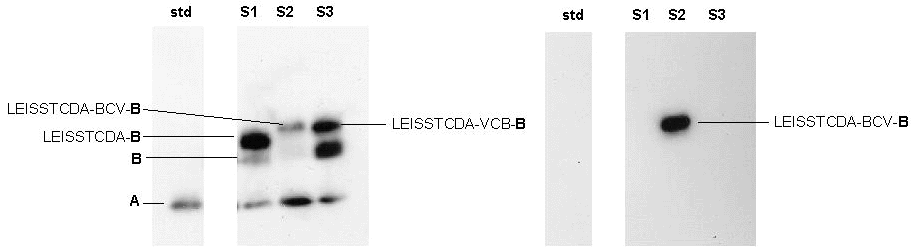Abstract
Lactic acid bacteria (LAB) are Gram-positive bacteria and are generally regarded as safe (GRAS) organisms. Therefore, LAB could be used for heterologous protein secretion and they are good potential candidates as antigen delivery vehicles. To develop such live vaccines, a better control of protein secretion is required. We developed an efficient secretion system in the model LAB, Lactococcus lactis. Staphylococcal nuclease (Nuc) was used as the reporter protein. We first observed that the quantity of secreted Nuc correlated with the copy number of the cloning vector. The nuc gene was cloned on a high-copy number cloning vector and no perturbation of the metabolism of the secreting strain was observed. Replacement of nuc native promoter by a strong lactococcal one led to a significant increase of nuc expression. Secretion efficiency (SE) of Nuc in L. lactis was low, i.e., only 60% of the synthesized Nuc was secreted. Insertion of a synthetic propeptide between the signal peptide and the mature moiety of Nuc increased the SE of Nuc. On the basis of these results, we developed a secretion system and we applied it to the construction of an L. lactis strain which secretes a bovine coronavirus (BCV) epitope-protein fusion (BCV-Nuc). BCV-Nuc was recognized by both anti-BCV and anti-Nuc antibodies. Secretion of this antigenic fusion is the first step towards the development of a novel antigen delivery system based on LAB-secreting strains.
protein secretion; staphylococcal nuclease; Lactococcus lactis; propeptide; bovine coronavirus; epitope
Braz J Med Biol Res, February 1999, Volume 32(2) 191-198
Heterologous protein secretion in Lactococcus lactis: a novel antigen delivery system
P. Langella and Y. Le Loir
Laboratoire de Génétique Appliquée, Unité de Recherches Laitières et de Génétique Appliquée, Institut National de la Recherche Agronomique, Domaine de Vilvert, France
 Text
Text
 References
References
Abstract
 Correspondence and FootnotesCorrespondence and FootnotesCorrespondence and Footnotes
Correspondence and FootnotesCorrespondence and FootnotesCorrespondence and Footnotes

Lactic acid bacteria (LAB) are Gram-positive bacteria and are generally regarded as safe (GRAS) organisms. Therefore, LAB could be used for heterologous protein secretion and they are good potential candidates as antigen delivery vehicles. To develop such live vaccines, a better control of protein secretion is required. We developed an efficient secretion system in the model LAB, Lactococcus lactis. Staphylococcal nuclease (Nuc) was used as the reporter protein. We first observed that the quantity of secreted Nuc correlated with the copy number of the cloning vector. The nuc gene was cloned on a high-copy number cloning vector and no perturbation of the metabolism of the secreting strain was observed. Replacement of nuc native promoter by a strong lactococcal one led to a significant increase of nuc expression. Secretion efficiency (SE) of Nuc in L. lactis was low, i.e., only 60% of the synthesized Nuc was secreted. Insertion of a synthetic propeptide between the signal peptide and the mature moiety of Nuc increased the SE of Nuc. On the basis of these results, we developed a secretion system and we applied it to the construction of an L. lactis strain which secretes a bovine coronavirus (BCV) epitope-protein fusion (BCV-Nuc). BCV-Nuc was recognized by both anti-BCV and anti-Nuc antibodies. Secretion of this antigenic fusion is the first step towards the development of a novel antigen delivery system based on LAB-secreting strains.
Key words: protein secretion, staphylococcal nuclease, Lactococcus lactis, propeptide, bovine coronavirus, epitope
Introduction
Lactic acid bacteria (LAB) are widely used in the food industry for the production and preservation of fermented products. Thus, they are considered as GRAS (generally regarded as safe) organisms. Since the 80's, many efforts have been made to better understand the molecular basis of the technological properties of LAB for a better control of industrial processes involving LAB.
Lactococcus lactis is the model LAB. Many genetic tools have been developed and its genome was recently sequenced in full (Bolotine S, personal communication). Several studies have investigated the potential of using LAB for new applications such as the production of heterologous proteins in fermenters, in fermented food products or in the digestive tract of humans or animals (for a review, see Ref. 1). In this area, our team is particularly interested in heterologous protein secretion and exportation in L. lactis (2-4). L. lactis is a good candidate for heterologous protein secretion since plasmidless strains do not secrete proteases. Secretion of a protein by this GRAS bacterium would allow production directly in a fermented food product and thus facilitate the interaction between the secreted protein (enzyme or antigen) and the environment (the food product itself or the digestive tract). To date, relatively few proteins are known to be secreted by L. lactis. Only one, Usp45, a protein of unknown function, is present in detectable quantities after Coomassie gel staining (5). Other secreted proteins are present only in trace amounts.
Until now, no detailed investigations of optimization of protein secretion in L. lactis have been reported. In a previous report (2), we investigated the secretion capacity of L. lactis using staphylococcal nuclease (Nuc) as the reporter protein. Nuc is a small and stable protein secreted by Staphylococcus aureus encoded by the nuc gene. It is genetically and biochemically well characterized (6). Its enzymatic activity can be easily detected on Petri dishes (7) as well as in PAGE gel zymograms (8). Translational fusions to the N-terminal end and to the C-terminal end of the mature protein do not abolish enzymatic activity (9). Although L. lactis does not secrete large amounts of proteins, we showed that it was able to secrete large quantities of heterologous proteins. On the basis of our previous work, we decided to develop tools for heterologous protein secretion allowing different levels of expression of the gene of interest and high secretion efficiency (SE) of the desired protein. Since LAB are food grade and ingested by humans and animals, they could be used to deliver antigens against diseases of digestive tropism and to induce mucosal immunity.
Here, we describe how we increased expression of the nuc gene by using a high-copy number cloning vector and a lactococcal promoter. We also improved the SE of Nuc in L. lactis by using a synthetic (secretion enhancer) propeptide. These results have been extended to construct one L. lactis strain able to secrete an epitope-protein fusion. The antigen chosen was an epitope of the bovine coronavirus (BCV) which provokes high mortality in young cattle.
Results
Secretion of Nuc is gene dose dependent in L. lactis
Heterologous expression could lead to intracellular accumulation, aggregation, or degradation of the protein in the cytoplasm (10). These phenomena can be deleterious to the cell physiology. To prevent a putative altering effect caused by high production of Nuc in L. lactis, we first cloned the nuc gene on pIL252 (11), a low-copy number plasmid, resulting in plasmid pNuc1. The growth rate of L. lactis (pNuc1) was similar to that of L. lactis (pIL252). Nuc activity was found in the supernatant and Nuc release was not due to cell lysis as no lactate dehydrogenase activity was found in the culture supernatant. These results demonstrate that L. lactis recognizes nuc expression and secretion signals and it is able to secrete Nuc.
We also cloned nuc on pGK, a medium-copy number plasmid, resulting in pNuc2, and on pVE3556, a high-copy number plasmid, resulting in pNuc3 (12). The growth rates of lactococcal strains carrying pNuc2 and pNuc3 were not altered. A Nuc activity test on Petri dishes was performed on the three constructed strains and showed that the halos around colonies of the strain carrying pNuc3 were significantly larger than those obtained with pNuc2 and pNuc1 (Figure 1). This was confirmed by Western blot analysis of supernatant fractions of overnight cultures of the corresponding strains (data not shown). A clear positive correlation between plasmid copy number and secretion of Nuc was observed: a 40-fold increase of Nuc secretion was measured with pNuc3 compared to pNuc1. This observation led us to choose pVE3556 as the backbone for our secretion vector.
- Halo size around secreting colonies is proportional to the number of nuc gene copies. Three Lactococcus lactis strains (MG1363(pNuc1), MG1363(pNuc2) and MG1363(pNuc3)) were plated onto BHI agar culture medium (Brain Heart Infusion, Difco, Grand Island, NY) containing erythromycin (5 µg/ml) for selection. After overnight incubation, Petri dishes were overlaid with TBD-agar (toluidine blue dye) detection medium as previously described (12). Halos appeared around secreting colonies after a 1- to 2-h incubation time at 37oC. Plasmid extractions were performed on several colonies for each of the three halo sizes (small, medium and large). Each halo size can be correlated to one plasmid DNA, i.e., pNuc1 for small halos, pNuc2 for medium halos and pNuc3 for large halos.
The secretion efficiency of Nuc is low in L. lactis
To develop our secretion vector, we considered two parameters: i) the quantity of secreted Nuc expressed as milligrams per liter (mg/l) of culture supernatant, and ii) SE which is the ratio of secreted Nuc forms/secreted and intracellular Nuc forms. To evaluate SE, Western blot experiments and immunorevelation of Nuc were performed on cellular and supernatant fractions of overnight cultures. Three Nuc forms were detected in L. lactis (Figure 2). A precursor (preNuc) is found in the cell fraction and migrates at the expected size (25.1 kDa (13)) and two mature forms are found in the supernatant: NucB (18.8 kDa) and NucA (16.5 kDa). In S. aureus, NucB is transiently associated with the cell envelope and undergoes the cleavage of a 19-amino acid residue (aa) propeptide. The resulting form, NucA, is the major form and is released in the supernatant (14). In L. lactis, both Nuc mature forms are found active in the supernatant and the secondary proteolytic cleavage is weak, probably because of the scarcity of extracellular proteases. We determined by microsequencing that NucA results in the cleavage of a 21-aa propeptide instead of a 19-aa propeptide in S. aureus (2).
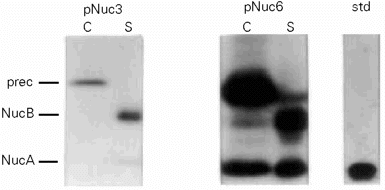
- Replacement of the native nuc promoter by a strong lactococcal one dramatically increases Nuc production in L. lactis. Nuc secretion was analyzed in L. lactis MG1363(pNuc3) and MG1363(pNuc6) cultures. Protein samples were prepared on cell and supernatant fractions of overnight cultures. After SDS-PAGE, samples were analyzed by Western blot and by immunorevelation with anti-Nuc antibodies. prec: precursor; B: NucB; A: NucA; std: standard commercial NucA; C: cell fraction; S: supernatant fraction.
Zymogram experiments were performed on SDS-PAGE renatured gels: both NucB and A were active in lactococcal culture supernatants, whereas preNuc had no detectable nuclease activity under zymogram conditions (data not shown).
L. lactis (pNuc3) secreted 0.5 mg of Nuc/l with an SE of 70% (Figure 2) without alteration of its growth (1). The remaining 30% were detected as preNuc in the cell fraction. Even with a plasmid of lower copy number (pNuc2), preNuc was found in comparable proportions in the cell fraction (data not shown). These results suggest that preNuc accumulation was not due to the saturation of the lactococcal secretion machinery.
The use of a strong lactococcal promoter enhances nuc expression in L. lactis
To enhance nuc expression in L. lactis, we replaced the native nuc promoter with P59, a strong lactococcal promoter (15). To combine high copy number and strong promoter effects, the P59nuc cassette was cloned in L. lactis on pVE3556, resulting in pNuc6. Analysis by Western blot experiments of L. lactis (pNuc6) supernatants revealed a 10-fold increase in Nuc secretion compared to L. lactis (pNuc3) (5 mg/l; Figure 2). Northern analysis confirmed that this result was due to a better transcription (data not shown). The SE was estimated at 60% and remained low and comparable to that obtained with lower nuc expressions (Figure 2).
Using Usp45 as a control for homologous protein secretion, we looked for a possible perturbation of the lactococcal secretion machinery due to the high level of Nuc secretion and to the accumulation of preNuc. No significant difference was observed for growth and Usp45 secretion between L. lactis (pVE3556) and L. lactis (pNuc6). These results suggest i) that high level of Nuc secretion did not alter the growth of L. lactis, and ii) that the low SE of Nuc in L. lactis was due to intrinsic features of preNuc (for instance, Nuc signal peptide (SPNuc) poorly recognized) rather than to the saturation of the lactococcal secretion machinery.
Deletion of the Nuc propeptide leads to a dramatic decrease of Nuc SE in L. lactis
Some secreted bacterial enzymes are synthesized as preproprotein containing an SP and a propeptide which, in some cases, may act as an intramolecular chaperone or as a translocation enhancer (16). In L. lactis, both NucB and A are found active in the supernatant. Thus, Nuc propeptide does not seem to play a role either in mature Nuc activity or in Nuc localization. To determine the role of Nuc propeptide, we deleted 17 of the 21-aa propeptide cleaved in L. lactis. Secretion of the resulting NucT mature form was analyzed by Western blotting and compared to that of NucB (Figure 3A). We observed a dramatic decrease of SE from 60% for NucB to 30% for NucT. A similar observation was made by Suciu and Inouye (17) in E. coli. Statistical analysis revealed that, in E. coli, exported proteins have a neutral or negative global charge on the first ten aa of the mature part (18). Furthermore, the insertion of positively charged aa into the N-terminus of the mature part drastically impairs maturation (19). The net charge of the first ten aa of NucB and NucT is 0 and +3, respectively (Table 1). This difference could explain the decrease of NucT SE.
- A, Deletion of the Nuc propeptide leads to a lower efficiency of Nuc secretion in L. lactis. Nuc secretion was analyzed for L. lactis MG1363(pNuc6) and MG1363(pNuc9) cultures. Protein samples were prepared in cell and supernatant fractions of overnight cultures. After SDS-PAGE, samples were analyzed by Western blot and by immunorevelation with anti-Nuc antibodies. Secretion efficiencies were estimated after scanning the autoradiograms. prec: precursor; B: NucB; A: NucA; T: NucT; C: cell fraction; S: supernatant fraction; SE: secretion efficiency. B, Insertion of a LEISSTCDA synthetic propeptide enhances NucB and NucT secretion. Nuc secretion was analyzed in L. lactis MG1363(pNuc7) and MG1363(pNuc10) cultures. LEISSTCDA-B: LEISSTCDA-NucB fusion; LEISSTCDA-T: LEISSTCDA-NucT.
Insertion of a synthetic propeptide leads to high Nuc SE in L. lactis
The low SE for NucT (30%) was probably due to the presence of positively charged aa close to the SP cleavage site (20). This result led us to test the effect of negatively charged aa inserted downstream of the SP cleavage site. For this purpose, we cloned a synthetic oligonucleotide between Nuc SP and NucB- or NucT-mature proteins. This oligonucleotide was described in a previous work and was originally designed to introduce restriction sites (multicloning site, MCS) just downstream of the SPNuc (12). This MCS was designed in order to keep a negative net charge in the first ten aa of the mature part. It encodes a 9-residue oligopeptide with the following sequence: Leu-Glu-Ile-Ser-Ser-Thr-Cys-Asp-Ala (LEISSTCDA, hereafter called LEISS). The net charges of the first ten residues of Nuc and LEISS-Nuc are 0 and -2, respectively (Table 2). Two major differences were observed in the secretion of Nuc and LEISS-Nuc: i) the proportion of the unprocessed preLEISS-Nuc form was estimated to be about 20% compared to 40% for preNuc; ii) the concentration of mature LEISS-Nuc in the supernatant was about 2- to 5-fold higher (10 to 25 mg/l) than that of Nuc (5 mg/l). Whole cell lysates and Northern analysis of these two strains showed that the plasmid copy number and the quantity of messengers were equivalent for both the pNuc6 and pNuc7 constructions (data not shown). Therefore, this increase cannot be explained at the DNA level or at the RNA level. In a previous report, we showed by pulse-chase experiments that preLEISS-Nuc is more rapidly processed than preNuc (2). The precise mechanism of this LEISS effect has not yet been determined but the negative net charge of the first 10 aa in LEISS-Nuc could explain this positive effect (Table 2).
L. lactis can secrete an epitope-protein fusion
One of our long-term goals is the development of live vaccines based on LAB as delivery vehicles. One example of veterinary applications is the development of a live vaccine against BCV. BCV leads to diarrhea in young calves and is one of the major causes of mortality and morbidity in cattle breeding. Several BCV epitopes have been identified and are available from the INRA colleague team (21).
To validate our secretion system, the pNuc7 construction was used to fuse a 9-residue BCV epitope between the LEISS propeptide and the Nuc mature part. The goal was to use LEISS-Nuc as a carrier protein to drive the secretion of this BCV-epitope. With our Nuc system, this epitope-protein fusion was expected to be detected by its activity and its antigenic properties. We designed a 39-mer oligonucleotide encoding a 13-residue oligopeptide with the following sequence: Tyr-Arg-Phe-Thr-Asn-Phe-Glu-Pro-Phe-Arg-Ile-Pro-Ala (called BCV). The underlined aa correspond to the BCV epitope and the last four aa correspond to two restriction sites used to detect recombinant clones and to fuse the oligonucleotide between the LEISS propeptide and the Nuc mature part. This oligonucleotide was cloned in two orientations: the antigenic one, BCV (resulting in pNuc12) and the non-antigenic one, VCB (resulting in pNuc11). The culture supernatant of L. lactis (pNuc12) was analyzed by Western blotting experiments and compared to that of L. lactis (pNuc7) and L. lactis (pNuc11), both taken as negative controls (Figure 4).
As expected, the negative control VCB-Nuc protein fusion was non-antigenic, i.e., was not recognized by anti-BCV antibodies. Both BCV-Nuc and VCB-Nuc proteins were secreted by L. lactis and have conserved their nuclease activity. Only BCV-Nuc was recognized by both anti-BCV and anti-Nuc antibodies. Secretion of this antigenic fusion protein is the first step towards the development of "oral live vaccines" based on LAB strains and on our secretion system.
- Secretion of an epitope-protein fusion recognized by both anti-Nuc and anti-BCV antibodies. Culture supernatants were prepared on overnight cultures of L. lactis strains MG1363(pNuc7), MG1363(pNuc12) and MG1363(pNuc11) and immunorevelations were performed with anti-Nuc antibodies (left panel) and anti-BCV antibodies (right panel). S1: MG1363(pNuc7) supernatant; S2: MG1363(pNuc12); S3: MG1363(pNuc11); std: standard commercial NucA.
Concluding remarks
In this work we identified parameters that affect Nuc secretion efficiency and we showed that L. lactis can secrete an antigenic protein such as the BCV-Nuc fusion. Our results demonstrate that Nuc is a good reporter to optimize heterologous protein secretion in L. lactis. In our laboratory, J.C. Piard recently developed an elegant system, using Nuc as a reporter protein, to target a given protein into three cellular localizations (intracellular, cell-wall anchored or secreted) (9; Piard JC, personal communication). This work is of great interest to evaluate the effect of the antigen localization on the immune response development. Nuc was also used to identify new lactococcal exported proteins (4). Furthermore, we currently use the Nuc phenotype to screen for secretion mutants in L. lactis.
The development of a live vaccine requires three preliminary steps: i) construction of the recombinant strain and in vitro control of the antigenicity of the protein produced, ii) oral or nasal administration of the recombinant strain to animals (murine model) and measurement of its immunogenicity (i.e., its capacity to induce an immune response), and iii) challenge against the pathogen to check whether the immune response is protective. The present study shows that L. lactis is a good candidate for the development of a live vaccine based on LAB as delivery vehicles. We showed here that the bacterium can secrete an antigenic recombinant protein. Although there is no murine model to perform a challenge against BCV, this result is encouraging. Indeed, L. lactis has been successfully tested as a vaccine vector (22) and recent studies have shown that its metabolism is still active during transit through the intestinal tract (23). The latter result shows that recombinant strains are likely to express the same features in vitro (i.e., in an assay tube) and in vivo (i.e., in the intestinal tract).
Plasmid-based constructions often carry antibiotic resistance markers for selection. The use of these markers is a major limitation to the application of genetically modified organisms. To avoid this, antigenic constructions can be integrated into the bacterial chromosome. This stabilizes the new genetic information in the recombinant strain and thus does not require any antibiotic resistance marker.
Further development of vaccinal lactococcal strains will be carried out with antigens originating from pathogenic organisms such as Brucella abortus. This bacterium continues to be the cause of a major zoonosis worldwide. Until now no satisfactory vaccines against this disease are available and new vaccinal strategies (such as oral live vaccines) should be explored. Recent work demonstrated a protective immunization against Brucella abortus in mice using a ribosomal protein (24). We think that a lactococcal strain secreting this ribosomal protein (or a part of it) would be a promising vector to induce a protective immune response against brucellosis.

We thank Jean-François Vautherot and Denis Rasschaert (INRA, Tours) for their generous gifts of anti-bovine coronavirus antiserum. We thank Jacqueline Commissaire, Aimé Jacquet, and Laurent Brétigny (INRA, Jouy en Josas) for technical assistance, and Alexandra Gruss (INRA, Jouy en Josas) for her support during this work. We are very grateful to Emmanuelle Maguin and Pascale Serror (INRA, Jouy en Josas) for constant discussion during the course of this work and to Jean-Christophe Piard (INRA, Jouy en Josas) for permitting us to cite his unpublished data.
Address for correspondence: P. Langella, Laboratoire de Génétique Appliquée, Unité de Recherches Laitières et de Génétique Appliquée, Institut National de la Recherche Agronomique, Domaine de Vilvert, 78352 Jouy en Josas Cedex, France. Fax: +33-1-3465-2065. E-mail: langella@biotec.jouy.inra.fr
Presented at the International Symposium "The Third Revolution on Vaccines: DNA Vaccines", Belo Horizonte, MG, Brasil, November 3-7, 1997. Received November 24, 1998. Accepted December 9, 1998.
Acknowledgments
- 1. de Vos WM & Simons GFM (1994). Gene cloning and expression in Lactococci. In: Gasson MJ & de Vos WM (Editors), Genetics and Biotechnology of Lactic Acid Bacteria Chapman and Hall, London, 52-105.
- 2. Le Loir Y, Gruss A, Ehrlich SD & Langella P (1998). A nine-residue synthetic propeptide enhances secretion efficiency of heterologous proteins in Lactococcus lactis Journal of Bacteriology, 180: 1895-1903.
- 3. Piard JC, Hautefort I, Fischetti VA, Ehrlich SD, Fons M & Gruss A (1997). Cell-wall anchoring of the Streptococcus pyogenes M6 protein in various lactic acid bacteria. Journal of Bacteriology, 179: 3068-3072.
- 4. Poquet I, Ehrlich SD & Gruss A (1997). An export-specific reporter designed for gram-positive bacteria: application to Lactococcus lactis Journal of Bacteriology, 180: 1904-1912.
- 5. van Asseldonk M, Rutten G, Oteman M, Siezen RJ, de Vos WM & Simons G (1990). Cloning, expression in Escherichia coli and characterization of usp45, a gene encoding a highly secreted protein from Lactococcus lactis MG1363. Gene, 95: 155-160.
- 6. Shortle D (1983). A genetic system analysis of staphylococcal nuclease. Gene, 22: 181-189.
- 7. Lachica RVF, Genigeorgis C & Hoeprich PD (1971). Metachromatic agar-diffusion methods for detecting staphylococcal nuclease activity. Applied Microbiology, 21: 585-587.
- 8. Liebl W, Sinskey AJ & Schleifer KH (1992). Expression, secretion, and processing of staphylococcal nuclease by Corynebacterium glutamicum Journal of Bacteriology, 174: 1854-1861.
- 9. Piard JC, Jimenez-Diaz R, Ehrlich SD, Fischetti VA & Gruss A (1997). The M6 protein of Streptococcus pyogenes and its potential as a tool to anchor biologically active molecules at the surface of lactic acid bacteria. In: Horaud T (Editor), Streptococci and the Host Plenum Press, New York, 545-550.
- 10. Makrides SC (1996). Strategies for achieving high-level expression of genes in Escherichia coli Microbiological Reviews, 60: 512-538.
- 11. Simon D & Chopin A (1988). Construction of a vector plasmid family and its use for molecular cloning in Streptococcus lactis. Biochimie, 70: 559-567.
- 12. Le Loir Y, Gruss A, Ehrlich SD & Langella P (1994). Direct screening of recombinants in Gram-positive bacteria using the secreted staphylococcal nuclease as a reporter. Journal of Bacteriology, 176: 5135-5139.
- 13. Miller JR, Kovacevic S & Veal LE (1987). Secretion and processing of staphylococcal nuclease by Bacillus subtilis Journal of Bacteriology, 169: 3508-3514.
- 14. Davis A, Moore IB, Parker DS & Taniuchi H (1977). Nuclease B: a possible precursor of nuclease A, an extracellular nuclease of Staphylococcus aureus Journal of Biological Chemistry, 252: 6544-6553.
- 15. van der Vossen JMBM, van der Lelie D & Venema G (1987). Isolation and characterization of Streptococcus cremoris Wg-2 specific promoters. Applied and Environmental Microbiology, 53: 2452-2457.
- 16. Simonen M & Palva I (1993). Protein secretion in Bacillus species. Microbiological Reviews, 57: 109-137.
- 17. Suciu D & Inouye M (1996). The 19-residue propeptide of staphylococcal nuclease has a profound secretion-enhancing ability in Escherichia coli Molecular Microbiology, 21: 181-195.
- 18. von Heijne G (1986). Net N-C charge imbalance may be important for signal sequence function in bacteria. Journal of Molecular Biology, 192: 287-290.
- 19. Li PJ, Beckwith J & Inouye H (1988). Alteration of the amino terminus of the mature sequence of a periplasmic protein can severely affect protein export in Escherichia coli Proceedings of the National Academy of Sciences, USA, 85: 7685-7689.
- 20. von Heijne G (1990). The signal peptide. Journal of Membrane Biology, 115: 195-201.
- 21. Vautherot JF, Laporte J & Boireau P (1992). Bovine coronavirus spike glycoprotein: localization of an immunodominant region at the amino-terminal end of S2. Journal of General Virology, 73: 3289-3294.
- 22. Robinson K, Chamberlain LM, Schofield KM, Wells JM & Le Page RW (1997). Oral vaccination of mice against tetanus using recombinant Lactococcus lactis Nature Biotechnology, 15: 653-657.
- 23. Corthier G, Delorme C, Ehrlich SD & Renault P (1998). Use of luciferase genes as biosensors to study bacterial physiology in the digestive tract. Applied and Environmental Microbiology, 64: 2721-2725.
- 24. Oliveira SC & Splitter GA (1996). Immunization of mice with recombinant L7/L12 ribosomal protein confers protection against Brucella abortus infection. Vaccine, 14: 959-962.
Correspondence and Footnotes
Publication Dates
-
Publication in this collection
04 Feb 1999 -
Date of issue
Feb 1999
History
-
Accepted
09 Dec 1998 -
Received
24 Nov 1998