteria. Mol Microbiol 45: 267-276.
- Bi E, Lutkenhaus J 1991. FstZ ring structure associated with division in Escherichia coli Nature 354: 161-164.
- Blakemore RP 1975. Magnetotactic bacteria. Science 190: 377-379.
- Bobik TA 2006. Polyhedral organelles compartmenting bacterial metabolic processes. App Microbiol Biotechnol 70: 517-525.
- Bramhill D, Thompson CM 1994. GTP-dependent polymerization of Escherichia coli FtsZ protein to form tubules. Proc Natl Acad Sci USA 91: 5813-5817.
- Carballido-Lopez R, Errington J 2003. The bacterial cytoskeleton: in vivo dynamics of the actin-like protein Mb1 of Bacillus subtilis Dev Cell 4: 19-28.
- Chen P, Anderson DI, Roth JR 1994. The control region of the pdulcob regulon in Salmonella typhymurium J Bacteriol 176: 5474-5482.
- Cohen-Bazire G, Kunisawa R 1963. The fine structure of Rhodospirillum rubrum J Cell Biol 16: 401-419.
- Dai K, Luthkenhaus J 1992. The proper ratio of FtsZ to FtsA is required for cell division to occur in Escherichia coli. J Bacteriol 174: 6145-6151.
- Daniel RA, Errington J 2003. Control of cell morphogenesis in bacteria: two distinct ways to make a rod-shaped cell. Cell 113: 767-776.
- Davis DM, Sowinski S 2008. Membrane nanotubes: dynamic long-distance connections between animal cells. Nature Rev Mol Cell Biol 9: 431-436.
- de Boer P, Crossley R, Rothfield L 1992. The essential bacteria cell-division protein FstZ is a GTPase. Nature 359: 254-256.
- Defeu-Soufo H-J, Graumann PL 2004. Dynamic movement of actin-like proteins within bacterial cells. EMBO Rep 5: 789-794.
- Docampo R, de Souza W, Miranda K, Rohloff P, Moreno SN 2005. Acidocalcisomes-conserved from bacteria to man. Nature Rev Microbiol 3: 251-261.
- Drews G, Niklowitz W 1956. Beitrage zur cytologie der Blaualgen. II. Zentroplasm und granulare Einschlusse vom Phormidium uncinatum Arch Mikrobiol 24: 147-162.
- Dubey GP, Ben-Yehuda S 2011. Intercellular nanotubes mediate bacterial communication. Cell 144: 590-600.
- Erickson HP 1997. FtsZ, a tubulin homologue in prokaryote cell division. Trends Cell Biol 9: 362-367.
- Erickson HP 2000. Dynamin and ftsZ: missing links in mitochondrial and bacterial division. J Cell Biol 148: 1103-1105.
- Erickson HP, Taylor DW, Taylor KA, Bramhill D 1996. Bacterial cell division protein FstZ assembles into protofilament sheets and minirings, structural homologs of tubulin polymer. Proc Natl Acad Sci USA 93: 519-523.
- Esue O, Wirtz D, Tseng Y 2006. GTPase activity, structure and mechanical properties of filaments assembled from bacterial cytoskeleton protein MreB. J Bacteriol 188: 968-976.
- Farina M, Esquivel DM, Lins de Barros H 1990. Magnetic iron-sulphur crystals from a magnetotactic microorganism. Nature 343: 256-258.
- Figge RM, Divakaruni AV, Gober JW 2004. MreB, the cell shape-determinant bacterial actin homologue, co-ordinates cell wall morphogenesis in Caulobacter crescentus Mol Microbiol 51: 1321-1332.
- Frankel RB, Bazylinski DA 2006. How magnetotactic bacteria make magnetosomes queue up. Trends Microbiol 14: 329-331.
- Frankel RB, Blakemore RP, Wolfe RS 1979. Magnetite in freshwater magnetotactic bacteria. Science 203: 1355-1356.
- Gerdes HH, Carvalho NR 2008. Intercellular transfer mediated by tunneling nanotubes. Curr Op Cell Biol 20: 470-475.
- Gitai Z 2005. The new bacterial cell biology: moving pats and subcellular architecture. Cell 120: 577-586.
- Gorby YA, Beveridge TJ, Blakemore RP 1988. Characterization of the bacterial magnetosome membrane. J Bacteriol 170: 834-841.
- Gousset K, Schiff E, Longevin C, Marijanovic Z, Caputo A, Brownan DT, Chenovard N, de Chaumont F, Martino A, Enninga J, Olivo-Marin JC, Mannel D, Zurzolo C 2009. Prions hijack tunneling nanotubes for intercellular spread. Nature Cell Biol 11: 328-336.
- Graumann PL 2004. Cytoskeletal elements in bacteria. Curr Op Microbiol 7: 565-571.
- Graumann PL 2007. Cytoskeletal elements in bacteria. Annu Rev Microbiol 61: 589-618.
- Hale CA, de Boer PAJ 1997. Direct binding of FtsZ to ZipA, an essential component of the septal ring structure that mediates cell division in E. coli Cell 88: 175-185.
- Hale CA, de Boer PAJ 1999. Recruitment of ZipA to the septal ring of Escherichia coli is dependent on FstZ and independent of FstA. J Bacteriol 181: 167-176.
- Hegermann J, Herrmann R, Mayer F 2002. Cytoskeletal elements in the bacterium Mycoplasma pneumoniae Naturwissenschaften 89: 453-458.
- Heuser JE 2011. The origins and evolution of freeze-etch electron microscopy. J Elec Microscop 60 (Suppl.): S3-S29.
- Heywood BR, Bazylinski DA, Garrat-Reed AJ, Mann S, Frankel RB 1990. Controlled biosynthesis of greigite (Fe3S4) in magnetotactic bacteria. Naturwiss 77: 536-538.
- Hoenger A, Bouchet-Marquis C 2011. Cellular tomography. Adv Protein Chem Struct Biol 82: 67-90.
- Hu B, Xie G, Lo CC, Starkenburg SR, Chain PS 2011. Pathogen comparative genomics in the next-generation sequencing era: genome alignments, pangenomics and metagenomics.
- Hurbain I, Sachse M 2011. The future is cold: cryopreparation methods for transmission electron microscopy of cells. Biol Cell 103: 405-420.
- Izard J 2006. Cytoskeletal cytoplasmic filaments ribbon of Treponema: a member of an intermediate-like filament protein family. J Mol Microbiol Biotechnol 11: 159-166.
- Izard J, McEwen BF, Bernard RM, Portuese T, Samsonoff WA, Limberger RJ 2004. Tomographic reconstruction of treponemal cytoplasmic filaments reveals novel bridging and anchoring components. Mol Microbiol 51: 609-615.
- Izard J, Samsonoff WA, Kinoshita MB, Limberger RJ 1999. Genetic and structural analyses of cytoplasmic filaments of wild-type Treponema phagedensis and a flagellar filament-deficient mutant. J Bacteriol 181: 6739-6746.
- Jennings PC, Cox GC, Monahan LG, Harry EJ 2011. Super-resolution imaging of the bacterial cytokinetic protein FtsZ. Micron 42: 336-341.
- Jensen TE 1993. Cyanobacterial ultrastructure. In T Berner (ed.), Ultrastructure of microalgae, CRC Press, Boca Raton, p. 7-51.
- Jones LJ, Carballido-Lopez R, Errington J 2001. Control of cell shape in bacteria: helical, actin-like filaments in Bacillus subtilis Cell 104: 913-922.
- Kalman S, Mitchell W, Marathe R, Lammel C, Fan J, Hyman RW, Olinger L, Grimwood J, Davis RW, Stephens RS 1999. Comparative genomes of Chlamydia pneumoniae and C. trachomatis Nature Genet 21: 385-389.
- Keim CN, Abreu F, Lins U, Lins de Barros H, Farina M 2004. Cell organization and ultrastructure of a magnetotactic multicellular organism. J Struct Biol 145: 254-262.
- Komeili A, Li Z, Newmann DK, Jensen GJ 2006. Magnetosomes are cell membrane invaginations organized by the actin-like protein MamK. Science 311: 242-245.
- Kurner J, Frangakis AS, Baumeister W 2005. Cryo-electron tomography reveals the cytoskeletal structure of Spiroplasma melliferum Science 307: 456-438.
- Lefévre CT, Menguy N, Abreu F, Lins U, Pósfai M, Prozorov T, Pignot D, Frankel RB, Bazylinski DA 2011. A cultured greigite-producing magnetotactic bacterium in a novel group of sulfate-reducing bacteria. Science 334: 1720-1723.
- Levin PA, Losick R 1996. Transcription factor SpoOa switches the localization of the cell division protein FstZ from a medial to a bipolar pattern in Bacillus subtilis Genes Dev 10: 478-488.
- Lindsay MR, Webb RY, Strous M, Jetten MS, Butler MK, Forde RJ, Fuerst JA 2001. Cell compartmentalisation in planctomycetes: novel types of structural organisation for the bacterial cell. Arch Microbiol 175: 413-429.
- Liu Z, Mukherjee A, Lutkenhaus J 1999. Recruitment of ZipA to the division site by interaction with TtsZ. Mol Microbiol 31: 1853-1861.
- Lowe J, Amos LA 1998. Crystal structure of the bacterial cell-division protein FtsZ. Nature 391: 203-206.
- Lowe J, Amos LA 1999. Tubulin-like protofilament in Ca2+-induced FtsZ sheets. EMBO J 18: 2364-2371.
- Lowe J, Amos LA 2009. Evolution of cytomotive filaments: the cytoskeleton from prokaryotes to eukaryotes. Int J Biochem Cell Biol 41: 323-329.
- Lu C, Reedy M, Erickson HP 2000. Straight and curved conformations of FtsZ are regulated by GTP hydrolysis. J Bacteriol 182: 164-170.
- Ma X, Ehrhardt DW, Margolin W 1996. Colocalization of cell division proteins FstZ and FtsA to cytoskeletal structures in living Escherichia coli cells by using green fluorescent protein. Proc Natl Acad Sci USA 93: 12998-13003.
- Mann S, Sparks NHC, Frankel RB, Bazylinski DA, Jannasch HW 1990. Biomineralization of ferrimagnetic greigite (Fe3 O4) and iron pyrite (FeS2) in a magnetotactic bacterium. Nature 343: 258-260.
- Margolin W 2005. FtsZ and the division of prokaryotic cells and organelles. Nature Rev Mol Cell Biol 6: 862-872.
- Martin-Galiano AJ, Oliva MA, Sanz L, Bhattacharyya A, Serna M, Yebenes H, Valpuesta JM, Rev JM 2011. Bacterial tubulin distinct loop sequences and primitive assembly properties support its origin from a eucaryotic tubulin ancestor. J Biol Chem 286: 19789-19803.
- Masuda K, Kawata T 1989. Isolation and characterization of cytoplasmic fibrils from treponemes. Microbiol Immunol 33: 619-630.
- Moller-Jensen J, Borch J, Dam M, Jensen RB, Ropstroff P, Gerdes K 2003. Bacterial mitosis: ParM of plasmid R1 moves plasmid DNA by an actin-like insertional polymerization mechanism. Mol Cell Biol 12: 1477-1487.
- Moller-Jensen J, Lowe J 2005. Increasing complexity of the bacterial cytoskeleton. Curr Op Cell Biol 17: 75-81.
- Mukherjee A, Dai K, Lutkenhaus J 1993. Escherichia coli cell division protein FstZ is a guanine nucleotide binding protein. Proc Natl Acad Sci USA 90: 1053-1057.
- Mukherje A, Lutkenhaus J 1994. Guanine nucleotide-dependent assembly of FtsZ into filaments. J Bacteriol 176: 2754-2758.
- Murphy DJ 2011. The dynamic roles of intracellular lipid droplets: from archaea to mammals. Protoplasma: doi: 10.1007/s00709-011-0329-7.
- Nogales EK, Downing H, Amos IA, Lowe J 1998. Tubulin and FtsZ form a distinct family of GTPases. Nature Struct Biol 5: 451-458.
- Okamura Y, Takeyama H, Matsunaga T 2001. A magnetosome specific GTPase from the magnetic bacteria Magnetospirillum magneticum AMB-1. J Biol Chem 276: 48183-48188.
- Oliva MA, Cordell SC, Lowe J 2004. Structural insights into the FstZ protofilament formation. Nature Struct Mol Biol 11: 1243-1250.
- Onfelt B, Nedvetzki S, Benninger RK, Purbhoo MA, Sowinsnki S, Hume AN, Seabra MC, Neil MA, French PM, Davis DM 2006. Structurally distinct membrane nanotubes between human macrophages support long-distance vesicular traffic or surfing of bacteria.
- Osteryoung KW, Stokes KD, Rutheford SM, Percival AL, Lee WY 1998. Chloroplast division in higher plants requires members of two functionally divergent gene families with homology to bacterial FtsZ. Plant Cell 10: 1991-2004.
- Osteryoung KW, Vierling E 1995. Conserved cell and organelle division. Nature 376: 473-474.
- Posfai M, Buseck PR, Bazylinski DA, Frenkel RB 1998. Reaction sequence of iron sulfide minerals and bacteria and their use as biomarkers. Science 280: 880-883.
- Ptacin JL, Lee SF, Garner EC, Toro E, Eckart M, Comolli LR, Moerner WE, Shapiro L 2010. A spindle-like apparatus guides bacterial chromosome segregation. Nature Cell Biol 12: 791-798.
- Ramirez-Weber FA, Kornberg TB 2000. Signaling reaches to new dimensions in Drosophila marginal disc. Cell 103: 189-192.
- Raychaudhuri D, Park JT 1992. Escherichia coli cell division gene ftsZ encodes a novel GTP-binding protein. Nature 359: 251-254.
- Regula JT, Boguth G, Gorg A, Hegermann J, Mayer F, Frank R, Hermann R 2001. Defining the mycoplasma "cytoskeleton": the protein composition of the Triton X-100 insoluble fraction of the bacterium Mycoplasma pneumoniae determined by 2-D gel electrophoresis and mass spectrometry. Microbiology 147: 1045-1057.
- Sabidó E, Selevsek N, Aebersold R 2011. Mass spectrometry-based proteomics for systems biology. Curr Opin Biotechnol 23: 1-7.
- Sako Y, Nomura N, Uchida A, Ishida Y, Morii H, Koga Y, Hoaki T, Maruyama T 1996. Aeropyrum pernix gen. nov., sp. nov., a novel aerobic hypertermophilic archeon growing at temperatures up to 100 degrees C. Int J Syst Bacteriol 46: 1070-1077.
- Schaffer C, Graninger M, Messner P 2001. Prokaryotic glycosylation. Proteomics 1: 248-261.
- Schlieper DM, Oliva MA, Andreu JM, Lowe J 2005. Structure of the bacterial tubulin BtubA/B: evidence for horizontal gene transfer. Proc Natl Acad Sci USA 102: 9170-9175.
- Schmid MF, Paredes AM, Khant HA, Soyer F, Aldrich HC, Chiu W, Shiveley JM 2006. Structure of Halothiobacillus neapolitanus carboxysomes by cryo-electron tomography. J Mol Biol 364: 526-535.
- Schuller D 2008. Genetics and cell biology of magnetosomes formation in magnetotactic bacteria. FEMS Microbiol Rev 32: 654-672.
- Seufferheld M, Vieira MCF, Ruiz FA, Rodrigues CO, Moreno SNJ, Docampo R 2003. Identification of organelles in bacteria similar to acidocalcisomes of unicellular eukaryotes. J Biol Chem 278: 22971-22978.
- Sherer NM, Lehmann MJ, Jimenez-Soto LF, Horensavitz C, Pypaert M, Mothes W 2007. Retroviruses can establish filopodial bridges for efficient cell-to-cell transmission. Nature Cell Biol 9: 310-315.
- Sherer NM, Mothes W 2008. Cytonemes and tunneling nanotubules in cell-cell communication and viral pathogenesis. Trends Cell Biol 18: 414-420.
- Shih Y-L, Rothfield L 2006. The bacterial cytoskeleton. Microbiol Mol Biol Rev 70: 729-754.
- Shively JM, Ball F, Brown DH, Saunders RE 1973. Functional organelles in prokaryotes: polyhedral inclusions (carboxysomes) of Thiobacillus neopolitanus Science 182: 6584-586.
- Souto HJ, Graumann PL 2003. Actin-like proteins MreB and Mb1 from Bacilus subtilis are required for bipolar positioning of replication origins. Curr Biol 13: 1916-1920.
- Stephens RS, Kalman S, Lammel C, Fan J, Marathe R, Aravind L, Mitchell W, Olinger L, Tatusov RL, Zhao Q 1998. Genome sequence of an obligate intracellular pathogen of humans: Chlamydia trachomatis Science 282: 754-759.
- Terfve C, Saez-Rodriguez J 2012. Modeling signaling networks using high-throughput phosphoproteomics. Ad Exp Med Biol 736: 19-57.
- Townsend R, Burgess J, Plaskitt KA 1980. Morphology and ultrastructure of helical and nonhelical strains of Spiroplasma citri. J Bacteriol 142: 973-984.
- Trachtenberg S 1998. Mollicutes-wall-less bacteria with internal cytoskeleton 1998. J Struct Biol 124: 244-256.
- Van den Ent F, Amos LA, Lowe J 2001. Prokaryotic origin of the actin cytoskeleton. Nature 413: 39-44.
- Van den Ent F, Moller-Jensen J, Amos LA, Gerdes K, Lowe J 2002. F-actin-like filaments formed by plasmid segregation protein ParM. EMBO J 21: 6935-6943.
- Vats P, Rothfield L 2007. Duplication and segregation of the actin (MreB) cytoskeleton during the prokaryotic cell cycle. Proc Natl Acad Sci USA 104: 17795-17800.
- Volmer W 2006. The prokaryotic cytoskeleton: a putative target for inhibitors and antibiotics? App Microbiol Biotechnol 73: 37-47.
- Waltermann M, Steinbuchel A 2005. Neutral lipid bodies in prokaryotes: recent insights into structure, formation and relationship to eukaryotic lipid depots. J Bacteriol 187: 3607-3619.
- Wang J, Galgoci A, Kodali S, Herath KB, Jayasuriya H, Dorso K, Vicente F, Gonzalez A, Cully D, Bramhill D, Singh S 2003. Discovery of a small molecule that inhibits cell division by blocking FstZ, a novel therapeutic target antibiotics. J Biol Chem 278: 44424-44428.
- Yang R, Bartle S, Otto R, Stassinopoulos A, Rogers M, Plamann L, Hartzell P 2004. AgIZ is a filament-forming coiled-coil protein required for adventurous gliding motility of Mycoplasma xanthus J Bacteriol 186: 6168-6178.
- Yeates TO, Kerfeld CA, Heinhorst S, Cannon GC, Shively JM 2008. Protein-based organelles in bacteria: carboxysomes and related microcompartments. Nature Rev Microbiol 6: 681-691.
- Yeates TO, Tsai Y, Tanaka S, Sawaya MR, Kerfeld CA 2007. Self-assembly in the carboxysome: a viral capsid-like protein shell in bacterial cells. Biochem Soc Trans 35: 508-511.
- You Y, Elmore S, Cotton LL, McKenzie C, Stoops JK, Wenstock GM, Norris SJ 1996. Characterizationnn of the cytoplasmic filament protein gene (cfpA) of Treponema pallidum subsp. Pallidum. J Bacteriol 178: 3177-3187.
Publication Dates
-
Publication in this collection
13 Apr 2012 -
Date of issue
May 2012
History
-
Received
23 Aug 2011 -
Accepted
23 Feb 2012