Abstracts
Resistance to nearly all pathogens occurs abundantly in our crops. Much of the resistance exploited by breeders is of the major gene type. Polygenic resistance, although used much less, is even more abundantly available. Many types of resistance are highly elusive, the pathogen apparently adapting very easily them. Other types of resistance, the so-called durable resistance, remain effective much longer. The elusive resistance is invariably of the monogenic type and usually of the hypersensitive type directed against specialised pathogens. Race-specificity is not the cause of elusive resistance but the consequence of it. Understanding acquired resistance may open interesting approaches to control pathogens. This is even truer for molecular techniques, which already represent an enourmously wide range of possibilities. Resistance obtained through transformation is often of the quantitative type and may be durable in most cases.
types of resistance; genetics of resistance; acquired resistance
Na natureza a resistência à maioria das doenças ocorre nas culturas. A maior parte da resistência explorada pelos melhoristas envolve genes maiores ou principais. Resistência poligênica, embora seja muito menos utilizada nos programas de melhoramento, encontra-se em maior disponibilidade nos cultivares. Muitos tipos de resistência são altamente temporária; o patógeno aparentemente adapta-se muito facilmente a estes tipos de resistência. Outros tipos de resistência permanecem efetivos por muito mais tempo e é chamado resistência durável. A resistência temporária é invariavelmente do tipo monogênica, e usualmente é do tipo hipersensível e atua sobre patógenos especializados. Raça-específica não é a causa da resistência temporária mas sua conseqüência. A compreensão sobre a resistência adquirida abre interessante caminho para o controle de fitopatógenos. Isto é mais válido ainda para técnicas moleculares, que já apresentam enorme gama de possibilidades. Resistência obtida através da transformação é sempre do tipo quantitativa e pode ser durável em muitos casos.
REVISÃO / REVIEW
CONCEPTS IN PLANT DISEASE RESISTANCE
FRANCISCO XAVIER RIBEIRO DO VALE1, J. E. PARLEVLIET2 & LAÉRCIO ZAMBOLIM1
1Departamento de Fitopatologia, Universidade Federal de Viçosa, CEP 36571-000, Viçosa, MG, Brasil, e-mail: dovale@mail.ufv.br; 2Department of Plant Breeding (IVP), Wageningen Agricultural University, P.O. Box 386, 6700 AJ Wageningen, The Netherlands
(Aceito para publicação em 13/08/2001)
Autor para correspondência: Francisco Xavier Ribeiro do Vale
ABSTRACT
Resistance to nearly all pathogens occurs abundantly in our crops. Much of the resistance exploited by breeders is of the major gene type. Polygenic resistance, although used much less, is even more abundantly available. Many types of resistance are highly elusive, the pathogen apparently adapting very easily them. Other types of resistance, the so-called durable resistance, remain effective much longer. The elusive resistance is invariably of the monogenic type and usually of the hypersensitive type directed against specialised pathogens. Race-specificity is not the cause of elusive resistance but the consequence of it. Understanding acquired resistance may open interesting approaches to control pathogens. This is even truer for molecular techniques, which already represent an enourmously wide range of possibilities. Resistance obtained through transformation is often of the quantitative type and may be durable in most cases.
Key words: types of resistance; genetics of resistance; acquired resistance.
RESUMO
Conceitos em resistência de plantas a doenças
Na natureza a resistência à maioria das doenças ocorre nas culturas. A maior parte da resistência explorada pelos melhoristas envolve genes maiores ou principais. Resistência poligênica, embora seja muito menos utilizada nos programas de melhoramento, encontra-se em maior disponibilidade nos cultivares. Muitos tipos de resistência são altamente temporária; o patógeno aparentemente adapta-se muito facilmente a estes tipos de resistência. Outros tipos de resistência permanecem efetivos por muito mais tempo e é chamado resistência durável. A resistência temporária é invariavelmente do tipo monogênica, e usualmente é do tipo hipersensível e atua sobre patógenos especializados. Raça-específica não é a causa da resistência temporária mas sua conseqüência. A compreensão sobre a resistência adquirida abre interessante caminho para o controle de fitopatógenos. Isto é mais válido ainda para técnicas moleculares, que já apresentam enorme gama de possibilidades. Resistência obtida através da transformação é sempre do tipo quantitativa e pode ser durável em muitos casos.
INTRODUCTION
In nature organisms are classified as producers, green plants, consumers (organisms exploiting other organisms), and decomposers (organisms using dead organisms). Green plants, including our crops, are used by a multitude of consumers of almost every kind, from various types of herbivores (mammals, snails, insects) to typical parasites (insects, mites, fungi, bacteria). In order to survive green plants developed a broad range of defence mechanisms to ward off most of these consumers. These defence mechanisms are principally based on avoidance, resistance or tolerance. Avoidance operates before parasitic contact between host and parasite is established and decreases the frequency of incidence. After parasitic contact has been established the host may resist the parasite by decreasing its growth, or tolerate its presence by suffering relatively little damage.
Avoidance is mainly active against animal parasites and includes such diverse mechanisms as volatile repellents, mimicry and morphological features like hairs, thorns and resin ducts. Resistance is usually of a chemical nature. Little is known of tolerance; it is very difficult to measure and is usually confounded with quantitative forms of resistance.
Parasites classified as fungi, bacteria, viruses or viroids are considered disease inciting parasites or pathogens. Resistance mechanisms are by far the most important defence mechanisms employed by host plants, including our crops, against pathogens. Avoidance and tolerance play a minor role here. In the never-ending arms race between plant and pathogen, the latter have developed widely different host ranges. Pathogens such as some Pythium species, Rhizoctonia solani Kühn, and Sclerotinia sclerotiorum (Lib.) de Bary have a wide host range; they are non-specialized, polyphagous pathogens or generalists. The latter one, for instance, has been reported to attack hundreds of plant species belonging to at least 64 families of flowering plants and gymnosperms (Parlevliet, 1989). A large proportion of the pathogens, however, have a narrow host range restricted to a few closely related plant species; they are specialized, monophagous pathogens or specialists. Puccinia hordei Otth. and Phytophthora phaseoli, pathogenic on barley (Hordeum vulgare L.) and lima beans (Phaseolus lunatus L.) respectively, are typical specialists.
As resistance is by far the most important defence of plants against pathogens this chapter discusses the various aspects of disease resistance.
MEASURING RESISTANCE
Selection for resistance implies measurements of plant- resistance. Ideally one should measure the amount of pathogen present at a given moment compared with the amount present on or in a extremely susceptible cultivar. The larger the difference in amount the larger the difference in susceptibility/resistance. It is normally not possible to measure the amount of pathogen, because the pathogen is either not visible or only partially so. However, one can evaluate the direct or indirect effects of the pathogen on the host even if the pathogen itself is not visible (Parlevliet, 1993).
The quantitative or partial resistance of a host cultivar cannot be assessed in absolute terms; it is always a relative measure compared with that of a well-known standard cultivar. This standard cultivar is often the most susceptible cultivar available (Parlevliet, 1989).
The amount of tissue affected is, in general, a good estimator of the amount of pathogen present. The amount of pathogen present, however, is not jsut dependent on the level of resistance of the host cultivar. Other factors may and do interfere with it such as:
Interplot interference
Van der Plank (1963) stated: "Plots in the experiment are meant to represent farmers' fields receiving the same treatment as these fields receive". But plots represent fields only when the plots within an experimental area do not interfere with one another. The representational error - the error of taking plots to represent fields when they do not can be large.
The most frequent causes of errors in interpreting results are experimental interplot interference (Bainbridge & Jenkyn, 1976; Jenkyn et al.,1983; Parlevliet & Van Ommeren, 1984; Parlevliet & Van Ommeren, 1975) and/or interactions between host or pathogen with the environment (Van der Plank, 1968; Colhoun, 1973; Jeger et al., 1983; Fraser, 1985; Hunter et al.,1986; Falkhof et al. 1988). This applies especially to partial resistance, which, together with the aggressivity of the pathogen in general, is a quantitative characteristic and is influenced by changes in the environment.
Parlevliet & Van Ommeren (1975) observed a very strong interplot interference in barley evaluated for partial resistance to leaf rust. In plots well isolated from one another the most susceptible cultivar had more than 2,000 times more uredosori than the least susceptible one. In plots of less than 1m2 and adjacent to each other the difference between the extremes was not more than 25 times.
The interplot interference may not only underestimate the partial resistance, but it may cause the ranking order of cultivars compared to the one in large isolated plots (Norgaard Knudsen et al., 1986). The authors concluded that for a reliable selection of partial resistance, plots of some 1,4 m2 were advisable.
However, not all airborne pathogens cause interplot interference. In wheat (Triticum aestivum L.) against stripe rust no interplot interference of any significance could be observed, and even single hill plots neighbouring spreader rows give a representative assessment of resistance (Parlevliet & Daniel, 1992). Interplot interference in airborne pathogens appears to vary greatly. It seems that this interference is greater the more the total disease is based on more and smaller individual infections.
Relation between disease symptoms and amount of the pathogen
True disease symptoms are observed with several pathogens such as wilting caused by vascular pathogens, and leaf rolling, mottling, stunting, etc., caused by viruses. These symptoms tend to be rather unreliable for assessing resistance, since the relationship between the amount of pathogen present and the severity of symptoms is often poor.
In other cases the pathogen itself is observed, making assessment much easier and far more reliable. The ecoparasitic powdery mildews are good examples; their mycelia remain on the surface of the host epidermis and are visible as white to grey spots. In rust diseases the pathogen becomes visible when the sporulating infections rupture the epidermis, exposing powdery masses of spores.
A large number of pathogens fall between these two types. The pathogen itself is not visible, but the symptoms of its presence are more or less easily discernible and restricted to the parts of the host tissue invaded. Discoloration of the invaded tissue and immediately adjacent tissue is the most general indication of the pathogen. The reliability of this measure of pathogen presence or inversely, host resistance, varies with host and pathogen but tends to be fairly good in many cases.
Inoculum Density
This factor may obscure real differences in quantitative resistance. In order to prevent escape of genotypes from infection, there is a tendency to apply high inoculum densities. Complete resistance in such cases is easily detectable, but small differences in susceptibility tend to disappear. The optimal inoculum density is the density whereby escapes are largely prevented while only the most susceptible cultivars are strongly affected (Parlevliet, 1989).
Earliness
If the entries differ considerably in earliness the period of exposure to the pathogen varies greatly as the assessment is usually done at the same moment for all entries. Resistance to head blight caused by Fusarium in wheat is considerably overestimated in late cultivars due to this aspect (Parlevliet, 1993).
Plant Habitat
In dense crops and short plants the amount of tissue affected tends to increase. In loose crops and tall plants it tends to decrease. This is probably due to micro-climatic effects. Short wheat cultivars are more affected than tall cultivars by Septoria leaf and glume blotch (Parlevliet, 1993).
GENETICS OF RESISTANCE
If published research is representative of the resistance present it is most often controlled by major genes. These major genes are often inherited dominantly, less frequently recessively. Polygenic inheritance of resistance has been reported as well, but its much lower frequency is most likely due to the more difficult nature of the research than to a truly lower frequency.
Major resistance genes often occur in a surprisingly high numbers. In coffee (Coffea arabica L.) - Hemileia vastatrix Berk. & Br., maize (Zea mays L.) - Puccinia sorghi Schw., oats (Avena sativa L.) - P. coronata Cda., wheat [Triticum aestivum (L.) Thell.] - P. graminis f. sp. tritici Pers., wheat - P. recondita, wheat - P. striiformis Westend, barley - Erysiphe graminis D.C. f. sp. hordei, flax (Linum usitatissimun L.)- Melampsora lini (Ehrenb.) Desmaz. from over 20 to over 50 major resistance genes are known. These resistance genes often cluster together in certain chromosome arms, sometimes so tightly that they can be considered as complex loci, and true allelic series also occur. In the flax-flax rust pathosystem 34 R-genes have been identified in seven regions - K(2), L(14), M(7), N(3), P(6), D(1), Q(1). Regions N and P are linked, as well as regions N and K. The N region consists of at least two closely linked loci. The M region, together with seven resistance alleles, also consists of some closely linked loci. The L region, with 14 resistance alleles, behaves as a locus with an allelic series, but intra-allelic recombination has been reported (Islam and Shepherd, 1991). In barley most of the resistance genes to powdery mildew are located on one arm of chromosome 5 and one arm of chromosome 4 (Jorgensen, 1990). On the short arm of chromosome 10 in maize, at least 16 resistance genes to P. sorghi are found on the complex locus Rp1 and the loci Rp5 and Rp6 within three centimorgans of each other (Saxena and Hooker, 1968). The three downy mildew [Peronospora effusa (Grev.) Tul.] resistance genes known in spinach are tightly linked.
Minor or polygenic resistance has been reported fairly often. Typical examples are the quantitative resistance in maize to the northern [Setosphaeria turcica (Luttr.) Leonard & Suggs] and southern (Cochliobolus heterostrophus Drechsler) leaf blight (Jenkins and Robert, 1961; Burnette, and White, 1985) in barley to Puccinia hordei Otth. (Parlevliet, 1978), in wheat to P. recondita f. sp. tritici (Broers and Jacobs, 1989) and in rice to bacterial blight (Koch and Parlevliet, 1991). Minor genic or polygenic resistance is almost certainly ubiquitous as is discussed in the sub-chapter on quantitative resistance.
The expression of resistance genes can be modified by the action of other genes (epistasis), the development stage or tissue of the plant or the environment. The major resistance gene Pa7 in barley to P. hordei gives complete resistance in 'Cebada Capa', but incomplete resistance in the cultivars L94, Zephyr and Vada. In cereals, adult plant resistance genes to the various rusts occur frequently; they give resistance only in the adult plant stages. In the seedling stage the resistance is not expressed. In potatoes (Solanum tuberosum L.) the quantitative resistance in the foliage to late blight, Phytophthora infestans (Mont.) de Bary is poorly correlated with the resistance in the tubers, indicating that different genes are involved or expressed in the different tissues of the plant (Anonymous, 1958-1998). Of the environmental factors temperature plays a major role. Many scientists have reported that the expression of certain resistance genes depends on the temperature to which they are exposed, such as L1, L3, L7, L8, L10 and L11 in flax to flax rust (Islam and Mayo, 1990), Sr6 in wheat to stem rust, Lr16 and Lr17 in wheat to leaf rust and many other genes (Browder, 1985).
The gene-for-gene concept
Many major resistance genes operate in a gene-for-gene way. For each resistance gene in the host there is a corresponding avirulence gene in the pathogen (Flor, 1971), and only the corresponding avirulence gene can initiate the hypersensitive reaction (HR) leading to incompatibility. Resistance and avirulence inherit in most cases in a dominant manner, susceptibility and virulence in a recessive way. The HR is now known to result from the specific interaction at the cellular level of the product of the resistance gene and the product of the avirulence gene. If one of the two products is absent, there is no incompatibility; the normal pathogenicity of the pathogen results in a compatible reaction (the host appears susceptible). What is normally meant with virulence is actually the normal pathogenicity shown in the absence of avirulence. Virulence is absence of avirulence, it is genetically seen as an empty concept; there are no virulence genes.
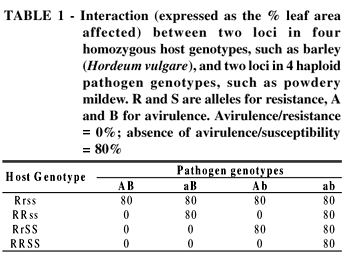
Host and pathogen genotypes interact with each other. Each of the four pathogen genotypes shows a different reaction pattern with the four host cultivars (Table 1). This makes it QR is, except for a few cases of monogenic, incomplete inheritance, inherited oligogenically or polygenically. Examples of the former are the QR of maize to P. sorghi (Kim and Brewbaker, 1977) and of wheat to P. recondita possible to differentiate between the four pathogen genotypes (races), but only because the interaction effects, which are equal to the gene effects, are so large (difference between 80% and 0%). The four races are identifiable through the avirulence genes they carry. The resistance of the host cultivar is dependent on the race; this is called race-specific resistance.
This gene-for-gene system appears to be widespread. It occurs most clearly and frequently in pathosystems where a biotrophic, highly specialized (in terms of host range) pathogen is involved such as the cereals with the various rusts, smuts and bunts, and with the powdery mildew. The resistance in these systems is typically race-specific and also very easily neutralized by new races of the pathogen; the resistance is highly elusive, with the exception of pathosystems involving viruses.
QUANTITATIVE RESISTANCE
Resistance, like other traits, occurs in a qualitative or in a quantitative way. With the former the different genotypes in a population occur as discernible phenotypes; it is usually controlled by a major gene. Quantitative resistance (QR) is defined as a resistance that varies in a continuous way between the various phenotypes of the host population, from almost imperceptible (only a slight reduction in the growth of the pathogen) to quite strong (little growth of the pathogen). This resistance is often indicated with other terms such as partial, residual and field resistance or even (wrongly) with tolerance. QR occurs at various levels to nearly all important pathogens in most cultivars of our crops.
i) Barley and barley leaf rust, P. hordei. The QR to this pathogen inherits polygenically (Parlevliet, 1978) and all cultivars in Western Europe, including the very susceptible cultivars, carry at least some QR. Most cultivars, though, carry considerable levels of QR, preventing the barley leaf rust from becoming a major pathogen in Western Europe (Parlevliet, 1989). In Ethiopia the barley landraces represent a centre of diversity. Barley genotypes without any QR are very rare (Fekadu and Parlevliet, 1996).
ii) Rice (Oryza sativa L.) and bacterial blight, Xanthomonas campestris pv. oryzae (Ishiyama) Dye. From a cross between two very susceptible cultivars some lines were obtained that were considerably more susceptible than either parent, while a few other progeny lines were moderately resistant. This clear transgression beyond the parental values meant that both highly susceptible cultivars carried minor genes for QR that differed from each other (Koch & Parlevliet, 1991). Thus, even so called very susceptible genotypes may harbour some QR confirming the experience with barley leaf rust.
iii) Similar observations were reported from quantitative trait loci(QTL)-analyses done in the pathosystems maize/Cercospora zeae-maydis Tehon & Daniels, pea Pisum sativum L./Ascochyta pisi Lib. and tomato (Lycopersicon esculentum Mill.)/Ralstonia solanacearum (Smith 1896) Yabuuchi et al. 1996. In crosses between a susceptible and a QR parent QTLs for QR were found that originated not only from the QR parent, but also from the susceptible parent (Young, 1996).
iv) Wheat/yellow rust, wheat/leaf rust, and barley/powdery mildew. Breeding in Western Europe is directed at selecting major genes of the non-durable type to protect against these three major pathogens. In this way, new recommended cultivars tend to enter the recommended cultivar list with high scores for resistance. After a number of years these scores are much lower as the major gene resistance is not effective any more. After the resistance "breaks down", QR becomes visible if present. All cultivars selected for their major gene resistance appear to carry moderate to fair levels of QR hidden behind that major gene (Anonymous 1958-1998). This hidden QR is sometimes indicated as residual resistance.
v) Potatoes have a range of viruses that may affect them. Apart from major resistance genes QR also exists to those viruses. This QR is often expressed through a reduced frequency of infected plants (incidence). The Dutch recommended list of potato cultivars discerns between major gene resistance and QR. All potato cultivars listed in the period 1958 to present carry low to high levels of QR to each of the four viruses assessed, Potato virus X, Potato virus Y and Potato virus A and Leaf roll virus (Anonymous, 1958-1998).
Therefore, QR is present almost everywhere. Cultivars without any QR are very rare. For this type of resistance breeders do not need to look for primitive genotypes from centres of diversity nor to related wild species. The resistance is near at hand in adapted cultivars, a fortunate situation as it makes breeding easier. McIntosh (1997) concluded that the ideal sources of resistance are those present in closely related, commercial genotypes, and any effort to transfer resistance from related species and genera should be considered long term.
QR is, except for a few cases of monogenic, incomplete inheritance, inherited oligogenically or polygenically. Examples of the former are the QR of maize to P. sorghi (Kim and Brewbaker, 1977) and of wheat to P. recondita (Broers and Jacobs, 1989), based on a few (two or three) additive genes. Polygenic QR is exemplified by the field resistance of potato to P. infestans (Black, 1970), the QR of maize to Cochliobolus heterostrophus Drechsler and Setosphaeria turcica (Lutrell) Leonard & Sugg (Leonard, 1993), in rice to bacterial blight (Koch and Parlevliet, 1991a) and of barley to Rhynchosporium secalis (Oud.) Davis. (Habgood, 1974, 1976) and to P. hordei (Parlevliet, 1978a), and of rice to Magnaporthe grisea (T.T. Hebert) Yagashi & Udagawa (Roumen, 1994). Polygenic QR is usually supposed to be non race specific, but does not appear to be so. In six of the seven polygenic pathosystems mentioned above, small race-specific effects have been reported (Parlevliet, 1997), and it is probable that polygenic resistance to specialized pathogens often goes together with small race-specific effects. Parlevliet and Zadoks (1977) described this in the following way: When resistance in the host and aggressiveness in the pathogen interact on a polygene-for-polygene basis and several host cultivars are tested against a series of pathogen isolates the general impression is that of non-race-specificity. Most variation is between cultivars and between isolates. If the accuracy of the experiment is sufficiently high small but significant race-specific effects can be observed.
Components of partial resistance
QR is expressed as a reduced amount of tissue in the invaded or affected host compared with that of a highly susceptible standard. When the total amount of disease is the collective result of a large number of discrete lesions, it is possible to identify a number of components contributing to the amount of tissue affected, as in the case of the cereal rusts (Parlevliet,1992). QR may reduce the chance of infection, resulting in fewer lesions. It may reduce the growth of the pathogen once the infection is successfull, causing smaller lesions that may sporulate less. It is possible to discern at least three components of QR against pathogens that are not systemic; infection frequency, lesion size and sporulation rate per lesion. Associated with lesion size and sporulation rate is the latency period, the period between infection and first spore production. Also, the effective life length of lesions may play a role (the infectious period), albeit a small one, as the epidemic development of the first spores produced by a lesion are far more important than the ones produced later. This also means that in the case of polycyclic pathogens, the latency period is a highly important component. A short latency period is essential for the pathogen, a long one for good QR. For the same reason the infectious period is much less important. Also, for this reason total spore production measured over a period of time, considered by Johnson & Taylor (1976) as an accurate measure of QR, and assumed to reflect the sum of the components effects, cannot be the right approach. Another reason why their conclusion cannot be right is the fact that spore production, except at very low pathogen densities, is determined by the host tissue area itself and not by the host tissue area invaded because of the strong interference between spore production per lesion and the lesion density (Mehta & Zadoks, 1970; Teng & Close, 1978).
Components tend to be associated with one another in most pathosystems. This association varies from very strong in barley/P. hordei to less strong in potato/P. infestans to no association in rice/X. campestris pv oryzae (Parlevliet, 1992).
Assessing QR through one or more of its components is usually not advisable if the aim is selection for QR. The major reason for this is that assessing components is more laborious and not always more accurate than assessing QR directly in the field (Parlevliet, 1992). Measurements of spore production per unit host area, and infection frequency are especially laborious.
Epidemiology of quantitative resistance
In polycyclic pathogens the epidemic builds up each season; the higher the level of QR, the lower the rate of build-up. Van der Plank (1963, 1968) expressed the rate of build-up with the value r, which gets smaller QR increases. He therefore called this resistance rate-reducing resistance. Unfortunately, r not only depends on the level of QR but also on several other factors, such as the development stage of the epidemic and the growth habit of the cultivar (short or tall, early or late, loose or dense foliage). The value r is, therefore, a very inaccurate measure for QR and certainly of no value for breeders as its assessment is quite laborious.
A much more often used method is measuring the area according to the disease progress curve. This estimates the accumulated disease severity (DS) by assessing the DS several times. This method is only marginally better and much more laborious than measuring the DS at one moment, when the most diseased genotype is well affected. Measuring DS is, as a whole, the best method to assess QR, but one has to realize that various factors may interfere with it, as described in "measuring resistance".
Some scientists use the degree of yield reduction to measure resistance. This, however does not measure resistance alone because it also includes the consequence of tolerance. At the same time, it is a very inaccurate way of measuring as yield is very sensitive to genotype (g) x environment (e) interactions, while the trait DS is much less sensitive to g x e interactions.
In monocyclic pathogens, often of a soil borne nature, the epidemic builds up over the seasons (Van der Plank, 1963). The higher the QR, the slower the build up will be over time. Measuring QR is done in a similar way. The most important requirement is, as with polycyclic pathogens, a uniform and not too severe exposure to the pathogens of the entries to be tested. This is not always easy with soil borne pathogens, which are usually distributed quite heterogeneously in infested fields.
With systemic pathogens, such as most viral pathogens, the DS is often expressed as the percentage of plants diseased, or the incidence. QR reduces the incidence both within the growing season and measured over the seasons (see potato viruses in "quantitative resistance").
Therefore, in all situations QR reduces the rate of epidemic build up. This allows for more time to use other control methods. QR is, therefore, well suited to use in integrated disease management programmes; it enhances the effects of other control methods.
Being a quantitative trait, it is often thought that QR will be sensitive to genotype x environmental interactions (Geiger & Heun, 1989). This, however, is most likely not true. The partial resistance in barley-to-barley leaf rust and in wheat to leaf rust is expressed under a wide range of conditions. The quantitative traits that are sensitive to genotype x environment interactions are in fact complex traits, traits that are the accumulated result of a number of other traits, such as yield. Yield is the result of a large number of traits, such as earliness, plant length, disease resistance, drought tolerance, etc. and each of these contributing traits may react differently to different environments, hence resulting in strong g x e interactions. QR, however, is not a complex trait; the QR genes only affect the trait QR, which is the reason why QR is not more sensitive to g x e interactions than qualitative resistance.
NON-DURABLE RESISTANCE
In nature there is a constant race of arms between the attacking parasite and the defending host, and in the evolutionary sense, all resistance is transitory. But large differences exist in the ease by which parasites can overcome a resistance. In agriculture, too the durability of a resistance varies greatly. Resistance may already be neutralized in the last stages of the breeding program (at zero years) and may, still be effective after more than 130 years and wide exposure, as the case of the Phylloxera aphid resistance of grape (Vitis vinifera L.) rootstocks (Niks et al. 1993). And even among the non-durable types of resistance clear differences exist (Table 2).
Resistance is considered durable if it remains effective when used for many years over a substantial area (Johnson, 1981). Much of the resistance used by breeders has not broken down; multiple major gene resistance and all QR based on some to several genes appeared to be durable (Parlevliet, 1997).
But it is possible to discern some patterns. There are many types of resistance that appeared highly elusive, the effective time ranging from less than one year to several years. A major gene including the hypersensitive reaction almost invariably controls this clearly non-durable resistance. These genes operate in a gene-for-gene way with avirulence genes in the pathogen (see "genetics of resistance"). The pathogens are mostly specialized fungi or bacteria such as the cereal rusts, the cereal powdery mildew, downy mildews of lettuce and of lima bean, flax rust of flax, rusts of maize, late blight of potato, blast of rice, bean rust of common bean (Phaseolus vulgaris L.), anthracnose of common bean and bacterial blight of rice (Parlevliet, 1993). The host population invariably carries many different resistance genes (see "genetics of resistance" and Table 3) and the pathogen population is characterized by many races.
However, not all major gene resistance of the hypersensitive type is so elusive, and some major gene resistance of the non-hypersensitive type appeared highly elusive. Hypersensitive resistance to viruses is often fairly durable even if corresponding strains occur (Fraser, 1992; Parlevliet, 1993; 1997), while the non-hypersensitive major resistance gene Tm-1 of tomato to tobacco mosaic virus (Fraser, 1992) and some resistance to the non-specialized root knot nematode Meloidogyne incognita (Kofoid & White) Chitwood (Roberts, 1995) were highly elusive.
QR based on some to several genes with additive effects has never been reported to be elusive (Parlevliet, 1997). Between the highly elusive resistance (the hypersensitive resistance to specialized fungi and bacteria) and the resistance that remains effective even after a long period there is resistance that lasted for a considerable time but ultimately became ineffective. The resistance of barley governed by gene Rpg1 to the wheat stem rust in North America is a good example. The resistance remained effective for over 50 years, despite the fact that resistant barley was grown yearly over a million ha and over the years several races were observed that appeared virulent on Rpg1, but which did not establish. It broke down effectively in 1989 when race QCC appeared (Steffenson, 1992).
From the above it is clear that it is not possible to clearly discern two groups of resistance as non-durable and durable. Between the typical non-durable resistance and the typical durable resistance there are types of resistance difficult to classify. Are the resistances of Tadorna, Arminda and Minerva (Table 2) to be considered as durable? They were considered to be at the time until the resistance broke down. The lasting resistance of Tadorna and of Arminda appeared to be based on the presence of two resistance genes, unintentionally came together in a single cultivar; these genes are also present individually in other cultivars that typically behaved in a non-durable manner. Minerva's resistance, however, was based on a single gene.
DURABLE RESISTANCE
Resistance is considered durable when it remains effective for a considerable time, despite wide exposure. In this sense, it is a quantitative concept. The Rpg1 gene discussed above was durable, but did not last forever. And in the evolutionary sense, no resistance will last forever. It is possible to discern three groups of resistances that are predominantly durable.
1 - Resistance to pathogens with a wide host range, generalists
Are usually of a quantitative nature (Bruehl, 1983) and nearly always durable (Parlevliet, 1993). But there are exceptions, such as the major resistance genes Mi in tomato and Rk in cow pea against the root knot nematode, M. incognita (Roberts, 1995).
2 - QR against specialists and based on some to several genes with additive effects seems durable.
In the few cases where reported QR broke down, the resistance appeared to be monogenic, like the field resistance against rice blast, M. grisea, in the rice cultivar St-1. The resistance became ineffective within a few years and appeared to be based on a single dominant gene Pi-f (Toriyama, 1975).
3 - Monogenic resistance against specialists of a non-hypersensitive nature.
Such resistance is often quite durable. The non-hypersensitive resistance genes Rpg2 (sr-2) and Rpr34 (Lr-34) of wheat to stem rust and leaf rust respectively and the mlo-gene of barley to powdery mildew have already lasted for a considerable time.
Usually, the presence of race-specific resistance effects is considered as evidence of non-durable resistance. This idea, however, cannot be maintained. There are too many examples of race-specific resistance that lasted a long time or that are still effective (Parlevliet, 1993;1997). The Rpg1 gene of barley discussed above is such an example. The hypersensitive resistance genes Nx and Nb to virus X in potato are still effective in Europe, despite the fact that the corresponding virulent strains have been present for many years but at very low frequencies (Cockerham, 1955; Parlevliet, 1993). Meiners (1981) concluded that all known resistance to pea viruses is race-specific in nature but this resistance appears to last for a long time.
Polygenic resistance is usually considered to be non-race-specific and durable. However, in several host-pathogen systems with demonstrated durability, polygenic resistance race-specific effects were demonstrated (Parlevliet, 1997). Parlevliet and Zadoks (1997) realized that in gene-for-gene relationships the race-specific effects are of the same size as the gene effects. Major resistance genes therefore are associated with clear, identifiable races, while polygenes result in only small race-specific effects, insufficient for unambiguously identifying races. In this latter case, one gets a general impression of non-race-specificity. Parlevliet and Zadoks (1977) also compared two models of polygenic pathosystems with one another. In system I the host polygenes operated in a gene-for-gene interaction with the pathogen polygenes, causing small race-specific effects. In system II the host polygenes did not operate in a gene-for-gene way with the pathogen polygenes, representing true horizontal resistance. The two models showed that the polygene-for-polygene system was more durable (less liable to adaptation in the pathogen population) than the system where the polygenes were not operating in a gene-for-gene system, assuming all other variables, such as resistance mechanisms, are the same.
The durability of resistance also may depend on the circumstances. The major gene resistance of flax to flax rust was not durable in the USA, but it has been durable in The Netherlands (Parlevliet, 1993). In the latter situation, flax has been a small crop and all flax grown is completely resistant. The pathogen population was, therefore, reduced so strongly that new races have little chance to emerge.
WHAT RESISTANCE IS OPERATING IN OUR CROPS?
Conscious breeding for disease resistance started early in this century and concentrated on major resistance genes. Soon after resistant cultivars were released farmers were confronted with the versatility of the pathogen; new forms of the pathogen neutralized the introduced resistance (the resistance was said to be "broken"). This did not stop breeders from using such non-durable resistance. On the contrary, the rapid increase of man's effort to produce resistant crops based on major resistance genes was accompanied by an equally rapid increase in "broken" resistance. Even today the situation has not changed much. Screening and selection methods favouring major gene resistance are still widely used by breeders. As a result of the widespread use of such screening and selection methods QR has been used considerably less than major gene resistance, although abundantly present to most pathogens (Parlevliet, 1997).
ACQUIRED RESISTANCE
Some susceptible plants become systemically resistant in response to localized infections, a phenomenon known as acquired resistance. This is best known in cucurbits and tobacco. When a lower leaf is infected, the whole plant becomes resistant to the same and to other pathogens and remains so for weeks. Plants with acquired resistance have high levels of pathogenesis - related proteins, salicylic acid, peroxidase, and other factors (Scheffer, 1997). Obviously, there is a signalling mechanism that carries information to distant parts of the plant, but the nature of the signals is unknown (Hammerschmidt and Kuc, 1995).
Sequeira (1983) pointed out that the terminology used in this field is confusing. The terms acquired resistance, activated resistance, acquired immunity, pre-munity, immunization, sensitisation as well as cross protection have been used as synonym for induced resistance to describe a range of related phenomena.
Induced resistance means the enhancement of resistance in plants that are otherwise susceptible, in response to an extrinsic stimulus without alterations of the genome. The inducing agents can be of biotic or abiotic nature (Schonbeck & Steiner, 1997).
Induced resistance exists in two different forms. It may be localized at the site of the inducing treatment or it may be systemic. The latter is commonly called SAR (Systemic acquired or activated resistance) and is effective in some, all or newly emerging plant parts distant from the site of induction (Schonbeck & Steiner, 1997).
The early pioneer research with phytoalexins established many basic principles for induced resistance. Plants respond to pathogens and non-pathogens by synthesizing low molecular weight, generally lipophilic compounds which inhibit the growth of fungi and bacteria in vitro and accumulate at the sites of infection to levels that inhibit the development of some pathogens (Kuc, 1995).
It is clear that phytoalexin structures and activity do not explain the often high specificity in plant-pathogen interaction. Biotic and abiotic agents cause phytoalexin synthesis and accumulation. The genes for the synthesis of phytoalexins are present in resistant and susceptible plants, even those reported to lack genes for resistance to pathogen (Kuc & Strobel, 1992).
Certain fungal pathogens, especially in the genera Alternaria and Cochliobolus, are known to produce host-specific toxins or host-selective toxins (HSTs) as agents of virulence or pathogenicity. In the genus Alternaria, at least nine examples of HST-producing pathogens have been reported ( Kohmoto & Itabum 1991; Otani et al., 1995).
Extensive studies on the mode of action of these HSTs has been done, leading to an understanding of their importance, not only to induce necrosis, but to suppress host defences, preparing the plant for infection by fungus (Kohmoto & Otani, 1991; Otani et al., 1995; Nakashima, et al., 1985). Subsequently with the advance of molecular biology, one of the main interests in HST research is the molecular analysis of the genes involved in HST biosynthesis, because these genes correspond to genes for pathogenicity or virulence. However, investigations into the genetic control of HST biosynthesis and pathogenicity in Alternaria pathogens have been limited due to their lack of a sexual cycle (Kodama et al., 1998).
Gene tagging utilizing the heterologous integration of plasmids in now widely used to clone genes where little biochemical is available. The development of the so-called REMI (restriction enzyme mediated integration) method has substantially improved this technique and some genes important for fungal pathogenicity have already been tagged and cloned (Kodama, 1998).
Advances have been made in the discovery of host-specific toxins, their chemistry, their sites of action and physiological effects on host plants, their roles in fungal pathogenesis, and the genetics of toxin production by fungi (Yoder, 1998). Small molecules have been implicated in systemic induced resistance, although their role appears to be more important for signal transduction than for directly inhibiting pathogens. The best studied is salicylic acid (SA), a compound derived from cinnamic acid (Stermer, 1995)
RESISTANCE PRODUCED THROUGH TRANSFORMATION: WHAT KIND IS IT?
In the past 80 years resistance breeding has increased greatly to become a highly important, if not indispensable part of crop improvement. Screening and selection methods have been were refined and have became more efficient. Our scientific knowledge has increased even more. Recently the molecular techniques have revolutionized our technical possibilities. With these new transformation techniques the questions arise what new perspectives are there for disease resistance breeding and how promising are they? For discussion purposes, they have been classified into three categories:
i) Transfer between plant species.
ii) Transfer from pathogens.
iii) Production of new genetic constructs.
i) Transfer between plant species
For decades now, breeders have been transferring major resistance genes across the species barrier by crossing them with related species. Bread wheat (Triticum aestivum L.) is an excellent example. Several Triticum species and species of at least six other genera from the tribe Triticeae have been used successfully as donors of resistance genes (Jones et al., 1995). Most of these transferred resistance genes were of the hypersensitive, non-durable type. However, the transfer of alien genes is often far from easy, always time consuming, some other alien DNA is always transferred in the process as well. When fully developed, transformation will not only be less time consuming, it will also be much more efficient than the conventional procedure, as in principal, no additional donor material will be transferred with the transformed gene. The molecular transformation techniques not only allow the transfer of genes from related species, they can transfer genes from any organism to the crop species to be improved. Transformation of resistance genes of the non-durable type from one species to another does not seem very useful, as the non-durable character will not change. The transformation of potentially durable resistance genes, however, is highly interesting. For example, the transfer of genes controlling the production of a phytoalexin in a given species to an unrelated crop species could make this latter species resistant to several of its pathogens, which are not adapted to this foreign phytoalexin. The genes for resveratrol, a phytoalexin of grapevine (Vitis viniferaL.) have been transformed to tobacco (Nicotiana tabacum L.), which became more resistant Botrytis cinerea Pers. ex Fr. (Hain et al, 1993).
ii) Transfer from pathogens
Transformation of plants with a gene of a viral genome often gives rise to plants resistant to the virus from which the gene was derived; this is known as pathogen-derived resistance (Lomonossoff, 1995). This pathogen-derived resistance (PDR) varies greatly from protecting against very high levels to very low levels of inoculum. The specificity also varies from highly specific, resistant only to the strain from which the gene was derived, to moderately specific, providing resistance to the virus from which the gene originated and to related viruses.
Initially, all attempts to use PDR were directed at transferring the viral genes coding for the coat (nucleocapsid) protein to the host plants. Such coat protein mediated resistance has been demonstrated against many viruses of widely different groups (Singh et al., 1995). The resistance level obtained varies greatly from only slight to fairly strong.
Later other viral genes or parts of viral genes were transformed to host plants. From the diversity of results published so far, the conclusion seems justified that, through transformation to the host, any part of the viral genome can potentially give rise to PDR (Lomonossoff, 1995). All resulting resistance had to be classified as quantitative.
It is likely that PDR can be obtained from other pathogens. However, pathogens such as fungi have a much larger genome than viruses, where many genes are not directly involved in the pathogenicity process. It is quite possible that only the transformation of pathogenicity genes may result in PDR, which means that as a first step pathogenicity genes have to be identified, localized, marked and cloned, a considerably more complex affair than with the simple genomes of viruses.
iii) Production of new genetic constructs
With the possibility of isolating specific genes it becomes possible to make genetic constructs by combining genes from different origins or even by changing the isolated genes. The possibilities are almost infinite and each construct has to be tested.
One could bring together two or three non-durable resistance genes, which are individually still effective, together in a construct to be introduced into a cultivar. Experience has taught us that a barrier of two or three such genes increases the durability greatly, provided these genes are not used individually by breeders (Pederson & Leath 1988; Parlevliet, 1997).
Constructs can also change the resistance expression from being induced upon attack to that of a constitutive expression. The invading pathogen is thus exposed to the resistance at an earlier stage. The first success was reported in 1991. An endo-chitinase gene of bean, its production normally induced after pathogen attack, was transformed with a constitutive promoter to tobacco. This increased the resistance to Rhizoctonia solani. It was ineffective to Pythium aphanidermatum, a fungus lacking a chitin-containing cell wall (Broglie et al., 1991).
In summary one can conclude that molecular techniques of transferring genetic material with the aim of obtaining resistance are enormously diverse and very promising. The resistance obtained so far is largely of a quantitative nature.
BREEDING FOR RESISTANCE
In order to reduce costs and to increase the efficiency of identifying resistant plants or lines in segregating populations, breeders developed screening methods in which plants as young as possible were exposed to high concentrations of, preferably, a specified inoculum. This efficiently identifies complete resistance based on major genes but is inadequate for recognizing small differences in resistance. These screening approaches, together with the belief that polygenic resistance is difficult to select for and might not give a good level of resistance, led to the present situation where major gene resistance has been exploited very well, while QR has been used only sparingly. This is unfortunate as there is so much QR available. Quantitative Resistance occurs to most of our important pathogens at various levels in nearly all our crops as discussed in the chapter "quantitative resistance". Since this QR does occur in the cultivars grown, it is genetic material that is related to what the breeders' desire. For this type of resistance breeders do not need to look for primitive genotypes from centres of diversity nor to related wild species. The resistance is near at hand in adapted cultivars, a fortunate situation as it makes breeding easier. McIntosh (1997) concluded that the ideal sources of resistance are those present in closely related, commercial genotypes, and any effort to transfer resistance from related species and genera should be considered long term.
To select for QR means accumulating QR in much the same way as selecting for higher yields. The breeder selects the plants or lines with the lower levels of disease severity and by doing that continuously over the seasons, the level of QR will increase fairly rapidly as Parlevliet and Van Ommeren (1988) showed. There is, however, one complication. If there is also non-durable major gene resistance around, it has to be taken into account. The QR is not visible when such an effective major gene is present. By using, preferably, local material, the frequency of such non-durable still effective major genes is often low, as the local pathogen population has adapted to these genes. Introducing plant material from elsewhere, especially from the centres of diversity, increases the frequency of such non-durable effective major genes considerably, as the local pathogen population has not yet adapted to the newly introduced resistance. Therefore, to select QR stick as much as possible to local material as they will almost certainly carry QR. One can also avoid ending up with non-durable major resistance in the selected material by selecting against susceptibility, i.e. removing the most susceptible plants and lines all the time (Parlevliet and Van Ommeren, 1988). Plants or lines with complete resistance should also be removed in case of resistance breeding against specialized fungal pathogens, as such resistances can be assumed to be non-durable. In case of non-specialized pathogen and viruses one may use any resistance.
CONCLUDING REMARKS
In the coming 50 to 60 years the world population will about double and hopefully also become more prosperous. This demands large yield increases in our food crops, which have to be grown in more sustainable agricultural systems. The need for durable disease resistance, therefore, can be expected to grow enormously. This need can be met technically by exploiting two sources that are largely untapped at present. These sources are the QR already present in our crops and the possibilities of transforming genes or gene constructs encoded for resistance into our crops.
Quantitative Resistance at present is poorly exploited. If the same large effort that went into breeding for the hypersensitive, major gene type had gone into QR, most cultivars of our major crops would now carry high levels of it. With respect to sustainable agriculture and integrated forms of crop protection quantitative, durable resistance is a more desirable form of resistance than the non-durable type.
Much of the resistance obtained after transformation is of a quantitative nature. This view should be consequential in modern genetic engineering activities. A considerable part of successful molecular manipulation leads to the type of resistance in which there is no shortage in most crops to most pathogens, and which is poorly used by the breeders
GLOSSARY
Acquired resistance: See "induced resistance".
Adult plant resistance: Resistance expressed in the adult plant stage only. This resistance is often governed by race-specific major genes and is therefore not of a more durable nature than seedling/overall resistance.
Aggressiveness: Counterpart of race-non-specific resistance; the ability of the isolate to grow vigorously on or in its host. The more aggressive an isolate of a pathogen is the more of the host tissue it can invade in a given time.
Avirulence: The (near) absence of pathogenicity of a pathogen genotype when it comes into contact with a host genotype that carries a race-specific resistance gene corresponding to an avirulence gene of the pathogen genotype.
Biotroph: A pathogen that obtains its nutrient supply only from living host tissue.
Complete resistance: Resistance, that does not allow growth of the pathogen. There are no signs of disease or of the presence of the pathogen.
Constitutive resistance: Resistance which is present when exposed to the pathogen. Many resistances are induced (see "induced resistance") when exposed to a pathogen.
Durable resistance: Resistance that remains effective for long periods when widely exposed to the pathogen under the prevailing growing conditions.
Field resistance: Resistance that is expressed best in the field; it is usually a QR (Quantitative Resistance).
General resistance: It is sometimes used as an equivalent to race-non-specific or horizontal resistance. The term should be avoided as there are forms of resistance that are truly general, being effective to a wide range of pathogens. Phytoalexins belong to this class of resistance.
Generalist: Pathogen that has a wide host range including species from various families.
Horizontal resistance: Equivalent to "race-non-specific resistance".
Immunity: Extreme form of resistance; after exposure to a pathogen it is not possible to demonstrate its presence.
Hypersensitivity: Response to infection in which the invaded cells and neighbouring cells die rapidly and the pathogen is prevented to spread further. The result is strongly localized necrosis.
Incomplete resistance: Any resistance that is not complete; there is some growth of the pathogen. Some major gene and all QR can be seen as forms of incomplete resistance.
Induced resistance: Enhancement of resistance of a susceptible plant in response to an extrinsic stimulus, the stimulus being of a biotic or abiotic nature. The enhanced resistance can be localized at the site of the inducing treatment or it can be systemic. The latter commonly indicated with acquired resistance.
Isolate: A population of a micro-organism obtained by isolating it from a host or substrate and establishing it in pure culture.
Major gene resistance: Resistance governed by one or more genes with large effects; large enough to produce a discontinuous character in segregating populations, see qualitative resistance.
Mature tissue resistance: In some plant species only the the young tissue is suscetible to the pathogen, the mature tissue is fully resistant (Apple/Venturia inaequalis, Rice/Magnapothe grisea)
Minor gene resistance: Resistance governed by genes with small effects; too small to identify the individual genes. It gives a continuous character in segregating populations; see quantitative and polygenic resistance.
Monogenic resistance: Resistance controlled by one gene, usually a major gene; as a minor gene is very hard to discern.
Partial resistance: Equivalent to QR. In crops against biotrophic pathogens (rusts, powdery mildews) it means QR associated with susceptible infection types.
Pathogenicity: The ability of the pathogen to grow and to develop on or in its host and at the costs of the host.
Pathotype: Equivalent to "Race".
Polygenic resistance: Equivalent to minor genes. The effect of the minor genes together can be large as the small effects are additive.
Qualitative resistance: Host genotypes show a discontinuous range of variation in resistance. Susceptible and resistant genotypes can be easily discerned; see major gene resistance.
Quantitative resistance (QR): Host genotypes show a continuous range of variation in resistance from extremely susceptible to fairly resistant; see minor gene and polygenic resistance.
Race: All genotypes of a pathogen that carry the same set of avirulence genes.
Race-non-specific or pathotype-non-specific resistance: Resistance effective to all genotypes of the pathogen. There are no cultivar x race interactions.
Race-specific or pathotype-specific resistance: Resistance effective to certain races or pathotypes of the pathogen, but not to others. There are cultivar x race interactions.
Resistance: Mechanisms, which interfere with and so reduce the growth and/or development of the parasite.
Seedling or overall resistance: Resistance expressed in all stages of the plant. Selection for it is often done in the seedling stage. It is often controlled by race-specific major genes and considered to be non-durable.
Specialist: Pathogen that has a narrow host range, one species only or species of one genus or of a few related genera.
Stable resistance: Sometimes wrongly used as an equivalent for durable resistance. The right meaning is: Resistance that is expressed under a wide range of growing conditions. Various resistance genes in cereals to rusts are temperature sensitive, they come to expression at certain temperatures but not at others. This can be seen as unstable resistance.
Strain: It is vaguely used to indicate a group of similar genotypes within a pathogen species.
Vertical resistance: Equivalent to "race-specific resistance".
Virulence: Counterpart of race-specific resistance; the ability of a race to be pathogenic on certain host genotypes only. It lacks the functional avirulence genes corresponding with the resistance genes in these host genotypes.
- ALEMAYEHU, F. & PARLEVLIET, J.E. Variation for resistance to Puccinia Hordei in Ethiopian barley landraces. Euphytica 90:365-370. 1996.
- ANONYMOUS. Beschrijvende rassenlijst voor landbouwgewassen (Descriptive lists of varieties for arable crops) nrs 29-73. Leiter-Nypels, Maastricht. 1954-1998.
- BAINBRIDGE, A.& JENKYN, J.F. Mildew reinfection in adjacent and separated plots of sprayed barley. Annals. Applied Biology 82:477-487. 1973.
- BLACK, W. The nature and inheritance of field resistance to late blight (Phytopththora infestans) in potatoes. American Potato Journal 47:279-288. 1970.
- BROERS, L.H.M. & JACOBS, TH. The inheritance of host plant effect on latency period of wheat leaf rust in spring wheat. II Number of segregating factors and evidence for transgressive segregation in F3 and F5 generations. Euphytica 44:207-214. 1989.
- BROGLIE, K., CHET, I., HOLLIDAY, M., CRESSMAN, R., BIDDLE, P., KNOWLTON, S., MAUVAIS, C.J. & BROGLIE, R. Transgenic plants with enhanced resistance to the fungal pathogen Rhizoctonia solani Science 254:1194-1197. 1991.
- BROWDER, L.E. 1985. Parasite: host: environment specificity in the cereal rusts. Annual Review Phytopathology 23:201- 222.
- BRUEHL, G.W. Nonspecific genetic resistance to soilborne fungi. Phytopathology 73:948-951. 1983
- BURNETTE, D.C. & WHITE, D.G. Inheritance of resistance to Bipolaris maydis race 0 in crosses derived from nine inbred lines of maize. Phytopathology 75:1195-1200. 1985.
- COCKERHAM, G. Strains of potato virus X. In: Proceeding, 2nd, Conference of Potato virus diseases, June 1954. pp 89-90. 1955.
- COLHOUN, J. Effects of environmental factors on plant disease. Annual Review Phytopathology 11:343-364. 1973
- EETERS, J.P., ALBRECT, J.C., GAWEY, N.W., GILES, R.J., JESTIN, L. & VAN SOEST, L.J.M. Variation over time and environments in resistance to Erysiphe graminis hordei in samples from barley germplam collection. Euphytica 46:43-50. 1990.
- FALKHOF, A.G., DEHNE, H.W. & SCHONBECK, F. Dependence of the effectiveness of induced resistance on environmental conditions. Journal Phytopathology 123:311-321. 1988
- FLOR, H.H. Current status of the gene-for-gene concept, Annual Review Phytopathology 9:275-296. 971.
- FRASER, R.S.S. Mechanisms of resistance to plant diseases. Kluwer Academic Publishers, Dordrecht, Boston and London. 1985
- FRASER, R.S.S. The genetics of plant-virus interactions: implications for plant breeding. Euphytica 63: 175-185. 1992.
- GEIGER, H.H. & HEUN, M. Genetics of quantitative resistance to fungal diseases. Annual Review Phytopathology 27: 317-341. 1989.
- HABGOOD, R.M. Differential aggressiveness to Rhynchosporium secalis isolates towards specified barley genotypes. Trans British. Mycological Society 66:201-204. 1976
- HABGOOD, R.M. The inheritance to Rhynchosporium secalis in some European spring barley, Annals Applied Biology 77:191-200. 1974.
- HAIN, R., REIF, H.J., KRAUSE, E., LANGEBARTELS, R.,. KINDL, V.B, WEISE, W., SCHMELZER, E., SCHREIER, P.H. & STOCKER, S.K. Disease resistance results from foreign phytoalexin expression in a novel plant. Nature 361:153-156. 1993.
- HAMMERSCHMIDT, R. & J. KUC. Induced Resistance to disease in Plants. Kluwer Academic Publishers, Dordrecht, Boston and London. 1995.
- HUNTER, R.B., ATLIN, G.N. & MULDOON, J.F. Genotype X Environmental interactions for ear mold resistance and its sub-components in maize hybrids Canadian Plant Science 66:291-297. 1986
- ISLAM, M.R. & MAYO, G.M.E. A compendium on host genes in flax conferring resistance to flax rust. Plant Breeding 104:89-100. 1990.
- ISLAM, M.R. & SHEPHERD, K.W. Present status of genetics of resistance in flax. Euphytica 55:255-267. 1991.
- JEGER, M.J., GRIFFITHS, E. & GARETH JONES, D. Seasonal variation in the components of partial resistance of seedlings of winter wheat cultivars. Plant Pathology 32:187-195. 1983.
- JENKINS, M.T. & ROBERT, A.L. Further genetic studies of resistance to Helminthosporium turcicum Pass. in maize by means of chromosomal translocations. Crop Science 1:450-455. 1961.
- JENKYN, J.F., BAINBRIDGE, A., DYKE, G.V. & TOLD, A.D. An investigation into inter-plot interactions, in experiments with mildew on barley, using balanced deigns. Annals Applied Biology 92:11-28. 1983
- JOHNSON, R & TAYLOR, A.J. Spore yield of pathogens in investigations of the race-specificity of host resistance. Annual Review Phytopathology 14: 97-119. 1976.
- JOHNSON, R. Durable resistance: Definition of genetic control, and attainment in plant breeding. Phytopathology 71:567-568. 1981.
- JONES, S.S., MURRAY, T.D. & ALLEN, R.E. Use of alien genes for the development of disease resistance in wheat. Annual Review Phytopathology 33:429-443. 1995.
- JØRGENSEN, J.H. Disease and pest resistance genes. Coordinator's report. Barley Genetics Newsletter 20:89. 1990.
- KHOMOTO, K. & OTANI, H. Host recognition by toxigenic plant pathogen. Experientia 47:755-764. 1991.
- KIM, S.K. & BREWBAKER, J.L. Inheritance of general resistance in maize to Puccinia sorghi Schw. Crop Science 17:456-461. 1977
- KOCH, M.F. & PARLEVLIET, J.E. Genetic analysis of, and selection for, factors affecting quantitative resistance to Xanthomonas campestris pv oryzae in rice. Euphytica 53:235-245. 1991.
- KODAMA, M., AKAMATSU, H., ITOH, Y., NARUSAKA, Y., SANEKATA, T., OTANI, H. & KOHMOTO, K. Host- specific toxin deficient mutants of the tomato pathotype of Alternaria alternata obtained by restriction enzyme - mediated integration. In: Kohmoto, K. & Yoder, O.C. (Eds.): Molecular Genetics of host - specific toxins in plant disease. Kluwer Academic Publishers. 1998. pp. 35-40.
- KUC, J. & STROBEL, N. Induced resistance using pathogens and nonpathogens. In: Jajamos, E.G. & Papavizas, R.C. (Eds.). Biological control of plant diseases. NATO ASI Series, Plenum, NY. 1992. pp. 295-303.
- KUC, J. Induced systemic resistance an Overview. In: Hammerschmidt, R, & Kuc, J. (Eds.): Induced Resistance to disease in Plants. Kluwer Academic Publishers, Dordrecht, Boston and London. 1995. pp. 169-175.
- KULKARNI, R.N. & CHOPRA, V.L. Environment as the cause of differential interaction between host cultivars and pathogenic races. Phytopathology 72:1384-1386. 1982
- LEONARD, K.J. Durable resistance in the pathosystems: maize-Northern and Southern leaf blights. In: Jacobs,Th. & Parlevliet, J.E. (Eds.). Durability of Disease Resistance. Kluwer Academic Publishers, Dordrecht, The Netherlands. 1993. pp. 99-114.
- LOMONOSSOFF, G.P. Pathogen-derived resistance to plant viruses. Annual Review Phytopathology 33: 323-343. 1995.
- MCINTOSH, R.A. Breeding wheat for resistance to biotic stress. In: Braun, H.J., Altay, F., Kronstad, W.E., Beniwal, S.P.S. & McNab, A. (Eds.). Wheat: prospects for global improvement. Proc. 5th Int. Wheat Conf. 1996, Ankara, Turkey. Kluwer Acad. Publ. Dordrecht. The Netherlands. 1997. pp 71-86.
- MEHTA, Y.R. & ZADOKS, J.C. Uredopspore production and sporulation period of Puccinia recondita f. sp. triticina on primary leaves of wheat. Netherlands Journal Plant Pathology 76:267-176. 1970.
- MEINERS, J.P. Genetics of disease resistance in edible legumes. Annual Review Phytopathology 19:189-209. 1981.
- NAKASHIMA, T., UENO, T., FUKAMI, H., TAGA, T., MASUDA, H., OSAKI, K., OTANI, H., KOHMOTO, K. & NISHIMURA, S. Isolation and structures of AK - toxin I and II, host specific phytotoxic metabolites produced by Alternaria alternata Japonese pear pathotype, Agriculture Biological Chemistry 49:807-815. 1985.
- NIKS, R.E., ELLIS, P.R. & PARLEVLIET, J.E. Resistance to parasites. In: Hayward, M.D., Bosemark, N.O. & Romagosa, I. (Eds.). Plant Breeding, Principles and Prospects. Chapman & Hall, London. 1993. pp. 422-447.
- NORGAARD KNUDSEN, J.C., DALSGAARD, H.H. & JORGENSEN, J.J. Field assessment of partial resistance to powdery mildew in spring barley. Euphytica 35. 233-243. 1986.
- OTANI, H. KOHMOTO, K. & KODAMA, M. Alternaria toxins and their effects on host plants. Canadian Journal Botany 73:453-458. 1995.
- PARLEVLIET, J.E. & DANIAL, D.L. How does interplot interference affect the field assessment for resistance in cereals to rusts and powdery mildew? Vortr. Pflanzenzüchtg. 24:289-291. 1992.
- PARLEVLIET, J.E. & VAN OMMEREN, A. Accumulation of partial resistance in barley to barley leaf rust and powdery mildew through recurrent selection against susceptibility. Euphytica 37:261-274. 1988.
- PARLEVLIET, J.E. & VAN OMMEREN, A. Interplot interference an the assessmentof barley cultivars for partial resistance to leaf rust, Puccinia hordei Euphytica 33:685-6971984
- PARLEVLIET, J.E. & VAN OMMEREN, A. Partial resistance of barley leaf rust, Puccinia hordei II Relationship between field trials, micro - plot tests and latent period. Euphytica 24:293-303. 1975.
- PARLEVLIET, J.E. & ZADOKS, J.C. The integrated concept of disease resistance; a new view including horizontal and vertical resistance in plants. Euphytica 26:5-21. 1977.
- PARLEVLIET, J.E. Durable resistance. . In: H. Hartleb, H., Heitefuss, R. & Hoppe, H.H. (Eds.). Resistance of crop plants against fungi. Gustav Fisher, Jena, Germany. 1997. pp. 238-253.
- PARLEVLIET, J.E. Further evidence of polygenic inheritance of partial resistance in barley to leaf rust, Puccinia hordei Euphytica 27:369-379. 1978.
- PARLEVLIET, J.E. Genetic and breeding aspects of durable resistance of crops to pathogens. African Crop Science Journal 3:1-13. 1995.
- PARLEVLIET, J.E. Identification and evaluation of quantitative resistance. In: Leonard, K.J. & Fry, W.E. (Eds.). Plant disease epidemiology - Genetics, Resistance and Management. Vol. 2 McGraw - Hill Publishing Company, New York, 1989. pp. 215-248.
- PARLEVLIET, J.E. Selecting components of partial resistance. In: Stalker, H.T. & Murphy, J.P. (Eds.). Plant Breeding in the 1990's. CAB, Intern. Wallingford, UK. 1992. pp. 281-302.
- PARLEVLIET, J.E. What is durable resistance, a general outline. In: Jacobs, Th. & Parlevliet, J.E. (Eds.) Durability of disease resistance. Kluwe Academic Publisher. 1993. pp. 23-39.
- PAYSOUR, R.E. & FRY, W.E. Interplot inference: A model for planning field experiments with aerially disseminated pathogens. Phytopathology 73:1014-1020. 1983.
- PEDERSON, W., & LEATH, S. Pyramiding major genes for resistance to maintain residual effects. Annual Review Phytopathology 26:369-378. 1988.
- ROBERTS, P.A. Conceptual and practical aspects of variability in root-knot nematodes related to host plant resistance. Annual Review Phytopathology 33:199-221. 1995.
- ROUMEN, E.C. The inheritance of host plant resistance and its effect on the relative infection efficiency of Magnaporthe grisea in rice. Theoretical Applied Genetics 89:489-503. 1994.
- SAXENA, K.M.S. & HOOKER, A.L. On the structure of a gene for disease resistance in maize. Proceedings Nature Academic Science. USA 61:1300-1303. 1968.
- SCHEFFER, R.P. The nature of Disease in Plants. Cambridge University Press. 1997.
- SCHONBECK, F. & STEINER, U. Induced resistance. Pages In: Hartleb, H., Heitefuss, R. & Hoppe, H.H. (Eds.) Resistance of Crop plant against fungi. Gustav Fischer, Jena, Stuttgart. 1997. pp. 272-297.
- SEQUEIRA, L. Mechanisms of induced resistance in plants. Annual Review Microbiology 37: 51-79. 1983.
- SINGH, R.P., SINGH, U.S. & KOHMOTO, K. Pathogenesis and host specificity in plant diseases. Vol. III. Viruses & viroids. Elsevier Science, Oxford, UK. 1995.
- STEFFENSON, B.J. Analysis of durable resistance to stem rust in barley. Euphytica 63:153-167. 1992.
- STERMER, B.A. Molecular regulation of systemic induced resistance. In: Hammerschemidt, R. & Kuc, J. (Eds.). Induced Resistance to disease in Plants. Kluwer Academic Publishers, Dordrecht, Boston and London. 1995. pp. 111-140.
- TENG, P.S. & CLOSE, R.C. Effect of temperature and uredinium density on urediniospore production, latent period and infectious period of Puccinia hordei Otth. N.Z.J. Agriculture Research 21:287-296. 1978.
- TORYAMA, K. Recent progress of studies on horizontal resistance in rice breeding for blast resistance in Japan. In: Proceedings of Seminar on Horizontal Resistance to Blast Disease of Rice. CIAT Series CE9, Colombia. 1975. pp. 65-100.
- TURKESTEEN, L.J. Durable resistance of potatoes against Phytophthora infestans. In: Jacobs Th. & Parlevliet, J.E. (Eds.). Durability of disease resistance. Kluwe Academic Publisher, Dordrecht, The Netherlands. 1993. pp. 115-124.
- VAN DER PLANK, J.E. Disease resistance in plants. Academic Press, New York, London. 1968.
- VAN DER PLANK, J.E. Plant disease: epidemic and control. Academic Press, New York, London. 1963.
- WELZ, H.G. & KRANZ, J. How resistance affects disease epidemics. In: Hartleb, H., Heitefuss, R. & Hoppe, H.H. (Eds.). Resistance of crop plants against fungi. Gustav Fischer, Jena, Stuttgart 1997. pp. 327-348.
- YODER, O.C. A mechanistic view of the fungal / plant interaction based on host - specific toxin studies. In: Kohmoto, K. & Yoder, O.C. (Eds.). Molecular Genetics of host - specific toxins in plant disease. Kluwer Academic Publishers. 1998. pp. 3-15.
- YOUNG, N.D. QTL mapping and quantitative disease resistance in plants. Annual Review Phytopathology 34:479-501. 1996.
Publication Dates
-
Publication in this collection
27 Nov 2001 -
Date of issue
Sept 2001
History
-
Accepted
13 Aug 2001