Abstracts
The aim of the present study was to evaluate a species-specific nested PCR based on a previously described species-specific PCR for detection of B. ovis in semen and urine samples of experimentally infected rams. The performance of the species-specific nested PCR was compared with the results of a genus-specific PCR. Fourteen rams were experimentally infected with the Brucella ovis REO 198 strain and samples of semen and urine were collected every week up to 180 days post infection. Out of 83 semen samples collected, 42 (50.6%) were positive for the species-specific nested PCR, and 23 (27.7%) were positive for the genus-specific PCR. Out of 75 urine samples, 49 (65.3%) were positive for the species-specific nested PCR, whereas 11 (14.6%) were genus-specific PCR positive. Species-specific nested PCR was significantly more sensitive (P<0.001) than the genus-specific PCR in semen and urine from experimentally infected rams. In conclusion, the species-specific nested PCR developed in this study may be used as a diagnostic tool for the detection of B. ovis in semen and urine samples from suspected rams.
Brucella ovis; species-specific; nested PCR; semen; urine
O presente estudo objetivou avaliar uma técnica de nested PCR espécie-específica delineada a partir de PCR espécie-específica descrita anteriormente para detecção de B. ovis em sêmen e urina de carneiros infectados experimentalmente. O desempenho da nested PCR espécie-específica foi comparado com os resultados de uma PCR gênero-específica. Quatorze carneiros foram infectados experimentalmente com Brucella ovis REO 198 e amostras de sêmen e de urina foram colhidas semanalmente até 180 dias após a infecção. De 83 amostras de sêmen, 42 (50,6%) foram positivas pela nested PCR espécie-específica, e 23 (27,7%) foram positivas pela PCR gênero-específica. De 75 amostras de urina, 49 (65,3%) foram positivas pela nested PCR espécie-específica, enquanto 11 (14,6%) foram positivas em PCR gênero-específica. A técnica de nested PCR espécie-específica foi significativamente mais sensível (P<0,001) do que a PCR gênero-específica no sêmen e na urina de carneiros infectados experimentalmente. Em conclusão, a nested PCR espécie-específica desenvolvida neste estudo pode ser utilizada como ferramenta de diagnóstico para detecção de B. ovis em sêmen e urina de carneiros suspeitos.
Brucella ovis; espécie-específica; nested PCR; sêmen; urina
VETERINARY MEDICINE
Species-specific nested PCR as a diagnostic tool for Brucella ovis infection in rams
Nested PCR espécie-específica para o diagnóstico da infecção por Brucella ovis em carneiros
L.F. CostaI; C.N. NozakiI; N.S.C. LiraI; J.M.A.P. AntunesI; M.N. XavierII; E.A. CostaII; T.A. PaixãoII; R.L. SantosII; J. MegidI.* * Autor para correspondência ( corresponding author) E-mail: jane@fmvz.unesp.br
IFaculdade de Medicina Veterinária e Zootecnia - UNESP - Botucatu, SP
IIEscola de Veterinária - Universidade Federal de Minas Gerais - Belo Horizonte, MG
ABSTRACT
The aim of the present study was to evaluate a species-specific nested PCR based on a previously described species-specific PCR for detection of B. ovis in semen and urine samples of experimentally infected rams. The performance of the species-specific nested PCR was compared with the results of a genus-specific PCR. Fourteen rams were experimentally infected with the Brucella ovis REO 198 strain and samples of semen and urine were collected every week up to 180 days post infection. Out of 83 semen samples collected, 42 (50.6%) were positive for the species-specific nested PCR, and 23 (27.7%) were positive for the genus-specific PCR. Out of 75 urine samples, 49 (65.3%) were positive for the species-specific nested PCR, whereas 11 (14.6%) were genus-specific PCR positive. Species-specific nested PCR was significantly more sensitive (P<0.001) than the genus-specific PCR in semen and urine from experimentally infected rams. In conclusion, the species-specific nested PCR developed in this study may be used as a diagnostic tool for the detection of B. ovis in semen and urine samples from suspected rams.
Keywords:Brucella ovis, species-specific, nested PCR, semen, urine
RESUMO
O presente estudo objetivou avaliar uma técnica de nested PCR espécie-específica delineada a partir de PCR espécie-específica descrita anteriormente para detecção de B. ovis em sêmen e urina de carneiros infectados experimentalmente. O desempenho da nested PCR espécie-específica foi comparado com os resultados de uma PCR gênero-específica. Quatorze carneiros foram infectados experimentalmente com Brucella ovis REO 198 e amostras de sêmen e de urina foram colhidas semanalmente até 180 dias após a infecção. De 83 amostras de sêmen, 42 (50,6%) foram positivas pela nested PCR espécie-específica, e 23 (27,7%) foram positivas pela PCR gênero-específica. De 75 amostras de urina, 49 (65,3%) foram positivas pela nested PCR espécie-específica, enquanto 11 (14,6%) foram positivas em PCR gênero-específica. A técnica de nested PCR espécie-específica foi significativamente mais sensível (P<0,001) do que a PCR gênero-específica no sêmen e na urina de carneiros infectados experimentalmente. Em conclusão, a nested PCR espécie-específica desenvolvida neste estudo pode ser utilizada como ferramenta de diagnóstico para detecção de B. ovis em sêmen e urina de carneiros suspeitos.
Palavras-chave:Brucella ovis, espécie-específica, nested PCR, sêmen, urina
INTRODUCTION
Ovine brucellosis caused by the Brucella ovis is a contagious disease, with venereal and oral transmission, which causes reproductive failure in rams. B. ovis, a Gram negative coccobacilli of α2 Proteobacteriacea family, can cause chronic epididymitis, orchitis and infertility in rams; whereas in ewes the infection uncommonly causes abortion and placentitis or birth of weak lambs (Xavier et al., 2009). In addition to B. ovis, which has no zoonotic potential, ovine brucellosis can also be caused by B. melitensis, which is the most pathogenic Brucella species for humans (Blasco, 1990; Xavier et al., 2009). Therefore, a precise differential diagnosis between these two species of Brucella in rams is highly important (Xavier et al., 2010).
Currently, routine diagnostic tests for ovine brucellosis caused by B. ovis include clinical examination, serological tests, and bacteriology (Alton et al., 1988; Grilló et al., 1999). Some infected rams do not develop palpable lesions (Webb et al., 1980) or, conversely, have genital disease compatible with infection but have been exposed to different bacteria (Hughes e Claxton, 1968). The bacteriology of B. ovis is time consuming and not always efficient, because bacteria can be excreted intermittently in the semen of infected animals (Worthington et al., 1985; Baigun et al., 2000). Regarding serological tests, they generally fail in the sensitivity displayed. Due to the limitations exposed, the diagnosis of B. ovis infection is usually obtained by the association of two or more of these techniques.
More recently, PCR has been used as a diagnostic tool (Bricker, 2002). The diagnosis of B. ovis infection has been obtained through molecular amplification of respective DNA in samples. This technique demonstrated similar sensitivity when compared to bacteriology for B. ovis isolation (Manterola et al., 2003; Xavier et al., 2010). Furthermore, PCR allows a rapid identification of the pathogen in semen (Saunders et al., 2007; Xavier et al., 2010) or urine samples (Xavier et al., 2010) collected from infected rams. Xavier et al. (2010) recently developed a PCR based on the amplification of an open reading frame (ORF) specific for chromosome II of B. ovis (Tsolis et al., 2009) which has a remarkably good sensitivity when compared to bacteriological culture and has proven to be highly specific for detecting B. ovis infection. The authors demonstrated that the target sequence was conserved in 18 different B. ovis field strains and absent in other classical species of Brucella, in bacteria species that can cause epididimitis in rams, and in those microorganisms phylogenetically related to B. ovis (Xavier et al., 2010), confirming the specificity of the technique developed. Nevertheless, its sensitivity may be lower than currently available genus-specific PCR protocols. Bricker and Halling (1994) developed a PCR based on the amplification of 711 insertion elements (IS711 or IS6501) present in multiple copies in the Brucella spp. genomes. The possibility of high sensitivity obtained by a molecular technique in which there are several copies of the target sequence of amplification led Manterola et al. (2003) to develop a diagnostic test for biological samples from sheep experimentally infected with Brucella ovis based on PCR assay with primers derived from the insertion sequence IS6501. Considering the specificity on the method described by Xavier et al. (2010), a pair of internal primers was delineated and the sensibility obtained through this species-specific nested PCR evaluated comparatively to the genus-specific PCR described by Manterola et al. (2003), with the goal of developing a highly sensitive species-specific nested PCR for the detection of B. ovis genomic DNA in semen and urine from rams experimentally infected with B. ovis.
MATERIAL AND METHODS
Fourteen 1-2 year-old rams were experimentally inoculated with B. ovis REO 198 strain, obtained from the "Instituto de Pesquisas Veterinárias Desidério Finamor" (Eldorado do Sul, RS, Brazil). Prior to inoculation, blood samples were collected for serological analysis, as well as semen and urine for culture and PCR, to confirm that the rams were free of B. ovis infection. All rams were purchased form a farm with no previous history of B. ovis infection. A suspension of 2x109 CFU/mL B. ovis was inoculated (2mL intraprepucially and 50µL through the conjunctival route). After inoculation, semen, urine and blood samples were collected every 7 days, for 180 days for PCR and serology. Urine samples were collected by blocking the breath of the animals for thirty seconds and semen from each animal was collected by eletroejaculation. This experimental protocol was approved by the School of Veterinary Medicine and Animal Science - UNESP Committee in Animal Experimentation (Protocol 69/2008).
The semen and urine cultures from rams were carried out prior to inoculation and on the same days of collection. Samples of all animals were cultured in plates containing modified Thayer-Martin agar and Brucella blood agar. Plates were incubated at 37ºC with 5% CO2 for approximately 3-7 days. Suspect colonies were identified by Gram staining, positive urease test and catalase test, negative oxidase test, absence of H2S production, nitrate reduction test and growth in the presence of thionine and basic fucsine.
Serum samples from experimentally infected rams were tested with agar gel immunodifusion (AGID). A kit was made up of bacterial cell wall (Instituto de Pesquisas Veterinárias Desidério Finamor, Eldorado do Sul, RS, Brazil). Gel was prepared using 1% agarose gel (Invitrogen, Brazil) diluted in 0.1M borate buffer. The reaction was carried out at 25ºC in a dark chamber, and the reading was performed at 24, 48 and 72 hours.
Semen and urine samples were collected every week from each experimentally infected ram and a monthly pool was done to perform the genus-specific and species-specific nested PCRs. Semen and urine were thawed and diluted in 500µL of Tris-EDTA (TE) buffer, pH 8.0 (10mM Tris-HCL pH 8.0; 1mM EDTA pH 8.0), incubated at 80ºC for 10 minutes, and centrifuged at 13,000 g for 15 minutes. The supernatant was discarded, the pellet obtained was consecutively washed 2-3 times and then eluted in 500µL of buffer solution containing TE, pH 8.0 (10mM Tris-HCl pH 8.0, 25mM EDTA, pH 8.0, 100mM NaCl), 50µL of SDS 10% and 10µL of proteinase K (20mg/mL) and incubated at 37ºC for 24hours. The remaining extraction was made using the phenol/chloroform/isoamyl alcohol protocol (Cortez et al., 2001), and the final precipitate was resuspended in 30µL of TE, pH 8.0, and incubated at 56ºC for 30 minutes.
For positive control, cultures of Brucella ovis were extracted, adding a colony of bacteria in 400µL of TE pH 8.0 and following the protocol described for semen and urine extraction. The pellet obtained was then resuspended in 100µL of TE, pH 8.0.
The present study used PCR with a primer pair targeting the IS711 sequence (ISP1: 5'-GGTTGTTAAAGGAGAACAGC-3' and ISP2: 5'-GACGATAGCGTTTCAACTTG -3') present in multiple copies in the Brucella spp. genomes (Manterola et al., 2003). PCR reaction and cycle parameters were performed as previously described (Manterola et al., 2003) and a volume of 2.5µL of template DNA was added per PCR reaction. PCR products were resolved by 2% agarose gel electrophoresis. Reactions were considered positive when they yielded products of 700 bp.
Nested PCR amplified products were obtained from species-specific PCR. Species-specific PCR used a primer pair targeting the ORF AO503 (F: 5'-ATCCCCCCATCACCATAACCGAAG-3'and R: 5'-GCCTACGCTGAAACTTGCTTTTG-3') located in the B. ovis-pathogenicity island 1 (BOPI-1) (Tsolis et al., 2009; Silva et al., 2011) as previously described (Xavier et al., 2010). PCR reaction and cycle parameters were performed as previously described (Xavier et al., 2010) and a volume of 2.5µL of template DNA was added per reaction. Nested PCR used a primer pair targeting an internal sequence of the ORF 503 (F NO503: 5'-CAACCACCCAAGTGTCAGTG-3' and R NO503: 5'-ATGGGATTTGCGATCCTGA-3'). Nested PCR reactions were performed using 12,5pmols of each primer at 25µM, 1.25U of Taq Polymerase (Invitrogen, Brazil), 23µL of a commercial PCR mix (PCR Supermix, Invitrogen), containing 22mM Tris-HCl (pH 8.4), 55mM KCl, 1.65mM MgCl2, 220µM dGTP, 220µM dATP, 220µM dTTP, 220µM dCTP, 22U recombinant Taq DNA Polymerase/mL, and stabilizers, and adding 2.5µL of the previous PCR product. Cycling parameters were denaturation at 95ºC for 5 minutes; 35 cycles of denaturation (95ºC for 1 minute), annealing (55ºC for 1 minute), and extension (72ºC for 1 minute); and a final extension at 72ºC for 5 minutes. Nested PCR products were resolved by 2% agarose gel electrophoresis and reactions were considered positive when they yielded products of 171 bp. Positive and negative controls were used by adding B. ovis DNA genome and TE, respectively, in PCR and Nested PCR.
The frequencies of B. ovis detection by genus-specific PCR and species-specific nested PCR were compared by McNemar chi-square. The proportion of the agreement between diagnosis techniques was assessed with the Kappa test.
RESULTS AND DISCUSSION
To test the sanitary status of rams, culture and PCR of semen and urine, and serology were performed prior to inoculation, and all results were negative.
Each one of the experimentally infected rams was serologically positive in at least one time-point post infection. Antibody titers were detected by AGID in all animals in the third week after inoculation, decreasing soon after the fifth week, remaining variable as time passed, which confirms that the experimental challenge resulted in infection in all rams. The serological results are in accordance with previous reports, in which antibody titers were detected between 2 to 9 weeks after intraprepucial inoculation of B. ovis, and then became intermitent (Webb et al., 1980; Xavier et al., 2010)
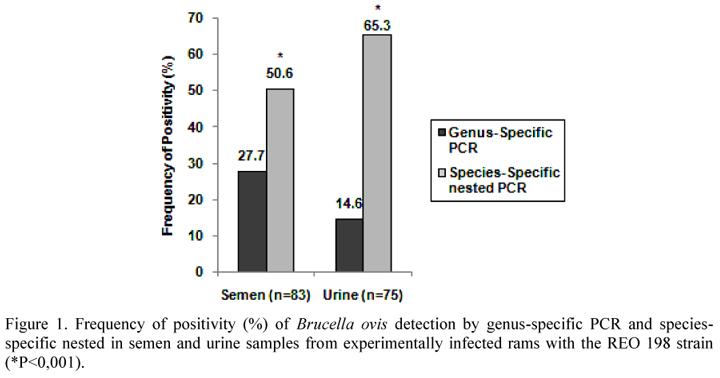
In semen samples, the positivity of the species-specific nested PCR was 50.6% (42/83), which was significantly different from the genus-specific PCR (P<0.001) which had a positivity rate of 27.7% (23/83) (Figure 1). Similarly, when the urine samples were subjected to species-specific nested PCR, the percentage of positive samples was 65.3% (49/75), which was significantly higher (P<0.001) than that obtained by genus-specific PCR, which detected 14.6% (11/75) of positive samples (Figure 1). Xavier at al. (2010) detected B. ovis DNA by species-specific PCR in 17.9% and 19.7% semen and urine samples, respectively, from experimentally infected rams. Thus, the species-specific nested PCR demonstrated higher positivity when compared to genus-specific PCR and improved the sensitivity of the species-specific PCR method.
Species-specific nested PCR and genus-specific PCR had an agreement of 65% and 46.6%, for semen and urine samples, respectively. Kappa test values were 0.3 for semen and 0.1 for urine, considered fair and poor agreements, since nested PCR detected a higher number of positive samples when compared to the genus-specific method (Table 1).
In this study, considering species-specific nested PCR results for semen and urine collected from the same experimental infected rams at a given time-point, the detection of B. ovis DNA in at least sample (i.e. semen and/or urine), reached 81.4% (59/72) (Figure 2). As semen samples are usually the specimen of choice for detection of B. ovis DNA by PCR (Manterola et al., 2003; Saunders et al., 2007), and B. ovis excretion in semen from rams is known to be intermittent (Hughes and Claxton, 1968; Manterola et al., 2003; Saunders et al., 2007, Xavier et al., 2010), urine is a valuable specimen from rams suspected to have the infection. Our results are in agreement with those obtained by Xavier et al. (2010), who demonstrated that the frequency of positive rams was higher when more than one biological sample was considered for diagnosis.
Six and seven experimentally infected rams were positive by bacteriology in at least one time point during the course of the infection in semen cultures and urine samples, respectively. All experimentally infected rams had B. ovis DNA detected by analysis of specie-specific nested PCR in semen in at least one time point of infection. Regarding urine, 13 animals were positive using this technique. By genus-specific PCR, 11 and 7 experimentally infected rams were considered positive for infection based on the results obtained from semen and urine samples, respectively. In this study all PCR were conducted including negative controls, which were never positive, therefore indicating the absence of cross contamination, which supports the notion that the nested PCR can be routinely used in a diagnosis laboratory. The results of this study are in agreement with previous reports that nested PCR has an increased sensitivity for the diagnosis of infections caused by Brucella spp. (Baily et al., 1992; Herman, 1992; Romero et al., 1995; Da Costa et al., 1996).
CONCLUSIONS
The species-specific nested PCR developed in this study allowed the detection of B. ovis genomic DNA in semen and urine of rams and demonstrated improvement in the sensitivity of diagnosis. Thus, the species-specific nested PCR is a tool that can be implemented routinely as a specific and sensitive diagnostic method for Brucella ovis infection in rams.
ACKNOWLEDGEMENTS
This work was supported by FAPESP (Fundação de Amparo a Pesquisa do Estado de São Paulo, São Paulo, Brazil). LFC, RLS and JM are recipients of fellowships from CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico, Brasília, Brazil). Work in the RLS lab is supported by CNPq and FAPEMIG (Fundação de Amparo a Pesquisa do Estado de Minas Gerais, Belo Horizonte, Brazil). Work in the JM lab is supported by CNPq and FAPESP. RLS is currently a fellow of the John Simon Guggenheim Memorial Foundation.
Recebido em 17 de maio de 2011
Aceito em 25 de setembro de 2012
- ALTON, G.G.; JONES, L.M.; ANGUS, R.D. et al Techniques for the Brucellosis Laboratory. INRA, 1988.
- BAIGUN, R.; CONIGLIARO, A.S.; LUNA, F. Aislamiento de Brucella ovis y control de reaccionantes serológicos en epididimitis ovina. Vet. Argent., v.17, p.103-107, 2000.
- BAILY, G.G.; KRAHN, J.B.; DRASAR, B.S. et al Detection of Brucella melitensis and Brucella abortus by DNA amplification. J. Trop. Med. Hyg., v.95, p.271-275, 1992.
- BLASCO, J.M. Brucella ovis In: NIELSEN, K.; DUNCAN, J.R. Animal Brucellosis. CRC Press, 1990. p.351-378.
- BRICKER, B.J. PCR as a diagnostic tool for brucellosis. Vet. Microbiol., v.90, p.435-446, 2002.
- BRICKER, B.J.; HALLING, S.M. Differentiation of Brucella abortus bv. 1, 2 and 4, Brucella melitensis, Brucella ovis and Brucella suis bv. 1 by PCR. J. Clin. Microbiol., v.33, p.1640-1642, 1994.
- CORTEZ, A.; SCARCELLI, E.; SOARES, R.M. et al. Detection of Brucella DNA from aborted bovine foetus by polymerase chain reaction. Aust. Vet. J., v.79, p.500-501, 2001.
- DA COSTA, M.; GUILLOU, J.P.; GARIN-BASTUJI, B. et al Specificity of six gene sequences for the detection of genus Brucella by DNA amplification. J. Appl. Microbiol., v.81, p.267-275, 1996.
- GRILLÓ, M.J.; MARÍN, C.M.; BARBERAN, M. et al Experimental Brucella ovis infection in pregnant ewes. Vet. Rec., v.144, p.555-558, 1999.
- HERMAN, L.; DE RIDDER, H. Identification of Brucella spp. by using the polymerase chain reaction. Appl. Environ. Microbiol, v.58, p.2099-2101, 1992.
- HUGHES, K.L.; CLAXTON, P.D. Brucella ovis infection. 1. An evaluation of microbiological, serological and clinical methods of diagnosis in the rams. Aus. Vet. J., v.30, p.201-203, 1968.
- MANTEROLA, L.; TEJERO-GARCES, A.; FICAPAL, A. et al Evaluation of a PCR test for the diagnosis of Brucella ovis infection in sêmen samples from rams. Vet. Microbiol., v.92, p.65-72, 2003.
- MATRONE, M.; KEID, L.B.; ROCHA, V.C.M. et al Evaluation of DNA extraction protocols for Brucella abortus pcr detection in aborted fetuses or calves born from cows experimentally infected with strain 2308. Braz. J. Microbiol., v.40, p.480-489, 2009.
- ROMERO, C.; GAMAZO, C.; PARDO, M. et al Specific detection of Brucella DNA by PCR. J. Clin. Microbiol., v.33, p.615-617, 1995.
- SAUNDERS, V.F.; REDDACLIFF, L.A.; BERG, T. et al Multiplex PCR for the detection of Brucella ovis, Actinobacillus seminis and Histophilus somni in ram semen. Aust. Vet. J., v.85, p.72-77, 2007.
- SILVA, T.M.A.; PAIXÃO, T.A.; COSTA, E.A. et al. Putative ATP-binding cassette transporter is essential for Brucella ovis pathogenesis in mice. Infection and Immunity, v.79, p.1706-1717, 2011.
- TSOLIS, R.M.; SESHADRI, R.; SANTOS, R.L. et al Genome degradation in Brucella ovis, corresponds with narrowing of its host range and tissue tropism. PLoS One, v.4, p.1-9, 2009.
- WEBB, R.F.; QUINN, C.A.; COCKRAM, F.A. et al Evaluation of procedures for the diagnosis of Brucella ovis infection in rams. Aust. Vet. J., v.56, p.172-175, 1980.
- WORTHINGTON, R.W.; STEVENSON, B.J.; LISLE, G.W. Serology and semen culture for the diagnosis of Brucella ovis infection in chronically infected rams. N. Z. Vet. J., v.33, p.84-86, 1985.
- XAVIER, M.N.; COSTA, E.A.; PAIXÃO, T.A. et al The genus Brucella and clinical manifestations of brucellosis. Cienc. Rural., v.39, p.2252-2260, 2009.
- XAVIER, M.N.; SILVA, T.M.A.; COSTA, E.A. et al Development and evaluation of a species-specific PCR assay for the detection of Brucella ovis infection in rams. Vet. Microbiol, v.145, p.158-164, 2010.
Publication Dates
-
Publication in this collection
05 Mar 2013 -
Date of issue
Feb 2013
History
-
Received
17 May 2011 -
Accepted
25 Sept 2012