Abstract
Molecular oxygen (O2) is the premier biological electron acceptor that serves vital roles in fundamental cellular functions. However, with the beneficial properties of O2 comes the inadvertent formation of reactive oxygen species (ROS) such as superoxide (O2<FONT FACE=Symbol>·-</FONT>), hydrogen peroxide, and hydroxyl radical (OH<FONT FACE=Symbol>·</FONT>). If unabated, ROS pose a serious threat to or cause the death of aerobic cells. To minimize the damaging effects of ROS, aerobic organisms evolved non-enzymatic and enzymatic antioxidant defenses. The latter include catalases, peroxidases, superoxide dismutases, and glutathione S-transferases (GST). Cellular ROS-sensing mechanisms are not well understood, but a number of transcription factors that regulate the expression of antioxidant genes are well characterized in prokaryotes and in yeast. In higher eukaryotes, oxidative stress responses are more complex and modulated by several regulators. In mammalian systems, two classes of transcription factors, nuclear factor kB and activator protein-1, are involved in the oxidative stress response. Antioxidant-specific gene induction, involved in xenobiotic metabolism, is mediated by the "antioxidant responsive element" (ARE) commonly found in the promoter region of such genes. ARE is present in mammalian GST, metallothioneine-I and MnSod genes, but has not been found in plant Gst genes. However, ARE is present in the promoter region of the three maize catalase (Cat) genes. In plants, ROS have been implicated in the damaging effects of various environmental stress conditions. Many plant defense genes are activated in response to these conditions, including the three maize Cat and some of the superoxide dismutase (Sod) genes.
Catalase; Aging; Telomeres; Gene regulation; Superoxide dismutase; Genomics
Braz J Med Biol Res, July 2005, Volume 38(7) 995-1014 (Review)
Oxidative stress: molecular perception and transduction of signals triggering antioxidant gene defenses
Department of Genetics, North Carolina State University, Raleigh, NC, USA
 Text
Text
 References
References
 Correspondence and Footnotes
Correspondence and Footnotes
Correspondence and Footnotes
Correspondence and Footnotes
Correspondence and Footnotes
Correspondence and Footnotes
Abstract
Molecular oxygen (O2) is the premier biological electron acceptor that serves vital roles in fundamental cellular functions. However, with the beneficial properties of O2 comes the inadvertent formation of reactive oxygen species (ROS) such as superoxide (O2·-), hydrogen peroxide, and hydroxyl radical (OH·). If unabated, ROS pose a serious threat to or cause the death of aerobic cells. To minimize the damaging effects of ROS, aerobic organisms evolved non-enzymatic and enzymatic antioxidant defenses. The latter include catalases, peroxidases, superoxide dismutases, and glutathione S-transferases (GST). Cellular ROS-sensing mechanisms are not well understood, but a number of transcription factors that regulate the expression of antioxidant genes are well characterized in prokaryotes and in yeast. In higher eukaryotes, oxidative stress responses are more complex and modulated by several regulators. In mammalian systems, two classes of transcription factors, nuclear factor kB and activator protein-1, are involved in the oxidative stress response. Antioxidant-specific gene induction, involved in xenobiotic metabolism, is mediated by the "antioxidant responsive element" (ARE) commonly found in the promoter region of such genes. ARE is present in mammalian GST, metallothioneine-I and MnSod genes, but has not been found in plant Gst genes. However, ARE is present in the promoter region of the three maize catalase (Cat) genes. In plants, ROS have been implicated in the damaging effects of various environmental stress conditions. Many plant defense genes are activated in response to these conditions, including the three maize Cat and some of the superoxide dismutase (Sod) genes.
Key words: Catalase, Aging, Telomeres, Gene regulation, Superoxide dismutase, Genomics
Introduction
The environment in which they exist affects all living organisms. Whether internal or external to the organism, the environment is continually changing, and the organism must adapt if it is to survive. An organism apparently well adapted to its environment at any one time may be poorly adapted only a short time later if it cannot modify its physiology or behavior in response to changing environmental or metabolic conditions. Organisms that can adjust to changes in the environment are likely to exhibit a greater degree of adaptiveness than those that cannot. Environmental changes, irrespective of source, cause a variety of "stresses" or "shocks" that a cell must face repeatedly, and to which its genome must respond in a programmed manner for the organism to survive. How the genome perceives and transduces environmental signals to effect the expression and/or repression of pertinent genes in a selective manner remains a key question. Examples are responses to light, oxidative stress, pathogenicity, wounding, anaerobiosis, thermal shock, and the "SOS" response in microorganisms (1). Some sensing mechanism(s) must be present to alert the cell to imminent danger, and to trigger the orderly sequence of events that will mitigate this danger. In addition, there are genomic responses to unanticipated, unprogrammed challenges for which the genome is unprepared, but to which it responds in discernible though initially unforeseen and unpredictable ways (2). Many, though not all, signals are perceived at the cell surface by plasma membrane receptors. Activation of such receptors by mechanisms such as ligand binding may lead to alterations in other cellular components, ultimately resulting in alterations in cell shape, ion conductivity, gene activity, and other cellular functions (3). Identification and isolation of mutants that are unable to respond, or that respond abnormally to a particular signal, may provide ways to decipher the mechanisms by which a particular signal is transduced into a given response. Long before humans began manipulating and altering their environment, organisms from the simplest to the most complex began evolving methods to cope with stressful stimuli. Consequently, most living cells possess an amazing capacity to cope with a wide diversity of environmental challenges, including natural and synthetic toxins, pathogens, extreme temperatures, high metal levels, and radiation. Many studies in the past have demonstrated clear "cause-effect" relationships upon exposure of a given organism or cell to a particular environmental factor or stressor. But only recently have certain environmental insults been shown to elicit specific genomic responses (4). At present, relatively little is known of the underlying molecular mechanisms by which the genome perceives environmental signals and mobilizes the organism to respond. Such information is not only interesting in and of itself, but is also essential in any future attempts to engineer organisms for increased tolerance to environmental adversity. Recent dramatic advances in molecular biology and genomics have made it possible to investigate the underlying mechanisms utilized by organisms to cope with environmental stresses. Investigations of genomic responses to challenge are shedding light on unique DNA sequences capable of perceiving stress signals, thus allowing the cell or organism to mobilize its defenses (5). The general picture emerging from recent studies involves the sensing of a signal and the transduction of the signal to the transcription apparatus to catalyze transcription initiation. A signal transduction pathway contains elements that enable a signal to be transmitted within and between cells and to be translated into an appropriate response. Cells can respond to a variety of environmental, physical, and chemical stimuli using a diverse range of transduction and response mechanisms. The essential features of a signal transduction pathway comprise a receptor (recognition element) capable of detecting a stimulus, second messengers (transmission elements) such as calcium or phosphorylation cascades, and response elements (e.g., gene transcription). Such signaling networks are now amenable to study and dissection by biochemical and genetic approaches that may elucidate the underlying mechanisms and lead to the identification of the molecules responsible. A thorough understanding of how organisms perceive, respond, and adapt to changing environmental and developmental stimuli is now attainable.
Gene responses
Terminally differentiated cells express an array of genes required for their stable functioning and precise metabolic roles. A genome can respond in a rapid and specific manner by selectively decreasing or increasing the expression of specific genes. Genes whose expression is increased during times of stress presumably are critical to the organism's survival under adverse conditions. Examination and study of such "stress-responsive" genes have implications for human health and well being, for agricultural productivity, and for furthering basic biological knowledge. In addition to aiding the organism under stress, genomes that are modified by stress can be utilized to study the molecular events that occur during periods of increased or decreased gene expression. The mechanisms by which an organism recognizes a signal to alter gene expression and responds to fill that need are important physiologically and render possible the examination of gene regulation under various environmental regimens. The mechanisms of induction of stress response genes are similar among various organisms examined. Similarities in stress-induced changes in gene expression have been observed for a variety of stressors (3,6-8). Some that have been studied in detail include radiation, thermal shock, pathogenic infections, anaerobiosis, trauma, photostress, physical wounding, oxidative stress, water stress, and heavy metals. In all cases, specific changes in transcript and/or protein expression have been observed in various organisms subjected to such challenges.
Recent studies by several laboratories, using a variety of organisms, indicate that oxidative stress is a common denominator underlying many diseases and environmental insults which can lead to cell death in virtually all aerobes (9). It is also becoming clear that a variety of different biotic and abiotic stresses cause their deleterious effects, directly or indirectly, via reactive oxygen species (ROS) generation.
For example, numerous toxic environmental chemicals such as xenobiotics, pesticides, herbicides, fungicides, ozone, cigarette smoke, and radiation cause their harmful environmental effects via generation of free radicals and other ROS.
Herein, I will focus on oxidative stress, its causes and consequences, and mechanisms employed by organisms to cope with it.
Oxygenation of the Earth
Earth is the only planet in our solar system known to contain molecular oxygen (O2) in its atmosphere and to support aerobic life. However, when Earth was formed about 4.5 billion years ago, its atmosphere was unlike the present, being primarily reducing and essentially free of oxygen. Most likely, the earliest living organisms were anaerobic heterotrophs living in the primitive ocean depths, shielded from the damaging effects of solar ionizing radiation. The earliest relatively low levels of oxygen were probably the result of photolytic dissociation of water by the sun's ionizing radiation. The bulk of Earth's present oxygen concentration (21% O2) is derived from the photosynthetic activities of cyanobacteria and plants. It has been estimated that Earth contains about 410 x 103 Erda moles (Emol = 108 mol) of oxygen, and of this, 38.4 x 103 Emol is in the hydrosphere as water. Molecular oxygen is present in the atmosphere (37 Emol) and in the hydrosphere (0.4 Emol) and undergoes continuous turnover, with the total oxygen exchange estimated at ~15 x 103 Emol/106 years. Aerobic life is responsible for the major portion of oxygen turnover, with photosynthesis being the main input into the oxygen reservoir, and respiration the main output. The two processes are in approximate equilibrium, and fossil fuel combustion is the major source of oxygen loss from the reservoir (10,11).
The accumulation of dioxygen (O2) in Earth's atmosphere permitted the evolution of the enormous variety of aerobic organisms that use O2 as the electron acceptor, thus providing a higher yield of energy compared with fermentation and anaerobic respiration.
O 2, ROS, and oxygen toxicity
In its ground state (its normal configuration, O2) molecular oxygen is relatively unreactive. However, during normal metabolic activity, and as a consequence of various environmental perturbations (e.g., extreme temperatures, radiation, xenobiotics, toxins, air pollutants, various biotic and abiotic stresses, and diseases) O2 is capable of giving rise to frightfully reactive excited states such as free radicals and derivatives (9,12).
The oxidation powers of O2 are restricted because electrons can only be absorbed from another species whose electron spin is antiparallel to the two unpaired, parallel-spin electrons in diatomic oxygen. This spin restriction renders ground state molecular oxygen sufficiently unreactive, so that it cannot abstract electrons from other species. However, removal of the spin restriction by adding a single e-, or upon transfer of energy to oxygen from a photosensitizer (e.g., chlorophyll, flavin-containing compounds), increases the reactivity of oxygen. Photosensitizers can harvest light and energize O2 to form singlet oxygen (1DgO2), which can interact directly with another molecule, transferring the additional energy to the target molecule.
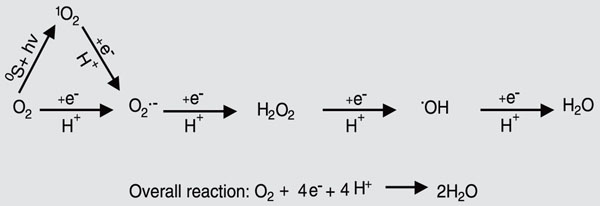
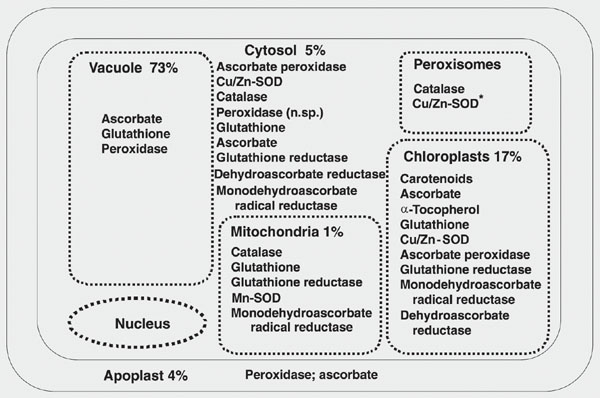
The complete reduction of O2 to water requires four electrons. O2 has a preference for a stepwise univalent pathway of reduction resulting in partially reduced intermediates (Figure 1). The reactive species of reduced dioxygen include the superoxide radical (O2·-), hydrogen peroxide (H2O2), and the hydroxyl radical (OH·). The latter can also be generated by the interaction of O2·-and H2O2 in the presence of metal ions. Both O2·- and OH· are extremely reactive and can cause molecular damage, leading to cell death. The hydroxyl radical reacts with virtually anything, inflicting indiscriminate and extensive intracellular damage. The O2·- is the conjugate base of a weak acid, the perhydroxyl radical (HO2), whose pKa is 4.69 ± 0.08. Thus, under acidic conditions, the very reactive perhydroxyl radical may predominate following a one electron reduction of dioxygen, while at higher pH values the O2·- is predominant. These and the physically energized form of dioxygen, singlet oxygen (1O2), are the biologically most important ROS (Table 1). An activation energy of ~22 kcal/mol is required to raise molecular oxygen (O2) from its ground state to its first singlet state. In higher plants, this energy is readily obtained from light quanta via such transfer molecules as chlorophyll. Unless abated, all of these intermediate oxygen species are extremely reactive and cytotoxic in all organisms (9,13). ROS can interact with proteins, lipids, and nucleic acids to cause severe molecular damage (Table 2). Thus, oxygen provides a paradox, in that it is essential for aerobic life, yet in its reduced forms is one of the most toxic substances with which life on Earth must cope. ROS are found in virtually all intracellular organelles or compartments as a consequence of normal metabolic activity. Each organelle or compartment has potential targets for oxidative damage, as well as mechanisms for the elimination of excess ROS accumulation (Figure 2).
Pathways in the univalent reduction of oxygen to water leading to generation of various intermediate reactive oxygen species (ROS).
Defenses against reactive oxygen
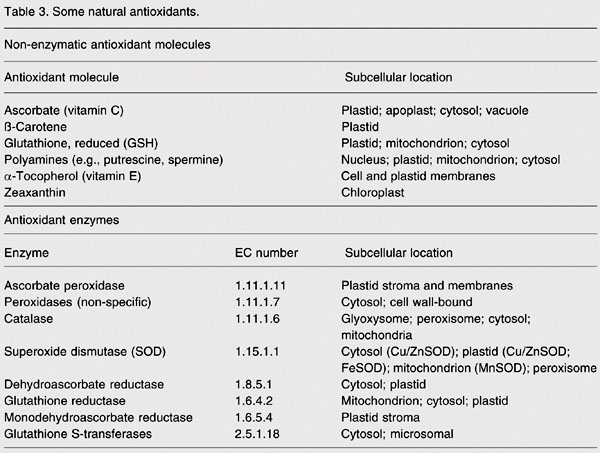
To minimize the damaging effects of ROS, aerobic organisms evolved both non-enzymatic and enzymatic antioxidant defenses (Table 3). Non-enzymatic defenses include compounds of intrinsic antioxidant properties, such as vitamins C and E, glutathione, and ß-carotene. Purely enzymatic defenses, such as superoxide dismutases (SOD), catalases (CAT) and peroxidases, protect by directly scavenging superoxide radicals and hydrogen peroxide, converting them to less reactive species. SODs catalyze the dismutation of O2·- to H2O2, and CAT and peroxidases reduce H2O2 to 2H2O. The similarity between the SOD and CAT reactions is that each is an oxidation-reduction in which the substrate, O2·- for SOD and H2O2 for CAT, is both reductant and oxidant, whereas different reductants are required for the peroxidases, depending upon their specificities. Under some conditions CAT can act as an efficient peroxidase. SODs deal with the first product of the univalent reduction of O2, converting it to H2O2, which must then be destroyed by CAT and/or peroxidases. Thus, the SOD and CAT serve, in tandem, as front-line antioxidant defenses:

Superoxide dismutase
SODs have been isolated and characterized from a wide variety of organisms. One class consists of SODs with Cu(II) plus Zn(II) at the active site (Cu/ZnSOD), another with Mn(III) (MnSOD), a third with Fe(III) (FeSOD), and a fourth with Ni(II/III) (NiSOD). Cu/ZnSODs are generally found in the cytosol of eukaryotic cells, in chloroplasts, and in some prokaryotes; MnSODs are found in prokaryotes and in mitochondria; FeSODs are generally found in prokaryotes, in algae and in some higher plant chloroplasts; NiSODs have been found in Streptomyces. Unlike most other organisms that have only one of each type of SOD in the various cellular compartments, plants have multiple forms of each type encoded by more than one gene (Figure 3), indicative that plants have far more complex antioxidant defenses (14,15). Plants also produce a large variety of small non-enzymatic antioxidant compounds as second tier defenses, such as glutathione, ascorbate, tocopherols, flavonoids, alkaloids, and carotenoids in high concentrations that are capable of quenching ROS. The dismutation of O2·- to O2 + H2O2 by SOD is hardly a bargain, as the resulting H2O2 can react with metal ions, giving rise to the highly toxic OH·. Fortunately, CAT come to the rescue by degrading H2O2 to O2 and H2O. Most aerobes, including mammals, possess at least one form of homotetrameric CAT with ferriheme at the active sites.
Zymograms showing multiplicity of superoxide dismutase (SOD, left) and catalase (CAT, right) in maize. The cytosolic CuZnSOD-4 and SOD-4A co-migrate to the same position, as do all members of the Sod3 multigene family (14). Intracellular location is indicated in parentheses. Catalases are tissue-specific in their expression: LE = milky endosperm; COL = coleoptile; SC = scutellum; ALEU = aleurone; PER = pericarp; LF = green leaf. When CAT-1 and CAT-2 are co-expressed (ALEU) their subunits interact to generate intergenic heterotetramers. CAT-3 does not form heterotetramers in vivo when co-expressed with either CAT-1 or CAT-2 (16). Arrow shows direction of migration in gel.
Catalase
CAT is largely, but not exclusively, localized in peroxisomes, wherein many H2O2-producing enzymes reside. Thus CAT, which exhibits a high Km for H2O2, can act upon the H2O2 produced before it diffuses to other parts of the cell. CAT is a tetrameric heme containing enzyme that is found in all aerobic organisms. Because of its wide distribution, evolutionary conservation, and capacity to rapidly degrade hydrogen peroxide, it has been proposed that CAT plays an important role in systems which have evolved to allow organisms to live in aerobic environments.
CAT is one of the most active catalysts produced by nature. It decomposes hydrogen peroxide at an extremely rapid rate, corresponding to a catalytic center activity of about 107 min1. Depending upon the concentration of H2O2, it exerts a dual function. At low concentrations (<1µM) of H2O2, it acts "peroxidatically", i.e., a variety of hydrogen donors (e.g., ethanol, ascorbic acid) can be oxidized in the following manner: RH2 + H2O2® R + 2H2O.
At high concentrations of substrate, CAT decomposes toxic hydrogen peroxide at an extremely rapid rate using the "catalatic" reaction in which H2O2 acts as both an acceptor and donor of hydrogen molecules: 2H2O2® 2H2O + O2.
CAT is unique among H2O2 degrading enzymes in that it degrades H2O2 without consuming cellular reducing equivalents. Hence, CAT provides the cell with a very energy efficient mechanism to remove hydrogen peroxide. Therefore, when cells are stressed for energy and are rapidly generating H2O2 through "emergency" catabolic processes, H2O2 is degraded by CAT in an energyefficient manner. This results in a net gain of reducing equivalents and, therefore, cellular energy. It has been proposed that CAT may be uniquely suited to regulate the homeostasis of H2O2 in the cell. In the catalatic mode, CAT has a very high apparent Michaelis constant and, therefore, is not easily saturated with substrate. Thus, the enzyme activity increases linearly over a wide range of H2O2 concentrations, thereby maintaining a controlled intracellular H2O2 concentration. In mammalian systems, organs with high concentrations of CAT (i.e., liver and kidney) have low levels of endogenous H2O2, and organs with low concentrations of CAT (i.e., lung and heart) have high endogenous levels of H2O2. Further, if CAT activity is inhibited, H2O2 concentrations rise in the liver. As in the case of SOD, multiple CATs (isozymes; Figure 3) encoded by specific genes are found in plants, whereas animals exhibit one form of CAT (16,17). Both Cat and Sod genes respond differentially to various stresses known to generate ROS (Figure 4).
Response to acute ozone exposure of the Sod (top-left) and Cat (top-right) genes in maize leaves after 6 h of O3 fumigation. As noted, some of the transcripts are upregulated (e.g., Sod3, Sod4, Cat1, Cat3), while others are downregulated (e.g., Sod1, Cat2). The 18S rRNA is a loading control. The production of H2O2 in maize leaves (bottom-left) in response to wounding. Seven-day-old plants were excised at the base of the stems and supplied with diaminobenzidine (DAB) for 6 h. The plants then were wounded and continuously supplied with DAB for 4 h. The production of H2O2 can be visualized by the deposition of the brown-red (dark areas) color products in the leaves. A, Control and wounded leaves (wounding was conducted near the main vein). B, Control and wounded leaves (wounding was conducted at the main vein and cut at the leaf edges). Cat transcript accumulation in response to wounding (bottom-right). RNA was isolated from similarly treated leaves, indicative that differential transcript accumulation is effected by induction of H2O2.
ROS, telomeres, and aging
Many hypotheses have been proposed to explain the root cause of aging. One broad-based hypothesis is that generalized homeostatic failure leads to age-related decline. Another is the "free-radical theory of aging" suggesting that endogenous ROS continually damage cellular macromolecules, including DNA (18). Incomplete repair of such damage would lead to its accumulation over time resulting in age-related deterioration.
The evidence implicating the generation of ROS as key factors in determining longevity is accumulating (19). Much of the earlier evidence was correlative. However, recent evidence has identified longevity-influencing genes responsive to ROS. It has further been shown that continual induction of DNA damage during normal aging results in genomic instability due to persistent DNA lesions, mutations, stalled repair, and transcription interference. The involvement of ROS in limiting lifespan has been suggested by analyses of transgenic Drosophila, which systematically overexpress both Sod and Cat genes. Some strains of these transgenic flies live up to 30% longer than their natural counterparts, whereas flies carrying only one of these constitutively expressed transgenes do not live longer, suggesting that ROS toxicity plays an important role in longevity (20,21).
Much attention has also recently been given to the role of telomere shortening as a causative factor in aging/senescence (22,23). Telomeres are stretches of repetitive DNA of tandem short sequence repeats (TTAGGG) that "cap" the ends of eukaryotic chromosomes to protect against degradation. Telomeres in most human cells shorten with each round of DNA replication, because they lack the enzyme telomerase. Telomerase is a specialized reverse transcriptase, which helps to replicate the telomere ends of chromosomes. However, telomerase is normally expressed only in germ-line cells and is derepressed in tumor cells in which telomeres are stabilized. It has recently been demonstrated that oxidative stress is the main reason for telomere shortening (24). It has been observed that H2O2 plus Cu(II) induced 8-oxo-7,8-dihydro-2'-deoxyguanosine formation in the telomere sequences more efficiently than in non-telomere sequences. Oxidative damage is not repaired as well in telomeric DNA as elsewhere in the chromosome. Oxidative stress accelerates telomere loss, while antioxidants decelerate it. Thus, oxidative stress is a critical modulator of telomere loss and telomere-driven replicative senescence is primarily a stress response. In a recent study, individuals under generalized stress were found to have shorter telomeres and less telomerase activity, and more oxidative stress (25). Given the available information, aging is likely to be a multifactorial process; whether ROS are peripheral targets that correlate with longevity or central regulators of human aging remains to be resolved.
Oxidative stress
ROS such as O2·-, H2O2, and OH· are produced in all aerobic organisms and normally exist in the cell in balance with antioxidant molecules. Oxidative stress occurs when this critical balance is disrupted due to depletion of antioxidants or excess accumulation of ROS, or both (Figure 5). That is, when antioxidants are depleted and/or if the formation of ROS increases beyond the ability of the defenses to cope, then oxidative stress and its detrimental consequences ensue. Such stress occurs when severely adverse environments or physiologic conditions overwhelm biological systems. One rapid and clear indicator of oxidative stress is the induction of antioxidant defenses and/or increases in endogenous ROS levels. The formation of ROS can be accelerated as a consequence of various environmental stress conditions, including UV-radiation, high light intensities, exposure to herbicides, extreme temperatures, toxins such as cercosporin and aflatoxin, air pollutants, metals, wounding, and xenobiotics. Many inducers of oxidative stress are known carcinogens, mutagens, and toxins. ROS production and accumulation is a common denominator in many diseases and environmental insults and can lead to severe cellular damage leading to physiological dysfunction and cell death in virtually all aerobes (Figure 6).
When oxidative stress occurs, cells function to counteract the oxidant effects and to restore redox balance by resetting critical homeostatic parameters. Such cellular activity leads to activation or silencing of genes encoding defensive enzymes, transcription factors, and structural proteins (26,27).
Oxidative stress results from imbalance between the levels of antioxidants (AOX) and reactive oxygen species (ROS). Cells are normally able to balance the production of oxidants and antioxidants to maintain redox equilibrium. Oxidative stress occurs when this equilibrium is upset by excess levels of ROS, or depletion of antioxidant defenses.
Scheme showing some of the initiators (stressors) of reactive oxygen species (ROS) and the biological consequences leading to a variety of physiological dysfunctions that can lead to cell death.
ROS perform essential cellular functions
It has recently become apparent that ROS are not always harmful metabolic byproducts as generally believed (27). When tightly regulated, ROS perform critical functions in the cell. In fact, a significant body of evidence indicates that ROS, particularly O2·- and H2O2, act as intracellular signaling molecules. In bacteria, the transcription factor OxyR activates a number of genes inducible by H2O2, while the transcription factors SoxR/SoxS mediate responses to O2·- (28,29). Yeast also has two distinct adaptive stress responses, one directed towards H2O2 and one towards O2·- (30-32). In higher eukaryotes, both animals and plants, oxidative stress responses are more complex and are modulated by several regulators (9). ROS-dependent redox cycling of cysteinyl thiols is critical for establishing protein-protein and protein-DNA interactions that determine many aspects of signal transduction pathways by regulating the activity of many transcription factors (26,32). For example, activation of nuclear factor kB (NF-kB) and activator protein-1 (AP-1), known to have critical roles in proliferation, differentiation and morphogenesis, can result from stimulation by diverse agents proceeding through a common pathway involving ROS generation. The pathway leading to H2O2 production and subsequent redox activation of NF-kB has been shown to involve the Rho family of small GTP-binding proteins (33).
Both intracellular and extracellular sources of ROS are capable of modulating gene expression. Low doses of H2O2 (<20 µM) can elicit changes in phosphorylation of specific regulatory proteins including protein kinase B. Direct signaling action of H2O2 in the differential regulation of antioxidant genes in plants likely occurs via protein-DNA interactions in the region of the antioxidant-responsive element (ARE; TGACTCA), NF-kB, and abscisic acid-responsive element 2 (ACGT) in the promoters of these genes. In addition to induction of defense gene expression, other roles of ROS in plants include direct killing of pathogens, involvement in cell wall structure, and promotion of programmed cell death (9). In yeast and animal cells, ROS have been shown to arrest cell division, and cell cycle progression is under negative ROS control (34). A clear and powerful example of how ROS are put to constructive uses was the observation that O2·- plays an important role against invading microbes, in effect serving as a broad-spectrum antibiotic (35). In response to invasion by pathogens, plants also mount a broad range of defense responses, including a rapid and transient production of large amounts of ROS ("oxidative burst") (36).
Thus, ROS play different, or even opposing roles, during different cellular processes (Figure 7). For example, under physiologic conditions, H2O2 may play an important role in signal transduction pathways and in activation of the transcription factor NF-kB, while under pathologic (stress) conditions, H2O2 can lead to apoptosis or necrosis. Consequently, the steady-state level of ROS within cells is critical and is determined by the interplay between ROS-generating mechanisms and the subtle modulating roles played by cellular antioxidants. In plants, H2O2 is generated under a diverse range of biotic and abiotic conditions, and its accumulation in specific tissues at specific developmental times, in the appropriate quantities, benefits plants and can mediate cross-tolerance to other stresses (37). H2O2 affects gene expression and activates MAP kinases (MAPK), which in turn function as regulators of transcription (3).
Scheme showing some of the useful pleiotropic roles of reactive oxygen species (ROS) known to occur in most higher organisms, indicative of the fact that ROS are not always harmful to cells. MAPK = mitogen-activated protein kinase.
ROS and gene expression
Numerous studies indicate that cells have the means to sense ROS and to induce specific responses, but the underlying mechanisms are not fully understood (9,26). The transcriptional network that responds to ROS in eukaryotes is currently being deciphered, whereas the prokaryotic system is better understood.
Nearly three decades ago, it was shown that the expression of ~30 proteins was induced by H2O2 in bacteria (38). Of these 30 proteins, 12 were maximally induced within 10 min and 18 between 10-30 min. The OxyR regulatory protein was subsequently shown to regulate expression of 9 of the 12 rapidly induced proteins. The tetrameric OxyR protein is a member of the LysR family of transcription activators and exists in two forms, reduced and oxidized; only the oxidized form is able to activate transcription. Further studies led to the identification of a number of OxyR-activated genes (39). Similarly, the SoxRS regulatory proteins were found to regulate expression of O2·--responsive proteins in bacteria (40). Regulation of the SoxRS regulon occurs by conversion of SoxR to an active form that enhances soxS transcription. The enhanced levels of SoxS in turn activate expression of the regulon (40). In addition to SoxR and OxyR, several other transcriptional regulators modulate the expression of antioxidant genes in bacteria, indicative of the complexity and connectivity of overlapping regulatory networks. No apparent homologs of OxyR, SoxR, or SoxS have been found in eukaryotes (28), but a number of other transcription factors have been found to play a role in regulating the expression of antioxidant genes in eukaryotes. In yeast, transcription regulators of antioxidant genes include ACE1, MAC1, YAP1, YAP2, HAP1, and HAP2/3/4 (30). In higher eukaryotes, oxidative stress responses are more complex and are modulated by several different regulators. In mammalian systems, NF-kB and AP-1 are involved in regulating the oxidative stress response. The ARE, present in the promoter region of mammalian glutathione S-transferase, metallothionein-I, and MnSod genes, causes induction of these genes in response to oxidants (41). NF-kB, AP-1, and ARE have also been found in promoters of antioxidant genes in higher plants (Figure 8) (9). The role of these factors is not unique to activation of antioxidant genes, as they are known, particularly NF-kB, to play central roles in regulating cellular responses to other stresses as well as regulating normal growth and metabolism.
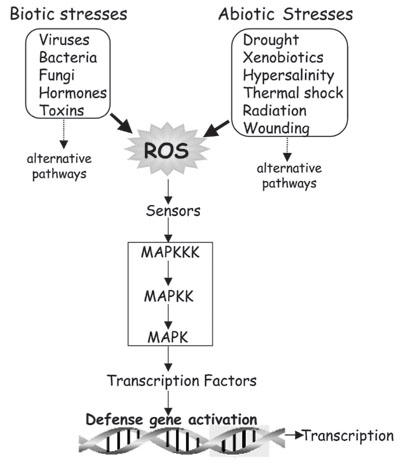
There is substantial evidence that a variety of biotic and abiotic stresses induce ROS, which serve as a common factor in regulating various signaling pathways (Figure 9) (9,42). Similar stresses also activate MAPKs with kinetics that either precede or parallel H2O2 production, indicating that MAPKs may be one of several converging points in the defense-signaling network (43). In addition, exogenous application of several plant hormones and toxins has been shown to induce O2·- and H2O2 synthesis, leading to differential induction of some antioxidant genes and isozymes (Figure 10) (9,37,41,44). Thus, the identification of all genes and proteins regulated by H2O2 is an important step toward treatments that might confer tolerance to multiple, but interrelated, stresses. In addition to induction/repression of antioxidant defense genes, ROS are known to similarly affect expression of a variety of other genes involved in different signaling pathways in microbes (28), yeast (45), plants (46), and animals (19).
Schematic representation of the promoter region of each of the three catalase (Cat) genes of maize indicating the locations of the NF-kB, ARE, AP-1, and ABRE motifs relative to the transcription start site (+1) of each gene. NF-kB = nuclear factor kB; ARE = antioxidant-responsive element: ABRE = abscisic acid-responsive element; AP-1 = activator protein-1.
Convergence of various biotic and abiotic stress stimuli onto mitogen-activated protein kinase (MAPK) pathways via reactive oxygen species (ROS) as a common factor, leading to activation of antioxidant defense genes. Alternative pathways have also been implicated. MAPKKK = MAPK kinase kinases.
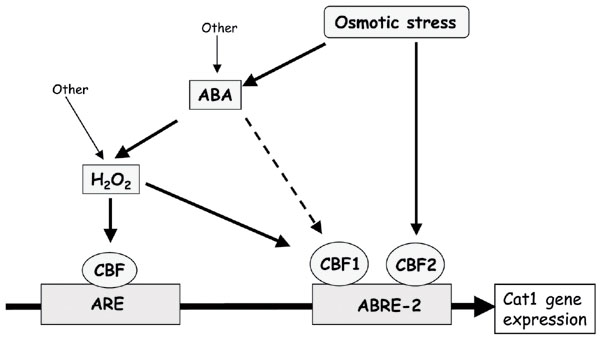
Schematic representation of the possible signal transduction pathways of Cat1 gene expression in response to the plant hormone abscisic acid (ABA) and osmotic stress in maize. ARE = antioxidant-responsive element; ABRE-2 = ABA responsive element 2; CBF1 = Cat1 binding factor-1; CBF2 = Cat1 binding factor-2.
Genomic scale ROS-responsive gene expression
The advent of microarray expression analysis makes possible the assessment of gene expression on a genomic scale, rendering tens of thousands of genes assayable in a single experiment (47). Thus, identification of ROS-responsive genes on a global scale is now tenable. DNA microarrays can comprehensively examine gene expression networks during oxidative stress. There is now significant progress being made in surveying gene expression in response to H2O2 in Escherichia coli (48), yeast (4,45), animals (19), and higher plants (46).
A genome-wide transcription profile of E. coli cells exposed to H2O2 was examined with a DNA microarray composed of 4169 E. coli open reading frames (48). Gene expression was measured in isogenic wild-type and oxyR deletion mutants (DoxyR) to confirm that the H2O2-response regulator OxyR activates most of the H2O2-inducible genes. A very rapid and strong induction was observed of a set of OxyR-regulated genes in the wild type but not in the DoxyR, providing internal validation of the experiment and confirmation for the induction of the oxidative stress genes identified earlier by other means (38). Several new H2O2-inducible genes were also identified: some were members of the OxyR regulon and some induced by an OxyR-independent mechanism suggestive of other H2O2 sensors and regulators in E. coli (38). Several genes repressed by OxyR were highly expressed in the DoxyR mutant. Overall, the mRNA of 140 genes in the wild type, and 167 genes in the DoxyR were significantly induced after H2O2 treatment. It was also found that soxS was induced by H2O2, indicating an overlap with other regulatory pathways. Two genes, Fpr and sodA, known to be members of the SoxRS regulon were also highly induced by H2O2 in both wild type and DoxyR. The microarray data also showed an overlap between oxidative stress, heat shock and SOS responses (38). The results from the E. coli microarrays clearly indicate that the activities of transcription factors in addition to OxyR and SoxRS are likely modulated by oxidative stress.
Whole-genome expression patterns in Saccharomyces cerevisiae cells exposed to H2O2, in addition to other stresses, indicated that ~2/3 of the genome is involved in the response to environmental changes, and the global set of genes induced/repressed by each environmental signal were identified (4,45). The response to oxidative stress involves ~1/3 of the yeast genome and the maximal effects on gene expression occur slightly later relative to other stresses examined during similar time-courses, with most of the transcriptome returning to prestress levels within 2 h following exposure to H2O2 (4). Genes that are repressed for ~60 min after exposure to H2O2 are only transiently repressed in other stress time courses. Thus, genes encoding the translation apparatus and its regulators are remarkably coordinated for each environmental change, although the dynamics of each response are different. The expression programs following H2O2 or O2·- treatment were essentially identical despite the fact that different ROS are involved. There was strong induction of genes known to be involved in detoxification of both H2O2 and O2·-, such as CAT, SOD, and glutathione peroxidase, as well as genes involved in oxidative and reductive reactions (e.g., thioredoxin, glutathione reductase, glutaredoxin). The genes most strongly induced in response to H2O2 and O2·- were dependent on the transcription factor Yap1p for their induction. Genes moderately induced by ROS and other signals are regulated by different transcription factors, depending on the conditions, and different upstream signaling pathways may govern their response. It has also been demonstrated that in Schizosaccharomyces pombe H2O2 activates the Sty1 (stress-activated MAPK) pathway in a dose-dependent manner via two sensing mechanisms (49). At low H2O2 levels, a two-component signaling pathway, which feeds into either of the two (Wak1 or Win1) stress-activated MAPK kinase kinases, regulates Sty1. At high H2O2 levels, however, Sty1 activation is controlled mainly by an independent two-component mechanism, which requires the function of both Wak1 and Win1. In addition, the individual bZip transcription factors, Pap1 and Atf1, were found to function within a limited range of [H2O2]: Pap1 activates target genes at low [H2O2], whereas Atf1 controls transcriptional responses to high [H2O2], with some minor overlap. Some apparent cross talk among Sty1, Atf1, and Pap1 has been detected (50). Thus, S. pombe deploys a combination of stress-responsive regulatory proteins to gauge and trigger the appropriate transcriptional response to increasing H2O2 concentrations (49). This organism mounts two separate responses to oxidative stress: an adaptive response to low-level H2O2 exposure that protects it from subsequent exposures to higher [H2O2], and an acute response that allows the cell to survive a sudden, potentially lethal dose of H2O2.
Large-scale cDNA microarray analysis of the Arabidopsis transcriptome during oxidative stress identified 175 non-redundant expressed sequence tags from a sample of 11,000, which are regulated by H2O2. Of these, 62 are repressed and 113 are induced. In addition, RNA blots showed that some of the H2O2-regulated genes are also modulated by other signals known to involve oxidative stress (51). Furthermore, a substantial portion of these genes have predicted functions in defense response, cell rescue and signaling, and transcription, underscoring the pleiotropic effects of H2O2 in the response of plants to stress. Overall, the microarray used was estimated to represent only ~30% of the Arabidopsis genome, depending on redundancy, and 1 to 2% of the genes represented in the array are affected by H2O2-imposed oxidative stress, being comparable to the situation in yeast (4). Of the 175 genes identified as H2O2-responsive, most have no obvious direct role in oxidative stress but may be linked to oxidative stress indirectly, as a consequence of other biotic and abiotic stresses, explaining their sensitivity to H2O2. Among the genes induced by H2O2 were genes encoding transcription factors, suggesting that they may mediate downstream H2O2 responses. As in other organisms, expression of the MAPKs in Arabidopsis is induced by oxidative stress, which in turn can mediate the induction of oxidative stress-responsive genes (52).
Toward an integrated view of oxy-stress responses
The traditional view of ROS as mere indiscriminate reactive byproducts of cellular metabolism has recently undergone a metamorphosis. This view came about by the discovery that ROS may act as signal-transducing molecules and that activation of intracellular transcription factors such as OxyR, SoxRS, NF-kB, and AP-1 occur via interaction with ROS, leading to gene transcription (Figure 11).
Genome sequencing and expression profiling using DNA or oligonucleotide microarrays, and related technologies, have been used effectively in the study of global gene expression patterns in response to different growth and environmental conditions to which organisms are exposed. Subsequent hierarchical clustering methods allow for the allocation of genes, coregulated temporally or in response to a given signal, into specific expression groups or regulons (48, 53). The numbers of genes that can be detected by these methods in response to any given environmental or developmental signal far exceed the limited number that could have been detected only a few years ago. With the current methods the expression of tens of thousands of genes can be detected in a single experiment in response to ROS, or to any given signal. Since transcription of genes into mRNA is governed by transcription factors which bind to cis- regulatory regions of the DNA in the vicinity of the target gene, the question arises as to whether large co-regulated groups of genes share cis-regulatory elements that bind to common transcription factors. The data available with respect to oxidative stress seem to suggest this. Cis-acting elements within the promoters of ROS-activated genes are being defined as well as their cognate trans-acting factors. A comparative analysis of promoter sequences of genes with similar expression profiles should provide a basis for unraveling common regulatory sequences and overlapping gene expression networks modulating ROS-responsive genes. The antioxidant enzymes CAT and SOD play key roles in modulating the levels of endogenous H2O2 and O2·-, which in turn, at specific concentrations, act to modulate the expression of other ROS-responsive genes.
The use of microarrays and future derivations thereof, to examine global gene responses to ROS is assured. It has become clear that there are far more genes and gene clusters responding to ROS than previously thought, and that ROS likely play far more key roles in cellular activities than anticipated. As global gene responses to ROS are examined temporally and spatially along with different time-courses, and varied oxidant concentrations, even more complex regulons and regulatory cascades will emerge. It will also be instructive to examine gene expression profiles in specific mutants to help delineate the roles of specific regulators and of ROS. The identification of alternate pathways of ROS-dependent gene expression and characterization of the redox sensing mechanisms involved should point to new insights and directions. Some genes have been identified whose transcription is responsive to a variety of stresses in addition to oxystress, while others appear to be responsive only to ROS or other specific signals. Although some regulatory systems have been implicated in modulating such responses, the complete network of regulators of ROS responses that activate such genes remains unclear.
Microarray results from different organisms clearly underscore the fluidity of genomes to reorganize and respond to changes in the cellular and extracellular environment. However, characterization of global gene expression programs at the transcript level is only the first step toward defining the role or function of each ROS-responsive gene. It is not unlikely that small changes in gene expression could lead to large alterations in protein levels. Consequently, proteomic analyses are essential to correlate mRNA changes with protein levels. There are in fact ample data demonstrating that ROS can alter the activity of cellular proteins.
A large and growing number of sequenced genomes and emerging technologies present us with enormous opportunities to advance biological science, and our knowledge of how genomes perceive signals to respond to variable environments. Oxidative stress and the responses to it seem to be central to many of the key biological questions. However, as the euphoria of new sequencing and technological advances begins to fade, we must recognize the magnitude of the problems that still need to be solved. Knowing the sequences of tens of thousands of ROS-responsive genes only reminds us that we still do not know the many proteins they encode, nor the biochemical or biological function of the great majority of such proteins. How such proteins interact with ROS to drive the various physiological processes in aerobic organisms remains a great puzzle. For the future, the fundamental challenge will be to integrate the information now being obtained on gene expression patterns with structural, topological, and functional parameters and interactions of the various proteins encoded by ROS-responsive regulons, and to view the cell in which they function holistically.
The major signaling pathways activated in response to oxidative stress. Reactive oxygen species (ROS) originating from environmental signals or from metabolic activity are modulated by antioxidants to nontoxic levels, at which point they serve as signaling molecules. ROS can activate gene transcription in two ways: a) via transcription factors, such as NF-kB, AP-1, and ARE-binding proteins (ARE-BP) that can interact directly with specific DNA motifs on promoters of target genes, or b) via activation of MAPK cascades, which in turn activate transcription factors that trigger target gene transcription. The degree to which a given pathway is activated depends on the nature and duration of the stress, as well as on cell type and developmental stage. NF-kB = nuclear factor kB; AP-1 = activator protein-1; ARE-BP = antioxidant-responsive element binding proteins; HSF1 = heat shock transcription factor 1.
The paradox
The oxygen paradox is indeed the paradox of evolution itself. Evolutionary pressures have made the best of a bad situation by generating mechanisms to curtail the undesirable toxic effects of ROS, which are an unavoidable consequence of the aerobic lifestyle, and to put them to constructive uses. Indeed evolution has co-opted ROS to serve necessary and useful purposes in the maintenance of cellular homeostasis and in the communication of cells with the external environment. The focus must now be placed on a more thorough understanding of how ROS-mediated signals are perceived, transduced, and interpreted by the cell's genetic machinery. Perhaps the most noteworthy observation to date concerning oxidative stress and the negative and positive roles of ROS is their universality among aerobic organisms and the similarities emerging in the regulatory mechanisms underlying their roles in all species.
The classical view of ROS as villains that indiscriminately destroy biomolecules has undergone a shift, in which positive biological roles are considered as well. It is now accepted that ROS, particularly H2O2 and O2·-, are carefully regulated metabolites capable of signaling and communicating critical information to the cell's genetic machinery. Redox regulation of gene expression by oxidants and antioxidants is emerging as a vital mechanism in the health of all eukaryotes, including man.
It would indeed be interesting and challenging to identify all the changes in gene expression regulated by oxidative stress, and to determine their commonality among diverse species. Such a global analysis of the effects of H2O2 and O2·- on the transcriptome of eukaryotes has not yet been attained, but with the emergence of post-genomic technologies it will likely be not far off.
Conclusions
The survival of organisms on earth depends upon the interactions of their genomes with the environments in which they exist. In the course of evolution, organisms evolved a complex array of mechanisms for adapting to both minor and major fluctuations in the environment. The emergence of oxygenic photosynthesis presented early life forms with the greatest environmental challenge and an opportunity. The challenge was to develop antioxidant defenses in order to survive; the opportunity was to exploit the reactivity of oxygen for energy yielding and biosynthetic reactions. The opportunity led to the highly diversified life forms that evolved sufficient defenses and managed to exploit the aerobic lifestyle. Oxygen toxicity likely led to massive extinctions of those organisms unable to cope with it, unless they took refuge in isolated anaerobic niches. Thus, oxygen is a "double-edged sword" in that it makes life on earth possible, but in its reduced forms (ROS), it is highly toxic and lethal. Oxidative stress can arise from an imbalance between generation and elimination of ROS leading to excess ROS levels inflicting indiscriminate damage to virtually all biomolecules, leading in turn to various diseases and cell death. The notion that ROS are merely toxic byproducts of O2 metabolism has recently been altered by experimental evidence indicating that ROS are carefully regulated metabolites capable of signaling and communicating information to the cell's genetic machinery. Redox regulation of gene expression by oxidants and antioxidants is emerging as a vital mechanism in the health and disease of all organisms. Irrespective of how or when ROS are generated, an increase in intracellular oxidants results in two critical effects: damage to various cell components and activation of specific signaling pathways, influencing various cellular processes leading to proper cell functions or to cell death. Genomic tools are accelerating the discovery of ROS-responsive genes on a global scale and are expanding our understanding of the oxidative stress response and the pleiotropic roles of ROS in signaling and gene expression.
Acknowledgments
I gratefully acknowledge the generous, and continuous, support of my research over many years by the U.S. National Institutes of Health, Department of Energy, Environmental Protection Agency, National Science Foundation, and Department of Agriculture.
Address for correspondence: J.G. Scandalios, Department of Genetics, North Carolina State University, Raleigh, NC 27695, USA. E-mail: jgs@unity.ncsu.edu
Based on a Plenary Lecture presented at the XXXIII Annual Meeting of the Brazilian Society of Biochemistry and Molecular Biology, Caxambu, MG, Brazil, May 15-18, 2004. Received January 31, 2005. Accepted April 18, 2005.
- 1. Scandalios JG (2002). The rise of ROS. Trends in Biochemical Sciences, 27: 483-486.
- 2. McClintock B (1984). The significance of responses of the genome to challenge. Science, 226: 792-801.
- 3. Scandalios JG (2002). Oxidative stress responses: what have genome-scale studies taught us? Genome Biology, 3: 1019.1-1019.6.
- 4. Gasch A, Spellman P, Kao C, Harel O, Eisen M, Storz G, Botstein D & Brown P (2000). Genomic expression programs in the response of yeast cells to environmental changes. Molecular Biology of the Cell, 11: 4241-4257.
- 5. Scandalios JG & Guan LM (2001). Transcription factors regulating antioxidant gene expression in response to biotic and abiotic signals. In: Gene Families: Studies of DNA, RNA, Enzymes and Proteins World Scientific, Singapore, 287-301.
- 6. Laloi C, Apel K & Danon A (2004). Reactive oxygen signaling: the latest news. Current Opinion in Plant Biology, 7: 323-328.
- 7. Thorpe GW, Fong CS, Alic N, Higgins VJ & Dawes IW (2004). Cells have distinct mechanisms to maintain protection against different reactive oxygen species: Oxidative stress-response genes. Proceedings of the National Academy of Sciences, USA, 101: 6564-6569.
- 8. Hancock JT, Desikan R & Neill SJ (2001). Role of reactive oxygen species in cell signaling pathways. Biochemical Society Transactions, 29: 345-350.
- 9. Scandalios JG (1997). Oxidative Stress and the Molecular Biology of Antioxidant Defenses Cold Spring Harbor Laboratory Press, Plainview, NY, USA.
- 10. Elstner EF (1987). Metabolism of activated oxygen species. In: Davies DD (Editor), The Biochemistry of Plants Vol. 11. Academic Press, New York, 253-315.
- 11. Dismukes GC, Klimov VV, Baranov SV, Kozlov YN, DasGupta J & Tyryshkin A (2001). The origin of atmospheric oxygen on Earth: The innovation of oxygenic photosynthesis. Proceedings of the National Academy Sciences, USA, 98: 2170-2175.
- 12. Fridovich I (1995). Superoxide radical and superoxide dismutases. Annual Review of Biochemistry, 64: 87-112.
- 13. Halliwell B (1996). Free radicals in biochemistry and medicine. In: Meyers RA (Editor), Encyclopedia of Molecular Biology and Molecular Medicine Vol. 2. Wiley-VCH, Weinheim, Germany, 330-337.
- 14. Scandalios JG (1997). Molecular genetics of superoxide dismutases. In: Scandalios JG (Editor), Oxidative Stress and the Molecular Biology of Antioxidant Defenses Cold Spring Harbor Laboratory Press, Plainview, NY, USA, 527-568.
- 15. Fink RC & Scandalios JG (2002). Molecular evolution and structure function relationships of the superoxide dismutase gene families in angiosperms, and their relationship to other eukaryotic and prokaryotic superoxide dismutases. Archives of Biochemistry and Biophysics, 399: 19-35.
- 16. Scandalios JG, Guan L & Polidoros AN (1997). Catalases in plants: Gene structure, properties, regulation, and expression. In: Scandalios JG (Editor), Oxidative Stress and the Molecular Biology of Antioxidant Defenses Cold Spring Harbor Laboratory Press, Plainview, NY, USA, 343-406.
- 17. Guan L & Scandalios JG (1996). Molecular evolution of maize catalases and their relationship to other eukaryotic and prokaryotic catalases. Journal of Molecular Evolution, 42: 570-579.
- 18. Beckman KB & Ames BN (1998). The free radical theory of aging matures. Physiological Reviews, 78: 547-581.
- 19. Finkel T & Holbrook NJ (2000). Oxidants, oxidative stress and the biology of aging. Nature, 408: 239-247.
- 20. Beckman KB & Ames BN (1997). Oxidants, antioxidants, and aging. In: Scandalios JG (Editor), Oxidative Stress and the Molecular Biology of Antioxidant Defenses Cold Spring Harbor Laboratory Press, Plainview, NY, USA, 201-246.
- 21. Helfand SL & Rogina B (2003). Molecular genetics of aging in the fly: is this the end of the beginning? BioEssays, 25: 134-141.
- 22. Chan SRWL & Blackburn EH (2004). Telomeres and telomerases. Philosophical Transactions of the Royal Society of London, Series B. Biological Sciences, 359: 109-121.
- 23. McNight TD & Shippen DE (2004). Plant telomere biology. Plant Cell, 16: 794-803.
- 24. Von Zglincki T (2002). Oxidative stress shortens telomeres. Trends in Biochemical Sciences, 27: 339-344.
- 25. Sapolsky RM (2004). Organismal stress and telomeric aging: An unexpected connection. Proceedings of the National Academy of Sciences, USA, 101: 17323-17324.
- 26. Dalton TP, Shertzer HG & Puga A (1999). Regulation of gene expression by reactive oxygen. Annual Review of Pharmacology and Toxicology, 39: 67-101.
- 27. Scandalios JG (2004). Genomic responses to oxidative stress. In:Meyers RA (Editor), Encyclopedia of Molecular Cell Biology and Molecular Medicine Vol. 5. 2nd edn. Wiley-VCH, Weinheim, Germany, 489-512.
- 28. Storz G & Imlay JA (1999). Oxidative stress. Current Opinion in Microbiology, 2: 188-194.
- 29. Kiley PJ & Storz G (2004). Exploiting thiol modifications. PLoS Biology, 2: 1714-1717.
- 30. Ruis H & Schuller C (1995). Stress signaling in yeast. BioEssays, 17: 959-965.
- 31. Moradas-Ferreira P & Costa V (2000). Adaptive response of the yeast Saccharomyces cerevisiae to reactive oxygen species: defenses, damage and death. Redox Report, 5: 277-285.
- 32. Delaunay A, Isnard A & Toledano MB (2000). H2O2 sensing through oxidation of the Yap1 transcription factor. EMBO Journal, 19: 5157-5166.
- 33. Gabbita SP, Robinson K, Stewart C, Floyd R & Hensley K (2000). Redox regulatory mechanisms of cellular signal transduction. Archives of Biochemistry and Biophysics, 376: 1-13.
- 34. Paulovich AG, Toczysky DP & Hartwell LH (1997). When checkpoints fail. Cell, 88: 315-321.
- 35. Babior BM (1984). The respiratory burst of phagocytes. Journal of Clinical Investigation, 73: 599-601.
- 36. Doke N (1997). The oxidative burst: roles in signal transduction and plant stress. In: Scandalios JG (Editor), Oxidative Stress and the Molecular Biology of Antioxidant Defenses Cold Spring Harbor Laboratory Press, Plainview, NY, USA, 785-813.
- 37. Guan LM, Zhao J & Scandalios JG (2000). Cis-elements and trans-acting factors that regulate expression of the maize Cat1 antioxidant gene in response to ABA and osmotic stress: H2O2 is the likely intermediary signaling molecule for the response. Plant Journal, 22: 87-95.
- 38. Christman MF, Morgan RW, Jacobson FS & Ames BN (1985). Positive control of a regulon for defenses against oxidative stress and some heat-shock proteins in Salmonella typhimurium Cell, 41: 753-762.
- 39. Storz G & Zheng M (2000). Oxidative stress. In: Storz G & Henge-Aronis R (Editors), Bacterial Stress Responses ASM Press, Washington, DC, USA, 47-59.
- 40. Wu J & Weiss B (1991). Two divergently transcribed genes, soxR and SoxS, control a superoxide response regulon of Escherichia coli Journal of Bacteriology, 173: 2864-2871.
- 41. Scandalios JG (2001). Molecular responses to oxidative stress. In: Hawkesford MJ & Buchner P (Editors), Molecular Analysis of Plant Adaptation to the Environment Kluwer Academic Publishers, Dordrecht, The Netherlands, 181-208.
- 42. Somssich IE & Halbrock K (1998). Pathogen defence in plants - a paradigm of biological complexity. Trends in Plant Sciences, 3: 86-90.
- 43. Zhang S & Klessig D (2001). MAPK cascades in plant defense signaling. Trends in Plant Sciences, 6: 520-527.
- 44. Williamson J & Scandalios JG (1992). Differential response of maize catalases and superoxide dismutases to the photoactivated fungal toxin cercosporin. Plant Journal, 2: 351-358.
- 45. Causton H, Bing R, Koh S, Harbison C, Kanin E, Jennings E, Lee T, True H, Lander E & Young R (2001). Remodeling of yeast genome expression in response to environmental changes. Molecular Biology of the Cell, 12: 323-337.
- 46. Desikan R, Neill SJ & Hancock JT (2000). Hydrogen peroxide-induced gene expression in Arabidopsis thaliana Free Radical Biology and Medicine, 28: 773-778.
- 47. Schena M, Shalon D, Davis R & Brown P (1995). Quantitative monitoring of gene expression patterns with complementary DNA microarray. Science, 270: 467-470.
- 48. Zheng M, Wang X, Templeton L, Smulski D, LaRossa R & Storz G (2001). DNA microarray-mediated transcriptional profiling of the Escherichia coli response to hydrogen peroxide. Journal of Bacteriology, 183: 4562-4570.
- 49. Quin J, Findlay VJ, Dawson K, Millar JB, Jones N, Morgan BA & Toone WM (2002). Distinct regulatory proteins control the graded transcriptional response to increasing H2O2 levels in fission yeast Schizosaccharomyces pombe Molecular Biology of the Cell, 13: 805-816.
- 50. Nguyen AN, Lee A, Place W & Shiozaki K (2000). Multistep phosphorelay proteins transmit oxidative stress signals to the fission yeast stress-activated protein kinase. Molecular Biology of the Cell, 14: 1169-1181.
- 51. Desikan R, MacKerness A, Hancock JT & Neill SJ (2001). Regulation of the Arabidopsis transcriptome by oxidative stress. Plant Physiology, 127: 159-172.
- 52. Kovtun Y, Chiu WL, Tena G & Sheen J (2000). Functional analysis of oxidative stress-activated mitogen-activated protein kinases cascade in plants. Proceedings of the National Academy of Sciences, USA, 97: 2940-2945.
- 53. Frances M & Rao S (2002). Of chips and ChIPs. Science, 296: 666-669.
Correspondence and Footnotes
Publication Dates
-
Publication in this collection
04 July 2005 -
Date of issue
July 2005
History
-
Accepted
18 Apr 2005 -
Received
31 Jan 2005