ABSTRACT:
Areas affected by coal mining can be recovered by revegetation with leguminous plants associated with nitrogen-fixing bacteria. This study addressed the isolation and characterization of native nitrogen-fixing bacteria from coalmine wasteland under different vegetation restoration approaches using Macroptilium atropurpureum (DC) Urb and Vicia sativa L. as trap plants. The bacteria were characterized and identified on the basis of 16S rRNA sequences. Additionally, nitrogen-fixing strains were characterized for tolerance to high heavy metal and low pH levels, as well as for their effect on growth, nodulation, and symbiotic efficiency of M. atropurpureum and V. sativa. Soil samples were taken from the rhizosphere of eight areas, between 6 and 20 years under vegetation restoration, in the coal mining area of Candiota, RS-Brazil. The following properties were evaluated: colony characterization on solid “79” culture medium; pH (3.0-9.0) and heavy metal (Cr, Cd, Zn, Cu, and Ni) tolerance; partial sequencing of 16S rRNA gene; presence of nodA and nifH genes and symbiotic efficiency. A total of 115 isolates, i.e., 77 from M. atropurpureum and 38 from V. sativa, were obtained. The tolerance of these isolates is high for a wide range of pH levels and heavy metal contents, and 18 among them were selected for symbiotic efficiency and 16S rRNA sequencing. Inoculated with M. atropurpureum, the strains UFSM-B53, UFSM-B64, and UFSM-B74 had high symbiotic efficiency. The nitrogen-fixing bacteria were classified in the genera Rhizobium, Bradyrhizobium, and Burkholderia. The results indicate the potential of these native rhizospheric bacterial strains as inoculants and biofertilizers for legume species under pH and heavy metal stress in coal mining degraded areas in Southern Brazil.
Keywords:
diversity; 16S rRNA; Rhizobium; Bradyrhizobium; Burkholderia
INTRODUCTION
Coal provides two-thirds of the non-renewable energy resources in Brazil and reserves are 20 and 75 times higher than those of oil and natural gas, respectively. A total of 847.5 billion tons of coal is enough to maintain the current output for 130 years (Aneel, 2008Agência Nacional de Energia Elétrica - Aneel. Atlas de energia elétrica do Brasil. 3. ed. Brasília, DF: Aneel; 2008 [acesso em 15 mai 2012]. Available at: http://www2.aneel.gov.br/arquivos/pdf/atlas3ed.pdf.
http://www2.aneel.gov.br/arquivos/pdf/at...
). In Brazil, the major coal reserves are located in the region of Candiota, in the state of Rio Grande do Sul, with an estimated reserve of one billion tons, which can be extracted by open-cast mining down to a depth of 50 m (Stumpf et al., 2016Stumpf L, Pauletto EA, Pinto LFS. Soil aggregation and root growth of perennial grasses in a constructed clay minesoil. Soil Till Res. 2016;161:71-8. https://doi.org/10.1016/j.still.2016.03.005
https://doi.org/10.1016/j.still.2016.03....
). During mining and coal processing, a significant amount of sterile solid waste and sulfiderich (pyrite, marcasite, esfarelita, arseno-pyrite, galena, and chalcopyrite) tailings are produced. Upon exposure to oxygen, water, and Thiobacillus ferrooxidans, these materials are oxidized and cause acidification, inducing acid mine drainage (AMD). Due to AMD, dissolute minerals increase the solubility of some metals (As, Cd, Cu, Ni, Pb, Zn, Al, Cr, Mn, Mg etc) (Fungaro and Izidoro, 2006Fungaro DA, Izidoro JC. Remediação de drenagem ácida de mina usando zeólitas sintetizadas a partir de cinzas leves de carvão. Quim Nova. 2006;29:735-40. https://doi.org/10.1590/S0100-40422006000400019
https://doi.org/10.1590/S0100-4042200600...
). As a result of this process, the contamination risk of surface and groundwater as well as of the soil increases, and the restoration of these areas become more expensive.
Biological technologies, e.g., bacterial nitrogen fixation, represent efficient and low-cost alternatives for the recovery of mined areas (Valls et al., 2000Valls M, Atrian S, Lorenzo V, Fernández LA. Engineering a mouse metallothionein on the cell surface of Ralstonia eutropha CH34 for immobilization of heavy metals in soil. Nat Biotechnol. 2000;18:661-5. https://doi.org/10.1038/76516
https://doi.org/10.1038/76516...
; Sriprang et al., 2002Sriprang R, Hayashi M, Yamashita M, Ono H, Saeki K, Murooka Y. A novel bioremediation system for heavy metals using the symbiosis between leguminous plant and genetically engineered rhizobia. J Biotechnol. 2002;99:279-93. https://doi.org/10.1016/S0168-1656(02)00219-5
https://doi.org/10.1016/S0168-1656(02)00...
; Sriprang et al., 2003Sriprang R, Hayashi M, Ono H, Takagi M, Hirata K, Murooka Y. Enhanced accumulation of Cd2+ by a Mesorhizobium sp. transformed with a gene from Arabidopsis thaliana coding for phytochelatin synthase. Appl Environ Microb. 2003;69:1791-6. https://doi.org/10.1128/AEM.69.3.1791-1796.2003
https://doi.org/10.1128/AEM.69.3.1791-17...
; Vázquez et al., 2006Vázquez S, Agha R, Granado A, Sarro MJ, Esteban E, Peñalosa JM, Carpena RO. Use of white lupin plant for phytostabilization of Cd and As polluted acid soil. Water Air Soil Poll. 2006;177:349-65. https://doi.org/10.1007/s11270-006-9178-y
https://doi.org/10.1007/s11270-006-9178-...
; Wu et al., 2006Wu CH, Wood TK, Mulchandani A, Chen W. Engineering plant-microbe symbiosis for rhizoremediation of heavy metals. Appl Environ Microb. 2006;72:1129-34. https://doi.org/10.1128/AEM.72.2.1129-1134.2006
https://doi.org/10.1128/AEM.72.2.1129-11...
). Bioremediation technologies are usually based on the biochemical and genetic capacity of microorganisms to interact and survive under adverse environmental conditions (Vázquez et al., 2006Vázquez S, Agha R, Granado A, Sarro MJ, Esteban E, Peñalosa JM, Carpena RO. Use of white lupin plant for phytostabilization of Cd and As polluted acid soil. Water Air Soil Poll. 2006;177:349-65. https://doi.org/10.1007/s11270-006-9178-y
https://doi.org/10.1007/s11270-006-9178-...
). Therefore, the introduction of plants in association with nitrogen-fixing bacteria constitutes a promising biotechnological tool for the recovery of problematic environments and degraded ecosystems (Chen et al., 2003Chen W-M, Moulin L, Bontemps C, Vandamme P, Béna G, Boivin-Masson C. Legume symbiotic nitrogen fixation by β-proteobacteria is widespread in the nature. J Bacteriol. 2003;185:7266-72. https://doi.org/10.1128/JB.185.24.7266-7272.2003
https://doi.org/10.1128/JB.185.24.7266-7...
; Wani et al., 2007Wani PA, Khan MS, Zaidi A. Effect of metal tolerant plant growth promoting Bradyrhizobium sp. (vigna) on growth, symbiosis, seed yield and metal uptake by greengram plants. Chemosphere. 2007;70:36-45. https://doi.org/10.1016/j.chemosphere.2007.07.028
https://doi.org/10.1016/j.chemosphere.20...
; Ferreira et al., 2012Ferreira PAA, Bomfeti CA, Silva Júnior R, Soares BL, Soares CRFS, Moreira FMS. Eficiência simbiótica de estirpes de Cupriavidus necator tolerantes a zinco, cádmio, cobre e chumbo. Pesq Agropec Bras. 2012;47:85-95. https://doi.org/10.1590/S0100-204X2012000100012
https://doi.org/10.1590/S0100-204X201200...
).
Numerous studies have demonstrated the potential of nitrogen-fixing microorganisms isolated from copper, zinc, cadmium, lead, and arsenic mining areas to favor plant growth and enhance soil organic matter (Trannin et al., 2001Trannin ICB, Moreira FMS, Siqueira JO, Lima A. Tolerância de estirpes e isolados de Bradyrhizobium e de Azorhizobium a zinco, cádmio e cobre “in vitro”. Rev Bras Cienc Solo. 2001;25:305-16. https://doi.org/10.1590/S0100-06832001000200007
https://doi.org/10.1590/S0100-0683200100...
; Matsuda et al., 2002Matsuda A, Moreira FMS, Siqueira JO. Tolerância de rizóbios de diferentes procedências ao zinco, cobre e cádmio. Pesq Agropec Bras. 2002;37:343-55. https://doi.org/10.1590/S0100-204X2002000300016
https://doi.org/10.1590/S0100-204X200200...
; Carrasco et al., 2005Carrasco JA, Armario P, Pajuelo E, Burgos A, Caviedes MA, López R, Chamber MA, Palomares AJ. Isolation and characterisation of symbiotically effective Rhizobium resistant to arsenic and heavy metals after the toxic spill at the Aznalco'llar pyrite mine. Soil Biol Biochem. 2005;37:1131-40. https://doi.org/10.1016/j.soilbio.2004.11.015
https://doi.org/10.1016/j.soilbio.2004.1...
; Zhan and Sun, 2011Zhan J, Sun Q. Diversity of free-living nitrogen-fixing microorganisms in wastelands of copper mine tailings during the process of natural ecological restoration. J Environ Sci. 2011;23:476-87. https://doi.org/10.1016/S1001-0742(10)60433-0
https://doi.org/10.1016/S1001-0742(10)60...
; Ferreira et al., 2012Ferreira PAA, Bomfeti CA, Silva Júnior R, Soares BL, Soares CRFS, Moreira FMS. Eficiência simbiótica de estirpes de Cupriavidus necator tolerantes a zinco, cádmio, cobre e chumbo. Pesq Agropec Bras. 2012;47:85-95. https://doi.org/10.1590/S0100-204X2012000100012
https://doi.org/10.1590/S0100-204X201200...
; Klonowska et al., 2012Klonowska A, Chaintreuil C, Tisseyre P, Miché L, Melkonian R, Ducousso M, Laguerre G, Brunel B, Moulin L. Biodiversity of Mimosa pudica rhizobial symbionts (Cupriavidus taiwanensis, Rhizobium mesoamericanum) in New Caledonia and their adaptation to heavy metal-rich soils. FEMS Microbiol Ecol. 2012;81:618-35. https://doi.org/10.1111/j.1574-6941.2012.01393.x
https://doi.org/10.1111/j.1574-6941.2012...
). Native nitrogen-fixing bacteria in soils with high heavy metal contents and low pH are adapted to these stress types. Therefore, the isolation and selection of bacteria under these conditions can provide important information about the best-adapted genotypes for each scenario (Trannin et al., 2001Trannin ICB, Moreira FMS, Siqueira JO, Lima A. Tolerância de estirpes e isolados de Bradyrhizobium e de Azorhizobium a zinco, cádmio e cobre “in vitro”. Rev Bras Cienc Solo. 2001;25:305-16. https://doi.org/10.1590/S0100-06832001000200007
https://doi.org/10.1590/S0100-0683200100...
; Ferreira et al., 2012Ferreira PAA, Bomfeti CA, Silva Júnior R, Soares BL, Soares CRFS, Moreira FMS. Eficiência simbiótica de estirpes de Cupriavidus necator tolerantes a zinco, cádmio, cobre e chumbo. Pesq Agropec Bras. 2012;47:85-95. https://doi.org/10.1590/S0100-204X2012000100012
https://doi.org/10.1590/S0100-204X201200...
; Klonowska et al., 2012Klonowska A, Chaintreuil C, Tisseyre P, Miché L, Melkonian R, Ducousso M, Laguerre G, Brunel B, Moulin L. Biodiversity of Mimosa pudica rhizobial symbionts (Cupriavidus taiwanensis, Rhizobium mesoamericanum) in New Caledonia and their adaptation to heavy metal-rich soils. FEMS Microbiol Ecol. 2012;81:618-35. https://doi.org/10.1111/j.1574-6941.2012.01393.x
https://doi.org/10.1111/j.1574-6941.2012...
). In addition, legume inoculation with efficient strains, well-adapted to these conditions, is extremely important both from the ecological and economic point of view (Franco and Faria, 1997Franco AA, Faria SM. The contribution of N2-fixing tree legumes to land reclamation and sustainability in the tropics. Soil Biol Biochem. 1997;29:897-903. https://doi.org/10.1016/S0038-0717(96)00229-5
https://doi.org/10.1016/S0038-0717(96)00...
).
A wide range of prokaryotic microorganisms with broad morphological, physiological, genetic, biochemical, and phylogenetic diversity perform N fixation. This diversity not only ensures the resilience of biological N fixation in ecosystems, but also facilitates it in the most diverse terrestrial habitats (Moreira and Siqueira, 2006Moreira FMS, Siqueira JO. Microbiologia e bioquímica do solo. 2a ed atual e rev. Lavras: Editora UFLA; 2006.; Moreira et al., 2010MoreiraFMS, Huising EJ, Bignell DE. Manual de biologia dos solos tropicais: amostragem e caracterização da biodiversidade. Lavras: Editora UFLA; 2010.). Native nitrogen-fixing bacteria of mine-degraded areas are probably better adapted to the local soil and environmental conditions than non-native populations. However, only one study by Brazilian authors described the isolation, characterization, and symbiotic efficiency of nitrogen-fixing bacteria in another coal mining area, in the state of Santa Catarina, with heavy metal contamination and acidic pH (3.5) (Moura et al., 2016Moura GGD, Armas RD, Giachini AJ, Meyer E, Soares, CRFS. Rhizobia Isolated from coal mining areas in the nodulation and growth of leguminous trees. Rev Bras Cienc Solo. 2016;40:e0150091. http://doi.org/10.1590/18069657rbcs20150091
http://doi.org/10.1590/18069657rbcs20150...
). Nevertheless, a high diversity of these bacteria may favor symbiosis with various legume species and maximize biological N fixation in similarly degraded areas. Therefore, the diversity of these nitrogen-fixing bacteria must be investigated, in view of their important role for ecosystem sustainability.
Therefore, the hypothesis of this study was that bacteria isolated from coal mining areas with low pH and high heavy metal contents will also be tolerant to these conditions in vitro, and are efficient as biological N fixers. This study aimed to isolate nitrogen-fixing bacteria from coal mining areas under different vegetation restoration procedures to characterize their tolerance to low pH levels, high heavy metal contents, and their symbiotic efficiency.
MATERIALS AND METHODS
Soil sampling and analyses
Soil samples were collected from the coal mining area of Candiota (31° 33′ 55.5″ S and 53° 43′ 30.6″ W), in Rio Grande do Sul, Brazil. According to Köppen's classification system, the region has a humid subtropical (Cfa) climate, with a mean annual temperature of around 17.2 °C. The original soil of the mining area was classified as Lixisol (WRB, 2015IUSS Working Group WRB. World reference base for soil resources 2014, update 2015: International soil classification system for naming soils and creating legends for soil maps. Rome: Food and Agriculture Organization of the United Nations; 2015. (World Soil Resources Reports, 106).), an Argissolo Vermelho Eutrófico típico (Santos et al., 2013). The main mining area was subdivided into six areas in different recovery stages, and a native forest area was included as reference (Table 1). Soil from the 0.00-0.20 m layer was collected near the roots of species growing in the area. Six plants were randomly selected, with a minimum distance of 200 m between plants. Samples were composed of 12 sub-samples from under the canopy of the selected species, placed in sterile plastic bags, and identified according to the respective sampling point. Soil samples were then transported to the Laboratory of Soil Biology and Microbiology, Centro de Ciências Rurais at the Federal University of Santa Maria (CCR/UFSM) and stored at 4.0 °C until use. A portion of the samples was taken to the Laboratory of Soil Chemistry and Fertility to determine heavy metal availability (As, Cd, Cr, Cu, Ni, Mn, and Zn), as described by Silva et al. (2013)Silva LFO, Vallejuelo Sf-O, Martinez-Arkarazo I, Castro K, Oliveira MLS, Sampaio CH, Brum IAS, Leão FB, Taffarel SR, Madariaga JM. Study of environmental pollution and mineralogical characterization of sediment rivers from Brazilian coal mining acid drainage. Sci Total Environ. 2013;447:169-78. https://doi.org/10.1016/j.scitotenv.2012.12.013
https://doi.org/10.1016/j.scitotenv.2012...
. Inductively Coupled Plasma Optical Emission Spectrometry (ICP-OES, Perkin Elmer Optima 7000DV) was used to determine total cation concentrations.
Restoration history and soil chemical properties from the coalmine region of Candiota, RS-Brazil
Isolation and characterization of nitrogen-fixing bacteria
Macroptilium atropurpureum and V. sativa were used as trap plants for the isolation of nitrogen-fixing bacteria. These crops were chosen because of their well-known capacity to harbor a high diversity of symbiotic microorganisms (Lima et al., 2009Lima AS, Nóbrega RSA, Barberi A, Silva K, Ferreira DF, Moreira FMS. Nitrogen-fixing bacteria communities occurring in soils under different uses in the Western Amazon Region as indicated by nodulation of siratro (Macroptilium atropurpureum). Plant Soil. 2009;319:127-45. https://doi.org/10.1007/s11104-008-9855-2
https://doi.org/10.1007/s11104-008-9855-...
). Plants were grown from April to May 2012 in 110 cm3 plastic pots, in a greenhouse of the Department of Soil Science, UFSM. Pots were filled with a vermiculite - sand (ratio of 2:1) mixture and autoclaved for 1 h at 121 °C. A completely randomized design with a factorial arrangement (7 soil samples × 2 controls × 7 dilutions) with three replications was used. The controls were fertilized with 210 mg L-1 N and a low concentration of mineral nitrogen (5.25 mg L-1 N). Each trap plant species corresponded to one experiment.
To isolate nitrogen-fixing bacteria, the trap plants were inoculated with 1 mL suspension of soil sample collected from each of the seven areas. The soil suspensions were prepared by mixing 10 g of soil in 90 mL of 0.85 % NaCl solution, stirred for 30 min. In addition to the soil suspensions, two controls without inoculation (one with and the other without mineral N) were tested. All treatments were replicated twice.
To estimate the most probable number (MPN) of rhizobia in the seven areas under study, 1 mL aliquots of soil suspension per sample were serially diluted from 10-1 to 10-7 in saline solution (0.85 %) and inoculated on test plants (Table 2). They were categorized as positive or negative, based on the presence or absence of nodules in each dilution, and the MPN Enumeration System (Mpnes) software was used to estimate the MPN (Bennett et al., 1990Bennett JE, Woomer PL, Yost RS. User manual for MPNES most-probable-number enumeration system. Version 1.0. Hawaii: NifTAL project and University of Hawaii; 1990.).
Seeds were surface-sterilized with 70 % ethanol and 1 % sodium hypochlorite, for 1 min per solution, rinsed six times (1 min each) in sterile distilled water, and four inoculated seeds per pot were sown. Every two days, Hoagland and Arnon (1950)Hoagland DR, Arnon DI. The water-culture method for growing plants without soil. Berkeley: California Agricultural Experiment Station; 1950. nutrient solution (½ strength) was applied, while autoclaved distilled water with a low N concentration (10 mg L-1) was provided occasionally in-between when necessary.
Plants were harvested 50 days after emergence (DAE) for the isolation of root nodule bacteria. Five nodules were carefully removed from the roots and used for rhizobial isolation. The nodules were immersed in 95 % ethanol for 30 s, in H2O2 for 1 min, and then rinsed six times with sterile water to remove excess H2O2. Nodules were then crushed with sterile forceps and spread onto plates containing culture medium 79 (Fred and Waksman, 1928Fred EB, Waksman SA. Laboratory manual of general microbiology: with special reference to the microorganisms of the soil. New York: McGraw-Hill Book Company; 1928.). After obtaining pure isolate colonies, these were transferred to plates containing culture medium 79 and cultured for phenotypic characterization. Cultures were stored at 4 °C. The morphological characteristics of the isolates, including growth rate, determined by the time of appearance of isolated colonies (rapid growth – after 2 to 3 days; intermediate - 4 to 5 days; slow - more than 7 to 9 days) and the changes in pH (acid, alkaline, and neutral medium) were also recorded.
Acidic pH and heavy metal tolerance
To evaluate the acidic pH and heavy metal tolerance, isolated strains from nodules of M. atropurpureum (DC) Urb and V. sativa L., and the reference strains of Azorhizobium (A. caulinodans - ORS571T; A. doebereinerae - BR 5401T), Mesorhizobium (M. plurifarium - BR 3804), Rhizobium (R. tropici - CIAT 899T), Burkholderia (B. cepacia - LMG 1222T), and Bradyrhizobium (Bradyrhizobium sp. - BR 2001 and BR 2811) were grown in 30 mL culture medium 79 at pH 6.8, under orbital shaking (100 rpm) at 28 °C, until reaching an optical density (OD560 nm) of 0.5 at 560 nm. Subsequently, aliquots of 100 μL cell suspensions were inoculated into 10 mL liquid medium 79. Isolates were tested at pH 3.0, 4.0, 5.0, 7.0, and 9.0. To test the heavy metal tolerance, culture medium 79 was supplemented with different concentrations of each heavy metal and individually tested. The tested concentrations were: Zn (ZnSO4) 1.0, 3.0, and 5.0 mmol L-1; Cd (CdCl2) 0.1, 0.5, and 3.0 mmol L-1; Cu (CuCl2) 1.0, 2.5, and 5.0 mmol L-1; Ni (NiCl2) 3.0, 7.0, and 15 mmol L-1; and Cr (K2Cr2O7) 1.0, 2.5, and 5.0 mmol L-1. A control treatment without addition of heavy metals was also used. All treatments were tested in triplicate. After metal addition, the pH of the growing media was adjusted to 4.0 by adding 1.0 mol L-1 HCl. Measurements were performed after nine days (OD, 560 nm), and any particular strain considered tolerant only if the OD ≥0.5. Based on the results, strains with high growth at pH 3.0 and 4.0 and at concentrations of 0.86, 3.0, 5.0, 5.0, and 15.0 mg L-1 of Cr, Cd, Zn, Cu, and Ni, respectively, were selected as potential acidic pH and heavy metal-tolerant candidates.
Identification of acidic pH and heavy metal tolerant strains based on 16S rRNA, nodA, and nifH
The strains (with acidic pH and heavy metal tolerance) were identified by the similarity and phylogenetic analysis of the 16S rRNA gene partial sequence and the presence of nodA and nifH genes. All selected isolates were grown in liquid medium 79 at 28 °C for 3 and 7 days to obtain cells in the logarithmic growth phase. The extraction kit Zr Fungal/Bacterial DNA Miniprep® (Zymo Research Corp, USA) was used for genomic DNA extraction, following the manufacturer's instructions. The 16S rRNA gene was amplified using primer pair 27F and 1492R (Lane, 1991Lane DJ. 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M, editors. Nucleic acid techniques in bacterial systematics. New York: John Wiley & Sons; 1991. p. 115-75). Polymerase chain reaction (PCR) was performed with PCR buffer 1X, 2.5 mmol L-1 MgCl2, 0.2 mmol L-1 dNTP, 0.2 μmol L-1 primers, 0.02 U Taq DNA polymerase (Invitrogen), ultra-pure sterile H2O, and 1 μL DNA, for a final volume of 25 mL. Amplification was performed in a Techne® Endurance TC-312 thermal cycler. Thermal specifications were: one initial denaturation step at 94 °C for 5 min, 30 denaturation cycles at 94 °C for 40 s, annealing at 55 °C for 40 s, extension at 72 °C for 1.5 min, and a final extension at 72 °C for 7 min (Ferreira et al., 2012Ferreira PAA, Bomfeti CA, Silva Júnior R, Soares BL, Soares CRFS, Moreira FMS. Eficiência simbiótica de estirpes de Cupriavidus necator tolerantes a zinco, cádmio, cobre e chumbo. Pesq Agropec Bras. 2012;47:85-95. https://doi.org/10.1590/S0100-204X2012000100012
https://doi.org/10.1590/S0100-204X201200...
). The amplified product was resolved on 1 % agarose gel and visualized under UV light to confirm amplification. Subsequently, the amplified material was purified with a PureLink® Quick Gel Extraction Kit (Invitrogen, São Paulo, Brazil), and sequenced using an ABI PRISM® equipment-3100 Genetic Analyzer (Applied Biosystems, USA). Gene sequences were analyzed using the Staden Package 2.0.0b software (Staden et al., 2003Staden R, Judge DP, Bonfield JK. Analyzing sequences using the Staden package and EMBOSS. In: Krawetz SA, Womble DD, editors. Introduction to bioinformatics: a theoretical and practical approach. New York: Springer Science+Business Media; 2003. p. 393-412.) and then subjected to BLASTn (NCBI) for comparison with GenBank sequences. Alignment, editing, and phylogenetic analysis were performed using Mega 5.0 (Tamura et al., 2011Tamura K, Peterson D, Peterson N, Steche, G, Nei M, Kumar S. MEGA 5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol. 2011;28:2731-9. https://doi.org/10.1093/molbev/msr121
https://doi.org/10.1093/molbev/msr121...
). The neighbor-joining clustering method (Saitou and Nei, 1987Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406-25. https://doi.org/10.1093/oxfordjournals.molbev.a040454
https://doi.org/10.1093/oxfordjournals.m...
) and Kimura 2-parameter distance model (Kimura, 1980Kimura M. A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980;16:111-20. https://doi.org/10.1007/BF01731581
https://doi.org/10.1007/BF01731581...
) were used. Reference strains SEMIA 806 (Mesorhizobium sp.) and CIAT 889 (Rhizobium tropici) were also sequenced. The sequences were deposited in the GenBank database .
The nodA gene was PCR-amplified using the oligonucleotide primers nodABurkF and nodABurkR for both α and β-proteobacteria. The nifH gene was amplified with primers nifH-R and nifH-F for β-proteobacteria, and nifH-F and nifH-I for α-proteobacteria. The PCR products were separated on 1 % agarose gel and visualized under UV light using a 100 pb molecular DNA ladder marker (Invitrogen).
Plant nodulation and symbiotic efficiency tests
The symbiotic efficiency of the selected strains was evaluated on M. atropurpureum and V. sativa plants in a greenhouse. Each species was examined in one experiment. Plants were grown under the conditions described previously for the isolation of the strains. For inoculation, the selected strains were grown in liquid medium 79 (Fred and Waksman, 1928Fred EB, Waksman SA. Laboratory manual of general microbiology: with special reference to the microorganisms of the soil. New York: McGraw-Hill Book Company; 1928.) for five days under constant shaking at 28 °C. Each seed was inoculated with 1.0 mL of bacterial suspension (1 × 109 cells mL-1). The Hoagland and Arnon (1950)Hoagland DR, Arnon DI. The water-culture method for growing plants without soil. Berkeley: California Agricultural Experiment Station; 1950. nutrient solution (½ strength) in the plant pots was replaced every two days by fresh autoclaved solution, and the volume maintained by adding sterile distilled water. Ten days after germination only one plant per pot was left.
The experiment was arranged in a completely randomized design with three replications. The treatments consisted of inoculation with selected strains and two uninoculated controls treated with either 210 mg L-1 or 21 mg L-1 of mineral N.
Plants were harvested 50 days after emergence, and the morphological parameters root (RDM) and shoot dry matter (SDM), number of nodules per plant (NNP), nodule dry weight (NDW), and symbiotic efficiency (SE) were determined. The dry matter yield (dry matter of shoots, dry matter of roots, and nodule dry weight) was estimated after the biomass was dried in a forced-air oven (±65 °C) until reaching constant weight. The symbiotic efficiency of each isolate was calculated by the expression SE (%) = [(Shoot dry matter of inoculated treatments × 100) / (Shoot dry matter of control with 210 mg L-1 mineral N)].
Statistical analysis
The data were analyzed statistically with Sisvar, version 5.6 (Ferreira, 2011Ferreira DF. Sisvar: a computer statistical analysis system. Cienc agrotec. 2011;35:1039-42. https://doi.org/10.1590/S1413-70542011000600001
https://doi.org/10.1590/S1413-7054201100...
). All data were transformed when necessary to meet the assumptions of normality and homoscedasticity. Subsequently, the data were analyzed by Anova and the means compared by the Scott-Knott test (Scott and Knott, 1974Scott AJ, Knott MA. A cluster analysis method for grouping means in the analysis of variance. Biometrics. 1974;30:507-12. https://doi.org/10.2307/2529204
https://doi.org/10.2307/2529204...
), considering a nominal level of significance of 5 % probability (p<0.05). The NNP and NDM data were transformed to (x + 0.5) 0.5.
RESULTS
Isolation and characterization of nitrogen-fixing bacteria
The native rhizobia population was calculated by the serial dilution method using M. atropurpureum and V. sativa as trap plants (Table 2). One hundred and fifteen bacterial strains were obtained (77 strains from nodules on M. atropurpureum and 38 from V. sativa) (Figure 1a). Strains were obtained from all areas, but the soil with the longest restoration period (20 years re-vegetated with Acacia mearnsii + Eucalyptus sp.) had the highest number of strains (or isolates). The results indicated that the area with shortest time of revegetation (A2 - 6 years) and one area with Eucalyptus sp. (A4 – 20 years) had the lowest number of isolates.
Number of isolated bacterial strains from coal-mined areas (a). Group distribution of the 115 isolates based on colony growth rate (time) and pH variation (b). SAL = slow growth, and alkaline pH; SN = slow growth and neutral pH; SA = slow growth and acid pH; FAL = fast growth and alkaline pH; FN = fast growth and neutral pH; FA = fast growth and acid pH.
The 115 strains were phenotypically grouped, according to the growth rate and ability to change the pH of the culture medium (Figure 1b). Six phenotypic groups were observed: (i) slow growth, medium alkalinization (SAL); (ii) slow growth, no alteration of medium pH (SN); (iii) slow growth, medium acidification (SA); (iv) fast growth, medium alkalinization (FAL); (v) fast growth, no alteration of medium pH (FN); and (vi) fast-growth, medium acidification (FA). Only strains of M. atropurpureum showed slow growth without altering the medium pH (SN), and fast growth with medium alkalinization (FAL). On the other hand, isolates of V. sativa L. presented slow growth with acidifying capacity (SA).
Acidic pH and heavy metal tolerance assay
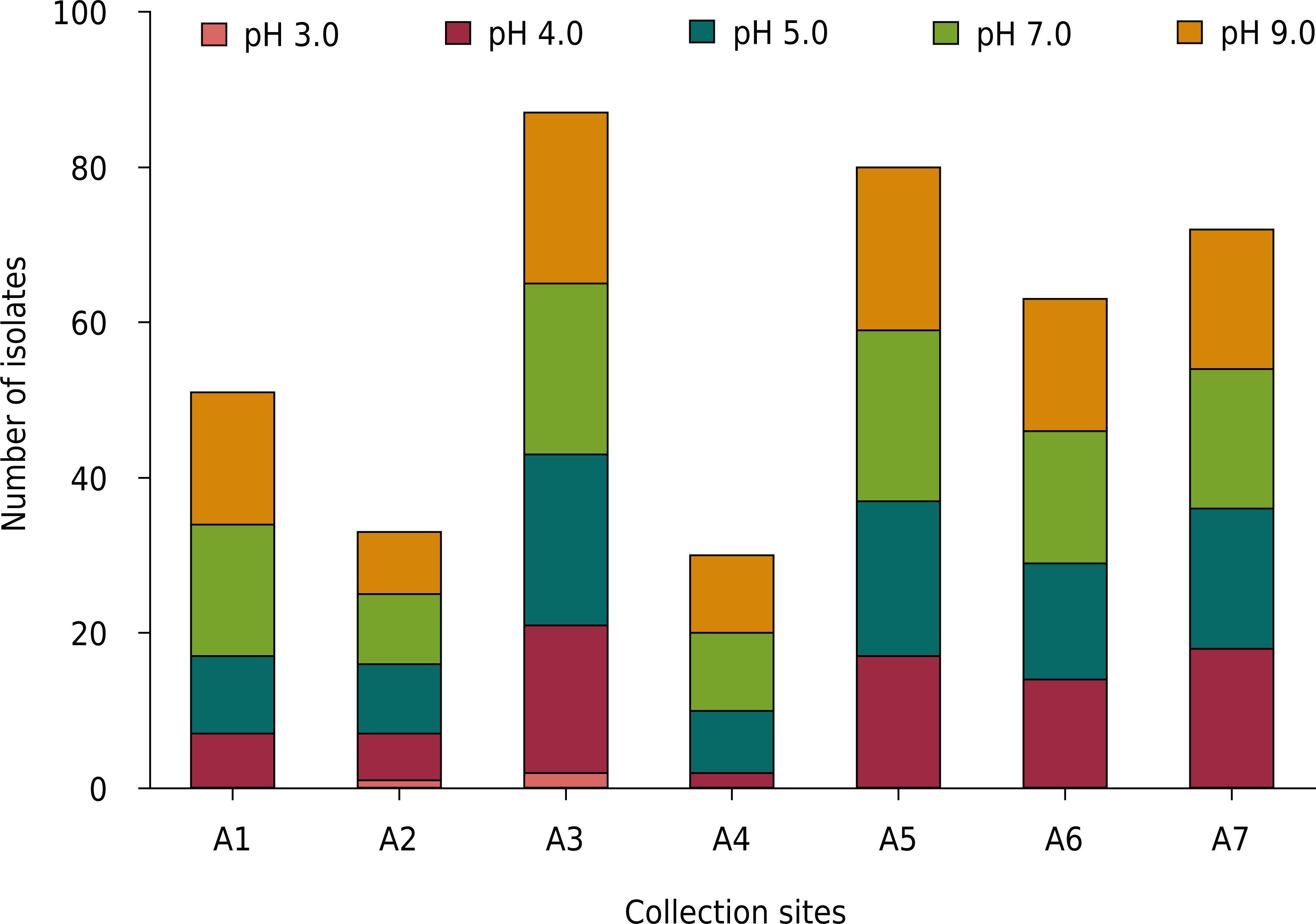
The results indicated differential growth and pH tolerance among the 115 isolated strains (Figure 2). Of these 115 strains, 98.2 % grew at pH 9.0, 100 % at pH 7.0, 88.6 % at pH 5.0, 72.2 % at pH 4.0, and only 2.6 % at pH 3.0. The isolates from areas A3, A5, A6, and A7 were more tolerant to pH 4.0. One strain from area A2 and two from area A3 maintained growth at pH 3.0. Strain Rhizobium tropici (CIAT 899T) grew at all evaluated pH whereas Burkolderia cepacia (LMG 1222T) and Bradyrhizobium spp. (BR 2001 and BR 2811) grew well at pH 4.0, 5.0, 7.0, and 9.0. Strains Azorhizobium caulinodans (ORS571T), Azorhizobium doebereinerae (BR 5401T), and Mesorhizobium plurifarium (BR 3804) grew only at pH >4.0.
For the heavy metal tolerance assay, a significant number of bacterial strains grew at higher metal concentrations, although a differential trend was observed (Figure 3). Forty-one strains were tolerant to 0.86 mmol L-1 Cr. No significant tolerance effects were observed for the strains M. plurifarium (BR 3804), R. tropici (CIAT 899T), B. cepacia (LMG 1222T), and Bradyrhizobium spp. (BR 2001 and BR 2811), which all responded similarly to Cr stress.
Number of heavy metal-resistant bacterial isolates at different Cr, Cd, Zn, Cu, and Ni concentrations in each representative area.
Twenty-two percent of the strains were tolerant to 3.0 mmol L-1 Cd, including all reference strains. The highest tolerance levels for Cu (5 mmol L-1) and Zn (5 mmol L-1) concentrations were observed for 8 and 17 % of the evaluated strains, respectively. Ten strains tolerated the highest Ni concentration (15 mmol L-1), including B. cepacia (LMG 1222T) and Bradyrhizobium sp. (BR 2811). In general, the strains showed the following order of tolerance to the heavy metals tested: Cr > Cd > Zn > Ni > Cu. Isolates from area A3 were more tolerant to all concentrations of metals, followed by isolates from areas A5, A6, and A7, respectively (Figure 3).
Phylogeny of 16S rRNA and phylogenetic identification of nodA and nifH symbiotic genes
The 18 strains (13 from M. atropurpureum and 5 from V. sativa) selected based on pH tolerance and resistance to high heavy metal concentration (Table 3) were identified by 16S rRNA gene sequence analysis (Table 4). The comparison of the 16S rRNA gene sequences revealed that the bacterial strains were phylogenetically related with nitrogen-fixing bacteria, belonging to genera such as Rhizobium, Bradyrhizobium, and Burkholderia (Table 4). All selected strains for acidic pH and heavy metal tolerance amplified DNA fragments related to the nifH (350 bp) and nodA (530 bp) genes (Table 4).
Values of pH and heavy metal concentration tolerated (mmol L-1) by 18 bacterial strains selected for sequencing the 16S rRNA
Origin, characteristics, and identification of bacterial isolated strains obtained from nodules of Macroptilium atropurpureum and Vicia sativa cultivated in the coal-mining region of Candiota, RS-Brazil
In the heavy metal assay to determine Cr, Cd, Zn, Cu, and Ni tolerance, all species of Bradyrhizobium, including strains UFSM-B21, UFSM-B51, UFSM-B53, UFSM-B55, and UFSM-B61, showed similar tolerance to that of M. plurifarium (BR 3804), R. tropici (CIAT 899T), B. cepacia (LMG 1222T), and Bradyrhizobium spp. (BR 2001 and BR 2811) when tested at 0.86 mmol L-1 Cr. As for Cd, tolerance at 3.0 mmol L-1 concentration was similar for UFSM-B55 (Bradyhrizobium sp.), UFSM-B33, UFSM-B34 (Burkholderia sp.), and reference strains. The most tolerant strains to 5 mmol L-1 Cu were UFSM-B11 (Rhizobium sp.), UFSM-B52, UFSM-B61 (Bradyrhizobium sp.), and UFSM-B65 (Rhizobium sp.), whereas the most tolerant species to 5 mmol L-1 Zn include UFSM-B33, UFSM-B34 (both Burkholderia sp.), and UFSM-B55 (Bradyrhizobium sp.). These strains showed similar growth to R. tropici (CIAT 899T), B. cepacia (LMG 1222T), and Bradyrhizobium sp. (BR 2001).
Plant nodulation and symbiotic efficiency tests
The results for the nodulation and symbiotic efficiency of the 18 selected strains with their respective host plants are shown in table 5. The 13 selected strains isolated from M. atropurpureum favored their growth and development. Strains UFSM-B53, UFSM-B64, and UFSM-B74 showed similar results for plant growth, with increases of 220, 240 and 250 % in dry matter, respectively, compared to the control without mineral N application. Inoculation with all strains, except UFSM-B32, provided better root development in both crops. The results also indicated the above strains not only for higher dry matter, but also the highest number of nodules, nodule dry weight and highest symbiotic efficiency of the inoculation treatments. For the other strains the results for nodulation potential and symbiotic efficiency were insignificant. Inoculation with UFSM-B11, UFSM-B12, UFSM-B33, UFSM-B34, and UFSM-B65 in V. sativa did not improve plant growth when compared to the treatment with mineral N (Table 5). These strains were able to induce higher nodulation but had a lower symbiotic efficiency than the treatment with mineral N.
Influence of the tested strains on shoot dry matter, root dry matter, number of nodules, nodule dry matter, and symbiotic efficiency (SE) of Atropurpureum macroptylium and Vicia sativa and two controls, C/N (with mineral nitrogen) and S/N (without mineral N)
DISCUSSION
The data of this study are not conclusive to support the hypothesis that bacteria isolated from coal mining areas with low pH and high heavy metal concentration are also more tolerant to these conditions in vitro and efficient as biological N fixers. There was no relationship between pH level or heavy metal concentration in the soil inhabited by the bacteria and their growth in culture medium at different acidity levels or metal concentrations. In addition, only three isolates stood out as good N fixers. Even in acid or heavy metal-contaminated soils, it is probable that neutrophilic and alkalophilic or heavy metal-sensitive bacteria can survive at microsites with specific environmental conditions (Vos et al., 2013Vos M, Wolf AB, Jennings SJ, Kowalchuk GA. Micro-scale determinants of bacterial diversity in soil. FEMS Microbial Rev. 2013;37:936-54. https://doi.org/10.1111/1574-6976.12023
https://doi.org/10.1111/1574-6976.12023...
).
Bacteria were isolated from all studied areas. However, higher numbers of bacteria were isolated from soils under plant establishment (A3 - eight years of spontaneous vegetation growth) and from tailing areas (A5 - 20 years under Acacia mearnsii + Eucalyptus sp. plantations) (Figure 1a). The high number of isolated strains may be related to the restoration period of the areas. In these environments, soil organic matter (SOM) is the main indicator of the sustainability of a cultivation system (Conceição et al., 2005Conceição PC, Amado TJC, Mielniczuk J, Spagnollo E. Qualidade do solo em sistemas de manejo avaliada pela dinâmica da matéria orgânica e atributos relacionados. Rev Bras Cienc Solo. 2005;29:777-88. https://doi.org/10.1590/S0100-06832005000500013
https://doi.org/10.1590/S0100-0683200500...
), mainly in tropical soils (Solomon et al., 2002Solomon D, Fritzsche F, Lehmann J, Tekalign M, Zech W. Soil organic matter dynamics in the subhumid agroecosystems of the Ethiopian Highlands. Soil Sci Soc Am J. 2002;66:969-78. https://doi.org/10.2136/sssaj2002.9690
https://doi.org/10.2136/sssaj2002.9690...
). An increase in SOM in these areas has a positive impact on soil biodiversity.
In coal-mining areas, all forest cover and the most fertile soil layer is removed to facilitate the access for mineral exploitation. This, in turn, results in the disruption of nutrients and energy flow. Under natural revegetation, N is one of the most limiting nutrients in the soil surface layers (Vitousek and Howarth, 1991Vitousek PM, Howarth RW. Nitrogen limitation on land and in the sea: how can it occur? Biogeochemistry. 1991;13:87-115. https://doi.org/10.1007/BF00002772
https://doi.org/10.1007/BF00002772...
). In degraded areas of tropical countries, soil N availability is low. Supplying N in such areas may favor the process of natural succession. Therefore, herbaceous and tree legume species that establish symbiosis with nitrogen-fixing bacteria can be a primary source of N, facilitating plant recolonization and biodiversity enrichment (Siddique et al., 2008Siddique I, Engel VL, Parrotta JA, Lamb D, Nardoto GB, Ometto JPHB, Martinelli LA, Schmidt S. Dominance of legume trees alters nutrient relations in mixed species forest restoration plantings within seven years. Biogeochemistry. 2008;88:89-101. https://doi.org/10.1007/s10533-008-9196-5
https://doi.org/10.1007/s10533-008-9196-...
). The addition of low C:N plant residues (Nardoto et al., 2008Nardoto GB, Ometto JPHB, Ehleringer JR, Higuchi N, Bastamante MMC, Martinelli LA. Understanding the influences of spatial patterns on N availability within the Brazilian Amazon Forest. Ecosystem. 2008;11:1234-46. https://doi.org/10.1007/s10021-008-9189-1
https://doi.org/10.1007/s10021-008-9189-...
) contributes to soil recovery, as well as to increase biological activity and SOM stabilization (Lavelle, 2000Lavelle P. Ecological challenges for soil science. Soil Sci. 2000;165:73-86. https://doi.org/10.1097/00010694-200001000-00009
https://doi.org/10.1097/00010694-2000010...
; Resh et al., 2002Resh SC, Binkley D, Parrotta JA. Greater soil carbon sequestration under nitrogen-fixing trees compared with Eucalyptus species. Ecosystems. 2002;5:217-31. https://doi.org/10.1007/s10021-001-0067-3
https://doi.org/10.1007/s10021-001-0067-...
).
Similarly to the variability identified by the phenotypic characterization, the number of strains varied as a function of the host plant (Figure 1b). When V. sativa was used as trap plant, there was a predominance of fast-growing strains with acidification capacity, which is a typical characteristic of the genus Rhizobium. The number of strains with greater phenotypic diversity was highest for M. atropurpureum. This legume is considered promiscuous in its ability to establish symbiosis with nitrogen-fixing bacteria, capable of associating with various genera of both α and β-rhizobia (Lima et al., 2009Lima AS, Nóbrega RSA, Barberi A, Silva K, Ferreira DF, Moreira FMS. Nitrogen-fixing bacteria communities occurring in soils under different uses in the Western Amazon Region as indicated by nodulation of siratro (Macroptilium atropurpureum). Plant Soil. 2009;319:127-45. https://doi.org/10.1007/s11104-008-9855-2
https://doi.org/10.1007/s11104-008-9855-...
).
The strains UFSM-B21 (Bradyrhizobium sp.) and UFSM-B33/UFSM-B34 (Burkholderia sp.) had better growth in culture with pH 3.0 (OD560 nm ≥0.5), similar to the pH of the soil from which they were isolated. Bacteria of very acidic sites must have efficient mechanisms to keep the cytoplasmic pH several units higher than the pH of the external medium. The reviews of Baker-Austin and Dopson (2007)Baker-Austin C, Dopson M. Life in acid: pH homeostasis in acidophiles. Trends Microbiol. 2007;15:165-71. https://doi.org/10.1016/j.tim.2007.02.005
https://doi.org/10.1016/j.tim.2007.02.00...
and Krulwich et al. (2011)Krulwich TA, Sachs G, Padan E. Molecular aspects of bacterial pH sensing and homeostasis. Nat Rev Microbiol. 2011;9:330-43. https://doi.org/10.1038/nrmicro2549
https://doi.org/10.1038/nrmicro2549...
on homeostasis in acidophilic bacteria indicate the following mechanisms: potassium-transporting ATPases, reduction of permeability of cell membranes, outflow of H+ out of the cell, expression of higher number of transporters, binding of H+ to molecules within the cytoplasm, more efficient DNA and polypeptide repair systems, and degradation of organic compounds decoupled from the electron transport chain. Although these are the most common mechanisms, not all acidophiles have all these mechanisms, because their occurrence is specific to certain bacterial groups. Several studies demonstrated the presence of nitrogen-fixing bacteria in soils contaminated with Cd, Zn, Cu, Pb, and Ni (Matsuda et al., 2002Matsuda A, Moreira FMS, Siqueira JO. Tolerância de rizóbios de diferentes procedências ao zinco, cobre e cádmio. Pesq Agropec Bras. 2002;37:343-55. https://doi.org/10.1590/S0100-204X2002000300016
https://doi.org/10.1590/S0100-204X200200...
; Melloni et al., 2004Melloni R, Nóbrega RSA, Moreira FMS, Siqueira JO. Densidade e diversidade fenotípica de bactérias diazotróficas endofíticas em solos de mineração de bauxita, em reabilitação. Rev Bras Cienc Solo. 2004;28:85-93. https://doi.org/10.1590/S0100-06832004000100009
https://doi.org/10.1590/S0100-0683200400...
; Carrasco et al., 2005Carrasco JA, Armario P, Pajuelo E, Burgos A, Caviedes MA, López R, Chamber MA, Palomares AJ. Isolation and characterisation of symbiotically effective Rhizobium resistant to arsenic and heavy metals after the toxic spill at the Aznalco'llar pyrite mine. Soil Biol Biochem. 2005;37:1131-40. https://doi.org/10.1016/j.soilbio.2004.11.015
https://doi.org/10.1016/j.soilbio.2004.1...
; Becerra-Castro et al., 2011Becerra-Castro C, Kidd PS, Prieto-Fernández A, Weyens N, Acea M-J, Vangronsveld J. Endophytic and rhizoplane bacteria associated with Cytisus striatus growing on hexachlorocyclohexane-contaminated soil: isolation and characterisation. Plant Soil. 2011;340:413-33. https://doi.org/10.1007/s11104-010-0613-x
https://doi.org/10.1007/s11104-010-0613-...
; Weyens et al., 2013Weyens N, Beckers B, Schellingen K, Ceulemans R, Croes S, Janssen J, Haenen S, Witters N, Vangronsveld J. Plant-associated bacteria and their role in the success failure of metal phytoextraction projects: firstobservations of a field-related experiment. Microb Biotechnol. 2013;6:288-99. https://doi.org/10.1111/1751-7915.12038
https://doi.org/10.1111/1751-7915.12038...
). However, this is the first report of nitrogen-fixing bacteria isolated from coal mining areas contaminated with heavy metals under extremely acid (pH = 3.5) soil conditions in Brazil. Strains UFSM-B55 (Bradyrhizobium sp.), UFSM-B65, and UFSM-B74 (both Rhizobium sp.) were grown at 15 mmol L-1 Ni, along with B. cepacia (LMG 1222T) and Bradyrhizobium sp. (BR 2811). The ability of some microorganisms to grow under high metal concentrations can result from intrinsic or induced mechanisms, as well as from other environmental factors in the culture medium, such as pH, redox potential, etc., which may reduce the metal toxicity (Zouboulis et al., 2004Zouboulis AI, Loukidou MX, Matis KA. Biosorption of toxic metals from aqueous solutions by bacteria strains isolated from metal-polluted soils. Process Biochem. 2004;39:909-16. https://doi.org/10.1016/S0032-9592(03)00200-0l
https://doi.org/10.1016/S0032-9592(03)00...
; Leedjärv et al., 2008Leedjärv A, Ivask A, Virta M. Interplay of different transporters in the mediation of divalent heavy metal resistance in Pseudomonas putida KT2440. J Bacteriol. 2008;190:2680-9. https://doi.org/10.1128/JB.01494-07
https://doi.org/10.1128/JB.01494-07...
; Xiao et al., 2010Xiao X, Luo S, Zeng G, Wei W, Wan Y, Chen L, Guo H, Cao Z, Yang L, Chen J, Xi Q. Biosorption of cadmium by endophytic fungus (EF) Microsphaeropsis sp. LSE10 isolated from cadmium hyperaccumulator Solanum nigrum L. Bioresource Technol. 2010;101:1668-74. https://doi.org/10.1016/j.biortech.2009.09.083
https://doi.org/10.1016/j.biortech.2009....
). Different mechanisms of metal tolerance of these bacteria are well-documented. They vary little between strains isolated from non-contaminated and contaminated environments (Barkay and Schaefer, 2001Barkay T, Schaefer J. Metal and radionuclide bioremediation: issues, considerations and potentials. Curr Opin Microbiol. 2001;4:318-23. https://doi.org/10.1016/S1369-5274(00)00210-1
https://doi.org/10.1016/S1369-5274(00)00...
; Matsuda et al., 2002Matsuda A, Moreira FMS, Siqueira JO. Tolerância de rizóbios de diferentes procedências ao zinco, cobre e cádmio. Pesq Agropec Bras. 2002;37:343-55. https://doi.org/10.1590/S0100-204X2002000300016
https://doi.org/10.1590/S0100-204X200200...
; Zouboulis et al., 2004Zouboulis AI, Loukidou MX, Matis KA. Biosorption of toxic metals from aqueous solutions by bacteria strains isolated from metal-polluted soils. Process Biochem. 2004;39:909-16. https://doi.org/10.1016/S0032-9592(03)00200-0l
https://doi.org/10.1016/S0032-9592(03)00...
; Ferreira et al., 2012Ferreira PAA, Bomfeti CA, Silva Júnior R, Soares BL, Soares CRFS, Moreira FMS. Eficiência simbiótica de estirpes de Cupriavidus necator tolerantes a zinco, cádmio, cobre e chumbo. Pesq Agropec Bras. 2012;47:85-95. https://doi.org/10.1590/S0100-204X2012000100012
https://doi.org/10.1590/S0100-204X201200...
). These characteristics explain the survival and persistence ability of certain strains examined in this study, unlike those isolated from environments with low heavy metal contents.
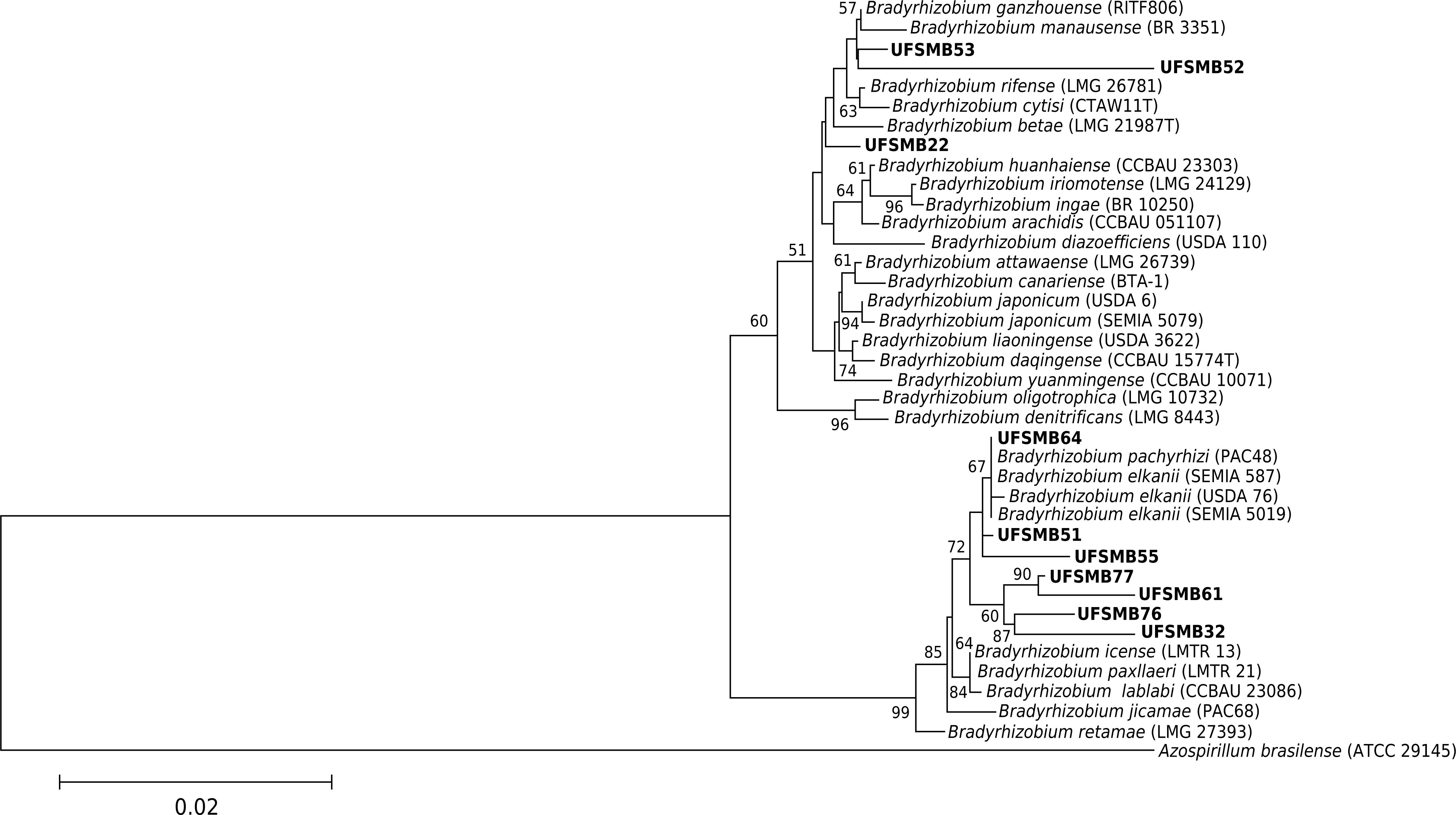
Out of the 13 selected strains obtained from M. atropurpureum, 12 belong to the genus Bradyrhizobium and one to genus Rhizobium (Table 4). The phylogenetic analysis of 10 strains revealed that seven of them belong to subgroup-II, in which the traditional species B. elkanii is also nested. Three belong to subgroup-I, one of which is B. japonicum (Figure 4). However, due to the high similarity of the 16S rRNA gene, further analyses are needed to better characterize these strains. Many studies that isolated nitrogen-fixing bacteria from V. sativa have reported the presence of Rhizobium leguminosarum bv. viciae (van Berkum et al., 1995van Berkum P, Beyene D, Vera FT, Keyser HH. Variability among Rhizobium strains originating from nodules of Vicia faba. Appl Environ Microb. 1995;61:2649-53.; Laguerre et al., 2003Laguerre G, Louvrier P, Allard M-R, Amarger N. Compatibility of rhizobial genotypes within natural populations of Rhizobium leguminosarum biovar viciae for nodulation of host legumes. Appl Environ Microb. 2003;69:2276-83. https://doi.org/10.1128/AEM.69.4.2276-2283.2003
https://doi.org/10.1128/AEM.69.4.2276-22...
; Moschetti et al., 2005Moschetti G, Peluso AL, Protopapa A, Anastasio M, Pepe O, Defez R. Use of nodulation pattern, stress tolerance, nodC gene amplification, RAPD-PCR and RFLP-16S rDNA analysis to discriminate genotypes of Rhizobium leguminosarum biovar viciae. Syst Appl Microbiol. 2005;28:619-31. https://doi.org/10.1016/j.syapm.2005.03.009
https://doi.org/10.1016/j.syapm.2005.03....
) and Rhizobium sp. (Lei et al., 2008Lei X, Wang ET, Chen WF, Sui XH, Chen WX. Diverse bacteria isolated from root nodules of wild Vicia species grown in temperate region of China. Arch Microbiol. 2008;190:657-71. https://doi.org/10.1007/s00203-008-0418-y
https://doi.org/10.1007/s00203-008-0418-...
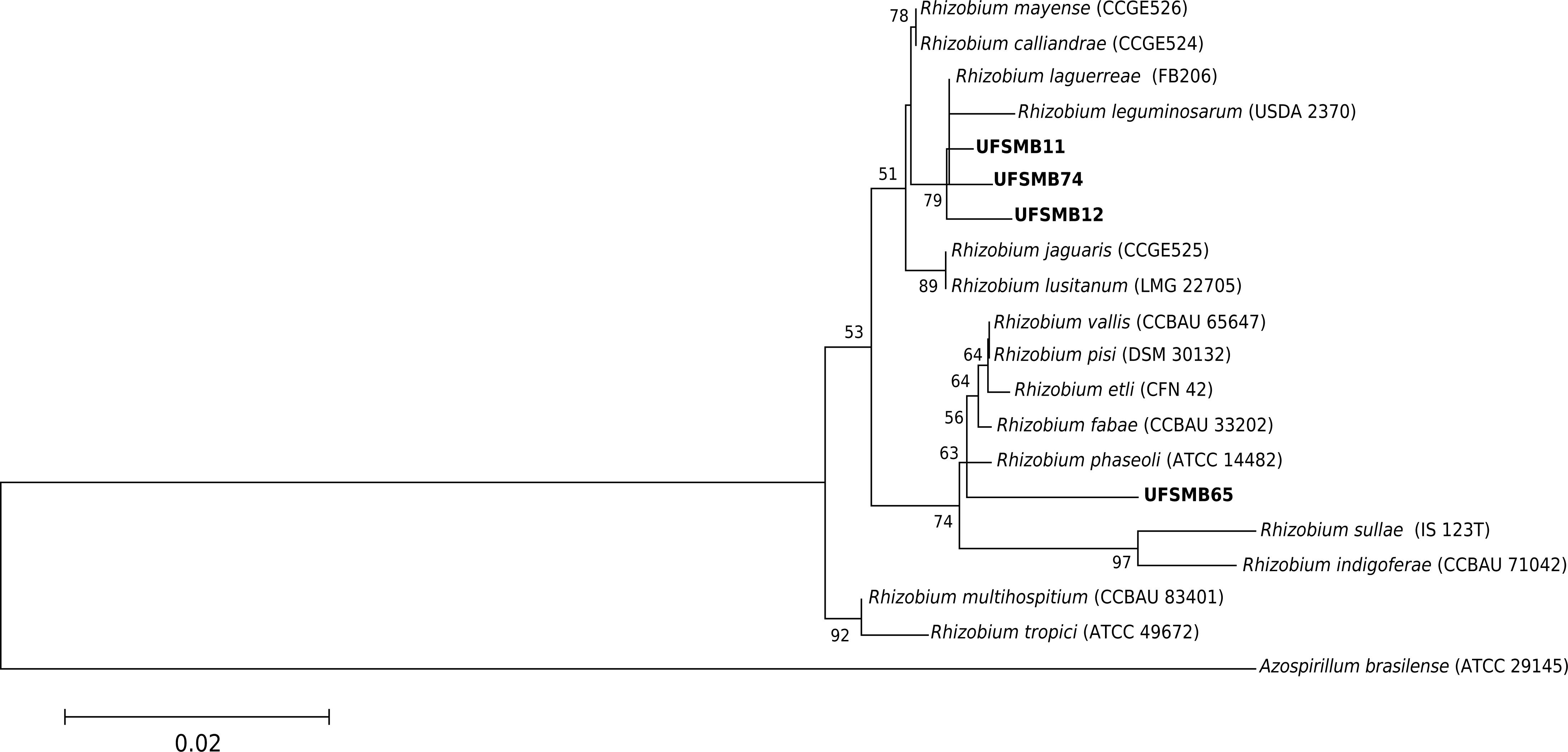
). Here, out of the five selected strains isolated from V. sativa and tolerant to acidic pH and heavy metals, three belong to Rhizobium and two to Burkholderia (Table 4). Phylogenetic analysis (Figure 5) showed that two strains were close to R. leguminosarum and one to R. phaseoli. This is the first report on the presence of Burkholderia on V. sativa. This bacterium, when re-inoculated, could induce nodule formation with a relative efficiency of 50 % (Table 5). The nodulation ability of V. sativa may be due to a lateral gene transfer from the symbiotic genus Rhizobium into Burkholderia, since symbiotic genes are not very different between α and β-rhizobia (Moulin et al., 2002Moulin L, Chen WM, Béna G, Dreyfus B, Boivin-Masson C. Rhizobia: the family is expanding. In: Finan TM, O'Brian MR, Layzell DB, Vessey JK, Newton W, editors. Nitrogen fixation: global perspectives. New York: CABI Publishing; 2002. p. 61-5.; Chen et al., 2003Chen W-M, Moulin L, Bontemps C, Vandamme P, Béna G, Boivin-Masson C. Legume symbiotic nitrogen fixation by β-proteobacteria is widespread in the nature. J Bacteriol. 2003;185:7266-72. https://doi.org/10.1128/JB.185.24.7266-7272.2003
https://doi.org/10.1128/JB.185.24.7266-7...
).
Phylogenetic tree based on 16S rRNA gene sequence similarity of Bradyrhizobium strains. The phylogenetic tree was built using neighbor-joining and Kimura 2-parameters (K2P). Numbers on branches indicate bootstrap values, with a total of 1000 replicates; only values greater than 50 are shown. Bar indicates 2 substitutions per 100 nucleotides.
Phylogenetic tree based on 16S rRNA gene sequence similarity of Rhizobium strains. The phylogenetic tree was built using neighbor-joining and Kimura 2-parameters (K2P). Numbers on branches indicate bootstrap values, with a total of 1000 replicates; only values greater than 50 are shown. Bar indicates 2 substitutions per 100 nucleotides.
In this regard, strains UFSM-B53 and UFSM-B54 (Bradyrhizobium sp.) and UFSM-B74 (Rhizobium) had the highest nitrogen-fixing efficiency in M. atropurpureum (Table 5). The selection of efficient nitrogen-fixing strains for the reforestation of degraded areas contributes to the adaptation of plant species. These strains can be an alternative inoculant for coal mining areas (tolerant to environmental stress), since M. atropurpureum may fix up to 181 kg ha-1 yr-1 N (Moreira and Siqueira, 2006Moreira FMS, Siqueira JO. Microbiologia e bioquímica do solo. 2a ed atual e rev. Lavras: Editora UFLA; 2006.).
This study describes some efficient nitrogen-fixing strains in acidic pH and heavy metal tolerance found in coal-mined soils in the state of Rio Grande do Sul, Southern Brazil. Even though the strains proved to be symbiotically efficient, tolerant to high acidity and elevated levels of heavy metals, these isolates still need to be tested under these conditions in field experiments. Efficient isolated strains can provide advances towards plant establishment and restoration of coal-mined degraded areas contaminated with high heavy metal levels. Furthermore, these potential strains must be propagated on a commercial basis to optimize their biological viability and activity for field applications.
CONCLUSIONS
Most of the 115 nitrogen-fixing bacteria isolated from the coal mining areas are tolerant to very acidic pH (4.0), with few tolerating extremely acidic pH levels (3.0).
The ability to tolerate high heavy metal concentrations is highly variable among the isolated nitrogen-fixing bacteria, but the following order of tolerance can be established: Cr > Cd> Zn > Ni > Cu.
The 18 selected nitrogen-fixing bacteria belong to the genera Rhizobium, Bradyrhizobium, and Burkholderia;
Rhizobium UFSM-B74 and Bradyrhizobium sp. UFSM-B53 and UFSM-B64 on M. atropurpureum have high symbiotic efficiency. For being tolerant to low pH and high heavy metal concentrations as well, they are promising for the production of specific inoculants for areas affected by mining.
ACKNOWLEDGEMENTS
We thank Fapergs, CNPq, and Capes, for the scholarships and the financial support of this project.
REFERENCES
- Agência Nacional de Energia Elétrica - Aneel. Atlas de energia elétrica do Brasil. 3. ed. Brasília, DF: Aneel; 2008 [acesso em 15 mai 2012]. Available at: http://www2.aneel.gov.br/arquivos/pdf/atlas3ed.pdf
» http://www2.aneel.gov.br/arquivos/pdf/atlas3ed.pdf - Baker-Austin C, Dopson M. Life in acid: pH homeostasis in acidophiles. Trends Microbiol. 2007;15:165-71. https://doi.org/10.1016/j.tim.2007.02.005
» https://doi.org/10.1016/j.tim.2007.02.005 - Barkay T, Schaefer J. Metal and radionuclide bioremediation: issues, considerations and potentials. Curr Opin Microbiol. 2001;4:318-23. https://doi.org/10.1016/S1369-5274(00)00210-1
» https://doi.org/10.1016/S1369-5274(00)00210-1 - Becerra-Castro C, Kidd PS, Prieto-Fernández A, Weyens N, Acea M-J, Vangronsveld J. Endophytic and rhizoplane bacteria associated with Cytisus striatus growing on hexachlorocyclohexane-contaminated soil: isolation and characterisation. Plant Soil. 2011;340:413-33. https://doi.org/10.1007/s11104-010-0613-x
» https://doi.org/10.1007/s11104-010-0613-x - Bennett JE, Woomer PL, Yost RS. User manual for MPNES most-probable-number enumeration system. Version 1.0. Hawaii: NifTAL project and University of Hawaii; 1990.
- Carrasco JA, Armario P, Pajuelo E, Burgos A, Caviedes MA, López R, Chamber MA, Palomares AJ. Isolation and characterisation of symbiotically effective Rhizobium resistant to arsenic and heavy metals after the toxic spill at the Aznalco'llar pyrite mine. Soil Biol Biochem. 2005;37:1131-40. https://doi.org/10.1016/j.soilbio.2004.11.015
» https://doi.org/10.1016/j.soilbio.2004.11.015 - Chen W-M, Moulin L, Bontemps C, Vandamme P, Béna G, Boivin-Masson C. Legume symbiotic nitrogen fixation by β-proteobacteria is widespread in the nature. J Bacteriol. 2003;185:7266-72. https://doi.org/10.1128/JB.185.24.7266-7272.2003
» https://doi.org/10.1128/JB.185.24.7266-7272.2003 - Conceição PC, Amado TJC, Mielniczuk J, Spagnollo E. Qualidade do solo em sistemas de manejo avaliada pela dinâmica da matéria orgânica e atributos relacionados. Rev Bras Cienc Solo. 2005;29:777-88. https://doi.org/10.1590/S0100-06832005000500013
» https://doi.org/10.1590/S0100-06832005000500013 - Ferreira DF. Sisvar: a computer statistical analysis system. Cienc agrotec. 2011;35:1039-42. https://doi.org/10.1590/S1413-70542011000600001
» https://doi.org/10.1590/S1413-70542011000600001 - Ferreira PAA, Bomfeti CA, Silva Júnior R, Soares BL, Soares CRFS, Moreira FMS. Eficiência simbiótica de estirpes de Cupriavidus necator tolerantes a zinco, cádmio, cobre e chumbo. Pesq Agropec Bras. 2012;47:85-95. https://doi.org/10.1590/S0100-204X2012000100012
» https://doi.org/10.1590/S0100-204X2012000100012 - Franco AA, Faria SM. The contribution of N2-fixing tree legumes to land reclamation and sustainability in the tropics. Soil Biol Biochem. 1997;29:897-903. https://doi.org/10.1016/S0038-0717(96)00229-5
» https://doi.org/10.1016/S0038-0717(96)00229-5 - Fred EB, Waksman SA. Laboratory manual of general microbiology: with special reference to the microorganisms of the soil. New York: McGraw-Hill Book Company; 1928.
- Fungaro DA, Izidoro JC. Remediação de drenagem ácida de mina usando zeólitas sintetizadas a partir de cinzas leves de carvão. Quim Nova. 2006;29:735-40. https://doi.org/10.1590/S0100-40422006000400019
» https://doi.org/10.1590/S0100-40422006000400019 - Hoagland DR, Arnon DI. The water-culture method for growing plants without soil. Berkeley: California Agricultural Experiment Station; 1950.
- IUSS Working Group WRB. World reference base for soil resources 2014, update 2015: International soil classification system for naming soils and creating legends for soil maps. Rome: Food and Agriculture Organization of the United Nations; 2015. (World Soil Resources Reports, 106).
- Kimura M. A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980;16:111-20. https://doi.org/10.1007/BF01731581
» https://doi.org/10.1007/BF01731581 - Klonowska A, Chaintreuil C, Tisseyre P, Miché L, Melkonian R, Ducousso M, Laguerre G, Brunel B, Moulin L. Biodiversity of Mimosa pudica rhizobial symbionts (Cupriavidus taiwanensis, Rhizobium mesoamericanum) in New Caledonia and their adaptation to heavy metal-rich soils. FEMS Microbiol Ecol. 2012;81:618-35. https://doi.org/10.1111/j.1574-6941.2012.01393.x
» https://doi.org/10.1111/j.1574-6941.2012.01393.x - Krulwich TA, Sachs G, Padan E. Molecular aspects of bacterial pH sensing and homeostasis. Nat Rev Microbiol. 2011;9:330-43. https://doi.org/10.1038/nrmicro2549
» https://doi.org/10.1038/nrmicro2549 - Lane DJ. 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M, editors. Nucleic acid techniques in bacterial systematics. New York: John Wiley & Sons; 1991. p. 115-75
- Laguerre G, Louvrier P, Allard M-R, Amarger N. Compatibility of rhizobial genotypes within natural populations of Rhizobium leguminosarum biovar viciae for nodulation of host legumes. Appl Environ Microb. 2003;69:2276-83. https://doi.org/10.1128/AEM.69.4.2276-2283.2003
» https://doi.org/10.1128/AEM.69.4.2276-2283.2003 - Lavelle P. Ecological challenges for soil science. Soil Sci. 2000;165:73-86. https://doi.org/10.1097/00010694-200001000-00009
» https://doi.org/10.1097/00010694-200001000-00009 - Leedjärv A, Ivask A, Virta M. Interplay of different transporters in the mediation of divalent heavy metal resistance in Pseudomonas putida KT2440. J Bacteriol. 2008;190:2680-9. https://doi.org/10.1128/JB.01494-07
» https://doi.org/10.1128/JB.01494-07 - Lei X, Wang ET, Chen WF, Sui XH, Chen WX. Diverse bacteria isolated from root nodules of wild Vicia species grown in temperate region of China. Arch Microbiol. 2008;190:657-71. https://doi.org/10.1007/s00203-008-0418-y
» https://doi.org/10.1007/s00203-008-0418-y - Lima AS, Nóbrega RSA, Barberi A, Silva K, Ferreira DF, Moreira FMS. Nitrogen-fixing bacteria communities occurring in soils under different uses in the Western Amazon Region as indicated by nodulation of siratro (Macroptilium atropurpureum). Plant Soil. 2009;319:127-45. https://doi.org/10.1007/s11104-008-9855-2
» https://doi.org/10.1007/s11104-008-9855-2 - Lourenzi CR, Ceretta CA, Brunetto G, Girotto E, Tiecher TL, Vieira RCB, Cancian A, Ferreira PAA. Pig slurry and nutrient accumulation and dry matter and grain yield in various crops. Rev Bras Cienc Solo. 2014;38:949-58. https://doi.org/10.1590/S0100-06832014000300027
» https://doi.org/10.1590/S0100-06832014000300027 - Matsuda A, Moreira FMS, Siqueira JO. Tolerância de rizóbios de diferentes procedências ao zinco, cobre e cádmio. Pesq Agropec Bras. 2002;37:343-55. https://doi.org/10.1590/S0100-204X2002000300016
» https://doi.org/10.1590/S0100-204X2002000300016 - Melloni R, Nóbrega RSA, Moreira FMS, Siqueira JO. Densidade e diversidade fenotípica de bactérias diazotróficas endofíticas em solos de mineração de bauxita, em reabilitação. Rev Bras Cienc Solo. 2004;28:85-93. https://doi.org/10.1590/S0100-06832004000100009
» https://doi.org/10.1590/S0100-06832004000100009 - MoreiraFMS, Huising EJ, Bignell DE. Manual de biologia dos solos tropicais: amostragem e caracterização da biodiversidade. Lavras: Editora UFLA; 2010.
- Moreira FMS, Siqueira JO. Microbiologia e bioquímica do solo. 2a ed atual e rev. Lavras: Editora UFLA; 2006.
- Moschetti G, Peluso AL, Protopapa A, Anastasio M, Pepe O, Defez R. Use of nodulation pattern, stress tolerance, nodC gene amplification, RAPD-PCR and RFLP-16S rDNA analysis to discriminate genotypes of Rhizobium leguminosarum biovar viciae. Syst Appl Microbiol. 2005;28:619-31. https://doi.org/10.1016/j.syapm.2005.03.009
» https://doi.org/10.1016/j.syapm.2005.03.009 - Moulin L, Chen WM, Béna G, Dreyfus B, Boivin-Masson C. Rhizobia: the family is expanding. In: Finan TM, O'Brian MR, Layzell DB, Vessey JK, Newton W, editors. Nitrogen fixation: global perspectives. New York: CABI Publishing; 2002. p. 61-5.
- Moura GGD, Armas RD, Giachini AJ, Meyer E, Soares, CRFS. Rhizobia Isolated from coal mining areas in the nodulation and growth of leguminous trees. Rev Bras Cienc Solo. 2016;40:e0150091. http://doi.org/10.1590/18069657rbcs20150091
» http://doi.org/10.1590/18069657rbcs20150091 - Nardoto GB, Ometto JPHB, Ehleringer JR, Higuchi N, Bastamante MMC, Martinelli LA. Understanding the influences of spatial patterns on N availability within the Brazilian Amazon Forest. Ecosystem. 2008;11:1234-46. https://doi.org/10.1007/s10021-008-9189-1
» https://doi.org/10.1007/s10021-008-9189-1 - Resh SC, Binkley D, Parrotta JA. Greater soil carbon sequestration under nitrogen-fixing trees compared with Eucalyptus species. Ecosystems. 2002;5:217-31. https://doi.org/10.1007/s10021-001-0067-3
» https://doi.org/10.1007/s10021-001-0067-3 - Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406-25. https://doi.org/10.1093/oxfordjournals.molbev.a040454
» https://doi.org/10.1093/oxfordjournals.molbev.a040454 - Scott AJ, Knott MA. A cluster analysis method for grouping means in the analysis of variance. Biometrics. 1974;30:507-12. https://doi.org/10.2307/2529204
» https://doi.org/10.2307/2529204 - Siddique I, Engel VL, Parrotta JA, Lamb D, Nardoto GB, Ometto JPHB, Martinelli LA, Schmidt S. Dominance of legume trees alters nutrient relations in mixed species forest restoration plantings within seven years. Biogeochemistry. 2008;88:89-101. https://doi.org/10.1007/s10533-008-9196-5
» https://doi.org/10.1007/s10533-008-9196-5 - Silva LFO, Vallejuelo Sf-O, Martinez-Arkarazo I, Castro K, Oliveira MLS, Sampaio CH, Brum IAS, Leão FB, Taffarel SR, Madariaga JM. Study of environmental pollution and mineralogical characterization of sediment rivers from Brazilian coal mining acid drainage. Sci Total Environ. 2013;447:169-78. https://doi.org/10.1016/j.scitotenv.2012.12.013
» https://doi.org/10.1016/j.scitotenv.2012.12.013 - Solomon D, Fritzsche F, Lehmann J, Tekalign M, Zech W. Soil organic matter dynamics in the subhumid agroecosystems of the Ethiopian Highlands. Soil Sci Soc Am J. 2002;66:969-78. https://doi.org/10.2136/sssaj2002.9690
» https://doi.org/10.2136/sssaj2002.9690 - Sriprang R, Hayashi M, Ono H, Takagi M, Hirata K, Murooka Y. Enhanced accumulation of Cd2+ by a Mesorhizobium sp. transformed with a gene from Arabidopsis thaliana coding for phytochelatin synthase. Appl Environ Microb. 2003;69:1791-6. https://doi.org/10.1128/AEM.69.3.1791-1796.2003
» https://doi.org/10.1128/AEM.69.3.1791-1796.2003 - Sriprang R, Hayashi M, Yamashita M, Ono H, Saeki K, Murooka Y. A novel bioremediation system for heavy metals using the symbiosis between leguminous plant and genetically engineered rhizobia. J Biotechnol. 2002;99:279-93. https://doi.org/10.1016/S0168-1656(02)00219-5
» https://doi.org/10.1016/S0168-1656(02)00219-5 - Staden R, Judge DP, Bonfield JK. Analyzing sequences using the Staden package and EMBOSS. In: Krawetz SA, Womble DD, editors. Introduction to bioinformatics: a theoretical and practical approach. New York: Springer Science+Business Media; 2003. p. 393-412.
- Stumpf L, Pauletto EA, Pinto LFS. Soil aggregation and root growth of perennial grasses in a constructed clay minesoil. Soil Till Res. 2016;161:71-8. https://doi.org/10.1016/j.still.2016.03.005
» https://doi.org/10.1016/j.still.2016.03.005 - Tamura K, Peterson D, Peterson N, Steche, G, Nei M, Kumar S. MEGA 5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol. 2011;28:2731-9. https://doi.org/10.1093/molbev/msr121
» https://doi.org/10.1093/molbev/msr121 - Trannin ICB, Moreira FMS, Siqueira JO, Lima A. Tolerância de estirpes e isolados de Bradyrhizobium e de Azorhizobium a zinco, cádmio e cobre “in vitro”. Rev Bras Cienc Solo. 2001;25:305-16. https://doi.org/10.1590/S0100-06832001000200007
» https://doi.org/10.1590/S0100-06832001000200007 - Valls M, Atrian S, Lorenzo V, Fernández LA. Engineering a mouse metallothionein on the cell surface of Ralstonia eutropha CH34 for immobilization of heavy metals in soil. Nat Biotechnol. 2000;18:661-5. https://doi.org/10.1038/76516
» https://doi.org/10.1038/76516 - van Berkum P, Beyene D, Vera FT, Keyser HH. Variability among Rhizobium strains originating from nodules of Vicia faba Appl Environ Microb. 1995;61:2649-53.
- Vázquez S, Agha R, Granado A, Sarro MJ, Esteban E, Peñalosa JM, Carpena RO. Use of white lupin plant for phytostabilization of Cd and As polluted acid soil. Water Air Soil Poll. 2006;177:349-65. https://doi.org/10.1007/s11270-006-9178-y
» https://doi.org/10.1007/s11270-006-9178-y - Vitousek PM, Howarth RW. Nitrogen limitation on land and in the sea: how can it occur? Biogeochemistry. 1991;13:87-115. https://doi.org/10.1007/BF00002772
» https://doi.org/10.1007/BF00002772 - Vos M, Wolf AB, Jennings SJ, Kowalchuk GA. Micro-scale determinants of bacterial diversity in soil. FEMS Microbial Rev. 2013;37:936-54. https://doi.org/10.1111/1574-6976.12023
» https://doi.org/10.1111/1574-6976.12023 - Xiao X, Luo S, Zeng G, Wei W, Wan Y, Chen L, Guo H, Cao Z, Yang L, Chen J, Xi Q. Biosorption of cadmium by endophytic fungus (EF) Microsphaeropsis sp. LSE10 isolated from cadmium hyperaccumulator Solanum nigrum L. Bioresource Technol. 2010;101:1668-74. https://doi.org/10.1016/j.biortech.2009.09.083
» https://doi.org/10.1016/j.biortech.2009.09.083 - Wani PA, Khan MS, Zaidi A. Effect of metal tolerant plant growth promoting Bradyrhizobium sp. (vigna) on growth, symbiosis, seed yield and metal uptake by greengram plants. Chemosphere. 2007;70:36-45. https://doi.org/10.1016/j.chemosphere.2007.07.028
» https://doi.org/10.1016/j.chemosphere.2007.07.028 - Weyens N, Beckers B, Schellingen K, Ceulemans R, Croes S, Janssen J, Haenen S, Witters N, Vangronsveld J. Plant-associated bacteria and their role in the success failure of metal phytoextraction projects: firstobservations of a field-related experiment. Microb Biotechnol. 2013;6:288-99. https://doi.org/10.1111/1751-7915.12038
» https://doi.org/10.1111/1751-7915.12038 - Wu CH, Wood TK, Mulchandani A, Chen W. Engineering plant-microbe symbiosis for rhizoremediation of heavy metals. Appl Environ Microb. 2006;72:1129-34. https://doi.org/10.1128/AEM.72.2.1129-1134.2006
» https://doi.org/10.1128/AEM.72.2.1129-1134.2006 - Zhan J, Sun Q. Diversity of free-living nitrogen-fixing microorganisms in wastelands of copper mine tailings during the process of natural ecological restoration. J Environ Sci. 2011;23:476-87. https://doi.org/10.1016/S1001-0742(10)60433-0
» https://doi.org/10.1016/S1001-0742(10)60433-0 - Zouboulis AI, Loukidou MX, Matis KA. Biosorption of toxic metals from aqueous solutions by bacteria strains isolated from metal-polluted soils. Process Biochem. 2004;39:909-16. https://doi.org/10.1016/S0032-9592(03)00200-0l
» https://doi.org/10.1016/S0032-9592(03)00200-0l
Publication Dates
-
Publication in this collection
13 Sept 2018 -
Date of issue
2018
History
-
Received
20 June 2017 -
Accepted
19 Feb 2018