ABSTRACT:
Chickens are considered to be potential reservoirs of Newcastle disease virus (NDV). In this study, six Newcastle disease virus strains were isolated and characterized in Tibetan chickens. The HN gene was sequenced, and phylogenetic relationship to reference strains was studied. The phylogenetic analysis demonstrated that these six isolated strains were closely related to NDV isolates of the reference strains GQ245823, KT002186, KU527561, KJ563939, AY225110, EU305607, KM056357, Y18898, GQ245832, AF077761 and lasota strain. Among them, EU305607, KJ563939 and KM056357 were isolated from India, while lasota strain came from attenuated vaccine widely used in China. Then, mean death time (MDT) and intracerebral pathogenicity index (ICPI) were used to estimate the pathogenicity of the isolates. Pathogenicity experiment showed HNH1 and HN17 to be virulent. Our results indicated that genetically diverse viruses circulate in Tibetan chickens, and based upon the phlogeographic analysis, we estimated the origin of ancestral viruses of the isolates and its sister strains located in India and China (lasota strain). It indicates the importance of continuous surveillance to enhance current understanding of the genetic evolution of the NDV strains.
INDEX TERMS:
Newcastle disease virus (NDV); Tibetan chickens; phylogenetic analysis; pathogenicity
Introduction
Newcastle disease (ND) is recognized as one of the highly contagious and lethal infectious disease of poultry industry globally (Munir et al. 2015Munir T., Aslam A., Zahid B., Ahmed I., Imranand M.S. & Ijaz M. 2015. Potential of commonly resident wild birds towards newcastle disease virus transmission. Pakistan Vet. J. 35(1):106-107.), mainly characterized by diarrhea, dyspnea, neurological symptoms, cloaca hemorrhage, focal glandular gastric bleeding or ulcers, intestinal mucosal bleeding and necrosis of the pancreas or the spleen (Zhang et al. 2011Zhang S., Wang X., Zhao C., Liu D., Hu Y., Zhao J. & Zhang G. 2011. Phylogenetic and pathotypical analysis of two virulent Newcastle disease viruses isolated from domestic ducks in China. PLoS One 6:e25000.). ND is caused by virulent strains of the ND virus (NDV), which belong to the genus Avulavirus, sub-family Paramyxoviridae, family Paramyxovirinae (Ge et al. 2015Ge M., Zhang W., Shi G., Xiao C., Zhao X. & Zhang R. 2015. Astragalus polysaccharide perseveres cytomembrane capacity against Newcastle disease virus infection. Pakistan Vet. J. 35(3):382-384.), with six transcriptional units encoding nucleocapsid protein (NP), phosphoprotein (P), matrix protein (M), fusion protein (F), haemagglutinin-neuraminidase protein (HN) and polymerase protein (L) (Mayo 2002aMayo M.A. 2002a. A summary of taxonomic changes recently approved by ICTV. Archs Virology 147:1655-1663., 2002bMayo M.A. 2002b. Virus taxonomy-Houston 2002. Archs Virol. 147:1071-1076.).
For many years, NDV has caused large-scale outbreaks in many countries around the world, especially in China (Qin et al. 2008Qin Z.M., Tan L.T., Xu H.Y., Ma B.C., Wang Y.L., Yuan X.Y. & Liu W.J. 2008. Pathotypical characterization and molecular epidemiology of Newcastle disease virus isolates from different hosts in China from 1996 to 2005. J. Clin. Microbiol. 46:601-611., Zhang et al. 2010Zhang R., Pu J. & Su J.L. 2010. Phylogenetic characterization of Newcastle disease virus isolated in the mainland of China during 2001-2009. Vet. Microbio. 141:246-257., Kang et al. 2016Kang Y., Xiang B., Yuan R., Zhao X., Feng M., Gao P., Li Y., Li Y., Ning Z. & Ren T. 2016. Phylogenetic and Pathotypic Characterization of Newcastle Disease Viruses Circulating in South China and Transmission in Different Birds. Front. Microbiol. 7:119.), and the infection types and the virulent strength vary with obvious regionality. In spite of extensive vaccination with the live attenuated vaccines, massive outbreaks are reported in China, and there is a long way to go to ultimately eradicate NDV (Bogoyavlenskiy et al. 2009Bogoyavlenskiy A., Berezin V., rilipov A., Usachev E., Lyapina O. & Korotetskiy I. 2009. Newcastle disease outbreaks in Kazakhstan and Kyrgyzstan during 1998, 2000, 2001, 2003, 2004, and 2005 were caused by viruses of the genotypes VIIb and VIId. Virus Genes 39:94-101., Kumar 2015Kumar U. & Kumar S. 2015. Molecular characterization of an apoptotic strain of Newcastle disease virus isolated from an outbreak in India. Cancer Gene Therapy 22:402-409.). Previously, the phylogenetic and pathotypic characterization of NDV has been reported in most breeds of chickens in China (Cao et al. 2001Cao D.J., Guo X., Liang R., Yan L.H., Liu P.X., Min P. & Chen J. 2001. Molecular epidemiology of newcastle disease virus isolates from part region of China. Chinese J. Prev. Vet. Med. 23:29-32. (In Chinese), Qin et al. 2008Qin Z.M., Tan L.T., Xu H.Y., Ma B.C., Wang Y.L., Yuan X.Y. & Liu W.J. 2008. Pathotypical characterization and molecular epidemiology of Newcastle disease virus isolates from different hosts in China from 1996 to 2005. J. Clin. Microbiol. 46:601-611., Zhang et al. 2010Zhang R., Pu J. & Su J.L. 2010. Phylogenetic characterization of Newcastle disease virus isolated in the mainland of China during 2001-2009. Vet. Microbio. 141:246-257.); however, none of them has been reported in Tibetan chickens.
The Tibetan chicken, aboriginal chicken breed, has a very wide distribution at altitudes of 2,200 to 4,100 m in the Qinghai-Tibet Plateau, with a history of domestication more than 1,000 years at high altitude, breeds successfully and has good resistance to hypobaric hypoxia and cold grazing conditions (Zhang et al. 2005Zhang H., Wu C., Chamba Y., Ma X., Tang X. & Po B. 2005. Curve analysis of embryonic mortality in chickens incubated at high altitude. J. China Agric. Univ. 10:109-114., Zhang et al. 2006Zhang H., Wu C., Chamba Y., Ling Y. & Luo Z. 2006. Adaptability to high altitude and NOS activity of lung in Tibetan chicken. J. China Agric. Univ. 11:35-38.). To the best of our knowledge, no information is available on the genetic characterization and pathogenicity of NDV in Tibetan chickens from the Tibetan Plateau of China. In this study, we investigated the presence of NDV in Tibetan chickens and selected six viruses to evaluate the genetic characterization and pathogenicity of NDV derived from Tibetan chickens.
Materials and Methods
The study site. The present study was carried out in Nyingchi Prefecture, in southeastern Tibet, China. As described by researcher (Zhang et al. 2014aZhang Y.Y., Shao M.Y., Yu X.H., Zhao J. & Zhang G.Z. 2014a. Molecular characterization of chicken-derived genotype VIId Newcastle disease virus isolates in China during 2005-2012 reveals a new length in hemagglutinin-neuraminidase. Infect. Genet. Evol. 21:359-366.), this area is geographically isolated from Tibet and Sichuan Provinces by Himalayas, and shares borders with Yunnan and Qinghai Provinces. Nepal in the southwest, and India and Myanmar in the south. The average elevation of the surveyed area is 3100 m above sea level.
Sample collection, NDV isolation and identification. A total of six NDVs were isolated from the organs (brain, heart, liver, spleen, lungs) of dead or killed moribund Tibetan chickens between 2015 and 2016. The virus isolation was carried out by using the guidelines and standards provided by the World Organization for Animal Health (Anonymous 2006Anonymous 2006. Newcastle disease, chapter 2.1.15. In: OIE Manual of Diagnostic Tests and Vaccines for Terrestrial Animals. World Organisation for Animal Health, Paris.). Briefly, homogenized and clarified organ suspensions were made in viral transportation media (VTM) containing penicillin (1000 U/mL) and streptomycin (1000 U/mL), at 4°C overnight, and the organ suspensions were plaque-purified three times by inoculating into the allantoic cavity of 9 to 11 day old specific pathogen free (SPF, Beijing Merial Vital Laboratory Technology Co., Ltd) chicken embryos (five eggs per sample) according to the standard methods of the virus isolation, and the chicken embryos were incubated at 37°C (Kumar & Kumar 2015Kumar S. 2015. Newcastle disease virus outbreaks in India: time to revisit the vaccine type and strategies. Vaccine 33:3268-3269.). After 3 days, allantoic fluid was harvested and after three series of blind passages, the NDV in the harvested allantoic fluid was tested using haemagglutination inhibition (HI) and haemagglutination (HA) tests. Inactivated NDV antigen strain (lasota strain) was used as a positive control. The virus stocks grown in allantoic fluids were stored at -70°C until subsequent use and further analysis.
Pathogenicity analysis. The pathogenic potential for the isolated viruses was evaluated using standard assay methods to determine the mean death time (MDT) using 9-days-old SPF embryonated chicken eggs and intracerebral pathogenicity index (ICPI) test using 1-day-old SPF chicks (Anonymous, 2006Anonymous 2006. Newcastle disease, chapter 2.1.15. In: OIE Manual of Diagnostic Tests and Vaccines for Terrestrial Animals. World Organisation for Animal Health, Paris., Kim et al. 2012Kim S.H., Xiao S., Shive H., Collins P.L. & Samal S.K. 2012. Replication, neurotropism, and pathogenicity of avian paramyxovirus serotypes 1-9 in chickens and ducks. PLoS One 7:34927.).
Molecular detection and characterization. Total RNA of the isolated strains was extracted using the TIANamp Virus DNA/RNA Kit (TianGen, China) according to the instructions of manufacture. A RT-PCR amplification approach was used to amplify HN and F genes. Based on the published F and HN gene sequences in the GenBank database, we designed two pairs of primers (Table 1).
For PCR, forward (5’-ATGGACCGCGCCGTTAGC-3’) and reverse (5’-CTAGCCAGACCTGGCTTCTCT-3’) primers were used. Polymerase chain reaction was used and reverse transcription was done by Quant One Step RT-PCR Kit (TianGen Co., LTD, China). The PCR was performed with the following cycling protocol: one cycle at 95°C for 40 s, and 35 amplification cycles at 56°C for 50s, and 72°C for 1.2 min after an initial hot start at 95°C and ending with 72°C each for 10 min. The PCR products were separated on agarose gel (1 %) along with ethidium bromide (at the rate of 0.5μg/ml) in the 0.5×TBE buffer at 5 V/cm for 30 minutes by electrophoresis. The RT-PCR products were analyzed on a 1.5% agarose gel stained with ethidium bromide following electrophoresis and the products were purified using a TaKaRa MiniBEST Agarose Gel DNA Extraction Kit Ver.4.0 (Takara Biotechnology Co., Ltd, Dalian, China) according to manufacturer’s instructions. The positive products were cloned into PGEM®-T Easy vector (You ni kang Co., Ltd, China) and the positive clones were identified by blue-white colony screening method. Further confirmation was carried out by PCR. Then the positive products were sequenced by a commercial company (Sangon Biosciences Co., Ltd, Wuhan, China).
Phylogenetic analysis. The nucleotide sequences editing of the HN gene of the six isolates in this study were compared with reference NDV sequences available at NCBI database using the maximum likelihood method with statistical analysis based on 1000 bootstrapping replicates. The alignments were conducted using Molecular Evolutionary Genetic Analysis (MEGA 6.0) and the DNASTAR (version 3.3.8; DNASTAR) software. Phylogenetic tree was constructed using the neighbor-joining method. The evolutionary distances were estimated using the Kimura two-parameter method (Tamura et al. 2013Tamura K., Stecher G., Peterson D., Filipski A. & Kumar S. 2013. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 30:2725-2729.).
Results and Discussion
The genetic characterization and pathogenicity of NDV has been reported in most parts of China; however, no such information is available in Tibetan chickens, China. In our study, six NDVs from Tibetan chickens were isolated on SPF embryonated chicken eggs in Tibet from 2013 to 2015, and the details of the NDV isolates are shown in Table 2. The identification and characterization was based on clinical symptoms, pathological observation and molecular investigations.
As shown in Table 2, HNH1 and HN17 strains were classified as velogenic NDV, which were consistent with the typical virulent motif 112 RRQKRF 117 at the F cleavage sites; while HN20, HN10-T, HN5-T and NHH3 were lentogenic with 112 GRQGRL 117 at the F cleavage sites. These amino acids changes in the protein may be related to change of pathogenicity because all six isolates presented high homology between themselves and against attenuated vaccine lasota, including HNH1 and HN17 isolates, which were classified as velogenic pathotype. Furthermore, the lentogenic NDV strain was reported in Tibet (Cao et al. 2001Cao D.J., Guo X., Liang R., Yan L.H., Liu P.X., Min P. & Chen J. 2001. Molecular epidemiology of newcastle disease virus isolates from part region of China. Chinese J. Prev. Vet. Med. 23:29-32. (In Chinese)), this was the first report to find the velogenic NDV. Altogether, these findings suggest that the homology between isolates and against attenuated vaccine strains may be due to possibly amino acids changes in the protein that resulted into the change in pathogenicity.
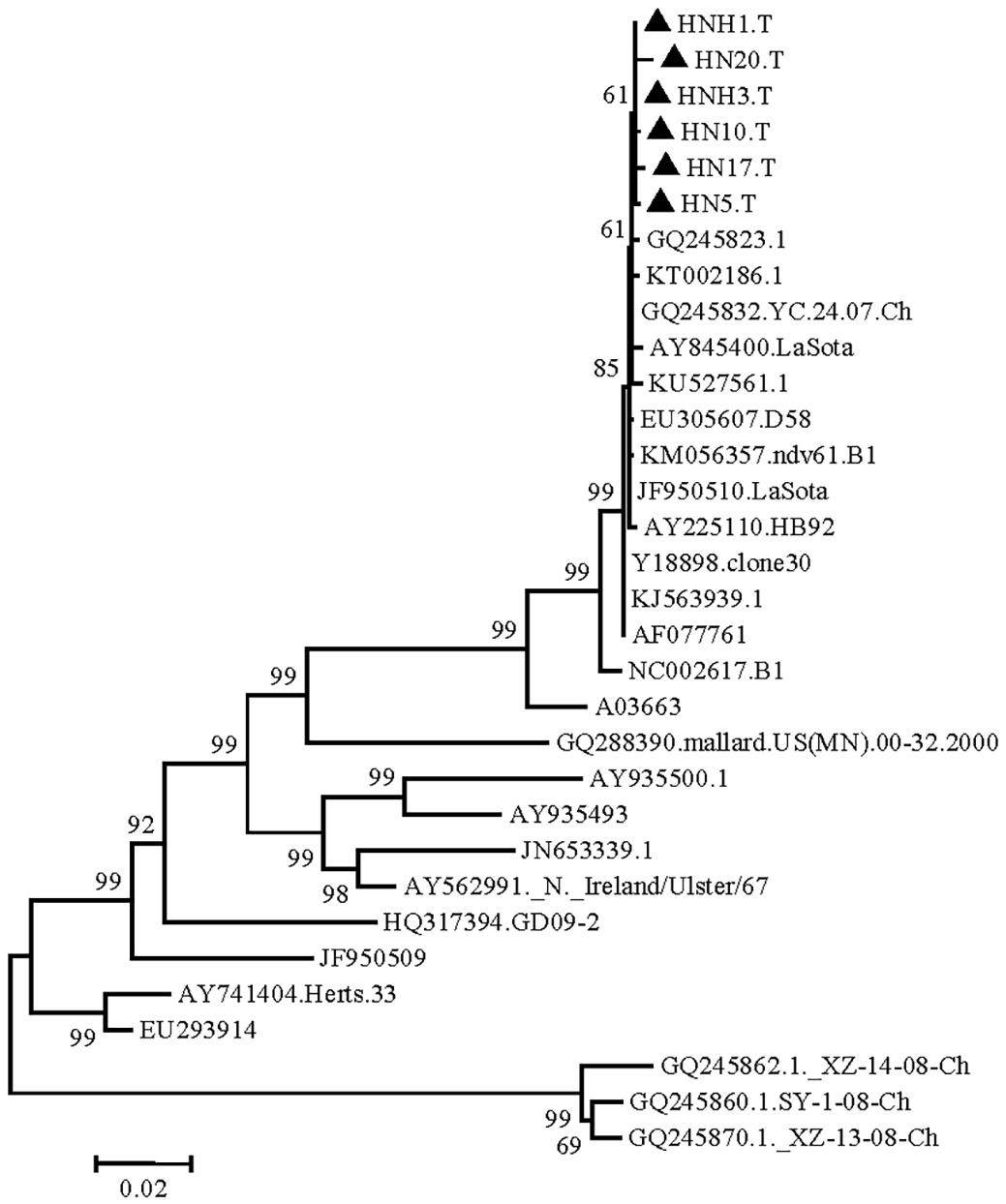
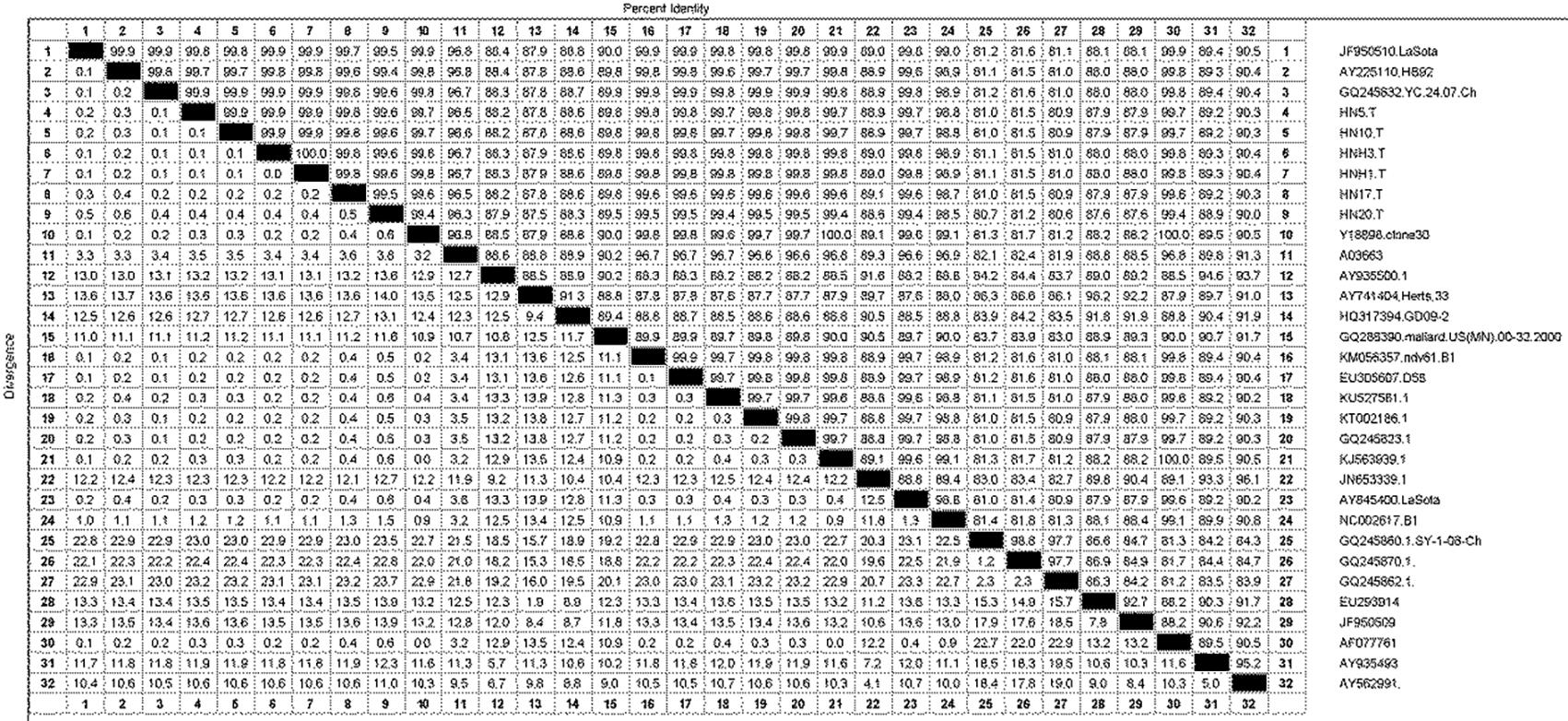
The isolated strains were phylogenetically compared to reference strains, based on the HN gene coding sequence (CDS). The phylogenetic analysis demonstrated that the strain had a genome size of 1713nt (Fig.1), and sharing highly similarity with Q245823, KT002186, KU527561, KJ563939, AY225110, EU305607, KM056357, Y18898, GQ245832, AF077761 and lasota strain isolated in India and China. Phylogeographic analysis indicated the ancestral state of isolate in Tibetan chickens located in India and China with high relationship (Fig.2). The nucleotide sequence of isolated strains shared 80.6%-100% sequence similarity with reference strains (Fig.3). The higher sequences (99.4%-99.9%) at the nucleotide level were identified with GQ245832, JF950510, Y18898 and AY225110 (GenBank accession number).
The RT-PCR amplification results of the NH gene of Newcastle disease virus isolated from Tibetan chickens (M = 2 kb Marker; Positive samples: Lane 2,3,6 = Negative: Lane 1,4,5).
Phylogenetic tree (Newcastle disease virus isolates from Tibetan chickens) constructed by the neighbor joining method using MEGA software.
Homology comparison of nucleotide sequence of Newcastle disease virus isolates with other strains (100%).
The prevalence of ND is determined by many factors, such as the geographical environment factors (altitude, mountains, etc.), migratory bird (density, activity, poisoned rate) and vaccine (wrong immunization approach; inadequate immunization dose; unreasonable vaccine dilutions; uneven dilutions, the interference of maternal antibody; poor quality vaccines). At present, lasota is widely and frequently used to prevent NDV infection in the poultry industry in China, and virulent NDV infections continue to occur in the well-vaccinated chicken flocks (Roy et al. 2004Roy P., Sundar N., Purushothaman V. & Balachandran C. 2004. Outbreak of Newcastle disease in a vaccinated flock. Indian J. Anim. Sci. 74(11):1116-1117., Qin et al. 2008Qin Z.M., Tan L.T., Xu H.Y., Ma B.C., Wang Y.L., Yuan X.Y. & Liu W.J. 2008. Pathotypical characterization and molecular epidemiology of Newcastle disease virus isolates from different hosts in China from 1996 to 2005. J. Clin. Microbiol. 46:601-611., Zhang et al. 2014a,Zhang Y.Y., Shao M.Y., Yu X.H., Zhao J. & Zhang G.Z. 2014a. Molecular characterization of chicken-derived genotype VIId Newcastle disease virus isolates in China during 2005-2012 reveals a new length in hemagglutinin-neuraminidase. Infect. Genet. Evol. 21:359-366. bZhang N.Z., Zhou D.H., Huang S.Y., Wang M., Shi X.C., Ciren D. & Zhu X.Q. 2014b. Seroprevalence and risk factors associated with Haemophilus parasuis infection in Tibetan pigs in Tibet. Acta Tropica 132:94-97.). Although this virus did not cause disease in chickens, the virus could effectively infect and replicate in these birds. Therefore, new vaccines to prevent the emerging infection in the chicken flocks should be developed as soon as possible, along with strict biosecurity measures to reduce the risk of Newcastle disease outbreaks. It suggests that irrational use of attenuated vaccines in Tibetan chickens may attribute to NDV infection (Palm et al. 2015Palm E.C., Newman S.H., Prosser D.J., Xiao X., Ze L., Batbayar N., Balachandran S. & Takekawa J.Y. 2015. Mapping migratory flyways in Asia using dynamic Brownian bridge movement models. Mov. Ecol. 3:1-10.). In addition, increase number of migratory birds to Tibet may be another possible explanation of ND in this region, as these birds play an important role in the transmission of NDV and act as main reservoirs of the virus.
In conclusion, this is the first report describing the occurrence and baseline genetic characteristics of NDV in high altitude region of China (Tibet). Our results revealed that Tibetan chickens carry NDVs of both velogenic and lentogenic types. Furthermore, genetic characterization showed clear evidence that irrational use of vaccines may be the key contributor to the occurrence of NDV in the study area. This report may help in the prevention and control strategies against NDV in Tibet, China.
Acknowledgments
This study was supported by Key laboratory of clinical veterinary medicine in Tibet and the National Natural Science Foundation of China (No. 31460682).
References
- Anonymous 2006. Newcastle disease, chapter 2.1.15. In: OIE Manual of Diagnostic Tests and Vaccines for Terrestrial Animals. World Organisation for Animal Health, Paris.
- Bogoyavlenskiy A., Berezin V., rilipov A., Usachev E., Lyapina O. & Korotetskiy I. 2009. Newcastle disease outbreaks in Kazakhstan and Kyrgyzstan during 1998, 2000, 2001, 2003, 2004, and 2005 were caused by viruses of the genotypes VIIb and VIId. Virus Genes 39:94-101.
- Cao D.J., Guo X., Liang R., Yan L.H., Liu P.X., Min P. & Chen J. 2001. Molecular epidemiology of newcastle disease virus isolates from part region of China. Chinese J. Prev. Vet. Med. 23:29-32. (In Chinese)
- Ge M., Zhang W., Shi G., Xiao C., Zhao X. & Zhang R. 2015. Astragalus polysaccharide perseveres cytomembrane capacity against Newcastle disease virus infection. Pakistan Vet. J. 35(3):382-384.
- Kang Y., Xiang B., Yuan R., Zhao X., Feng M., Gao P., Li Y., Li Y., Ning Z. & Ren T. 2016. Phylogenetic and Pathotypic Characterization of Newcastle Disease Viruses Circulating in South China and Transmission in Different Birds. Front. Microbiol. 7:119.
- Kim S.H., Xiao S., Shive H., Collins P.L. & Samal S.K. 2012. Replication, neurotropism, and pathogenicity of avian paramyxovirus serotypes 1-9 in chickens and ducks. PLoS One 7:34927.
- Kumar S. 2015. Newcastle disease virus outbreaks in India: time to revisit the vaccine type and strategies. Vaccine 33:3268-3269.
- Kumar U. & Kumar S. 2015. Molecular characterization of an apoptotic strain of Newcastle disease virus isolated from an outbreak in India. Cancer Gene Therapy 22:402-409.
- Mayo M.A. 2002a. A summary of taxonomic changes recently approved by ICTV. Archs Virology 147:1655-1663.
- Mayo M.A. 2002b. Virus taxonomy-Houston 2002. Archs Virol. 147:1071-1076.
- Munir T., Aslam A., Zahid B., Ahmed I., Imranand M.S. & Ijaz M. 2015. Potential of commonly resident wild birds towards newcastle disease virus transmission. Pakistan Vet. J. 35(1):106-107.
- Palm E.C., Newman S.H., Prosser D.J., Xiao X., Ze L., Batbayar N., Balachandran S. & Takekawa J.Y. 2015. Mapping migratory flyways in Asia using dynamic Brownian bridge movement models. Mov. Ecol. 3:1-10.
- Qin Z.M., Tan L.T., Xu H.Y., Ma B.C., Wang Y.L., Yuan X.Y. & Liu W.J. 2008. Pathotypical characterization and molecular epidemiology of Newcastle disease virus isolates from different hosts in China from 1996 to 2005. J. Clin. Microbiol. 46:601-611.
- Roy P., Sundar N., Purushothaman V. & Balachandran C. 2004. Outbreak of Newcastle disease in a vaccinated flock. Indian J. Anim. Sci. 74(11):1116-1117.
- Tamura K., Stecher G., Peterson D., Filipski A. & Kumar S. 2013. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 30:2725-2729.
- Zhang H., Wu C., Chamba Y., Ling Y. & Luo Z. 2006. Adaptability to high altitude and NOS activity of lung in Tibetan chicken. J. China Agric. Univ. 11:35-38.
- Zhang H., Wu C., Chamba Y., Ma X., Tang X. & Po B. 2005. Curve analysis of embryonic mortality in chickens incubated at high altitude. J. China Agric. Univ. 10:109-114.
- Zhang N.Z., Zhou D.H., Huang S.Y., Wang M., Shi X.C., Ciren D. & Zhu X.Q. 2014b. Seroprevalence and risk factors associated with Haemophilus parasuis infection in Tibetan pigs in Tibet. Acta Tropica 132:94-97.
- Zhang R., Pu J. & Su J.L. 2010. Phylogenetic characterization of Newcastle disease virus isolated in the mainland of China during 2001-2009. Vet. Microbio. 141:246-257.
- Zhang S., Wang X., Zhao C., Liu D., Hu Y., Zhao J. & Zhang G. 2011. Phylogenetic and pathotypical analysis of two virulent Newcastle disease viruses isolated from domestic ducks in China. PLoS One 6:e25000.
- Zhang Y.Y., Shao M.Y., Yu X.H., Zhao J. & Zhang G.Z. 2014a. Molecular characterization of chicken-derived genotype VIId Newcastle disease virus isolates in China during 2005-2012 reveals a new length in hemagglutinin-neuraminidase. Infect. Genet. Evol. 21:359-366.
Publication Dates
-
Publication in this collection
Jan 2018
History
-
Received
29 Sept 2016 -
Accepted
31 Mar 2017