Abstract
The study was carried out to screen and analyze the genetic characteristics of antimicrobial resistance in Campylobacter spp. from poultry sources. A total of 141 strains of Campylobacter isolated from samples of broilers of slaughterhouses in southern Brazil was identified by phenotypic and genotypic methods. Campylobacter isolates were evaluated for its antimicrobial susceptibility and the presence of resistance genes. The strains were investigated for antimicrobial susceptibility against two agents (ampicillin and tetracycline) by disk diffusion method. PCR assay was used to confirm the specie and the presence of ampicillin (blaOXA-61), tetracycline tet(O), and the energy-dependent multi-drug efflux pump (cmeB) genes. Campylobacter jejuni was the most ubiquitous; its presence was determined in 140 samples out of 141 (99.3%), whereas Campylobacter coli was found only in one of the contaminated samples (0.70%). The results obtained showed 65% and 35.5% of Campylobacter isolates resistant to β-lactams and tetracyclines, respectively. The cmeB gene responsible for multidrug resistance was detected in 26 isolates out 141 strains (18.5%). Moreover, 36 out of 141 Campylobacter strains (25.6%) were found to be resistant to at least two different antimicrobia resistance markers (β-lactams and tetracyclines).
Index Terms
Campylobacter; tetracycline; β-lactam; efflux pump; resistance genes; PCR
Resumo
O presente estudo foi realizado para examinar e analisar as características genéticas de resistência antimicrobiana de Campylobacter spp. a partir de fontes avícolas. Um total de 141 amostras de Campylobacter isolados em matadouros-frigoríficos de aves do estado do Rio Grande do Sul, Brasil, foi identificado por métodos fenotípicos e genotípicos. Foi analisada a susceptibilidade antimicrobiana e a presença de genes de resistência. As cepas foram testadas para detectar sensibilidade frente a dois antimicrobianos (ampicilina e tetraciclina) pelo método de difusão em disco. A seguir, usando a reação em cadeia da polimerase foi confirmada a espécie e a presença dos genes de resistência à ampicilina (blaOXA-61) e tetraciclina tet(O), assim como a detecção da bomba de efluxo (cmeB). Campylobacter jejuni foi a espécie mais isolada, sua presença foi determinada em 140 amostras (99,3%), e Campylobacter coli foi encontrada em uma única amostra (0,70%). Os resultados obtidos mostraram 65% e 35,5% de Campylobacter isolados resistentes a β-lactâmicos e tetraciclinas, respectivamente. O gene cmeB responsável pela resistência a múltiplos antimicrobianos foi detectado em 26 amostras (18,5%). Neste contexto, 36 das 141 amostras (25,6%) foram consideradas resistentes a dois grupos diferentes de antimicrobianos (β-lactâmicos e tetraciclinas).
Termos de Indexação
Campylobacter; tetraciclinas; β-lactâmicos; bomba de efluxo; genes de resistência; PCR
Introduction
Campylobacter is recognized as the leading causes of bacterial foodborne diarrheal disease throughout the worldwide (Park 2002Park S.F. 2002. The physiology of Campylobacter species and its relevance to their role as foodborne pathogens. Int. J. Food Microbiol.74:177-188.,Silva et al. 2011Silva J., Leite D., Fernandes M., Mena C., Gibbs P.A. & Teixeira P. 2011. Campylobacter spp. as a foodborne pathogen: a review. Frontiers Microbiol. 2:200.). Campylobacteriosis is estimated to cause about 1.3 million infections, 13,000 hospitalizations and 120 deaths each year in the United States (CDC 2013CDC 2013. Incidence and trends of infection with pathogens transmitted commonly through food. Foodborne Diseases Active Surveillance Network, 10 U.S. Sites, 1996-2012. Morbidity and Mortality Weekly Report 19:283-287.). It is also the most commonly reported antecedent infection in the development of Guillain Barré syndrome (GBS) and Miller Fisher syndrome (MFS) (Godschalk et al. 2004Godschalk P.C., Heikema A.P., Gilbert M., Komagamine T., Ang C.W., Glerum J., Brochu D., Li J., Yuki N., Jacobs B.C., van Belkum A. & Endtz H.P. 2004. The crucial role of Campylobacter jejuni genes in anti-ganglioside antibody induction in Guillain-Barré syndrome. J. Clin. Invest. 114:1659-1665.,Hardy et al. 2011Hardy C.G., Lackey L.G., Cannon J., Price L.B. & Silbergeld E.K. 2011. Prevalence of potentially neuropathic Campylobacter jejuni strains on commercial broiler chicken products. Int. J. Food Microbiol.145:395-399.,Van den Berg et al. 2014Van den Berg B., Walgaard C., Drenthen J., Fokke C., Jacobs B.C. & van Doorn P.A. 2014. Guillain-Barré syndrome: pathogenesis, diagnosis, treatment and prognosis. Nat. Rev. Neurol. 10:469-482.). A risk factor for human disease is the consumption of contaminated poultry products (Conlan et al. 2007Conlan A.J., Coward C., Grant A.J., Maskell D.J. & Gog J.R. 2007. Campylobacter jejuni colonization and transmission in broiler chickens: a modelling perspective. J. R. Soc. Interface 4:819-829.,Ellström et al. 2014Ellström P., Hansson I., Söderström C., Olsson Engvall E. & Rautelin H. 2014. A prospective follow-up study on transmission of Campylobacter from poultry to abattoir workers. Foodborne Pathog. Dis. 11:684-688.). Transmission to man usually results in sporadic infection, and is often associated with improper handling or cooking of food (Moore et al. 2005Moore J.E., Corcoran D., Dooley J.S., Fanning S., Lucey B., Matsuda M., McDowell D.A., Mégraud F., Millar B.C., O'Mahony R., O'Riordan L., O'Rourke M., Rao J.R., Rooney P.J., Sails A. & Whyte P. 2005. Campylobacter. Vet. Res. 36:351-382.). Campylobacter jejuni (C. jejuni) and Campylobacter coli (C. coli) are considered to be zoonotic pathogens, antimicrobial resistance among isolates in the animal reservoir has serious implications for the treatment in humans (Moore et al. 2006Moore J.E., Barton M.D., Blair I.S., Corcoran D., Dooley J.S., Fanning S., Kempt I., Lastovica A.J., Lowery C.J., Matsuda M., McDowell D.A., McMahon A., Millar B.C., Rao R.J., Rooney P.J., Seal B.S., Snelling W.J. & Tolba O. 2006. The epidemiology of antibiotic resistance in Campylobacter . Microbes Infect. 8:1955-1966.,EFSA 2011EFSA 2011. The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2009EFSA J. 9:2090-2468.). The majority of cases of clinical Campylobacter spp. enteritis are generally mild or self-limiting disease not to require of antimicrobial chemotherapy (Moore et al. 2005Moore J.E., Corcoran D., Dooley J.S., Fanning S., Lucey B., Matsuda M., McDowell D.A., Mégraud F., Millar B.C., O'Mahony R., O'Riordan L., O'Rourke M., Rao J.R., Rooney P.J., Sails A. & Whyte P. 2005. Campylobacter. Vet. Res. 36:351-382.). However, antimicrobial therapy may be used in a subset of patients with severe and prolonged systemic complications or to control infection (Avrain et al. 2003Avrain L., Humbert F., L'Hospitalier R., Sanders P., Vernozy-Rozand C. & Kempf I. 2003. Antimicrobial resistance in Campylobacter from broilers: association with production type and antimicrobial use. Vet. Microbiol. 96:267-276.,Janssen et al. 2008Janssen R., Krogfelt K.A., Cawthraw S.A., van Pelt W., Wagenaar J.A. & Owen R.J. 2008. Host-pathogen interactions in Campylobacter infections: the host perspective. Clin. Microbiol. Rev. 21:505-518.). Currently, macrolides and fluoroquinolones are the antimicrobial agents of choice when therapeutic intervention is warranted (Engberg et al. 2001Engberg J., Aarestrup F.M., Taylor D.E., Gerner-Smidt P. & Nachamkin I. 2001. Quinolone and macrolide resistance in Campylobacter jejuni and C. coli: Resistance mechanisms and trends in human isolates. Emerg. Infect. Dis. 7:24-34.,Moore et al. 2005Moore J.E., Corcoran D., Dooley J.S., Fanning S., Lucey B., Matsuda M., McDowell D.A., Mégraud F., Millar B.C., O'Mahony R., O'Riordan L., O'Rourke M., Rao J.R., Rooney P.J., Sails A. & Whyte P. 2005. Campylobacter. Vet. Res. 36:351-382.). Bacterial populations can respond to the threat of an antimicrobial agent by evolving some type of resistance mechanism(s) (Rowe-Magnus et al. 2002Rowe-Magnus D.A., Guerout A.M. & Mazel D. 2002. Bacterial resistance evolution by recruitment of super-integron gene cassettes. Mol. Microbiol. 43:1657-1669.,Luangtongkum et al. 2009Luangtongkum T., Jeon B., Han J., Plummer P., Logue C.M. & Zhang Q. 2009. Antibiotic resistance in Campylobacter: emergence, transmission and persistence. Future Microbiol. 4:189-200.). These resistant bacteria may be transferred to humans either through the food supply or by direct contact with animals (Khachatourians 1998Khachatourians G.G. 1998. Agricultural use of antibiotics and the evolution and transfer of antibiotic-resistant bacteria. Can. Med. Assoc. J. 159:1129-1136.,Angulo et al. 2004Angulo F.J., Nunnery J.A. & Bair H.D. 2004. Antimicrobial resistance in zoonotic enteric pathogens. Rev. Sci Tech. OIE 23:485-496.). Ampicillin and tetracycline have activity against Campylobacter , but in general, are not recommended for the treatment of Campylobacter infections (Blaser 1995Blaser M.J. 1995. Campylobacter and related species, p.1948-1956. In: Mandell G.L., Bennett J.E. & Dolin R. (Eds), Principles and Practice of Infectious Diseases. 4th ed. Churchill Livingstone, New York, NY.,Dasti et al. 2007Dasti J.I., Gross U., Pohl S., Lugert R., Weig M. & Schmidt-Ott R. 2007. Role of the plasmid-encoded tet (O) gene in tetracycline-resistant clinical isolates of Campylobacter jejuni and Campylobacter coli . J. Med. Microbiol. 56:833-837.). However, the increment of resistant strains to commonly used antimicrobials in clinical practices makes it necessary to consider alternatives therapies as well as the search of easy and reliable methods to study antimicrobial susceptibility (Luangtongkum et al. 2009Luangtongkum T., Jeon B., Han J., Plummer P., Logue C.M. & Zhang Q. 2009. Antibiotic resistance in Campylobacter: emergence, transmission and persistence. Future Microbiol. 4:189-200.). The aim of the present study was to determine the occurrence of Campylobacter spp. strains carrying resistance genes (tetracycline and β-lactam) and the energy-dependent multi-drug efflux pump through phenotypic and molecular analyses in poultry sources from slaughterhouses in Rio Grande do Sul state, Brazil.
Materials and Methods
Sample collection. Since January 2012 to December 2013, a total of 141 isolates from: carcasses through slaughter line (n=115); water collected from the chiller tank (n=18); and from swabs (cloacal and boxes of transport) (n=8) were obtained from broiler slaughterhouses of Rio Grande do Sul state, Brazil.
Antimicrobial susceptibility screening. Isolation was performed in accordance with the International Standards Organization guidelines (ISO 10272-1:2006ISO 2006. International Standard Organization 10272-1:2006 describes a horizontal method for the detection of Campylobacter spp.). Campylobacter isolates were analyzed for antimicrobial resistance using the agar disk diffusion method. The suspension was adjusted to match the 0.5 McFarland turbidity standards as recommended by the Clinical and Laboratory Standards Institute (CLSI 2010CLSI 2010. Methods for Antimicrobial Dilution and Disk Susceptibility Testing of Infrequently Isolated or Fastidious Bacteria: approved guideline. 2nd ed. Clinical and Laboratory Standards Institute (M45eA2), Wayne, PA.). Isolated cultures were analyzed for antimicrobial resistance using the disk diffusion assay on Mueller-Hinton agar plates (CM0337 Oxoid®, containing 5% sheep blood) incubated under microaerophilic conditions using a gas tank with a mixture (10% CO2, 2% H2, 5% O2, and 85% N2) for 48 hr at 41.5°C. Sheep blood agar plates were inoculated, and disks (Oxoid®) including tetracycline (30 μg), and ampicillin (10 μg) were added. Plates were incubated as described above. In view of lack of interpretative CLSI criteria for Campylobacter strains, the criteria used for the Enterobacteriaceae family were employed as breakpoints for Campylobacter resistance (CLSI 2012CLSI2012. Performance Standards for Antimicrobial Susceptibility Testing. 22nd Informational Supplement. Clinical and Laboratory Standards Institute (M100eS22), Wayne, PA.). C. jejuni ATCC 33560 strain was used as control throughout the testing period.
DNA extraction. Genomic DNA was extracted using an adapted protocol described byBorsoi et al. (2009)Borsoi A., Santin E., Santos L.R., Salle C.T.P., Moraes H.L.S. & Nascimento V.P. 2009. Inoculation of newly hatched broiler chicks with two Brazilian isolates of Salmonella Heidelberg strains with different virulence gene profiles, antimicrobial resistance, and pulsed field gel electrophoresis patterns to intestinal changes evaluation. Poult. Sci. 88:750-758.. Stored Campylobacter isolates were cultured on 5% sheep blood agar plates and incubated at 41.5oC for 48 hrs in microaerophilic conditions (10% CO2, 5% O2, and 85% N2). One milliliter of bacterial culture was centrifuged at 12.000 r.p.m. for 2 min (5415C Microcentrifuge, Eppendorf, Hamburg, Germany) and the supernatant was discarded. The pellet was suspended in 800 μL of sterile distilled water and the resulting mixture was centrifuged at 12.000 r.p.m. for 2 min. The pellet was again suspended in 200 μL of sterile distilled water. The sample was placed on a thermal block (Multi-Block Heater, Baxter, USA) at 95oC for 10 min. The mixture was centrifuged as describe above and the supernatants were transferred into fresh Eppendorf tubes to serve as a DNA template for subsequent processing.
Multiplex-PCR assay. The isolates were confirmed by Multiplex-PCR based detection of 16S rRNA, ceuE and mapA genes (Denis et al. 1999Denis M., Soumet C., Rivoal K., Ermel G., Blivet D., Salvat G. & Colin P. 1999. Development of a m-PCR assay for simultaneous identification of Campylobacter jejuni and C. coli. Lett. Appl. Microbiol. 29:406-410.).
Genotypic antimicrobial resistance. The confirmed C. jejuni isolates were screened for the presence of three genes: tetracycline (tetO), β-lactam (blaOXA-61), and the energy-dependent multi-drug efflux pump (cmeB). Primers, PCR conditions and lengths of products generated in this study are listed (Table 1). The PCR conditions were adapted (Pratt & Korolik 2005Pratt A. & Korolik V. 2005. Tetracycline resistance of Australian Campylobacter jejuni and Campylobacter coli isolates. J. Antimicrob. Chemother.55:452-460.,Obeng et al. 2012Obeng A.S., Rickard H., Sexton M., Pang Y., Peng H. & Barton M. 2012. Antimicrobial susceptibilities and resistance genes in Campylobacter strains isolated from poultry and pigs in Australia. J. Appl. Microbiol. 113:294-307.). All PCR amplifications were performed in a mixture (25 μL) consisting of 5 μL of 10X PCR Buffer [200 mM Tris-HCl (pH 8.4), 500 mM KCl], 0.25 μL (5U/μL) of Taq thermostable DNA polymerase (Invitrogen®), 2 μmol 1-l of MgCl2 (25 mM), 2 μL dNTPs (dATP, dCTP, dGTP and dTTP, each at 2.5 mM), 2 μL extracted template DNA and 0.5 μL (10 pmole 1-l) of each primer. Sterile Milli-Q water was added q.s.p 25 μL. All amplification reactions are performed in thermal cycler (Peltier Thermal Cycler Biocycler-MJ96+/MJ96G). The cycles were performed as described inTable 1. For visualization of PCR products, 10 μL aliquots were subjected to electrophoresis in a 2% agarose gel (Invitrogen®) in Tris-Acetated EDTA (TAE) buffer. DNA bands were stained with ethidium bromide for 2h at 100V, viewed under Ultraviolet (UV) transilluminator (ATTO®) and photographed (Fig. 1). The size of the PCR amplicons was compared to the 100 bp DNA ladder (Invitrogen®).
Agarose gel electrophoresis of PCR products from C. jejuni. Lanes: M, 100 bp marker; 1-2, blaOXA-61 (372 bp); 3-4, cmeB (241bp); 5, tetO (559 bp); 7, mapA (589 bp); 8, ceuE (462 bp); 9-10, Reaction control.
Statistical analysis. Statistical analyses were performed using Statistical Package for the Social Sciences (SPSS) v18 (IBM). Discrete variables were expressed as percentages, and proportions were compared using the Chi-square test with the significance level defined at P value <0.05.
Results and Discussion
All the isolates were confirmed by Multiplex-PCR based detection of 16SrRNA, ceuE and mapA genes. The most ubiquitous of the thermotolerant Campylobacter spp. was C. jejuni. It was found in 140 of the contaminated samples (99.3%), whereas C. coli was identified in the remaining sample (0.7%). The PCR-amplified products of Campylobacter species and three resistance associated genes in agarose gel are summarized inFigure 1. The most prevalent species in this study was C. jejuni detected in 99.3% of positive findings, similar to what has been observed by other researchers (Garin et al. 2012Garin B., Gouali M., Wouafo M., Perchec A.-M., Thu P.M., Ravaonindrina N., Urbès F., Gay M., Diawara A., Leclercq A., Rocourt J. & Pouillot R. 2012. Prevalence, quantification and antimicrobial resistance of Campylobacter spp. on chicken neck-skins at points of slaughter in 5 major cities located on 4 continents. Int. J. Food Microbiol. 157:102-107.,Hungaro et al. 2015Hungaro H.M., Mendonça R.C.S., Rosa V.O., Badaró A.C.L., Moreira M.A.S. & Chaves J.B.P. 2015. Low contamination of Campylobacter spp. on chicken carcasses in Minas Gerais state, Brazil: Molecular characterization and antimicrobial resistance. Food Control 51:15-22.).
An initial screening was performed to identify the antimicrobial susceptibility of 141 Campylobacter strains against ampicillin and tetracycline by using the disk diffusion test. Resistance of Campylobacter isolates among this method to ampicillin and tetracycline was detected in 65% and 35.5%, respectively (Fig. 2). The resistance we found to β-lactam among both our Campylobacter sppisolates was at the lower range of that reported for amoxicillin 87.5%) in the prior Brazilian study (De Moura et al. 2013De Moura H.M., Silva P.R., da Silva P.H.C., Souza N.R., Racanicci A.M.C. & Santana A.P. 2013. Antimicrobial resistance of Campylobacter jejuni isolated from chicken carcasses in the Federal District, Brazil. J. Food Prot. 76:691-693.). Expression of a penicillinase-type of β-lactamase in Campylobacter confers resistance to amoxicilin, ampicillin and ticarcilin (Iovine 2013Iovine N.M. 2013. Resistance mechanisms in Campylobacter jejuniVirulence 4:230-240.). This enzyme provided resistance to penicillins but not to cefotaxime and imipenem (Alfredson & Korolik 2005Alfredson D. & Korolik V. 2005. Isolation and expression of a novel molecular class D β-lactamase, OXA-61, from Campylobacter jejuni. Antimicrob. Agents Chemother. 49:2515-2518.). The level of resistance to Campylobacter spp. to tetracycline by phenotypic method was similar to that reported in other studies in Brazil (Kuana et al. 2008Kuana S.L., Santos L.R., Rodrigues L.B., Borsoi A., Moraes H.L.S., Salle C.T.P. & Nascimento V.P. 2008. Ocorrência de Campylobacter em lotes de frangos de corte e nas carcaças correspondentes. Ciênc. Anim. Bras. 9:480-486.). However, in contrast high presence of resistance (93.75%) was found byDe Moura et al. (2013)De Moura H.M., Silva P.R., da Silva P.H.C., Souza N.R., Racanicci A.M.C. & Santana A.P. 2013. Antimicrobial resistance of Campylobacter jejuni isolated from chicken carcasses in the Federal District, Brazil. J. Food Prot. 76:691-693..
The occurrence of Campylobacter spp. strains carrying resistance genes (tetracycline and β-lactam) and the energy-dependent multi-drug efflux pump.
Ampicillin is not recommended for the treatment of Campylobacter infections. However, the increasing resistance to fluoroquinolones, tetracycline and erythromycin of C. coli and C. jejuni strains, might compromise the effectiveness of treatment (Aarestrup & Engberg 2001Aarestrup F.M. & Engberg J. 2001. Antimicrobial resistance of thermophilic Campylobacter. Vet. Res. 32:311-321.,Engberg et al. 2001Engberg J., Aarestrup F.M., Taylor D.E., Gerner-Smidt P. & Nachamkin I. 2001. Quinolone and macrolide resistance in Campylobacter jejuni and C. coli: Resistance mechanisms and trends in human isolates. Emerg. Infect. Dis. 7:24-34.,Gibreel & Taylor 2006Gibreel A. & Taylor D.E. 2006. Macrolide resistance in Campylobacter jejuni and Campylobacter coli. J. Antimicrob. Chemother. 58:243-255.,Alfredson & Korolik 2007Alfredson D.A. & Korolik V. 2007. Antibiotic resistance and resistance mechanisms in Campylobacter jejuni and Campylobacter coli FEMS Microbiol. Lett. 277:123-132.,Silva et al. 2011Silva J., Leite D., Fernandes M., Mena C., Gibbs P.A. & Teixeira P. 2011. Campylobacter spp. as a foodborne pathogen: a review. Frontiers Microbiol. 2:200.). Recently, it was proposed that an oral β-lactam may provide an alternative therapy, although epidemiological studies have shown β-lactam resistance to be commonly observed in Campylobacter spp. (Griggs et al. 2009Griggs D.J., Peake L., Johnson M.M., Ghori S., Mott A. & Piddock L.J. 2009. β-Lactamase-mediated β-Lactam resistance in Campylobacter species: Prevalence of Cj0299 (blaOXA-61) and evidence for a novel β-Lactamase in C. jejuni. Antimicrob. Agents Chemother. 53:3357-3364.,Gormley et al. 2010Gormley F.J., Strachan N.J., Reay K., MacKenzie F.M., Ogden I.D., Dallas J.F. & Forbes K.J. 2010. Antimicrobial resistance profiles of Campylobacter from humans, retail chicken meat, and cattle feces. Foodborne Pathog. Dis.7:1129-1131.). β-lactam antimicrobials, which inhibit bacterial cell wall biosynthesis, have been the most commercially available antimicrobials in the market because of their high specificity and potent killing effect (Zeng et al. 2014Zeng X., Brown S., Gillespie B. & Lin J. 2014. A single nucleotide in the promoter region modulates the expression of the β-lactamase OXA-61 in Campylobacter jejuni. J. Antimicrob. Chemother.69:1215-1223.). Several major β-lactam resistance mechanisms have been characterized, including the production of β-lactamases (Lachance et al. 1991Lachance N., Gaudreau C., Lamothe F. & Lariviere L.A. 1991. Role of the β-lactamase of Campylobacter jejuni in resistance to β-lactam agents. Antimicrob. Agents Chemother. 35:813-818.,Griggs et al. 2009Griggs D.J., Peake L., Johnson M.M., Ghori S., Mott A. & Piddock L.J. 2009. β-Lactamase-mediated β-Lactam resistance in Campylobacter species: Prevalence of Cj0299 (blaOXA-61) and evidence for a novel β-Lactamase in C. jejuni. Antimicrob. Agents Chemother. 53:3357-3364.). A majority of C. jejuni and C. coli isolates are able to produce β-lactamases, which inactivate the β-lactam molecule by hydrolysis the structural lactam ring (Tajada et al. 1996Tajada P., Gomez-Garces J.-L., Alós J.-I., Balas D. & Cogollos R. 1996. Antimicrobial susceptibilities of Campylobacter jejuni and Campylobacter coli to 12 β-lactam agents and combinations with β-lactamase inhibitors. Antimicrob. Agents Chemother, 40:1924-1925.,Wieczorek & Osek 2013Wieczorek K. & Osek J. 2013. Antimicrobial resistance mechanisms among Campylobacter. BioMed Res. 2013:1-12.). In Campylobacter , ampicillin resistance and the associated β-lactamase production are chromosomally encoded (Alfredson & Korolik 2005Alfredson D. & Korolik V. 2005. Isolation and expression of a novel molecular class D β-lactamase, OXA-61, from Campylobacter jejuni. Antimicrob. Agents Chemother. 49:2515-2518.). A single nucleotide mutation (G→T transversion) upstream of blaOXA-61 was identified in the ampicillin-resistant derivative of C. jejuni NCTC 11168 (Zeng et al. 2014Zeng X., Brown S., Gillespie B. & Lin J. 2014. A single nucleotide in the promoter region modulates the expression of the β-lactamase OXA-61 in Campylobacter jejuni. J. Antimicrob. Chemother.69:1215-1223.). The results of phenotypic and genetic analyses of antimicrobial susceptibility were fully concordant. We showed that ampicillin resistance by disk diffusion in campylobacters isolated was associated with the presence of the blaOXA-61 gene carried on the chromosome.Griggs et al. (2009)Griggs D.J., Peake L., Johnson M.M., Ghori S., Mott A. & Piddock L.J. 2009. β-Lactamase-mediated β-Lactam resistance in Campylobacter species: Prevalence of Cj0299 (blaOXA-61) and evidence for a novel β-Lactamase in C. jejuni. Antimicrob. Agents Chemother. 53:3357-3364.reported that 91% of the ampicillin-resistant isolates carried the blaOXA-61 gene. This study found a high level of resistance to β -lactam gene in 65% (91/141) of isolates. Our results are similar to those described byObeng et al. (2012)Obeng A.S., Rickard H., Sexton M., Pang Y., Peng H. & Barton M. 2012. Antimicrobial susceptibilities and resistance genes in Campylobacter strains isolated from poultry and pigs in Australia. J. Appl. Microbiol. 113:294-307., who observed resistance of 59-65.4% for C. jejuni. In Brazilian studies,Hungaro et al. (2015)Hungaro H.M., Mendonça R.C.S., Rosa V.O., Badaró A.C.L., Moreira M.A.S. & Chaves J.B.P. 2015. Low contamination of Campylobacter spp. on chicken carcasses in Minas Gerais state, Brazil: Molecular characterization and antimicrobial resistance. Food Control 51:15-22.was found 100% resistant isolates to ampicillin.
An increase in tetracycline resistance in C. jejuni and C. coli strains has been observed in recent years (Luangtongkum et al. 2008Luangtongkum T., Morishita Y., Martin L., Choi I., Sahin O. & Zhang Q. 2008. Prevalence of tetracycline-resistant Campylobacter in organic broilers during a production cycle. Avian Dis. 52:487-490.). The tetO gene was detected in 35.5% (50/141) of isolates. Previous reports have indicated that tetO is present in 40% of C. jejuni isolates from chicken carcasses (Hungaro et al. 2015Hungaro H.M., Mendonça R.C.S., Rosa V.O., Badaró A.C.L., Moreira M.A.S. & Chaves J.B.P. 2015. Low contamination of Campylobacter spp. on chicken carcasses in Minas Gerais state, Brazil: Molecular characterization and antimicrobial resistance. Food Control 51:15-22.), and of C. jejuni from chickens 19.2-40.7% (Obeng et al. 2012Obeng A.S., Rickard H., Sexton M., Pang Y., Peng H. & Barton M. 2012. Antimicrobial susceptibilities and resistance genes in Campylobacter strains isolated from poultry and pigs in Australia. J. Appl. Microbiol. 113:294-307.). Tetracyclines have been suggested as an alternative choice in the treatment of clinical campylobacterosis, but in practice are rarely used (Moore et al. 2006Moore J.E., Barton M.D., Blair I.S., Corcoran D., Dooley J.S., Fanning S., Kempt I., Lastovica A.J., Lowery C.J., Matsuda M., McDowell D.A., McMahon A., Millar B.C., Rao R.J., Rooney P.J., Seal B.S., Snelling W.J. & Tolba O. 2006. The epidemiology of antibiotic resistance in Campylobacter . Microbes Infect. 8:1955-1966.). The tet(O) gene in Campylobacter encodes the Tet(O) protein that protects the ribosome from the inhibitory effect of tetracycline (Connell et al. 2003Connell S., Tracz D., Neirhaus K.H. & Taylor D.E. 2003. Ribosomal protection proteins and their mechanism of tetracycline resistance. Antimicrob. Agents Chemother. 47:3675-3681.). Its primary antimicrobial effect takes place by direct steric hindrance by binding to the A site in the 30S subunit, thus hindering the movement of transfer RNA and inhibits peptide elongation (Wieczorek & Osek 2013Wieczorek K. & Osek J. 2013. Antimicrobial resistance mechanisms among Campylobacter. BioMed Res. 2013:1-12.). Tet(O) is coded by a gene located on plasmids of different sizes and in some strains chromosomally (Lee et al. 1994Lee C., Tai C., Lin S. & Chen Y. 1994. Occurrence of plasmids and tetracycline resistance among Campylobacter jejuni and Campylobacter coli isolated from whole market chickens and clinical samples. Int. J. Food Microbiol.24:161-170.,Gibreel et al. 2004Gibreel A., Tracz D.M., Nonaka L., Ngo T.M., Connell S.R. & Taylor D.E. 2004. Incidence of antibiotic resistance in Campylobacter jejuni isolated in Alberta, Canada, from 1999 to 2002, with special reference to tet(O)-mediated tetracycline resistance.. Antimicrob. Agents Chemother 48:3442-3450.,Dasti et al. 2007Dasti J.I., Gross U., Pohl S., Lugert R., Weig M. & Schmidt-Ott R. 2007. Role of the plasmid-encoded tet (O) gene in tetracycline-resistant clinical isolates of Campylobacter jejuni and Campylobacter coli . J. Med. Microbiol. 56:833-837.). However, studies provide evidence that tetracycline-resistance Campylobacter have been recovered from organic and other production systems in which no antimicrobial have been used, which indicates that prior or current use may not be a defining attribution to resistance (Piddock et al. 2000Piddock L.J.V., Ricci V., Stanley K. & Jones K. 2000. Activity of antibiotics used in human medicine for Campylobacter jejuni isolated from farm animals and their environment in Lancashire, UK. J. Antimicrob. Chemother.46:303-306.,Luangtongkum et al. 2008Luangtongkum T., Morishita Y., Martin L., Choi I., Sahin O. & Zhang Q. 2008. Prevalence of tetracycline-resistant Campylobacter in organic broilers during a production cycle. Avian Dis. 52:487-490.,Cox et al. 2009Cox N.A., Richardson L.J., Buhr R.J. & Fedorka-Cray P.J. 2009. Campylobacter species occurrence within internal organs and tissues of commercial caged Leghorn laying hens. Poultry Sci. 88:2449-2456.).
In Campylobacter is described a multidrug efflux system (CmeABC), belonging to the Resistance Nodulation Division (RND) family of transporters that conferring intrinsic resistance to various antimicrobials and toxic compounds (Pumbwe & Piddock 2002Pumbwe L. & Piddock L.J. 2002. Identification and molecular characterization of CmeB, a Campylobacter jejuni multidrug efflux pump. FEMS Microbiol. Lett.206:185-189.,Lin et al. 2002Lin J., Michel L.O. & Zhang Q. 2002. CmeABC functions as a multidrug efflux system in Campylobacter jejuni. Antimicrob. Agents Chemother. 46:2124-2131.,Guo et al. 2010Guo B., Lin J., Reynolds D.L. & Zhang Q. 2010. Contribution of the multidrug efflux transporter CmeABC to antibiotic resistance in different Campylobacter species. Foodborne Path. Dis. 7:77-83.). This pump is widely distributed in Campylobacter, including C. coli, and is constitutively expressed (Payot et al. 2002Payot S., Cloeckaert A. & Chaslus-Dancla E. 2002. Selection and characterization of fluoroquinolone-resistant mutants of Campylobacter jejuni using enrofloxacin. Microb. Drug Resist. 8:335-343.).Lin et al. (2003)Lin J., Sahin O., Michel L.O. & Zhang Q. 2003. Critical role of multi-drug efflux pump CmeABC in bile resistance and in vivo colonisation of Campylobacter jejuni. Infect. Immun. 71:4250-4259.showed that this efflux pump is essential for the growth and survival of C. jejuni in chicken intestinal extracts, and their report indicates that, through mediating resistance to bile salts in the intestinal tract, CmeABC allows C. jejuni to colonize chickens successfully. It is also possible that, in some species, efflux pumps that export antimicrobial agents also export virulence determinants, such as adhesins, toxins or other proteins that are important for the colonization and infection of human and animal cells (Piddock 2006Piddock L.J. 2006. Multidrug-resistance efflux pumps? not just for resistance. Nat. Rev. Microbiol. 4:629-636.). The role of efflux pumps in biofilm formation, persistence, resistance to antimicrobials and biocides was largely hypothetical; until some recent studies have discovered that expression of efflux pumps contribute to antimicrobial resistance by biofilms (Zhang & Mah 2008Zhang L. & Mah T.F. 2008. Involvement of a novel efflux system in biofilm-specific resistance to antibiotics. J. Bacteriol. 190:4447-4452.,Andersen et al. 2015Andersen J.L., He G.X., Kakarla P., KC R., Kumar S., Lakra W.S., Mukherjee M.M., Ranaweera I., Shrestha U., Tran T. & Varela M.F. 2015. Multidrug efflux pumps from Enterobacteriaceae, Vibrio cholerae and Staphylococcus aureus bacterial food pathogens. Int. J. Environ. Res. Public Health 12:1487-1547.). The higher expression of efflux pumps offers physiological advantage in the form of expelling toxic wastes from the densely populated biofilms and in the process efflux antimicrobial compounds, which is manifested as enhanced antimicrobial resistances of biofilms (Andersen et al. 2015Andersen J.L., He G.X., Kakarla P., KC R., Kumar S., Lakra W.S., Mukherjee M.M., Ranaweera I., Shrestha U., Tran T. & Varela M.F. 2015. Multidrug efflux pumps from Enterobacteriaceae, Vibrio cholerae and Staphylococcus aureus bacterial food pathogens. Int. J. Environ. Res. Public Health 12:1487-1547.). In addition, adaptation to the environment, such as quorum sensing and biofilm formation can also contribute to bacterial persistence (Martins et al. 2013Martins M., McCusker M.P., Viveiros M., Couto I., Fanning S., Pagès J.M. & Amaral L. 2013. A simple method for assessment of MDR bacteria for over-expressed efflux pumps. Open Microbiol. J. 7:72-82.). In this study, twenty six isolates (18.5%) presents the gene that encode a multidrug efflux pumpThe CmeABC efflux pump may also contribute to β-lactam resistance (Iovine 2013Iovine N.M. 2013. Resistance mechanisms in Campylobacter jejuniVirulence 4:230-240.).Gibreel et al. (2007Gibreel A., Wetsch N.M. & Taylor D.E. 2007. Contribution of the CmeABC efflux pump to macrolide and tetracycline resistance in Campylobacter jejuni.. Antimicrob. Agents Chemother 51:3212-3216.) reported that inactivation of the cme B gene in the resistant isolates examined led to a 16 to 128-fold decrease in tetracycline minimum inhibitory concentration (MIC), resulting in the complete restoration of tetracycline susceptibility. Studies also suggested that when both CmeABC and tetO are functional, the impact on tetracycline resistance is synergistic (Lin et al. 2002Lin J., Michel L.O. & Zhang Q. 2002. CmeABC functions as a multidrug efflux system in Campylobacter jejuni. Antimicrob. Agents Chemother. 46:2124-2131.,Pumbwe & Piddock 2002Pumbwe L. & Piddock L.J. 2002. Identification and molecular characterization of CmeB, a Campylobacter jejuni multidrug efflux pump. FEMS Microbiol. Lett.206:185-189.).
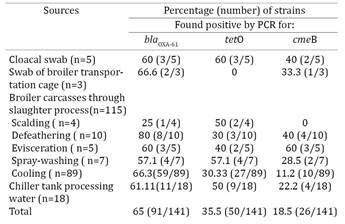
The obtained data revealed that 7.1% Campylobacter strains possess three resistance determinant (n=10), and 43 isolates (30.5%) presents two r esistanceassociated genes. A percentage 35.5% of isolates presents one markers (n = 50). No resistance markers were found in 38 isolates (27%). Moreover, 36 out of the 141 Campylobacter strains (25.6%) were found to be resistant to at least two different antimicrobial resistance groups (blaOXA-61 and tetO). The distribution of resistance markers among Campylobacter spp. tested according to the source of isolation are summarized inTable 2. This study has found a high genetic diversity of Campylobacter spp. isolated from broiler slaughterhouses. This observation suggests that such genotypes may be particularly adapted to survive cleaning and disinfection stress in poultry slaughterhouses and may contaminate the carcasses during processing (Peyrat et al. 2008Peyrat M.B., Soumet C., Maris P., Sanders P. 2008. Recovery of Campylobacter jejuni from surfaces of poultry slaughterhouses after cleaning and disinfection procedures: Analysis of a potential source of carcass contamination. Int. J. Food Microbiol.124:188-194.). Additionally, our results showed that Campylobacter may easily contaminate poultry carcasses at slaughter process.
Distribution of resistance markers among Campylobacter spp. tested according to the source of isolation
Antimicrobial resistance in both medicine and agriculture is documented by the World Health Organization (WHO), along with other various national authorities as a major emerging problem of public health importance, as well as consequences for animal treatment (Moore et al. 2006Moore J.E., Barton M.D., Blair I.S., Corcoran D., Dooley J.S., Fanning S., Kempt I., Lastovica A.J., Lowery C.J., Matsuda M., McDowell D.A., McMahon A., Millar B.C., Rao R.J., Rooney P.J., Seal B.S., Snelling W.J. & Tolba O. 2006. The epidemiology of antibiotic resistance in Campylobacter . Microbes Infect. 8:1955-1966.,Silva et al. 2011Silva J., Leite D., Fernandes M., Mena C., Gibbs P.A. & Teixeira P. 2011. Campylobacter spp. as a foodborne pathogen: a review. Frontiers Microbiol. 2:200.). Bacteria have developed multiple ways of becoming resistant to antimicrobials; in most cases bacteria are exposed to these substances, but have found a way to evade or resist the antimicrobial agent (Lin et al. 2003Lin J., Sahin O., Michel L.O. & Zhang Q. 2003. Critical role of multi-drug efflux pump CmeABC in bile resistance and in vivo colonisation of Campylobacter jejuni. Infect. Immun. 71:4250-4259.,Griggs et al. 2009Griggs D.J., Peake L., Johnson M.M., Ghori S., Mott A. & Piddock L.J. 2009. β-Lactamase-mediated β-Lactam resistance in Campylobacter species: Prevalence of Cj0299 (blaOXA-61) and evidence for a novel β-Lactamase in C. jejuni. Antimicrob. Agents Chemother. 53:3357-3364.,Wieczorek & Osek 2013Wieczorek K. & Osek J. 2013. Antimicrobial resistance mechanisms among Campylobacter. BioMed Res. 2013:1-12.). Further research in understanding the antimicrobial resistance mechanisms will facilitate the selection of antimicrobials for clinical treatment and the formulation of diagnostic media for various Campylobacter spp. To our knowledge, this is the first study to determine the frequency of the blaOXA-61, tet O and cmeB genes in campylobacters from poultry origin of Rio Grande do Sul state, Brazil. Our results also emphasize the need for a surveillance and monitoring system and risk analyzes for the prevalence and resistances of Campylobacter in poultry and other food animals.
References
- Aarestrup F.M. & Engberg J. 2001. Antimicrobial resistance of thermophilic Campylobacter Vet. Res. 32:311-321.
- Alfredson D.A. & Korolik V. 2007. Antibiotic resistance and resistance mechanisms in Campylobacter jejuni and Campylobacter coli FEMS Microbiol. Lett. 277:123-132.
- Alfredson D. & Korolik V. 2005. Isolation and expression of a novel molecular class D β-lactamase, OXA-61, from Campylobacter jejuni Antimicrob. Agents Chemother. 49:2515-2518.
- Andersen J.L., He G.X., Kakarla P., KC R., Kumar S., Lakra W.S., Mukherjee M.M., Ranaweera I., Shrestha U., Tran T. & Varela M.F. 2015. Multidrug efflux pumps from Enterobacteriaceae, Vibrio cholerae and Staphylococcus aureus bacterial food pathogens. Int. J. Environ. Res. Public Health 12:1487-1547.
- Angulo F.J., Nunnery J.A. & Bair H.D. 2004. Antimicrobial resistance in zoonotic enteric pathogens. Rev. Sci Tech. OIE 23:485-496.
- Avrain L., Humbert F., L'Hospitalier R., Sanders P., Vernozy-Rozand C. & Kempf I. 2003. Antimicrobial resistance in Campylobacter from broilers: association with production type and antimicrobial use. Vet. Microbiol. 96:267-276.
- Blaser M.J. 1995. Campylobacter and related species, p.1948-1956. In: Mandell G.L., Bennett J.E. & Dolin R. (Eds), Principles and Practice of Infectious Diseases. 4th ed. Churchill Livingstone, New York, NY.
- Borsoi A., Santin E., Santos L.R., Salle C.T.P., Moraes H.L.S. & Nascimento V.P. 2009. Inoculation of newly hatched broiler chicks with two Brazilian isolates of Salmonella Heidelberg strains with different virulence gene profiles, antimicrobial resistance, and pulsed field gel electrophoresis patterns to intestinal changes evaluation. Poult. Sci. 88:750-758.
- CDC 2013. Incidence and trends of infection with pathogens transmitted commonly through food. Foodborne Diseases Active Surveillance Network, 10 U.S. Sites, 1996-2012. Morbidity and Mortality Weekly Report 19:283-287.
- CLSI 2010. Methods for Antimicrobial Dilution and Disk Susceptibility Testing of Infrequently Isolated or Fastidious Bacteria: approved guideline. 2nd ed. Clinical and Laboratory Standards Institute (M45eA2), Wayne, PA.
- CLSI2012. Performance Standards for Antimicrobial Susceptibility Testing. 22nd Informational Supplement. Clinical and Laboratory Standards Institute (M100eS22), Wayne, PA.
- Conlan A.J., Coward C., Grant A.J., Maskell D.J. & Gog J.R. 2007. Campylobacter jejuni colonization and transmission in broiler chickens: a modelling perspective. J. R. Soc. Interface 4:819-829.
- Connell S., Tracz D., Neirhaus K.H. & Taylor D.E. 2003. Ribosomal protection proteins and their mechanism of tetracycline resistance. Antimicrob. Agents Chemother. 47:3675-3681.
- Cox N.A., Richardson L.J., Buhr R.J. & Fedorka-Cray P.J. 2009. Campylobacter species occurrence within internal organs and tissues of commercial caged Leghorn laying hens. Poultry Sci. 88:2449-2456.
- Dasti J.I., Gross U., Pohl S., Lugert R., Weig M. & Schmidt-Ott R. 2007. Role of the plasmid-encoded tet (O) gene in tetracycline-resistant clinical isolates of Campylobacter jejuni and Campylobacter coli . J. Med. Microbiol. 56:833-837.
- De Moura H.M., Silva P.R., da Silva P.H.C., Souza N.R., Racanicci A.M.C. & Santana A.P. 2013. Antimicrobial resistance of Campylobacter jejuni isolated from chicken carcasses in the Federal District, Brazil. J. Food Prot. 76:691-693.
- Denis M., Soumet C., Rivoal K., Ermel G., Blivet D., Salvat G. & Colin P. 1999. Development of a m-PCR assay for simultaneous identification of Campylobacter jejuni and C. coli Lett. Appl. Microbiol. 29:406-410.
- Ellström P., Hansson I., Söderström C., Olsson Engvall E. & Rautelin H. 2014. A prospective follow-up study on transmission of Campylobacter from poultry to abattoir workers. Foodborne Pathog. Dis. 11:684-688.
- Engberg J., Aarestrup F.M., Taylor D.E., Gerner-Smidt P. & Nachamkin I. 2001. Quinolone and macrolide resistance in Campylobacter jejuni and C. coli: Resistance mechanisms and trends in human isolates. Emerg. Infect. Dis. 7:24-34.
- EFSA 2011. The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2009EFSA J. 9:2090-2468.
- Garin B., Gouali M., Wouafo M., Perchec A.-M., Thu P.M., Ravaonindrina N., Urbès F., Gay M., Diawara A., Leclercq A., Rocourt J. & Pouillot R. 2012. Prevalence, quantification and antimicrobial resistance of Campylobacter spp. on chicken neck-skins at points of slaughter in 5 major cities located on 4 continents. Int. J. Food Microbiol. 157:102-107.
- Gibreel A. & Taylor D.E. 2006. Macrolide resistance in Campylobacter jejuni and Campylobacter coli J. Antimicrob. Chemother. 58:243-255.
- Gibreel A., Tracz D.M., Nonaka L., Ngo T.M., Connell S.R. & Taylor D.E. 2004. Incidence of antibiotic resistance in Campylobacter jejuni isolated in Alberta, Canada, from 1999 to 2002, with special reference to tet(O)-mediated tetracycline resistance.. Antimicrob. Agents Chemother 48:3442-3450.
- Gibreel A., Wetsch N.M. & Taylor D.E. 2007. Contribution of the CmeABC efflux pump to macrolide and tetracycline resistance in Campylobacter jejuni. Antimicrob. Agents Chemother 51:3212-3216.
- Godschalk P.C., Heikema A.P., Gilbert M., Komagamine T., Ang C.W., Glerum J., Brochu D., Li J., Yuki N., Jacobs B.C., van Belkum A. & Endtz H.P. 2004. The crucial role of Campylobacter jejuni genes in anti-ganglioside antibody induction in Guillain-Barré syndrome. J. Clin. Invest. 114:1659-1665.
- Gormley F.J., Strachan N.J., Reay K., MacKenzie F.M., Ogden I.D., Dallas J.F. & Forbes K.J. 2010. Antimicrobial resistance profiles of Campylobacter from humans, retail chicken meat, and cattle feces. Foodborne Pathog. Dis.7:1129-1131.
- Griggs D.J., Peake L., Johnson M.M., Ghori S., Mott A. & Piddock L.J. 2009. β-Lactamase-mediated β-Lactam resistance in Campylobacter species: Prevalence of Cj0299 (blaOXA-61) and evidence for a novel β-Lactamase in C. jejuni Antimicrob. Agents Chemother. 53:3357-3364.
- Guo B., Lin J., Reynolds D.L. & Zhang Q. 2010. Contribution of the multidrug efflux transporter CmeABC to antibiotic resistance in different Campylobacter species. Foodborne Path. Dis. 7:77-83.
- Hardy C.G., Lackey L.G., Cannon J., Price L.B. & Silbergeld E.K. 2011. Prevalence of potentially neuropathic Campylobacter jejuni strains on commercial broiler chicken products. Int. J. Food Microbiol.145:395-399.
- Hungaro H.M., Mendonça R.C.S., Rosa V.O., Badaró A.C.L., Moreira M.A.S. & Chaves J.B.P. 2015. Low contamination of Campylobacter spp. on chicken carcasses in Minas Gerais state, Brazil: Molecular characterization and antimicrobial resistance. Food Control 51:15-22.
- Iovine N.M. 2013. Resistance mechanisms in Campylobacter jejuniVirulence 4:230-240.
- ISO 2006. International Standard Organization 10272-1:2006 describes a horizontal method for the detection of Campylobacter spp.
- Janssen R., Krogfelt K.A., Cawthraw S.A., van Pelt W., Wagenaar J.A. & Owen R.J. 2008. Host-pathogen interactions in Campylobacter infections: the host perspective. Clin. Microbiol. Rev. 21:505-518.
- Khachatourians G.G. 1998. Agricultural use of antibiotics and the evolution and transfer of antibiotic-resistant bacteria. Can. Med. Assoc. J. 159:1129-1136.
- Kuana S.L., Santos L.R., Rodrigues L.B., Borsoi A., Moraes H.L.S., Salle C.T.P. & Nascimento V.P. 2008. Ocorrência de Campylobacter em lotes de frangos de corte e nas carcaças correspondentes. Ciênc. Anim. Bras. 9:480-486.
- Lachance N., Gaudreau C., Lamothe F. & Lariviere L.A. 1991. Role of the β-lactamase of Campylobacter jejuni in resistance to β-lactam agents. Antimicrob. Agents Chemother. 35:813-818.
- Lee C., Tai C., Lin S. & Chen Y. 1994. Occurrence of plasmids and tetracycline resistance among Campylobacter jejuni and Campylobacter coli isolated from whole market chickens and clinical samples. Int. J. Food Microbiol.24:161-170.
- Lin J., Michel L.O. & Zhang Q. 2002. CmeABC functions as a multidrug efflux system in Campylobacter jejuni Antimicrob. Agents Chemother. 46:2124-2131.
- Lin J., Sahin O., Michel L.O. & Zhang Q. 2003. Critical role of multi-drug efflux pump CmeABC in bile resistance and in vivo colonisation of Campylobacter jejuni Infect. Immun. 71:4250-4259.
- Luangtongkum T., Jeon B., Han J., Plummer P., Logue C.M. & Zhang Q. 2009. Antibiotic resistance in Campylobacter: emergence, transmission and persistence. Future Microbiol. 4:189-200.
- Luangtongkum T., Morishita Y., Martin L., Choi I., Sahin O. & Zhang Q. 2008. Prevalence of tetracycline-resistant Campylobacter in organic broilers during a production cycle. Avian Dis. 52:487-490.
- Martins M., McCusker M.P., Viveiros M., Couto I., Fanning S., Pagès J.M. & Amaral L. 2013. A simple method for assessment of MDR bacteria for over-expressed efflux pumps. Open Microbiol. J. 7:72-82.
- Moore J.E., Barton M.D., Blair I.S., Corcoran D., Dooley J.S., Fanning S., Kempt I., Lastovica A.J., Lowery C.J., Matsuda M., McDowell D.A., McMahon A., Millar B.C., Rao R.J., Rooney P.J., Seal B.S., Snelling W.J. & Tolba O. 2006. The epidemiology of antibiotic resistance in Campylobacter . Microbes Infect. 8:1955-1966.
- Moore J.E., Corcoran D., Dooley J.S., Fanning S., Lucey B., Matsuda M., McDowell D.A., Mégraud F., Millar B.C., O'Mahony R., O'Riordan L., O'Rourke M., Rao J.R., Rooney P.J., Sails A. & Whyte P. 2005. Campylobacter. Vet. Res. 36:351-382.
- Obeng A.S., Rickard H., Sexton M., Pang Y., Peng H. & Barton M. 2012. Antimicrobial susceptibilities and resistance genes in Campylobacter strains isolated from poultry and pigs in Australia. J. Appl. Microbiol. 113:294-307.
- Park S.F. 2002. The physiology of Campylobacter species and its relevance to their role as foodborne pathogens. Int. J. Food Microbiol.74:177-188.
- Payot S., Cloeckaert A. & Chaslus-Dancla E. 2002. Selection and characterization of fluoroquinolone-resistant mutants of Campylobacter jejuni using enrofloxacin. Microb. Drug Resist. 8:335-343.
- Peyrat M.B., Soumet C., Maris P., Sanders P. 2008. Recovery of Campylobacter jejuni from surfaces of poultry slaughterhouses after cleaning and disinfection procedures: Analysis of a potential source of carcass contamination. Int. J. Food Microbiol.124:188-194.
- Piddock L.J. 2006. Multidrug-resistance efflux pumps? not just for resistance. Nat. Rev. Microbiol. 4:629-636.
- Piddock L.J.V., Ricci V., Stanley K. & Jones K. 2000. Activity of antibiotics used in human medicine for Campylobacter jejuni isolated from farm animals and their environment in Lancashire, UK. J. Antimicrob. Chemother.46:303-306.
- Pratt A. & Korolik V. 2005. Tetracycline resistance of Australian Campylobacter jejuni and Campylobacter coli isolates. J. Antimicrob. Chemother.55:452-460.
- Pumbwe L. & Piddock L.J. 2002. Identification and molecular characterization of CmeB, a Campylobacter jejuni multidrug efflux pump. FEMS Microbiol. Lett.206:185-189.
- Rowe-Magnus D.A., Guerout A.M. & Mazel D. 2002. Bacterial resistance evolution by recruitment of super-integron gene cassettes. Mol. Microbiol. 43:1657-1669.
- Silva J., Leite D., Fernandes M., Mena C., Gibbs P.A. & Teixeira P. 2011. Campylobacter spp. as a foodborne pathogen: a review. Frontiers Microbiol. 2:200.
- Tajada P., Gomez-Garces J.-L., Alós J.-I., Balas D. & Cogollos R. 1996. Antimicrobial susceptibilities of Campylobacter jejuni and Campylobacter coli to 12 β-lactam agents and combinations with β-lactamase inhibitors. Antimicrob. Agents Chemother, 40:1924-1925.
- Van den Berg B., Walgaard C., Drenthen J., Fokke C., Jacobs B.C. & van Doorn P.A. 2014. Guillain-Barré syndrome: pathogenesis, diagnosis, treatment and prognosis. Nat. Rev. Neurol. 10:469-482.
- Wieczorek K. & Osek J. 2013. Antimicrobial resistance mechanisms among Campylobacter BioMed Res. 2013:1-12.
- Zeng X., Brown S., Gillespie B. & Lin J. 2014. A single nucleotide in the promoter region modulates the expression of the β-lactamase OXA-61 in Campylobacter jejuni J. Antimicrob. Chemother.69:1215-1223.
- Zhang L. & Mah T.F. 2008. Involvement of a novel efflux system in biofilm-specific resistance to antibiotics. J. Bacteriol. 190:4447-4452.
Publication Dates
-
Publication in this collection
July 2015
History
-
Received
08 Mar 2015 -
Accepted
21 July 2015