Abstract
American cutaneous leishmaniasis (ACL) is a vector-transmitted infectious disease with an estimated 1.5 million new cases per year. In Brazil, ACL represents a significant public health problem, with approximately 30,000 new reported cases annually, representing an incidence of 18.5 cases per 100,000 inhabitants. Corte de Pedra is in a region endemic for ACL in the state of Bahia (BA), northeastern Brazil, with 500-1,300 patients treated annually. Over the last decade, population and family-based candidate gene studies were conducted in Corte de Pedra, founded on previous knowledge from studies on mice and humans. Notwithstanding limitations related to sample size and power, these studies contribute important genetic biomarkers that identify novel pathways of disease pathogenesis and possible new therapeutic targets. The present paper is a narrative review about ACL immunogenetics in BA, highlighting in particular the interacting roles of the wound healing gene FLI1 with interleukin-6 and genes SMAD2 and SMAD3 of the transforming growth factor beta signalling pathway. This research highlights the need for well-powered genetic and functional studies on Leishmania braziliensis infection as essential to define and validate the role of host genes in determining resistance/susceptibility regarding this disease.
genetic biomarkers; American cutaneous leishmaniasis; wound healing genes
American cutaneous leishmaniasis (ACL) is a complex, multifactorial disease that results from environmental factors such as parasite polymorphism, phlebotomine sandfly components, as well as the host’s immune and genetic background. In northeastern Brazil, the endemic area of Corte de Pedra registers the highest incidence of ACL in the state of Bahia (BA). While the incidence of CL in BA varies from 1.5-3.2 per 10,000, the incidence in Corte de Pedra varies from 15-35 per 10,000. The predominant causative species is Leishmania braziliensis, which in most cases leads to CL, characterised by one or more ulcers with raised borders, most frequently located on the upper and lower extremities, but also on the head, face and trunk (Barral-Netto et al. 1997Barral-Netto M, Machado P, Bittencourt A, Barral A 1997. Recent advances in the pathophysiology and treatment of human cutaneous leishmaniasis. Curr Opin Dermatol 4: 51-58.). Although CL is a self-limiting disease, approximately 3-5% of subjects infected with L. braziliensis will eventually develop mucosal leishmaniasis (ML) or disseminated leishmaniasis (DL), considered now an emerging form of the disease in the area. Fig. 1 demonstrates these different clinical phenotypes, highlighting the sometimes disfiguring nature of the disease and the need to understand the variable disease pathology.
: the study area and spectrum of clinical disease caused by Leishmania braziliensis infection in Corte de Pedra, state of Bahia, Brazil. A: typical house and farm area; B: typical cutaneous leishmaniasis lesion characterised by granulomatous background and elevated borders; C: disseminated leishmaniasis, a form of disease that is increasing in the study area; D: mucosal leishmaniasis characterised by infiltrated ulcers that can cause extensive destruction of the nasal septum, columella and the upper lip.
A number of studies on ACL conducted in Corte de Pedra in the past 30 years have contributed enormously to the knowledge of ACL epidemiology and immune response (Carvalho et al. 2012Carvalho LP, Passos S, Schriefer A, Carvalho EM 2012. Protective and pathologic immune responses in human tegumentary leishmaniasis. Front Immunol 3: 301., de Oliveira & Brodskyn 2012de Oliveira CI, Brodskyn CI 2012. The immunobiology of Leishmania braziliensis infection. Front Immunol 3: 145.). Particularly in the last decade, a number of studies evaluating both parasite and host polymorphisms have demonstrated that genetic factors are associated to different clinical forms, revealing relevant biomarkers to understanding the disease pathogenesis (Schriefer et al. 2004Schriefer A, Schriefer AL, Goes-Neto A, Guimarães LH, Carvalho LP, Almeida RP, Machado PR, Lessa HA, de Jesus AR, Riley LW, Carvalho EM 2004. Multiclonal Leishmania braziliensis population structure and its clinical implication in a region of endemicity for American tegumentary leishmaniasis. Infect Immun 72: 508-514., Castellucci et al. 2006Castellucci L, Menezes E, Oliveira J, Magalhães A, Guimarães LH, Lessa M, Ribeiro S, Reale J, Noronha EF, Wilson ME, Duggal P, Beaty TH, Jeronimo S, Jamieson SE, Bales A, Blackwell JM, de Jesus AR, Carvalho EM 2006. IL6 -174 G/C promoter polymorphism influences susceptibility to mucosal but not localized cutaneous leishmaniasis in Brazil. J Infect Dis 194: 519-527., 2010Castellucci L, Jamieson SE, Miller EN, Menezes E, Oliveira J, Magalhães A, Guimarães LH, Lessa M, de Jesus AR, Carvalho EM, Blackwell JM 2010. CXCR1 and SLC11A1 polymorphisms affect susceptibility to cutaneous leishmaniasis in Brazil: a case-control and family-based study. BMC Med Genet 11: 10., 2011Castellucci L, Jamieson SE, Miller EN, de Almeida LF, Oliveira J, Magalhães A, Guimarães LH, Lessa M, Lago E, de Jesus AR, Carvalho EM, Blackwell JM 2011. FLI1 polymorphism affects susceptibility to cutaneous leishmaniasis in Brazil. Genes Immun 12: 589-594., 2012Castellucci L, Jamieson SE, Almeida L, Oliveira J, Guimarães LH, Lessa M, Fakiola M, de Jesus AR, Miller EN, Carvalho EM, Blackwell JM 2012. Wound healing genes and susceptibility to cutaneous leishmaniasis in Brazil. Infect Genet Evol 12: 1102-1110., Ramasawmy et al. 2010Ramasawmy R, Menezes E, Magalhães A, Oliveira J, Castellucci L, Almeida R, Rosa ME, Guimarães LH, Lessa M, Noronha E, Wilson ME, Jamieson SE, Kalil J, Blackwell JM, Carvalho EM, de Jesus AR 2010. The -2518 bp promoter polymorphism at CCL2/MCP1 influences susceptibility to mucosal but not localized cutaneous leishmaniasis in Brazil. Infect Genet Evol 10: 607-613., Queiroz et al. 2012Queiroz A, Sousa R, Heine C, Cardoso M, Guimarães LH, Machado PR, Carvalho EM, Riley LW, Wilson ME, Schriefer A 2012. Association between an emerging disseminated form of leishmaniasis and Leishmania (Viannia) braziliensis strain polymorphisms. J Clin Microbiol 50: 4028-4034.). Here we present a narrative review of host genetic studies of ACL conducted in Corte de Pedra over the last decade. Although there are a number of studies evaluating candidate genes in ACL (Table I), no genome-wide association studies have so far been reported that would provide a comprehensive map of genetic risk factors for this disease. This is in contrast to host genetic analysis of visceral leishmaniasis (VL), for which a well-powered genome-wide association study was recently reported (Fakiola et al. 2013Fakiola M, Strange A, Cordell HJ, Miller EN, Pirinen M, Su Z, Mishra A, Mehrotra S, Monteiro GR, Band G, Bellenguez C, Dronov S, Edkins S, Freeman C, Giannoulatou E, Gray E, Hunt SE, Lacerda HG, Langford C, Pearson R, Pontes NN, Rai M, Singh SP, Smith L, Sousa O, Vukcevic D, Bramon E, Brown MA, Casas JP, Corvin A, Duncanson A, Jankowski J, Markus HS, Mathew CG, Palmer CN, Plomin R, Rautanen A, Sawcer SJ, Trembath RC, Viswanathan AC, Wood NW, Wilson ME, Deloukas P, Peltonen L, Christiansen F, Witt C, Jeronimo SM, Sundar S, Spencer CC, Blackwell JM, Donnelly P 2013. Common variants in the HLA-DRB1-HLA-DQA1 HLA class II region are associated with susceptibility to visceral leishmaniasis. Nat Genet 45: 208-213.). Here we will focus on genetic susceptibility to ACL, beginning with the demonstration of familial aggregation of ACL disease in Corte de Pedra that led to analysis of specific candidate genes arising both from our knowledge of immune responses to human L. braziliensis infection, and through consideration of wound healing genes that was inspired initially by studies in mice (Sakthianandeswaren et al. 2005Sakthianandeswaren A, Elso CM, Simpson K, Curtis JM, Kumar B, Speed TP, Handman E, Foote SJ 2005. The wound repair response controls outcome to cutaneous leishmaniasis. Proc Natl Acad Sci USA 102: 15551-15556., 2009Sakthianandeswaren A, Foote SJ, Handman E 2009. The role of host genetics in leishmaniasis. Trends Parasitol 25: 383-391., 2010Sakthianandeswaren A, Curtis JM, Elso C, Kumar B, Baldwin TM, Lopaticki S, Kedzierski L, Smyth GK, Foote SJ, Handman E 2010. Fine mapping of Leishmania major susceptibility locus lmr2 and evidence of a role for Fli1 in disease and wound healing. Infect Immun 78: 2734-2744.). These data are further discussed in relation to studies of genetic susceptibility to CL in other geographic regions, as summarised along with all published (Barbier et al. 1987Barbier D, Demenais F, Lefait JF, David B, Blanc M, Hors J, Feingold N 1987. Susceptibility to human cutaneous leishmaniasis and HLA, Gm, Km markers. Tissue Antigens 30: 63-67., Lara et al. 1991Lara ML, Layrisse Z, Scorza JV, Garcia E, Stoikow Z, Granados J, Bias W 1991. Immunogenetics of human American cutaneous leishmaniasis. Study of HLA haplotypes in 24 families from Venezuela. Hum Immunol 30: 129-135., Petzl-Erler et al. 1991Petzl-Erler ML, Belich MP, Queiroz-Telles F 1991. Association of mucosal leishmaniasis with HLA. Human Immunol 32: 254-260., El-Mogy et al. 1993El-Mogy MH, Abdel-Hamid IA, Abdel-Razic MM, Rizk RA, Romia SA 1993. Histocompatibility antigens in Egyptians with cutaneous leishmaniasis: a preliminary study. J Dermatol Sci 5: 89-91., Cabrera et al. 1995Cabrera M, Shaw M-A, Sharples C, Williams H, Castes M, Convit J, Blackwell JM 1995. Polymorphism in TNF genes associated with mucocutaneous leishmaniasis. J Exp Med 182: 1259-1264., Karplus et al. 2002Karplus TM, Jeronimo SM, Chang H, Helms BK, Burns TL, Murray JC, Mitchell AA, Pugh EW, Braz RF, Bezerra FL, Wilson ME 2002. Association between the tumor necrosis factor locus and the clinical outcome of Leishmania chagasi infection. Infect Immun 70: 6919-6925., Olivo-Diaz et al. 2004Olivo-Diaz A, Debaz H, Alaez C, Islas VJ, Perez-Perez H, Hobart O, Gorodezky C 2004. Role of HLA class II alleles in susceptibility to and protection from localized cutaneous leishmaniasis. Hum Immunol 65: 255-261., Castellucci et al. 2006Castellucci L, Menezes E, Oliveira J, Magalhães A, Guimarães LH, Lessa M, Ribeiro S, Reale J, Noronha EF, Wilson ME, Duggal P, Beaty TH, Jeronimo S, Jamieson SE, Bales A, Blackwell JM, de Jesus AR, Carvalho EM 2006. IL6 -174 G/C promoter polymorphism influences susceptibility to mucosal but not localized cutaneous leishmaniasis in Brazil. J Infect Dis 194: 519-527., 2010Castellucci L, Jamieson SE, Miller EN, Menezes E, Oliveira J, Magalhães A, Guimarães LH, Lessa M, de Jesus AR, Carvalho EM, Blackwell JM 2010. CXCR1 and SLC11A1 polymorphisms affect susceptibility to cutaneous leishmaniasis in Brazil: a case-control and family-based study. BMC Med Genet 11: 10., 2011Castellucci L, Jamieson SE, Miller EN, de Almeida LF, Oliveira J, Magalhães A, Guimarães LH, Lessa M, Lago E, de Jesus AR, Carvalho EM, Blackwell JM 2011. FLI1 polymorphism affects susceptibility to cutaneous leishmaniasis in Brazil. Genes Immun 12: 589-594., 2012Castellucci L, Jamieson SE, Almeida L, Oliveira J, Guimarães LH, Lessa M, Fakiola M, de Jesus AR, Miller EN, Carvalho EM, Blackwell JM 2012. Wound healing genes and susceptibility to cutaneous leishmaniasis in Brazil. Infect Genet Evol 12: 1102-1110., Kamali-Sarvestani et al. 2006Kamali-Sarvestani E, Rasouli M, Mortazavi H, Gharesi-Fard B 2006. Cytokine gene polymorphisms and susceptibility to cutaneous leishmaniasis in Iranian patients. Cytokine 35: 159-165., Matos et al. 2007Matos GI, Covas CJ, Bittar RC, Gomes-Silva A, Marques F, Maniero VC, Amato VS, Oliveira-Neto MP, Mattos MS, Pirmez C, Sampaio EP, Moraes MO, da-Cruz AM 2007. IFNG +874T/A polymorphism is not associated with American tegumentary leishmaniasis susceptibility but can influence Leishmania induced IFN-gamma production. BMC Infect Dis 7: 33., Salhi et al. 2008Salhi A, Rodrigues Jr V, Santoro F, Dessein H, Romano A, Castellano LR, Sertorio M, Rafati S, Chevillard C, Prata A, Alcais A, Argiro L, Dessein A 2008. Immunological and genetic evidence for a crucial role of IL-10 in cutaneous lesions in humans infected with Leishmania braziliensis. J Immunol 180: 6139-6148., Ajdary et al. 2010Ajdary S, Ghamilouie MM, Alimohammadian MH, Hosseini M, Pakzad SR 2010. Lack of association of Toll-like receptor 2 Arg753Gln with cutaneous leishmaniasis. Parasitol Int 59: 466-468., 2011Ajdary S, Ghamilouie MM, Alimohammadian MH, Riazi-Rad F, Pakzad SR 2011. Toll-like receptor 4 polymorphisms predispose to cutaneous leishmaniasis. Microbes Infect 13: 226-231., Ramasawmy et al. 2010Ramasawmy R, Menezes E, Magalhães A, Oliveira J, Castellucci L, Almeida R, Rosa ME, Guimarães LH, Lessa M, Noronha E, Wilson ME, Jamieson SE, Kalil J, Blackwell JM, Carvalho EM, de Jesus AR 2010. The -2518 bp promoter polymorphism at CCL2/MCP1 influences susceptibility to mucosal but not localized cutaneous leishmaniasis in Brazil. Infect Genet Evol 10: 607-613., Samaranayake et al. 2010Samaranayake TN, Fernando SD, Dissanayake VH 2010. Candidate gene study of susceptibility to cutaneous leishmaniasis in Sri Lanka. Trop Med Int Health 15: 632-638., Oliveira et al. 2011Oliveira CR, Pereira LI, Pereira AJ, Ferreira AA, Crespo AM, Silveira LA 2011. Allelic polymorphism of human FcgammaRIIA-H/R131 receptor in American tegumentary leishmaniasis. Int J Immunogenet 38: 225-231., Fernández-Figueroa et al. 2012Fernández-Figueroa EA, Rangel-Escareno C, Espinosa-Mateos V, Carrillo-Sanchez K, Salaiza-Suazo N, Carrada-Figueroa G, March-Mifsut S, Becker I 2012. Disease severity in patients infected with Leishmania mexicana relates to IL-1beta. PLoS Negl Trop Dis 6: e1533., Covas et al. 2013Covas CJF, Cardoso CC, Gomes-Silva A, Oliveira JRS, da-Cruz AM, Moraes MO 2013. Candidate gene case-control and functional study shows macrophage inhibitory factor (MIF) polymorphism is associated with cutaneous leishmaniasis. Cytokine 61: 168-172.) data on susceptibility to CL in Table I. One factor that affects interpretation of all of these studies is the issue of sample size and power, which we will return to in our concluding remarks.
The endemic site of Corte de Pedra - Corte de Pedra, a village located in the southwestern region of BA, belongs to the municipality of Presidente Tancredo Neves, whose population is approximately 17,928 inhabitants (source: Brazilian Institute of Geography and Statistics). The endemic area of Corte de Pedra, however, extends far beyond the village, covering 20 municipalities in a total area of approximately 9,935 km2 around the site where a Health Post was established in 1980s as a reference centre for the treatment of leishmaniasis in the region. Currently, 430,347 people are distributed across these towns, for which the main economic activity is subsistence farming, particularly the cultivation of cocoa, cloves, guarana, banana, coffee, black pepper and rubber. The endemic area of Corte de Pedra is typically an area of rainforest that over the years has been reduced to isolated areas of secondary forest with agricultural activities providing the main source of income for the majority of its inhabitants. The occupational and domestic habits of these individuals, which involve work on farms and homes built in clearings in the woods, have increased the population’s exposure to L. braziliensis infection. From 2007-2012, 7,093 cases of ACL were recorded in the region, with 6,747 (95%) cases of CL, 138 (2%) cases of ML and 208 cases (3%) cases of DL.
A familial aggregation study - It is well known that the clinical outcome of parasitic infections is influenced by the complex interaction of parasite strain, host genetics and environmental factors. Leishmaniasis, in particular, has a broad clinical spectrum associated with variable profiles of immune response and different Leishmania species (Cabrera et al. 1995Cabrera M, Shaw M-A, Sharples C, Williams H, Castes M, Convit J, Blackwell JM 1995. Polymorphism in TNF genes associated with mucocutaneous leishmaniasis. J Exp Med 182: 1259-1264., Alcais et al. 1997Alcais A, Abel L, David C, Torrez ME, Flandre P, Dedet JP 1997. Evidence for a major gene controlling susceptibility to tegumentary leishmaniasis in a recently exposed Bolivian population. Am J Hum Genet 61: 968-979., Ribeiro-de-Jesus et al. 1998Ribeiro-de-Jesus A, Almeida RP, Lessa H, Bacellar O, Carvalho EM 1998. Cytokine profile and pathology in human leishmaniasis. Braz J Med Biol Res 31: 143-148.). Previous studies have described familial clustering of VL and CL (Alcais et al. 1997Alcais A, Abel L, David C, Torrez ME, Flandre P, Dedet JP 1997. Evidence for a major gene controlling susceptibility to tegumentary leishmaniasis in a recently exposed Bolivian population. Am J Hum Genet 61: 968-979., Blackwell et al. 1997Blackwell JM, Black GF, Peacock CS, Miller EN, Sibthorpe D, Gnananandha D, Shaw JJ, Silveira F, Lins-Lainson Z, Ramos F, Collins A, Shaw M-A 1997. Immunogenetics of leishmanial and mycobacterial infections: the Belém family study. Philos Trans R Soc Lond Biol Sci 352: 1331-1345., Jeronimo et al. 2000Jeronimo SM, Teixeira MJ, Sousa A, Thielking P, Pearson RD, Evans TG 2000. Natural history of Leishmania (Leishmania) chagasi infection in northeastern Brazil: long-term follow-up. Clin Infect Dis 30: 608-609.). Given that ML is a rare phenotype associated with a vigorous inflammatory response to parasite antigens (Bacellar et al. 2002Bacellar O, Lessa H, Schriefer A, Machado P, de Jesus AR, Dutra WO, Gollob KJ, Carvalho EM 2002. Up-regulation of Th1-type responses in mucosal leishmaniasis patients. Infect Immun 70: 6734-6740.), we conducted a study to address the hypothesis that familial clustering of ML would occur in the endemic area of Corte de Pedra. The study was a reconstructed cohort, a hybrid between a case-control and a retrospective cohort study. All members of 30 ML and 30 neighbourhood control families were assessed for history of exposure, as assessed by positive delayed type hypersensitivity (DTH) response and/or current or past disease confirmed from medical records or by clinical examination for presence of a scar in association with a positive DTH response. First-degree relatives of index cases were compared with those of index controls (Castellucci et al. 2005Castellucci L, Cheng LH, Araújo C, Guimarães LH, Lessa H, Machado P, Almeida MF, Oliveira A, Ko A, Johnson WD, Wilson ME, Carvalho EM, de Jesus AR 2005. Familial aggregation of mucosal leishmaniasis in Northeast Brazil. Am J Trop Med Hyg 73: 69-73.). There were significant differences between the frequencies of CL (37% vs. 20%) and ML (5% vs. 0%) when comparing case families and control families, respectively. Additionally, families with two cases of ML had a higher frequency (29.6%) of DTH-positive individuals than control families (9.4%). In this way we documented familial aggregation of CL and ML in a region where L. braziliensis is highly endemic. Although shared environment reflecting the rate of exposure to sandflies, the number of parasites inoculated by the infected sandflies, pre-existing immune responses to sandfly saliva products and variation between isolates of L. braziliensis (Grimaldi Jr & Tesh 1993Grimaldi Jr G, Tesh RB 1993. Leishmaniases of the New World: current concepts and implications for future research. Clin Microbiol Rev 6: 230-250., Gillespie et al. 2000Gillespie RD, Mbow ML, Titus RG 2000. The immunomodulatory factors of blood-feeding arthropod saliva. Parasite Immunol 22: 319-331.) could contribute to this familial aggregation, our data favoured the hypothesis that genetic background could be influencing a higher rate of infection and/or a propensity to develop or retain a positive skin test in family members. This was supported by our failure to detect differences between ML and neighbourhood control families for environmental factors evaluated in our study area. At the same time, other studies were already documenting (Table I) host genetic factors influencing the immune response and clinical outcome of leishmaniasis in mice and humans (Blackwell et al. 1997Blackwell JM, Black GF, Peacock CS, Miller EN, Sibthorpe D, Gnananandha D, Shaw JJ, Silveira F, Lins-Lainson Z, Ramos F, Collins A, Shaw M-A 1997. Immunogenetics of leishmanial and mycobacterial infections: the Belém family study. Philos Trans R Soc Lond Biol Sci 352: 1331-1345., Blackwell 1998Blackwell JM 1998. Genetics of host resistance and susceptibility to intramacrophage pathogens: a study of multicase families of tuberculosis, leprosy and leishmaniasis in north-eastern Brazil. Int J Parasitol 28: 21-28.). Based on these findings, we conducted a number of candidate gene studies in order to identify polymorphic markers associated with ACL in the Corte de Pedra population.
Analysis of candidate immune response genes - The first series of candidate gene studies undertaken in our study area were based on analysis of candidate immune response genes informed by our knowledge of the immunopathology of disease. These studies were initially based on a case-control study design, where possible supported by family-based analysis to control for ethnic admixture. Both cohorts were geographically and demographically equivalent. Table II describes the structure of case-control and family sample sets used as a resource for these candidate gene studies.
Characteristics of collections made during the primary (2000-2004) and secondary (2008-2010) sampling periods (A) and demographic data of the case-control groups (B)
Interleukin (IL)-6 - ML is a severe disease that normally follows localised CL. Immune pathology is created by a strong pro-inflammatory response with high levels of tumour necrosis factor (TNF) and failure of type 2 cytokines to regulate this response. IL-6 down-regulates T helper (Th) cell type 1 differentiation and drives Th2 cell differentiation. Previous studies have shown that pre-treatment with recombinant human IL-6 inhibits interferon (IFN)-γ and TNF mediated activation of human macrophages for killing of L. amazonensis (Hatzigeorgiou et al. 1993Hatzigeorgiou DE, He S, Sobel J, Grabstein KH, Hafner A, Ho JL 1993. IL-6 down-modulates the cytokine-enhanced antileishmanial activity in human macrophages. J Immunol 151: 3682-3692.) and IL-6 has been shown to down regulate the expression of TNF membrane receptors (Bermudez et al. 1992Bermudez LE, Wu M, Petrofsky M, Young LS 1992. Interleukin-6 antagonizes tumor necrosis factor-mediated mycobacteriostatic and mycobactericidal activities in macrophages. Infect Immun 60: 4245-4252.). We evaluated (Castellucci et al. 2006Castellucci L, Menezes E, Oliveira J, Magalhães A, Guimarães LH, Lessa M, Ribeiro S, Reale J, Noronha EF, Wilson ME, Duggal P, Beaty TH, Jeronimo S, Jamieson SE, Bales A, Blackwell JM, de Jesus AR, Carvalho EM 2006. IL6 -174 G/C promoter polymorphism influences susceptibility to mucosal but not localized cutaneous leishmaniasis in Brazil. J Infect Dis 194: 519-527.) the functional IL6-174 bp G/C promoter polymorphism, a single nucleotide polymorphism (SNP) associated with pro-inflammatory diseases and IL-6 regulation (Fishman et al. 1998Fishman D, Faulds G, Jeffery R, Mohamed-Ali V, Yudkin JS, Humphries S, Woo P 1998. The effect of novel polymorphisms in the interleukin-6 (IL-6) gene on IL-6 transcription and plasma IL-6 levels and an association with systemic-onset juvenile chronic arthritis. J Clin Invest 102: 1369-1376., Bidwell et al. 1999Bidwell J, Keen L, Gallagher G, Kimberly R, Huizinga T, Mcdermott MF, Oksenberg J, Mcnicholl J, Pociot F, Hardt C, D’Alfonso S 1999. Cytokine gene polymorphism in human disease: on-line databases. Genes Immun 1: 3-19., Terry et al. 2000Terry CF, Loukaci V, Green FR 2000. Cooperative influence of genetic polymorphisms on interleukin-6 transcriptional regulation. J Biol Chem 275: 18138-18144.). In addition, IL-6 levels were measured in macrophages with or without stimulation with soluble Leishmania antigen (SLA) from L. braziliensis. Our data (Castellucci et al. 2006Castellucci L, Menezes E, Oliveira J, Magalhães A, Guimarães LH, Lessa M, Ribeiro S, Reale J, Noronha EF, Wilson ME, Duggal P, Beaty TH, Jeronimo S, Jamieson SE, Bales A, Blackwell JM, de Jesus AR, Carvalho EM 2006. IL6 -174 G/C promoter polymorphism influences susceptibility to mucosal but not localized cutaneous leishmaniasis in Brazil. J Infect Dis 194: 519-527.) provide both population-based [odds ratio (OR) = 2.29, 95% confidence intervals (CI) = 1.40-3.77, p = 0.001] and family-based (z = 4.3, p = 1.5 x 10-5) evidence for an association between the C allele of the -174 bp SNP at IL6 and susceptibility to ML. The family-based analysis was important in confirming that the association was not due to population substructure that might have differed between case and control groups. In addition, we found that the C allele was associated with reduced baseline expression of IL-6 in unstimulated macrophages and in macrophages stimulated with SLA. There are inconsistencies among studies concerning the role of the IL6-174 bp G/C polymorphism, both in terms of which is the disease-associated allele, and when attempting to determine whether different genotypes are functionally associated with the production of differing IL-6 levels. The fact that IL-6 has many pleiotropic effects in regulating both type 1 and type 2 immune response pathways (Diehl & Rincon 2002Diehl S, Rincon M 2002. The two faces of IL-6 on Th1/Th2 differentiation. Mol Immunol 39: 531-536.), plus the complexities of the immunopathogenesis of these different diseases (Rincon et al. 1997Rincon M, Anguita J, Nakamura T, Fikrig E, Flavell RA 1997. Interleukin (IL)-6 directs the differentiation of IL-4-producing CD4+ T cells. J Exp Med 185: 461-469., Diehl et al. 2000Diehl S, Anguita J, Hoffmeyer A, Zapton T, Ihle JN, Fikrig E, Rincon M 2000. Inhibition of Th1 differentiation by IL-6 is mediated by SOCS1. Immunity 13: 805-815.), might explain such differences. Besides, it is important to bear in mind that the -174 bp SNP is not the sole polymorphic determinant of differential and cell type-specific promoter activity driving IL6 gene transcription (Fishman et al. 1998Fishman D, Faulds G, Jeffery R, Mohamed-Ali V, Yudkin JS, Humphries S, Woo P 1998. The effect of novel polymorphisms in the interleukin-6 (IL-6) gene on IL-6 transcription and plasma IL-6 levels and an association with systemic-onset juvenile chronic arthritis. J Clin Invest 102: 1369-1376., Terry et al. 2000Terry CF, Loukaci V, Green FR 2000. Cooperative influence of genetic polymorphisms on interleukin-6 transcriptional regulation. J Biol Chem 275: 18138-18144.). In relation to our own study, as macrophages are the primary site of infection, we hypothesise that low IL-6 production in carriers of the C allele may contribute to a reduced capacity to induce Th2 cell differentiation and regulate the activity of CD4+ Th1 cell-generated cytokines (such as IFN-γ and TNF) that contribute to the destructive pathological manifestations associated with ML.
CCL2/MCP1 - There are several reports for the putative roles of the CCL2-encoded monocyte chemoattractant protein-1 (MCP-1) in leishmaniasis from infection studies in vitro (Ritter & Moll 2000Ritter U, Moll H 2000. Monocyte chemotactic protein-1 stimulates the killing of Leishmania major by human monocytes, acts synergistically with IFN-gamma and is antagonized by IL-4. Eur J Immunol 30: 3111-3120., Bhattacharyya et al. 2002Bhattacharyya S, Ghosh S, Dasgupta B, Mazumder D, Roy S, Majumdar S 2002. Chemokine-induced leishmanicidal activity in murine macrophages via the generation of nitric oxide. J Infect Dis 185: 1704-1708.) as well as by analysis of human (Ritter et al. 1996Ritter U, Moll H, Laskay T, Brocker E, Velazco O, Becker I, Gillitzer R 1996. Differential expression of chemokines in patients with localized and diffuse cutaneous American leishmaniasis. J Infect Dis 173: 699-709.) and murine (de Moura et al. 2005de Moura TR, Novais FO, Oliveira F, Clarencio J, Noronha A, Barral A, Brodskyn C, de Oliveira CI 2005. Toward a novel experimental model of infection to study American cutaneous leishmaniasis caused by Leishmania braziliensis. Infect Immun 73: 5827-5834.) lesions. Previous studies have variably demonstrated increased risk or protection from pulmonary tuberculosis associated with single SNP variants and/or different haplotypes created by promoter region SNPs at -362 bp and at -2,518 bp (Flores-Villanueva et al. 2005Flores-Villanueva PO, Ruiz-Morales JA, Song CH, Flores LM, Jo EK, Montano M, Barnes PF, Selman M, Granados J 2005. A functional promoter polymorphism in monocyte chemoattractant protein-1 is associated with increased susceptibility to pulmonary tuberculosis. J Exp Med 202: 1649-1658., Thye et al. 2009Thye T, Nejentsev S, Intemann CD, Browne EN, Chinbuah MA, Gyapong J, Osei I, Owusu-Dabo E, Zeitels LR, Herb F, Horstmann RD, Meyer CG 2009. MCP-1 promoter variant -362C associated with protection from pulmonary tuberculosis in Ghana, West Africa. Hum Mol Genet 18: 381-388., Intemann et al. 2011Intemann CD, Thye T, Forster B, Owusu-Dabo E, Gyapong J, Horstmann RD, Meyer CG 2011. MCP1 haplotypes associated with protection from pulmonary tuberculosis. BMC Genet 12: 34.). One of these studies (Flores-Villanueva et al. 2005Flores-Villanueva PO, Ruiz-Morales JA, Song CH, Flores LM, Jo EK, Montano M, Barnes PF, Selman M, Granados J 2005. A functional promoter polymorphism in monocyte chemoattractant protein-1 is associated with increased susceptibility to pulmonary tuberculosis. J Exp Med 202: 1649-1658.) further showed that tuberculosis patients carrying the G allele for the SNP at -2,518 bp had the highest plasma levels of MCP-1 and the lowest plasma levels of IL-12p40, which was therefore interpreted as a secondary effect of MCP-1 in impairing the Th1 immune response against Mycobacterium tuberculosis. We also demonstrated (Ramasawmy et al. 2010Ramasawmy R, Menezes E, Magalhães A, Oliveira J, Castellucci L, Almeida R, Rosa ME, Guimarães LH, Lessa M, Noronha E, Wilson ME, Jamieson SE, Kalil J, Blackwell JM, Carvalho EM, de Jesus AR 2010. The -2518 bp promoter polymorphism at CCL2/MCP1 influences susceptibility to mucosal but not localized cutaneous leishmaniasis in Brazil. Infect Genet Evol 10: 607-613.) that the G allele at the regulatory CCL2 -2,518 bp promoter is a risk factor for ML using our population-based (OR = 4.4, 95% CI = 1.42-13.65, p = 0.01) and family-based (z = 2.68, p = 0.007) samples (Table II) from Corte de Pedra. A number of studies suggest a link between the leishmanicidal capacity of MCP-1 and lesion healing. Previous work has demonstrated that MCP-1 enhances the cytotoxic response via induction of reactive oxygen intermediates by infected macrophages (Ritter & Moll 2000Ritter U, Moll H 2000. Monocyte chemotactic protein-1 stimulates the killing of Leishmania major by human monocytes, acts synergistically with IFN-gamma and is antagonized by IL-4. Eur J Immunol 30: 3111-3120., Bhattacharyya et al. 2002Bhattacharyya S, Ghosh S, Dasgupta B, Mazumder D, Roy S, Majumdar S 2002. Chemokine-induced leishmanicidal activity in murine macrophages via the generation of nitric oxide. J Infect Dis 185: 1704-1708.). Moreover, in patients with self-healing CL, high levels of MCP-1 were detected in infected skin whereas, in the non-healing lesions of diffuse CL, MCP-1 expression was much lower with a predominance of another CC chemokine, CCL3 or macrophage inflammatory protein 1-α (MIP-1α) (Ritter et al. 1996Ritter U, Moll H, Laskay T, Brocker E, Velazco O, Becker I, Gillitzer R 1996. Differential expression of chemokines in patients with localized and diffuse cutaneous American leishmaniasis. J Infect Dis 173: 699-709.). In addition, it was demonstrated that the chemokines MCP-1, MIP-1α and CXCL1 were expressed in ears and draining lymph nodes of mice infected in the ear with L. braziliensis (de Moura et al. 2005de Moura TR, Novais FO, Oliveira F, Clarencio J, Noronha A, Barral A, Brodskyn C, de Oliveira CI 2005. Toward a novel experimental model of infection to study American cutaneous leishmaniasis caused by Leishmania braziliensis. Infect Immun 73: 5827-5834.). Our results suggest that high levels of MCP-1 appear to exacerbate ML disease. In contrast to previous data (Flores-Villanueva et al. 2005Flores-Villanueva PO, Ruiz-Morales JA, Song CH, Flores LM, Jo EK, Montano M, Barnes PF, Selman M, Granados J 2005. A functional promoter polymorphism in monocyte chemoattractant protein-1 is associated with increased susceptibility to pulmonary tuberculosis. J Exp Med 202: 1649-1658.), plasma levels of IL-12p40 and IL-12p70 did not differ significantly between our CCL2 -2,518 bp genotype groups. We also observed higher MCP-1 levels in the supernatants of macrophages from GG compared to AA genotypes both in un-stimulated as well as SLA and LPS stimulated cultures. Our data support the alternative view that the proinflammatory capacity of MCP-1 in recruiting host monocytes could provide both the environment for parasite replication and for tissue damage and lesion development. This could be due to a direct effect of MCP-1 in bringing fresh monocytes to the site of infection and/or to downstream events regulated by MCP-1 in macrophages and other cells.
CXCR1 and SLC11A1 - It has been hypothesised (Peters & Sacks 2009Peters NC, Sacks DL 2009. The impact of vector mediated neutrophil recruitment on cutaneous leishmaniasis. Cell Microbiol 11: 1290-1296.) that differences in the ability of macrophages and dendritic cells from different inbred mouse strains to respond to apoptotic vs. necrotic polymorphonuclear leukocytes (PMN), arising during the wound healing response to an infected sandfly bite, determines disease progression. The arrival and maintenance of infiltrating cells at bite sites is thought to be mediated by sandfly derived factors that either mimic a tissue damage signal or activate chemokine/chemokine receptor pathways (Teixeira et al. 2005aTeixeira CR, Teixeira MJ, Gomes RB, Santos CS, Andrade BB, Raffaele-Netto I, Silva JS, Guglielmotti A, Miranda JC, Barral A, Brodskyn C, Barral-Netto M 2005a. Saliva from Lutzomyia longipalpis induces CC chemokine ligand 2/monocyte chemoattractant protein-1 expression and macrophage recruitment. J Immunol 175: 8346-8353., bTeixeira MJ, Fernandes JD, Teixeira CR, Andrade BB, Pompeu ML, da Silva JS, Brodskyn CI, Barral-Netto M, Barral A 2005b. Distinct Leishmania braziliensis isolates induce different paces of chemokine expression patterns. Infect Immun 73: 1191-1195., 2006Teixeira MJ, Teixeira CR, Andrade BB, Barral-Netto M, Barral A 2006. Chemokines in host-parasite interactions in leishmaniasis. Trends Parasitol 22: 32-40.). Expression patterns for chemokines have been associated with the evolution of large and small lesions in mice following L. braziliensis infection, influenced by both the strain of parasite (Teixeira et al. 2005bTeixeira MJ, Fernandes JD, Teixeira CR, Andrade BB, Pompeu ML, da Silva JS, Brodskyn CI, Barral-Netto M, Barral A 2005b. Distinct Leishmania braziliensis isolates induce different paces of chemokine expression patterns. Infect Immun 73: 1191-1195.) and the mouse genetic background (Teixeira et al. 2005aTeixeira CR, Teixeira MJ, Gomes RB, Santos CS, Andrade BB, Raffaele-Netto I, Silva JS, Guglielmotti A, Miranda JC, Barral A, Brodskyn C, Barral-Netto M 2005a. Saliva from Lutzomyia longipalpis induces CC chemokine ligand 2/monocyte chemoattractant protein-1 expression and macrophage recruitment. J Immunol 175: 8346-8353.). One way to look at the interplay between PMN and macrophages in disease progression in humans is to determine whether polymorphisms at genes that regulate their infiltration or function are associated with different clinical phenotypes following infection with Leishmania spp. CXCR1 (IL8RA) and CXCR2 (IL8RB) are genes encoding receptors for chemokines that attract PMN to inflammatory sites. They lie on human chromosome 2q25 230-260 kb upstream of SLC11A1, a gene that regulates macrophage activation and resistance to VL (Blackwell et al. 2001Blackwell JM, Goswami T, Evans CAW, Sibthorpe D, Papo N, White JK, Searle S, Miller EN, Peacock CS, Mohammed H, Ibrahim M 2001. SLC11A1 (formerly NRAMP1) and disease. Cell Microbiol 3: 773-784.). In our studies (Castellucci et al. 2010Castellucci L, Jamieson SE, Miller EN, Menezes E, Oliveira J, Magalhães A, Guimarães LH, Lessa M, de Jesus AR, Carvalho EM, Blackwell JM 2010. CXCR1 and SLC11A1 polymorphisms affect susceptibility to cutaneous leishmaniasis in Brazil: a case-control and family-based study. BMC Med Genet 11: 10.), we showed an association between ACL and polymorphic variants at the CXCR1, specifically at SNP rs2854386 for both population-based (OR = 2.38, 95% CI = 1.23-4.57, p = 0.009) and family-based (z = 2.00, p = 0.045). Of interest, the common C allele (presumed to be the functional variant) was associated with CL, whereas the rare G allele was associated with ML (z = 2.00, p = 0.046). This suggested that, whereas high numbers of PMN might be detrimental in the context of CL disease, they may have an important positive role to play in preventing ML disease. In addition, in the family-based study CL was associated (z = 2.55, p = 0.011) with a 3’ insertion/deletion polymorphism at SLC11A1, a gene primarily known for its role in the regulation of macrophage activation. The association is also of interest in relation to the putative role of this molecule in regulating expression of secretory leukocyte protease inhibitor and hence affecting the wound healing response (Thuraisingam et al. 2006Thuraisingam T, Sam H, Moisan J, Zhang Y, Ding A, Radzioch D 2006. Delayed cutaneous wound healing in mice lacking solute carrier 11a1 (formerly Nramp1): correlation with decreased expression of secretory leukocyte protease inhibitor. J Invest Dermatol 126: 890-901.). Differences in lesion development have not been observed following subcutaneous needle injection of either Leishmania major (Alexander & Blackwell 1986Alexander J, Blackwell JM 1986. The immunological significance of genetically determined cross reactivity between taxonomically distinct Leishmania species. In JA Riou (ed.), Leishmania, taxonomie et phylogenese: application éco-épidemiologiques, Institut Méditerranée d’Etudes Epidémiologiques et Ecologiques, Montpellier, p. 185-191.) or Leishmania mexicana (Roberts et al. 1989Roberts M, Alexander J, Blackwell JM 1989. Influence of Lsh, H-2 and an H-11-linked gene on visceralization and metastasis associated with Leishmania mexicana infection in mice. Infect Immun 57: 875-881.) into Slc11a1 congenic mice, suggesting that the genetic influence of SLC11A1 on susceptibility to CL following natural infection in humans might be mediated by the effect on the wound healing response to the sandfly bite. This means that the mechanism by which SLC11A1 influences CL disease may be different to its influence on VL in mice following intravenous needle injection (Bradley & Kirkley 1977Bradley DJ, Kirkley J 1977. Regulation of Leishmania populations within the host. I. The variable course of Leishmania donovani infections in mice. Clin Exp Immunol 30: 119-129.) or in natural infection of dogs (Sanchez-Robert et al. 2005Sanchez-Robert E, Altet L, Sanchez A, Francino O 2005. Polymorphism of Slc11a1 (Nramp1) gene and canine leishmaniasis in a case-control study. J Hered 96: 755-758., 2008Sanchez-Robert E, Altet L, Utzet-Sadurni M, Giger U, Sanchez A, Francino O 2008. Slc11a1 (formerly Nramp1) and susceptibility to canine visceral leishmaniasis. Vet Res 39: 36.) and humans (Bucheton et al. 2003Bucheton B, Abel L, Kheir MM, Mirgani A, El-Safi SH, Chevillard C, Dessein A 2003. Genetic control of visceral leishmaniasis in a Sudanese population: candidate gene testing indicates a linkage to the NRAMP1 region. Genes Immun 4: 104-109., Mohamed et al. 2004Mohamed HS, Ibrahim ME, Miller EN, White JK, Cordell HJ, Howson JMM, Peacock CS, Khalil EAG, Elhassan AM, Blackwell JM 2004. SLC11A1 (formerly NRAMP1) and susceptibility to visceral leishmaniasis in The Sudan. Eur J Hum Genet 12: 66-74.). Our data supports roles for both CXCR1 and SLC11A1 in determining the outcome of L. braziliensis infection, providing interesting insight into the possible roles of PMN and macrophages in ACL.
The wound healing gene hypothesis: studies inspired by mice - Our observations on the possible role of wound healing genes in response to sandfly delivered parasites were not the first to suggest a possible role for wound healing genes in CL susceptibility. Indeed, our interpretation was based largely on the seminal mapping studies of susceptibility to CL carried out in mice (Sakthianandeswaren et al. 2005Sakthianandeswaren A, Elso CM, Simpson K, Curtis JM, Kumar B, Speed TP, Handman E, Foote SJ 2005. The wound repair response controls outcome to cutaneous leishmaniasis. Proc Natl Acad Sci USA 102: 15551-15556., 2009Sakthianandeswaren A, Foote SJ, Handman E 2009. The role of host genetics in leishmaniasis. Trends Parasitol 25: 383-391., 2010Sakthianandeswaren A, Curtis JM, Elso C, Kumar B, Baldwin TM, Lopaticki S, Kedzierski L, Smyth GK, Foote SJ, Handman E 2010. Fine mapping of Leishmania major susceptibility locus lmr2 and evidence of a role for Fli1 in disease and wound healing. Infect Immun 78: 2734-2744.), which inspired us to look for the possible role of these and other wound healing genes in susceptibility to ACL in Corte de Pedra.
FLI1 - Fine mapping in the region of chromosome 9 in mice (chromosome 11q24 in humans) identified Friend leukaemia virus integration 1 (Fli1) (FLI1 in humans) as a novel candidate influencing both resistance to L. major and an enhanced wound healing response (Sakthianandeswaren et al. 2010Sakthianandeswaren A, Curtis JM, Elso C, Kumar B, Baldwin TM, Lopaticki S, Kedzierski L, Smyth GK, Foote SJ, Handman E 2010. Fine mapping of Leishmania major susceptibility locus lmr2 and evidence of a role for Fli1 in disease and wound healing. Infect Immun 78: 2734-2744.). To determine whether polymorphisms at FLI1 were important in human disease, SNPs that tagged the first two major linkage disequilibrium blocks and the proximal promoter of the FLI1 gene were analysed in 325 endemic L. braziliensis families (Castellucci et al. 2011Castellucci L, Jamieson SE, Miller EN, de Almeida LF, Oliveira J, Magalhães A, Guimarães LH, Lessa M, Lago E, de Jesus AR, Carvalho EM, Blackwell JM 2011. FLI1 polymorphism affects susceptibility to cutaneous leishmaniasis in Brazil. Genes Immun 12: 589-594.). The proximal promoter region of FLI1 contains a functional GAn microsatellite, as well as a CpG island that spans the proximal promoter region and the 5′ region of intron. Using robust case-pseudocontrol conditional logistic regression analysis of discovery (OR = 1.65, 95% CI = 1.18-2.29, p = 0.003) and replication (OR = 1.60, 95% CI = 1.10-2.33, p = 0.014) family-based cohorts, we demonstrated association between FLI1 (rs7930515; Pcombined = 1.8 x 10-4) and susceptibility to CL caused by L. braziliensis (Castellucci et al. 2011Castellucci L, Jamieson SE, Miller EN, de Almeida LF, Oliveira J, Magalhães A, Guimarães LH, Lessa M, Lago E, de Jesus AR, Carvalho EM, Blackwell JM 2011. FLI1 polymorphism affects susceptibility to cutaneous leishmaniasis in Brazil. Genes Immun 12: 589-594.). In the murine study, resistance to L. major correlated with a wound-healing response that presented in congenic resistant mice as a large population of fibroblasts and an organised and abundant deposition of collagen bundles in the absence of inflammatory cells (Sakthianandeswaren et al. 2005Sakthianandeswaren A, Elso CM, Simpson K, Curtis JM, Kumar B, Speed TP, Handman E, Foote SJ 2005. The wound repair response controls outcome to cutaneous leishmaniasis. Proc Natl Acad Sci USA 102: 15551-15556.). Recent studies have shown an association between enhanced type I collagen expression and epigenetic repression of the FLI1 gene (Wang et al. 2006Wang Y, Fan PS, Kahaleh B 2006. Association between enhanced type I collagen expression and epigenetic repression of the FLI1 gene in scleroderma fibroblasts. Arthritis Rheum 54: 2271-2279.). As reviewed above, our group also reported an association between ML and the C allele at the IL6-174 bp G/C promoter polymorphism (Castellucci et al. 2006Castellucci L, Menezes E, Oliveira J, Magalhães A, Guimarães LH, Lessa M, Ribeiro S, Reale J, Noronha EF, Wilson ME, Duggal P, Beaty TH, Jeronimo S, Jamieson SE, Bales A, Blackwell JM, de Jesus AR, Carvalho EM 2006. IL6 -174 G/C promoter polymorphism influences susceptibility to mucosal but not localized cutaneous leishmaniasis in Brazil. J Infect Dis 194: 519-527.), which determines low levels of IL-6 release from macrophages. Homocysteine dependent stimulation of IL-6 has recently been reported (Thaler et al. 2011Thaler R, Agsten M, Spitzer S, Paschalis EP, Karlic H, Klaushofer K, Varga F 2011. Homocysteine suppresses the expression of the collagen cross-linker lysyl oxidase involving IL-6, Fli1 and epigenetic DNA methylation. J Biol Chem 286: 5578-5588.) to upregulate genes essential for epigenetic DNA methylation via expression of FLI1. Homocysteine increases the CpG methylation status (and hence represses gene expression) of the CpG-rich proximal promoter of the lysyl oxidase (LOX) gene (Thaler et al. 2011Thaler R, Agsten M, Spitzer S, Paschalis EP, Karlic H, Klaushofer K, Varga F 2011. Homocysteine suppresses the expression of the collagen cross-linker lysyl oxidase involving IL-6, Fli1 and epigenetic DNA methylation. J Biol Chem 286: 5578-5588.), an extra-cellular copper enzyme that initiates the cross-linking of collagens and elastins. Inhibition of IL-6 reverses this repression. Regulation of collagen expression and organisation may thus involve epigenetic regulation at both FLI1 and LOX genes, consistent with the presence of the CpG island across the region of the functional FLI1 promoter elements. This suggests that, although there are many immune-related functions for both IL-6 and FLI1 that could account for association with CL caused by L. braziliensis, there may be a direct functional link between these two genes that mediates resistance or susceptibility to infection through the wound-healing response. This, in turn, might provide novel therapeutic opportunities.
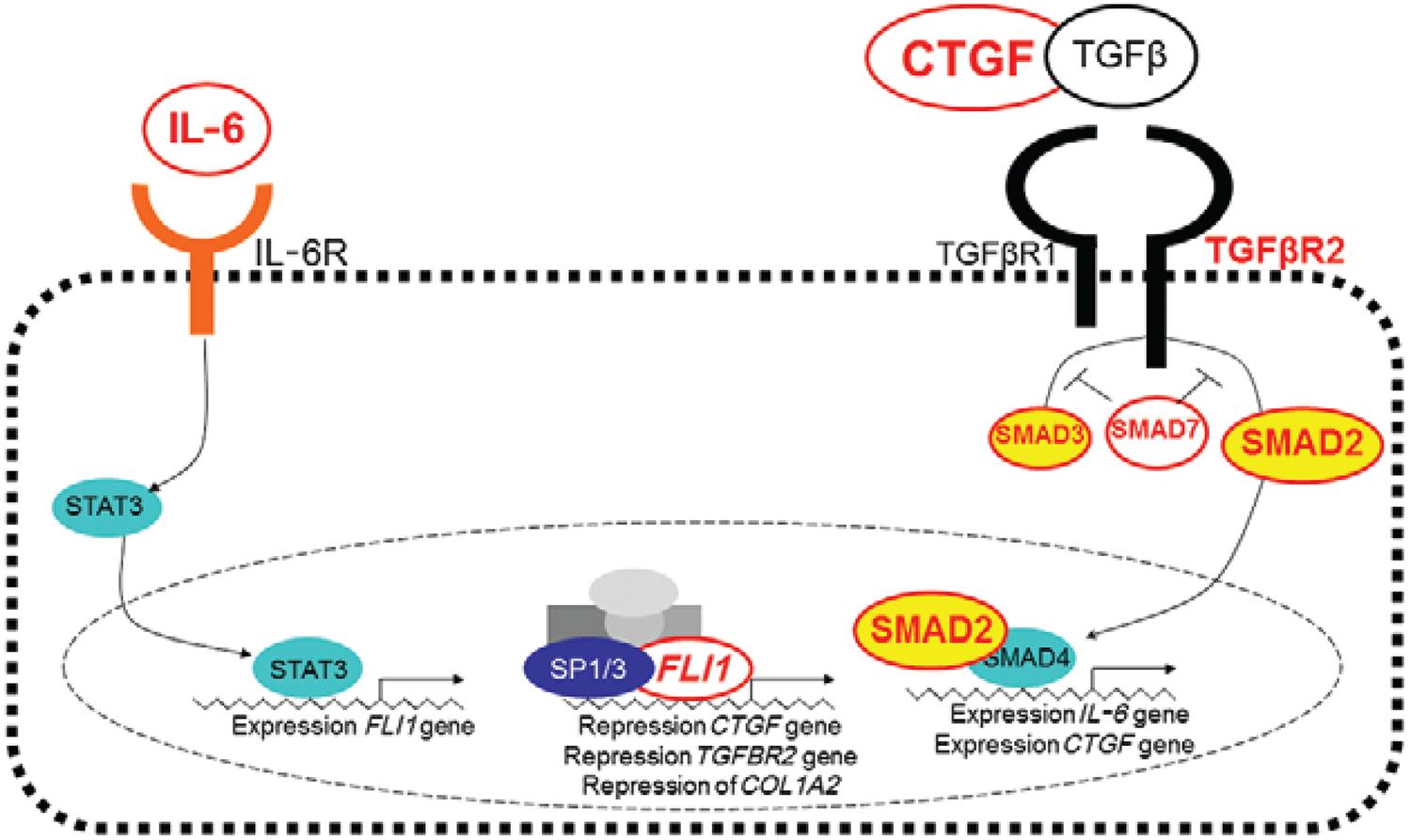
Transforming growth factor β (TGFβ) signalling pathway - IL-6 is known to increase expression of FLI1 (Thaler et al. 2011Thaler R, Agsten M, Spitzer S, Paschalis EP, Karlic H, Klaushofer K, Varga F 2011. Homocysteine suppresses the expression of the collagen cross-linker lysyl oxidase involving IL-6, Fli1 and epigenetic DNA methylation. J Biol Chem 286: 5578-5588.). In the wound healing response, both FLI1 (Nakerakanti et al. 2006Nakerakanti SS, Kapanadze B, Yamasaki M, Markiewicz M, Trojanowska M 2006. Fli1 and Ets1 have distinct roles in connective tissue growth factor/CCN2 gene regulation and induction of the profibrotic gene program. J Biol Chem 281: 25259-25269.) and IL-6 (Gressner et al. 2011Gressner OA, Peredniene I, Gressner AM 2011. Connective tissue growth factor reacts as an IL-6/STAT3-regulated hepatic negative acute phase protein. World J Gastroenterol 17: 151-163.) repress connective tissue growth factor (CTGF) and all three genes interact with the TGFβ pathway. We therefore interrogated further the possible roles of wound healing pathways in cutaneous forms of leishmaniasis caused by L. braziliensis by looking for genetic associations with polymorphisms in other genes through interaction with FLI1 and the TGFβ signalling pathway (Castellucci et al. 2012Castellucci L, Jamieson SE, Almeida L, Oliveira J, Guimarães LH, Lessa M, Fakiola M, de Jesus AR, Miller EN, Carvalho EM, Blackwell JM 2012. Wound healing genes and susceptibility to cutaneous leishmaniasis in Brazil. Infect Genet Evol 12: 1102-1110.). Robust case-pseudocontrol conditional logistic regression analysis showed associations between CL and SNPs at CTGF (rs6918698, OR = 1.67, 95% CI = 1.10-2.54, p = 0.016), TGFBR2 (rs1962859, OR = 1.50, 95% CI = 1.12-1.99, p = 0.005), SMAD2 (rs1792658, OR = 1.57, 95% CI = 1.04-2.38, p = 0.03), SMAD7 (rs4464148, OR = 2.80, 95% CI = 1.00-7.87, p = 0.05) and FLII (rs2071242, OR = 1.60, 95% CI = 1.14-2.24, p = 0.005) and between ML and SNPs at SMAD3 (rs1465841, OR = 2.15, 95% CI = 1.13-4.07, p = 0.018) and SMAD7 (rs2337107, OR = 3.70, 95% CI = 1.27-10.7, p = 0.016). There is a complex interplay between FLI1 and the TGFb signalling pathway in regulating collagen deposition and fibrosis during the wound healing process. In looking for genetic associations that might throw light on how those genes are influencing the wound healing processes important in CL vs. ML disease caused by L. braziliensis, our results indicate that CTGF regulated via the SMAD2 arm of the TGFβ signalling pathway is required for wound healing in CL disease. In contrast, ML disease was associated with polymorphism in SMAD3, suggesting that alternative regulation of gene expression via the TGFβ signalling pathway may lead to ML disease. Fig. 2 provides a model for how polymorphisms at genes regulating the different signalling pathways might influence CL and ML disease. Further functional data will be required to determine what the downstream events following signalling via SMAD3 in ML compared to signalling via SMAD2 for CL disease might be. Additionally, both forms of disease were influenced by polymorphisms in the negative regulator SMAD7 that blocks the TGFβ pathway upstream of both SMAD2 and SMAD3 emphasising the relevance of TGFβ signalling on ACL.
: diagram of genes that have been implicated in susceptibility to cutaneous leishmaniasis (CL) and mucosal leishmaniasis (ML) disease caused by Leishmania braziliensis in the area of Corte de Pedra, state of Bahia, Brazil, showing involvement of, and interaction with, the transforming growth factor β (TGFβ) pathway. Polymorphisms in genes annotated in red lettering have been associated with CL or ML disease. Turquoise circles indicate the pathway through which interleukin (IL)-6 influences SMAD4 via FLI1. SP1/3 are transcription factors that influence FLI1 expression. CTGF: connective tissue growth factor. Source: Castellucci et al. (2012).
Leishmania infection is associated with a broad spectrum of clinical phenotypes. L. braziliensis, in particular, causes debilitating and disfiguring CL, ML and DL that generally take a long time to heal. For over 50 years, pentavalent antimony (Sbv) given by the intramuscular or intravenous route remained the first-line drug for the treatment of ACL. This therapy can cause toxic side effects and is difficult to administer in poor rural areas (Machado et al. 2010Machado PR, Ampuero J, Guimarães LH, Villasboas L, Rocha AT, Schriefer A, Sousa RS, Talhari A, Penna G, Carvalho EM 2010. Miltefosine in the treatment of cutaneous leishmaniasis caused by Leishmania braziliensis in Brazil: a randomized and controlled trial. PLoS Negl Trop Dis 4: e912.). In Corte de Pedra, cure rates after Sbv therapy are becoming increasingly lower and vary from 50-90% (Romero et al. 2001Romero GA, Guerra MV, Paes MG, Macedo VO 2001. Comparison of cutaneous leishmaniasis due to Leishmania (Viannia) brazi- liensis and L. (V.) guyanensis in Brazil: therapeutic response to meglumine antimoniate. Am J Trop Med Hyg 65: 456-465., Unger et al. 2009Unger A, O’neal S, Machado PR, Guimarães LH, Morgan DJ, Schriefer A, Bacellar O, Glesby MJ, Carvalho EM 2009. Association of treatment of American cutaneous leishmaniasis prior to ulcer development with high rate of failure in northeastern Brazil. Am J Trop Med Hyg 80: 574-579.). In light of this, identifying important pathways/mechanisms of disease can lead to new therapeutic targets and more efficient intervention strategies that aim to increase adherence to treatment in areas with limited access to health services. Genetic studies in humans provide a potentially powerful route to understanding novel pathways of disease pathogenesis that could provide new chemotherapeutic targets.
Whilst broadly driven by parasite species, many studies have implicated host genetics in determining the outcome of infection within each species (El-Safi et al. 2006El-Safi S, Kheir MM, Bucheton B, Argiro L, Abel L, Dereure J, Dedet JP, Dessein A 2006. Genes and environment in susceptibility to visceral leishmaniasis. C R Biol 329: 863-870., Lipoldova & Demant 2006Lipoldova M, Demant P 2006. Genetic susceptibility to infectious disease: lessons from mouse models of leishmaniasis. Nat Rev Genet 7: 294-305., Blackwell et al. 2009Blackwell JM, Fakiola M, Ibrahim ME, Jamieson SE, Jeronimo SB, Miller EN, Mishra A, Mohamed HS, Peacock CS, Raju M, Sundar S, Wilson ME 2009. Genetics and visceral leishmaniasis: of mice and man. Parasite Immunol 31: 254-266., Sakthianandeswaren et al. 2009Sakthianandeswaren A, Foote SJ, Handman E 2009. The role of host genetics in leishmaniasis. Trends Parasitol 25: 383-391.). Nevertheless, the only definitive study carried out in humans to date was the recent genome-wide association study on VL (Fakiola et al. 2013Fakiola M, Strange A, Cordell HJ, Miller EN, Pirinen M, Su Z, Mishra A, Mehrotra S, Monteiro GR, Band G, Bellenguez C, Dronov S, Edkins S, Freeman C, Giannoulatou E, Gray E, Hunt SE, Lacerda HG, Langford C, Pearson R, Pontes NN, Rai M, Singh SP, Smith L, Sousa O, Vukcevic D, Bramon E, Brown MA, Casas JP, Corvin A, Duncanson A, Jankowski J, Markus HS, Mathew CG, Palmer CN, Plomin R, Rautanen A, Sawcer SJ, Trembath RC, Viswanathan AC, Wood NW, Wilson ME, Deloukas P, Peltonen L, Christiansen F, Witt C, Jeronimo SM, Sundar S, Spencer CC, Blackwell JM, Donnelly P 2013. Common variants in the HLA-DRB1-HLA-DQA1 HLA class II region are associated with susceptibility to visceral leishmaniasis. Nat Genet 45: 208-213.), which demonstrated that polymorphisms within the DRB1-DQA1 class II region of human leukocyte antigen were the only SNPs to attain genome-wide significance. Remarkably, this finding crossed the epidemiological divide of parasite species (Leishmania donovani and Leishmania chagasi) and geography (Indian and Brazil) and has important implications for the development of molecularly defined vaccines. While candidate gene studies (Table II) have implicated a broader array of genes in susceptibility to CL, these are compromised by lack of power and failure to obtain replication within and between populations. Large well-powered genome-wide studies with replication will be required to evaluate the real significance of these findings. It is of interest, nevertheless, that our studies of ACL have provided evidence in support of important roles for immune response genes involved in wound healing, which are underpinned by initial genetic studies in murine models of disease. These wound healing genes may provide novel therapeutic opportunities in ACL, not the least because there may already be great interest in the same genes as therapeutic targets for other skin disorders. For example, the use of imatinib mesylate has been proposed for treatment of systemic sclerosis (Asano 2010Asano Y 2010. Future treatments in systemic sclerosis. J Dermatol 37: 54-70., Asano et al. 2010Asano Y, Bujor AM, Trojanowska M 2010. The impact of Fli1 deficiency on the pathogenesis of systemic sclerosis. J Dermatol Sci 59: 153-162.), an autoimmune disorder similarly resulting from immune activation, fibrosis development and damage of small blood vessels, in which FLI1 is down regulated through an epigenetic mechanism (Asano et al. 2010Asano Y, Bujor AM, Trojanowska M 2010. The impact of Fli1 deficiency on the pathogenesis of systemic sclerosis. J Dermatol Sci 59: 153-162.). Imatinib mesylate reverses the expression levels of FLI1. Similar opportunities might apply in the case of other genes that we have demonstrated are associated with the spectrum of ACL disease. Work is in progress to analyse expression levels of FLI1 and other wound healing genes in tissue biopsies from L. braziliensis patients to determine their potential as therapeutic targets, along with plans to undertake well-powered genome-wide association studies to validate our genetic findings for this important tropical infectious disease.
REFERENCES
- Ajdary S, Ghamilouie MM, Alimohammadian MH, Hosseini M, Pakzad SR 2010. Lack of association of Toll-like receptor 2 Arg753Gln with cutaneous leishmaniasis. Parasitol Int 59: 466-468.
- Ajdary S, Ghamilouie MM, Alimohammadian MH, Riazi-Rad F, Pakzad SR 2011. Toll-like receptor 4 polymorphisms predispose to cutaneous leishmaniasis. Microbes Infect 13: 226-231.
- Alcais A, Abel L, David C, Torrez ME, Flandre P, Dedet JP 1997. Evidence for a major gene controlling susceptibility to tegumentary leishmaniasis in a recently exposed Bolivian population. Am J Hum Genet 61: 968-979.
- Alexander J, Blackwell JM 1986. The immunological significance of genetically determined cross reactivity between taxonomically distinct Leishmania species. In JA Riou (ed.), Leishmania, taxonomie et phylogenese: application éco-épidemiologiques, Institut Méditerranée d’Etudes Epidémiologiques et Ecologiques, Montpellier, p. 185-191.
- Asano Y 2010. Future treatments in systemic sclerosis. J Dermatol 37: 54-70.
- Asano Y, Bujor AM, Trojanowska M 2010. The impact of Fli1 deficiency on the pathogenesis of systemic sclerosis. J Dermatol Sci 59: 153-162.
- Bacellar O, Lessa H, Schriefer A, Machado P, de Jesus AR, Dutra WO, Gollob KJ, Carvalho EM 2002. Up-regulation of Th1-type responses in mucosal leishmaniasis patients. Infect Immun 70: 6734-6740.
- Barbier D, Demenais F, Lefait JF, David B, Blanc M, Hors J, Feingold N 1987. Susceptibility to human cutaneous leishmaniasis and HLA, Gm, Km markers. Tissue Antigens 30: 63-67.
- Barral-Netto M, Machado P, Bittencourt A, Barral A 1997. Recent advances in the pathophysiology and treatment of human cutaneous leishmaniasis. Curr Opin Dermatol 4: 51-58.
- Bermudez LE, Wu M, Petrofsky M, Young LS 1992. Interleukin-6 antagonizes tumor necrosis factor-mediated mycobacteriostatic and mycobactericidal activities in macrophages. Infect Immun 60: 4245-4252.
- Bhattacharyya S, Ghosh S, Dasgupta B, Mazumder D, Roy S, Majumdar S 2002. Chemokine-induced leishmanicidal activity in murine macrophages via the generation of nitric oxide. J Infect Dis 185: 1704-1708.
- Bidwell J, Keen L, Gallagher G, Kimberly R, Huizinga T, Mcdermott MF, Oksenberg J, Mcnicholl J, Pociot F, Hardt C, D’Alfonso S 1999. Cytokine gene polymorphism in human disease: on-line databases. Genes Immun 1: 3-19.
- Blackwell JM 1998. Genetics of host resistance and susceptibility to intramacrophage pathogens: a study of multicase families of tuberculosis, leprosy and leishmaniasis in north-eastern Brazil. Int J Parasitol 28: 21-28.
- Blackwell JM, Black GF, Peacock CS, Miller EN, Sibthorpe D, Gnananandha D, Shaw JJ, Silveira F, Lins-Lainson Z, Ramos F, Collins A, Shaw M-A 1997. Immunogenetics of leishmanial and mycobacterial infections: the Belém family study. Philos Trans R Soc Lond Biol Sci 352: 1331-1345.
- Blackwell JM, Fakiola M, Ibrahim ME, Jamieson SE, Jeronimo SB, Miller EN, Mishra A, Mohamed HS, Peacock CS, Raju M, Sundar S, Wilson ME 2009. Genetics and visceral leishmaniasis: of mice and man. Parasite Immunol 31: 254-266.
- Blackwell JM, Goswami T, Evans CAW, Sibthorpe D, Papo N, White JK, Searle S, Miller EN, Peacock CS, Mohammed H, Ibrahim M 2001. SLC11A1 (formerly NRAMP1) and disease. Cell Microbiol 3: 773-784.
- Bradley DJ, Kirkley J 1977. Regulation of Leishmania populations within the host. I. The variable course of Leishmania donovani infections in mice. Clin Exp Immunol 30: 119-129.
- Bucheton B, Abel L, Kheir MM, Mirgani A, El-Safi SH, Chevillard C, Dessein A 2003. Genetic control of visceral leishmaniasis in a Sudanese population: candidate gene testing indicates a linkage to the NRAMP1 region. Genes Immun 4: 104-109.
- Cabrera M, Shaw M-A, Sharples C, Williams H, Castes M, Convit J, Blackwell JM 1995. Polymorphism in TNF genes associated with mucocutaneous leishmaniasis. J Exp Med 182: 1259-1264.
- Carvalho LP, Passos S, Schriefer A, Carvalho EM 2012. Protective and pathologic immune responses in human tegumentary leishmaniasis. Front Immunol 3: 301.
- Castellucci L, Cheng LH, Araújo C, Guimarães LH, Lessa H, Machado P, Almeida MF, Oliveira A, Ko A, Johnson WD, Wilson ME, Carvalho EM, de Jesus AR 2005. Familial aggregation of mucosal leishmaniasis in Northeast Brazil. Am J Trop Med Hyg 73: 69-73.
- Castellucci L, Jamieson SE, Almeida L, Oliveira J, Guimarães LH, Lessa M, Fakiola M, de Jesus AR, Miller EN, Carvalho EM, Blackwell JM 2012. Wound healing genes and susceptibility to cutaneous leishmaniasis in Brazil. Infect Genet Evol 12: 1102-1110.
- Castellucci L, Jamieson SE, Miller EN, de Almeida LF, Oliveira J, Magalhães A, Guimarães LH, Lessa M, Lago E, de Jesus AR, Carvalho EM, Blackwell JM 2011. FLI1 polymorphism affects susceptibility to cutaneous leishmaniasis in Brazil. Genes Immun 12: 589-594.
- Castellucci L, Jamieson SE, Miller EN, Menezes E, Oliveira J, Magalhães A, Guimarães LH, Lessa M, de Jesus AR, Carvalho EM, Blackwell JM 2010. CXCR1 and SLC11A1 polymorphisms affect susceptibility to cutaneous leishmaniasis in Brazil: a case-control and family-based study. BMC Med Genet 11: 10.
- Castellucci L, Menezes E, Oliveira J, Magalhães A, Guimarães LH, Lessa M, Ribeiro S, Reale J, Noronha EF, Wilson ME, Duggal P, Beaty TH, Jeronimo S, Jamieson SE, Bales A, Blackwell JM, de Jesus AR, Carvalho EM 2006. IL6 -174 G/C promoter polymorphism influences susceptibility to mucosal but not localized cutaneous leishmaniasis in Brazil. J Infect Dis 194: 519-527.
- Covas CJF, Cardoso CC, Gomes-Silva A, Oliveira JRS, da-Cruz AM, Moraes MO 2013. Candidate gene case-control and functional study shows macrophage inhibitory factor (MIF) polymorphism is associated with cutaneous leishmaniasis. Cytokine 61: 168-172.
- de Moura TR, Novais FO, Oliveira F, Clarencio J, Noronha A, Barral A, Brodskyn C, de Oliveira CI 2005. Toward a novel experimental model of infection to study American cutaneous leishmaniasis caused by Leishmania braziliensis Infect Immun 73: 5827-5834.
- de Oliveira CI, Brodskyn CI 2012. The immunobiology of Leishmania braziliensis infection. Front Immunol 3: 145.
- Diehl S, Anguita J, Hoffmeyer A, Zapton T, Ihle JN, Fikrig E, Rincon M 2000. Inhibition of Th1 differentiation by IL-6 is mediated by SOCS1. Immunity 13: 805-815.
- Diehl S, Rincon M 2002. The two faces of IL-6 on Th1/Th2 differentiation. Mol Immunol 39: 531-536.
- El-Mogy MH, Abdel-Hamid IA, Abdel-Razic MM, Rizk RA, Romia SA 1993. Histocompatibility antigens in Egyptians with cutaneous leishmaniasis: a preliminary study. J Dermatol Sci 5: 89-91.
- El-Safi S, Kheir MM, Bucheton B, Argiro L, Abel L, Dereure J, Dedet JP, Dessein A 2006. Genes and environment in susceptibility to visceral leishmaniasis. C R Biol 329: 863-870.
- Fakiola M, Strange A, Cordell HJ, Miller EN, Pirinen M, Su Z, Mishra A, Mehrotra S, Monteiro GR, Band G, Bellenguez C, Dronov S, Edkins S, Freeman C, Giannoulatou E, Gray E, Hunt SE, Lacerda HG, Langford C, Pearson R, Pontes NN, Rai M, Singh SP, Smith L, Sousa O, Vukcevic D, Bramon E, Brown MA, Casas JP, Corvin A, Duncanson A, Jankowski J, Markus HS, Mathew CG, Palmer CN, Plomin R, Rautanen A, Sawcer SJ, Trembath RC, Viswanathan AC, Wood NW, Wilson ME, Deloukas P, Peltonen L, Christiansen F, Witt C, Jeronimo SM, Sundar S, Spencer CC, Blackwell JM, Donnelly P 2013. Common variants in the HLA-DRB1-HLA-DQA1 HLA class II region are associated with susceptibility to visceral leishmaniasis. Nat Genet 45: 208-213.
- Fernández-Figueroa EA, Rangel-Escareno C, Espinosa-Mateos V, Carrillo-Sanchez K, Salaiza-Suazo N, Carrada-Figueroa G, March-Mifsut S, Becker I 2012. Disease severity in patients infected with Leishmania mexicana relates to IL-1beta. PLoS Negl Trop Dis 6: e1533.
- Fishman D, Faulds G, Jeffery R, Mohamed-Ali V, Yudkin JS, Humphries S, Woo P 1998. The effect of novel polymorphisms in the interleukin-6 (IL-6) gene on IL-6 transcription and plasma IL-6 levels and an association with systemic-onset juvenile chronic arthritis. J Clin Invest 102: 1369-1376.
- Flores-Villanueva PO, Ruiz-Morales JA, Song CH, Flores LM, Jo EK, Montano M, Barnes PF, Selman M, Granados J 2005. A functional promoter polymorphism in monocyte chemoattractant protein-1 is associated with increased susceptibility to pulmonary tuberculosis. J Exp Med 202: 1649-1658.
- Gillespie RD, Mbow ML, Titus RG 2000. The immunomodulatory factors of blood-feeding arthropod saliva. Parasite Immunol 22: 319-331.
- Gressner OA, Peredniene I, Gressner AM 2011. Connective tissue growth factor reacts as an IL-6/STAT3-regulated hepatic negative acute phase protein. World J Gastroenterol 17: 151-163.
- Grimaldi Jr G, Tesh RB 1993. Leishmaniases of the New World: current concepts and implications for future research. Clin Microbiol Rev 6: 230-250.
- Hatzigeorgiou DE, He S, Sobel J, Grabstein KH, Hafner A, Ho JL 1993. IL-6 down-modulates the cytokine-enhanced antileishmanial activity in human macrophages. J Immunol 151: 3682-3692.
- Intemann CD, Thye T, Forster B, Owusu-Dabo E, Gyapong J, Horstmann RD, Meyer CG 2011. MCP1 haplotypes associated with protection from pulmonary tuberculosis. BMC Genet 12: 34.
- Jeronimo SM, Teixeira MJ, Sousa A, Thielking P, Pearson RD, Evans TG 2000. Natural history of Leishmania (Leishmania) chagasi infection in northeastern Brazil: long-term follow-up. Clin Infect Dis 30: 608-609.
- Kamali-Sarvestani E, Rasouli M, Mortazavi H, Gharesi-Fard B 2006. Cytokine gene polymorphisms and susceptibility to cutaneous leishmaniasis in Iranian patients. Cytokine 35: 159-165.
- Karplus TM, Jeronimo SM, Chang H, Helms BK, Burns TL, Murray JC, Mitchell AA, Pugh EW, Braz RF, Bezerra FL, Wilson ME 2002. Association between the tumor necrosis factor locus and the clinical outcome of Leishmania chagasi infection. Infect Immun 70: 6919-6925.
- Lara ML, Layrisse Z, Scorza JV, Garcia E, Stoikow Z, Granados J, Bias W 1991. Immunogenetics of human American cutaneous leishmaniasis. Study of HLA haplotypes in 24 families from Venezuela. Hum Immunol 30: 129-135.
- Lipoldova M, Demant P 2006. Genetic susceptibility to infectious disease: lessons from mouse models of leishmaniasis. Nat Rev Genet 7: 294-305.
- Machado PR, Ampuero J, Guimarães LH, Villasboas L, Rocha AT, Schriefer A, Sousa RS, Talhari A, Penna G, Carvalho EM 2010. Miltefosine in the treatment of cutaneous leishmaniasis caused by Leishmania braziliensis in Brazil: a randomized and controlled trial. PLoS Negl Trop Dis 4: e912.
- Matos GI, Covas CJ, Bittar RC, Gomes-Silva A, Marques F, Maniero VC, Amato VS, Oliveira-Neto MP, Mattos MS, Pirmez C, Sampaio EP, Moraes MO, da-Cruz AM 2007. IFNG +874T/A polymorphism is not associated with American tegumentary leishmaniasis susceptibility but can influence Leishmania induced IFN-gamma production. BMC Infect Dis 7: 33.
- Mohamed HS, Ibrahim ME, Miller EN, White JK, Cordell HJ, Howson JMM, Peacock CS, Khalil EAG, Elhassan AM, Blackwell JM 2004. SLC11A1 (formerly NRAMP1) and susceptibility to visceral leishmaniasis in The Sudan. Eur J Hum Genet 12: 66-74.
- Nakerakanti SS, Kapanadze B, Yamasaki M, Markiewicz M, Trojanowska M 2006. Fli1 and Ets1 have distinct roles in connective tissue growth factor/CCN2 gene regulation and induction of the profibrotic gene program. J Biol Chem 281: 25259-25269.
- Oliveira CR, Pereira LI, Pereira AJ, Ferreira AA, Crespo AM, Silveira LA 2011. Allelic polymorphism of human FcgammaRIIA-H/R131 receptor in American tegumentary leishmaniasis. Int J Immunogenet 38: 225-231.
- Olivo-Diaz A, Debaz H, Alaez C, Islas VJ, Perez-Perez H, Hobart O, Gorodezky C 2004. Role of HLA class II alleles in susceptibility to and protection from localized cutaneous leishmaniasis. Hum Immunol 65: 255-261.
- Peters NC, Sacks DL 2009. The impact of vector mediated neutrophil recruitment on cutaneous leishmaniasis. Cell Microbiol 11: 1290-1296.
- Petzl-Erler ML, Belich MP, Queiroz-Telles F 1991. Association of mucosal leishmaniasis with HLA. Human Immunol 32: 254-260.
- Queiroz A, Sousa R, Heine C, Cardoso M, Guimarães LH, Machado PR, Carvalho EM, Riley LW, Wilson ME, Schriefer A 2012. Association between an emerging disseminated form of leishmaniasis and Leishmania (Viannia) braziliensis strain polymorphisms. J Clin Microbiol 50: 4028-4034.
- Ramasawmy R, Menezes E, Magalhães A, Oliveira J, Castellucci L, Almeida R, Rosa ME, Guimarães LH, Lessa M, Noronha E, Wilson ME, Jamieson SE, Kalil J, Blackwell JM, Carvalho EM, de Jesus AR 2010. The -2518 bp promoter polymorphism at CCL2/MCP1 influences susceptibility to mucosal but not localized cutaneous leishmaniasis in Brazil. Infect Genet Evol 10: 607-613.
- Ribeiro-de-Jesus A, Almeida RP, Lessa H, Bacellar O, Carvalho EM 1998. Cytokine profile and pathology in human leishmaniasis. Braz J Med Biol Res 31: 143-148.
- Rincon M, Anguita J, Nakamura T, Fikrig E, Flavell RA 1997. Interleukin (IL)-6 directs the differentiation of IL-4-producing CD4+ T cells. J Exp Med 185: 461-469.
- Ritter U, Moll H 2000. Monocyte chemotactic protein-1 stimulates the killing of Leishmania major by human monocytes, acts synergistically with IFN-gamma and is antagonized by IL-4. Eur J Immunol 30: 3111-3120.
- Ritter U, Moll H, Laskay T, Brocker E, Velazco O, Becker I, Gillitzer R 1996. Differential expression of chemokines in patients with localized and diffuse cutaneous American leishmaniasis. J Infect Dis 173: 699-709.
- Roberts M, Alexander J, Blackwell JM 1989. Influence of Lsh, H-2 and an H-11-linked gene on visceralization and metastasis associated with Leishmania mexicana infection in mice. Infect Immun 57: 875-881.
- Romero GA, Guerra MV, Paes MG, Macedo VO 2001. Comparison of cutaneous leishmaniasis due to Leishmania (Viannia) brazi- liensis and L. (V.) guyanensis in Brazil: therapeutic response to meglumine antimoniate. Am J Trop Med Hyg 65: 456-465.
- Sakthianandeswaren A, Curtis JM, Elso C, Kumar B, Baldwin TM, Lopaticki S, Kedzierski L, Smyth GK, Foote SJ, Handman E 2010. Fine mapping of Leishmania major susceptibility locus lmr2 and evidence of a role for Fli1 in disease and wound healing. Infect Immun 78: 2734-2744.
- Sakthianandeswaren A, Elso CM, Simpson K, Curtis JM, Kumar B, Speed TP, Handman E, Foote SJ 2005. The wound repair response controls outcome to cutaneous leishmaniasis. Proc Natl Acad Sci USA 102: 15551-15556.
- Sakthianandeswaren A, Foote SJ, Handman E 2009. The role of host genetics in leishmaniasis. Trends Parasitol 25: 383-391.
- Salhi A, Rodrigues Jr V, Santoro F, Dessein H, Romano A, Castellano LR, Sertorio M, Rafati S, Chevillard C, Prata A, Alcais A, Argiro L, Dessein A 2008. Immunological and genetic evidence for a crucial role of IL-10 in cutaneous lesions in humans infected with Leishmania braziliensis J Immunol 180: 6139-6148.
- Samaranayake TN, Fernando SD, Dissanayake VH 2010. Candidate gene study of susceptibility to cutaneous leishmaniasis in Sri Lanka. Trop Med Int Health 15: 632-638.
- Sanchez-Robert E, Altet L, Sanchez A, Francino O 2005. Polymorphism of Slc11a1 (Nramp1) gene and canine leishmaniasis in a case-control study. J Hered 96: 755-758.
- Sanchez-Robert E, Altet L, Utzet-Sadurni M, Giger U, Sanchez A, Francino O 2008. Slc11a1 (formerly Nramp1) and susceptibility to canine visceral leishmaniasis. Vet Res 39: 36.
- Schriefer A, Schriefer AL, Goes-Neto A, Guimarães LH, Carvalho LP, Almeida RP, Machado PR, Lessa HA, de Jesus AR, Riley LW, Carvalho EM 2004. Multiclonal Leishmania braziliensis population structure and its clinical implication in a region of endemicity for American tegumentary leishmaniasis. Infect Immun 72: 508-514.
- Teixeira CR, Teixeira MJ, Gomes RB, Santos CS, Andrade BB, Raffaele-Netto I, Silva JS, Guglielmotti A, Miranda JC, Barral A, Brodskyn C, Barral-Netto M 2005a. Saliva from Lutzomyia longipalpis induces CC chemokine ligand 2/monocyte chemoattractant protein-1 expression and macrophage recruitment. J Immunol 175: 8346-8353.
- Teixeira MJ, Fernandes JD, Teixeira CR, Andrade BB, Pompeu ML, da Silva JS, Brodskyn CI, Barral-Netto M, Barral A 2005b. Distinct Leishmania braziliensis isolates induce different paces of chemokine expression patterns. Infect Immun 73: 1191-1195.
- Teixeira MJ, Teixeira CR, Andrade BB, Barral-Netto M, Barral A 2006. Chemokines in host-parasite interactions in leishmaniasis. Trends Parasitol 22: 32-40.
- Terry CF, Loukaci V, Green FR 2000. Cooperative influence of genetic polymorphisms on interleukin-6 transcriptional regulation. J Biol Chem 275: 18138-18144.
- Thaler R, Agsten M, Spitzer S, Paschalis EP, Karlic H, Klaushofer K, Varga F 2011. Homocysteine suppresses the expression of the collagen cross-linker lysyl oxidase involving IL-6, Fli1 and epigenetic DNA methylation. J Biol Chem 286: 5578-5588.
- Thuraisingam T, Sam H, Moisan J, Zhang Y, Ding A, Radzioch D 2006. Delayed cutaneous wound healing in mice lacking solute carrier 11a1 (formerly Nramp1): correlation with decreased expression of secretory leukocyte protease inhibitor. J Invest Dermatol 126: 890-901.
- Thye T, Nejentsev S, Intemann CD, Browne EN, Chinbuah MA, Gyapong J, Osei I, Owusu-Dabo E, Zeitels LR, Herb F, Horstmann RD, Meyer CG 2009. MCP-1 promoter variant -362C associated with protection from pulmonary tuberculosis in Ghana, West Africa. Hum Mol Genet 18: 381-388.
- Unger A, O’neal S, Machado PR, Guimarães LH, Morgan DJ, Schriefer A, Bacellar O, Glesby MJ, Carvalho EM 2009. Association of treatment of American cutaneous leishmaniasis prior to ulcer development with high rate of failure in northeastern Brazil. Am J Trop Med Hyg 80: 574-579.
- Wang Y, Fan PS, Kahaleh B 2006. Association between enhanced type I collagen expression and epigenetic repression of the FLI1 gene in scleroderma fibroblasts. Arthritis Rheum 54: 2271-2279.
-
Financial support: NIH (AI 30639), LC was supported by NIH/FIC 1 D43 TW007127-01 (UK).
Publication Dates
-
Publication in this collection
27 May 2014 -
Date of issue
June 2014
History
-
Received
23 Jan 2014 -
Accepted
7 Apr 2014