Abstracts
The affinities for human albumin (HSA) of five rhodium(II) complexes of general formula [Rh2(bridge)4] (bridge = acetate, propionate, butyrate, trifluoroacetate and trifluoroacetamidate) were determined by spectrophotometry. In the case of the alkylcarboxylates, an inverse correlation of affinity with their liposolubilities was observed. Diffusion of the free or protein-bound complexes into Ehrlich cells in vitro seems to be primarily governed by the hydrophobic character of the complex. The complex [Rh2(tfc)4] exhibited affinity towards the protein (K = 214.1) as well as cell partition both in the absence (32.1%) and presence (48.6%) of HSA. The compound HSA: [Rh2(tfc)4] has had its antitumoral action in tumor-bearing Balb-c mice investigated, showing that HSA can be a drug reservoir for the rhodium complex.
rhodium; human serum albumin; binding constant; antitumor
Cinco complexos de ródio(II) de fórmula geral [Rh2(ponte)4] (ponte = acetato, propionato, butirato, trifluoroacetato e trifluoroacetamidato) tiveram suas afinidades em relação à albumina humana (HSA) determinadas por espectrofotometria, observando-se no caso dos alquilcarboxilatos uma correlação inversa com suas lipossolubilidades. A difusão dos complexos livres ou ligados à proteína para células de Ehrlich in vitro parece primordialmente governada pelo caráter hidrofóbico do complexo. O complexo [Rh2(tfc)4] apresentou afinidade pela proteína (K = 214,1), além de partição celular tanto em ausência (32,1%) como na presença (48,6%) de HSA. Desta forma, o composto HSA: [Rh2(tfc)4] teve sua ação antitumoral investigada em camundongos Balb-c portadores de ascite de Ehrlich, mostrando que a HSA pode ser um reservatório para o complexo de ródio.
Article
Effects of Human Serun Albumin in Some Biological Properties of Rhodium(II) Complexes
Breno P. Espósitoa* * e-mail: bpesposi@iq.usp.br , Elisabeth de Oliveiraa, Szulim B. Zyngierb and Renato Najjara
a Departamento de Química Fundamental. Instituto de Química, Universidade de São Paulo,
CP 26077, 05513-970, São Paulo - SP, Brazil
b Departamento de Farmacologia, Instituto de Ciências Biomédicas I, Universidade de São Paulo,
Av. Lineu Prestes, 1524, 05508-900, São Paulo - SP, Brazil
Cinco complexos de ródio(II) de fórmula geral [Rh2(ponte)4] (ponte = acetato, propionato, butirato, trifluoroacetato e trifluoroacetamidato) tiveram suas afinidades em relação à albumina humana (HSA) determinadas por espectrofotometria, observando-se no caso dos alquilcarboxilatos uma correlação inversa com suas lipossolubilidades. A difusão dos complexos livres ou ligados à proteína para células de Ehrlich in vitro parece primordialmente governada pelo caráter hidrofóbico do complexo. O complexo [Rh2(tfc)4] apresentou afinidade pela proteína (K = 214,1), além de partição celular tanto em ausência (32,1%) como na presença (48,6%) de HSA. Desta forma, o composto HSA: [Rh2(tfc)4] teve sua ação antitumoral investigada em camundongos Balb-c portadores de ascite de Ehrlich, mostrando que a HSA pode ser um reservatório para o complexo de ródio.
The affinities for human albumin (HSA) of five rhodium(II) complexes of general formula [Rh2(bridge)4] (bridge = acetate, propionate, butyrate, trifluoroacetate and trifluoroacetamidate) were determined by spectrophotometry. In the case of the alkylcarboxylates, an inverse correlation of affinity with their liposolubilities was observed. Diffusion of the free or protein-bound complexes into Ehrlich cells in vitro seems to be primarily governed by the hydrophobic character of the complex. The complex [Rh2(tfc)4] exhibited affinity towards the protein (K = 214.1) as well as cell partition both in the absence (32.1%) and presence (48.6%) of HSA. The compound HSA: [Rh2(tfc)4] has had its antitumoral action in tumor-bearing Balb-c mice investigated, showing that HSA can be a drug reservoir for the rhodium complex.
Keywords: rhodium, human serum albumin, binding constant, antitumor
Introduction
In the field of metal based cancer chemotherapy, rhodium(II) compounds have been studied as alternatives to the platinum derivatives (cisplatin and carboplatin). Both their antibacterial and antitumoral actions to human and murine cell lines have been a target of research1-3. Moreover, the biodistribution of these complexes was studied with more detail recently4, meeting the current interest in the factors that could affect the availability of these drugs to tumor cells.
The pharmacokinetics and pharmacodynamics of any drug will depend, to a large extent, on the interaction it has with human serum albumin (HSA), the most abundant plasma protein. The hypothesis of HSA acting as a "drug reservoir" for the slow release of cisplatin has already been tested under clinical conditions5-7, though previous works had questioned this possibility8.
The three-dimensional structure of HSA has been published recently and the structural studies of its interactions with a number of ligands have been performed9-11. Like any protein, HSA has several possible coordination sites for metal ions but two of them deserve special attention, the Cys34 thiol-free residue and the amino-terminal sequence with a histidine in the third position9. To our knowledge, HSA has never been co-crystallized with antitumoral platinum-group metallodrugs or with any of the high-affinity metal binding sites occupied, so the evidences for these interactions are mainly spectroscopic. Cisplatin binding to HSA has been the subject of a recent work by Sadler and coworkers12. Contrary to previous assumptions, these authors found that Cys34 is not the preferred binding site for cisplatin. A S,N macrochelate, probably involving the surface-exposed Met298 residue, accounted well for the shifts observed in the NMR spectra, and this was speculated to be the main cisplatin binding site of HSA. On the other hand the anti-arthritic gold compound auranofin, triethylphosphine (2,3,4,6-tetra-O-acetylglucopyranosato -S-) gold(I) was found to bind mainly to HSA through formation of a Cys34-S-Au bond13. The reaction of rhodium(II) carboxylates with thiols or methionine lead to the disruption of the Rh-Rh bond with formation of square-planar monomeric rhodium(II) species and to bis-adducts, respectively, whose electronic absorption spectra have different patterns from both the parent complexes14 and the albumin-bound rhodium compounds. The permanence of the characteristic Rh-Rh absorption band at ca. 540 nm after the reaction of rhodium dimers with HSA has been understood as evidence that binding to sulfur-donating residues would not be taking place and that the complex remains intact in the reactional medium15. Histidine (His) residues were earlier suggested as important coordination sites for these complexes16. This interaction may account for the displacement of the Rh-Rh absorption band from ca. 580 nm to 540 nm when rhodium complexes react with HSA15,17,18. This is in accordance with the observations made for the reaction of HSA and ruthenium(III) antitumor complexes, where imidazole binding at His residues is usually suggested19 and verified through the decreased affinity of ruthenated albumin for dyethylpyrocarbonate, a common His modificator20. A similar study with rhodium-bound HSA should prove to be valuable in that this putative His binding could be unequivocally demonstrated.
The interactions between a number of rhodium(II) complexes and HSA have already been studied by means of spectroscopic and immunologic techniques17,18. A direct dependence has been noted between the liposolubility of the metal complexes and several properties, markedly their denaturing and Trp214 fluorescence quenching abilities15, as well as their biological activities21. In the current work, the relative affinities of the complexes [Rh2(ac)4], [Rh2(pr)4], [Rh2(but)4], [Rh2(tfa)4] and [Rh2(tfc)4] (ac = acetate, pr = propionate, but = butyrate, tfa = trifluoroacetate and tfc = trifluoroacetamidate) (Figure 1) for HSA and tumor cells were studied, as well as the in vivo biological activity of the compound HSA:[Rh2(tfc)4] compared to the free complex [Rh2(tfc)4].
Experimental
Reagents
HSA fraction V, A1653, was purchased from Sigma. Stock solutions of the protein were prepared in phosphate buffer (pH 7.4) and their concentrations were determined photometrically through the relation
HSA concentration (mg mL-1) = 1.55 A28022.
In all the calculations a Mr = 66439 for HSA was assumed23. Clinical grade HSA solution (20%, Grifols) was used in the experiments with animals. Rhodium acetate was purchased from Sigma and the other rhodium complexes were synthesized through methods already described24,25. The concentrations of the complexes were determined by means of atomic emission spectroscopy with argon plasma (ICP-AES, Spectroflame Modula) calibrated with rhodium atomic absorption standard from Aldrich, using the rhodium emission line at 343.489 nm.
Plasma binding of [Rh2(ac)4]
A 20 mL sample of blood was collected from a healthy human volunteer and centrifuged at 3500 rpm for 10 min after its clotting in test tubes. Three plasma samples (1.85 mL) were incubated with 0.5 mL of 1.0 ´ 10-3 mol L-1 [Rh2(ac)4] for 0, 2 and 24 h in a water bath at 37 oC, being kept under refrigeration. Albumin was fractionated through the salting-out effect with ammonium sulfate, allowing the sedimentation for 24 h at 4 oC. Albumin fractions with 0, 2 and 24 h incubation (henceforward denominated A0, A2 and A24) were resuspended in 2% NaCl and dialyzed against 2 L of distilled water for 24 h at room temperature. The dialysis bag contents were removed with Pasteur pipettes and aliquots were transferred to a mini-gel SDS-PAGE (Bio-Rad system) to assess protein purity. Protein concentrations were determined by the Lowry method26. Rhodium concentrations in the dialyzates were determined by atomic emission spectrometry.
Binding constants
The method originally described by Stephanos27 was employed for the determination of the binding constants through previously published photometric measurements15. It was assumed only one kind of interaction between the albumin and the rhodium complex [Rh2(bridge)4], according to the following equation:
 HSA:[Rh
HSA:[Rh
(1)
for which is valid

(2)
A plot of 1/(A-A0) versus 1/L0is linear, and the binding constant K can be estimated from the ratio of the intercept to the slope. A0 is the initial protein absorbance at the monitored wavelength (~500 nm), and therefore equal to zero. A is the recorded absorption for each increment in the complex concentration (L0). This method has been employed for the determination of K for several drug-protein interactions and specifically to estimate K for the binding of cisplatin to HSA28. The titrations were carried out in a Hitachi U3000 spectrophotometer, in the visible range (350 to 750 nm) at 28 oC, as already described15.
Partition studies
The partition of the rhodium complexes into tumor cells was determined both in the presence and absence of HSA. In the blank (protein-free) experiments, HSA was replaced by distilled water, with the further steps being identical. Five 0.5 mL HSA samples (3.7 mg mL-1) were incubated with 0.1 mL of each ca. 10-3 mol L-1 rhodium(II) complex solution for 5 min at 37 oC. Triplicate 0.1 mL samples of each mixture were transferred to Eppendorf tubes. Meanwhile a suspension of ascitic Ehrlich cells was prepared. A tumor-bearing mouse was sacrificed at the moment of the experiment. 1 mL of the ascitic fluid was collected, washed twice with 10 mL PBS and centrifuged. The pellet was resuspended in 25 mL PBS and the number of intact live and dead cells was counted. 0.9 mL of this suspension was transferred to Eppendorf tubes, and vortexed vigorously for 30 s. After this, the tubes were centrifuged for 5 min at 3000 rpm, and the supernatants were separated and diluted with 2 mL of distilled water. The rhodium concentration in the diluted supernatants was determined by means of the ICP-AES technique.
Antitumor in vivo activity
Thirty Balb-c mice weighting approximately 20g were inoculated with 5 ´ 105 Ehrlich cells, divided in two groups and maintained for 24 h. During this time, 0.0167g of [Rh2(tfc)4] was diluted in water (ca. 20 mL) and this pre-solution was dripped over 1.75 mL of HSA 20% to prepare HSA:[Rh2(tfc)4]. A characteristic color change from blue to reddish was observed, indicating the binding of the complex to the protein. After 10 min, this solution was filtered and transferred to a 25 mL volumetric flask. The final volume was completed with distilled water, and the rhodium concentration was determined by ICP-AES (concentration of [Rh2(tfc)4] = 1.12 ´ 10-4 mol L-1). In these conditions, nRh/nHSA = 0.5. This assured that all of the complex molecules would be bound to the protein. After 24 h, 15 animals were treated i.p. with 0.5 mL HSA solution (control group) and the other 15 received i.p. 0.5 mL of HSA:[Rh2(tfc)4], in order to reach a 2.8 mmol kg-1 dose of [Rh2(tfc)4]. Previous assays showed HSA to be innocuous to the mice. After a further 24 h, all the mice were submitted again to the same treatments. The doses have been chosen considering that 5.6 mmol kg-1 of the drug could lead to a relevant survival percentage, at the expense of important tissue damage29. The meaningfulness of the survival patterns of the two groups (displayed in Kaplan-Meier plots) was tested by the Mantel-Haenszel method30. A similar procedure was employed to assess the activity of the unbound [Rh2(tfc)4] complex (concentration of [Rh2(tfc)4] = 1.05 ´ 10-4 mol L-1).
Results and Discussion
The whole plasma binding experiment (Table 1) showed that albumin is indeed a serological target of the rhodium complexes. The simple salting-out procedure for protein separation afforded >95% HSA, checked electrophoretically. The fact that around 50% saturation is attained with no incubation confirms previous spectroscopic evidences that this reaction is fast15. In A2, an increase to 70% saturation can be observed, indicating that for biological purposes it is reasonable to assume that rhodium complexes would bind promptly to albumin as soon as they reach the bloodstream.
It is commonly assumed that the main binding site for rhodium(II) complexes in HSA is the histidine (His) residue16,17. It was noticed that the stability of rhodium(II) complexes and His was correlated with the lipophilic nature of the metal compounds. Binding constants to the formation of adducts of the kind [Rh2(bridge)4] (His) (bridge = ac, pr) were reported as 4.05 and 4.14, respectively, the difference being attributed to the lesser solvation and therefore the increased propionate binding to His31.
However, in this work an inverse correlation was observed between the lipophilic nature of the rhodium alkylcarboxylates and the value of K for binding with HSA. (Figure 2 and Table 2).
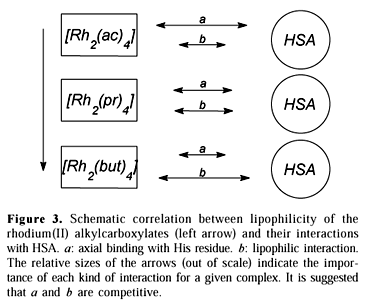
The interaction with histidine residues in the protein molecule seems to be favored for the more solvated alkylcarboxylates, which suggests an additional stabilization effect nonexistent in the reaction with the free aminoacid. This possibility has indeed been proposed31. In fact, one could consider that the overall affinity between rhodium(II) complexes and HSA results from two distinct phenomena: (a) axial coordination with His residues, and (b) hydrophobic interactions between the complex and the protein skeleton.
The affinity values obtained through spectrophotometry reflect specifically the case (a) above. It is suggested that the axial binding of the alkylcarboxylates is weaker for the more lipophilic complexes precisely because of the larger contribution of the hydrophobic interactions (case b). A glance at the scheme proposed in Figure 3 is elucidative. The relative sizes of the arrows indicate the importance of each interactive phenomenon for a given complex.
Even though exhibiting high hydrosolubility, the His coordination with [Rh2(tfa)4] should be of importance, given the higher hardness of the metal center. This effect apparently is lost when CF3COO bridging ligands are substituted by the correlated amidate, leading to a better p-donator complex to the axial ligand32.
The magnitude of the obtained binding constants is in accordance with the reported results for the interaction of cisplatin with HSA (K = 852)28. These affinities are not as high as those described for other drugs (K ~ 105-107) and thus possibly they are less of a hindrance to release from the protein. So it seems reasonable that the complexes transfer to the target cells by passive diffusion governed by lipophilicity (HSA-bound drugs are not available to cross the cell membrane33).
Turning to the partition results (Table 3), it can be seen that indeed there are different absorptions by the cell suspension correlated to the lipophilicity of the rhodium alkylcarboxylates.
In the partition into cells, the complex [Rh2(tfa)4] behaved similarly to [Rh2(ac)4], even though it is more lipophilic. Considering its already described affinity to HSA15, a lesser transference of [Rh2(tfa)4] from the protein to the cells would not be surprising. However, the explanation for the relatively small partition of the unbound complex into cells seems to involve other phenomena at the membrane level, which deserve further studies. The complex [Rh2(tfc)4] diffused into cells more readily than [Rh2(ac)4] (ca. 20%) when bound to the protein but this trend is reversed for the free complexes. A more favorable release from the protein could be carried out by the trifluoroacetamidate ligands.
The survival profiles of the mice treated with both HSA-bound and unbound [Rh2(tfc)4] complex are presented in Figures 4 and 5. [Rh2(tfc)4] has been elected to in vivo studies to give continuity to our previous research on the antitumoral action of rhodium(II) amidates29,34.
Some of the mice treated with HSA:[Rh2(tfc)4] which survived after the period of investigation (ca. 30 days) exhibited solid tumors, a physiologic response to the tumoral focus caused either by the small amount of inoculated tumor cells or by a therapeutic effect of the drug. The values of the Mantel-Haenszel test (6.8 and 19.4, respectively for HSA:[Rh2(tfc)4] and [Rh2(tfc)4]) show that the differences in the survival profiles between treated and untreated groups are statistically meaningful (p < 0.01).
These results, together with the above considerations relating to the affinity of the rhodium complexes with HSA and/or intact cells, indicate that albumin can be a drug reservoir for [Rh2(tfc)4], and probably for the other rhodium complexes as well, by means of capturing it in the bloodstream after injection and transferring it slowly to the tumor cell by passive diffusion. The possibility for some sort of active transport mechanism through the membrane, however, can not be ruled out with these results.
Acknowledgments
The authors would like to thank Prof. Dr. A. R. Souza (Universidade Federal de Goiás) for helpful comments and suggestions throughout this work, and Dr. A. C. Magnanelli and Mrs. Teresa Victorio (Fundação Pró-Sangue, Hemo-centro de São Paulo) for the the SDS-PAGE experiments and for a gift of clinical grade HSA. The authors acknowledge financial support from CNPq and FAPESP (Brazilian agencies).
Received: December 13, 1999
Published on the web: August 31, 2000
FAPESP helped in meeting the publication costs of this article.
- 1. Bien, M.; Pruchnik, F. P.; Seniuk, A.; Lachowicz, T. M.; Jakimowicz, P. J. Inorg. Biochem. 1999, 73, 49.
- 2. Sinisterra, R. D.; Shastri, V. P.; Najjar, R.; Langer, R. J. Pharm. Sci. 1999, 88, 574.
- 3. Pruchnik, F. P.; Kluczewska, G.; Wiczok, A.; Mazurek, U.; Wilczok, T. J. Inorg. Biochem. 1997, 65, 25.
- 4. Souza, A. R.; Najjar, R.; Oliveira, E.; Zyngier, S. B. Metal-Based Drugs 1997, 4, 39.
- 5. Vreeburg, G. C. M.; Stell, P. M.; Holding, J. D.; Lindup, W. E. J. Laryngol. Otol. 1992, 106, 832.
- 6. Holding, J. D.; Lindup, W. E.; Bowdler, D. A.; Siodlak, M. Z.; Stell, P. M. Br. J. Clin. Pharmac. 1992, 32, 173.
- 7. Holding, J. D.; Lindup, W. E.; Van Laer, C.; Vreeburg, G. C. M.; Schilling, V.; Wilson, J. A.; Stell, P. M. Br. J. Clin. Pharmac. 1992, 33, 75.
- 8. Cole, W. C.; Wolf, W. Chem.-Biol. Interactions 1980, 30, 223.
- 9. Carter, D. C.; Ho, J. X. Adv. Prot. Chem. 1994, 45, 153.
- 10. Sugio, S.; Kashima, A.; Mochizuki, S.; Noda, M.; Kobayashi, K. Protein Eng. 1999, 12, 439.
- 11. Curry, S.; Brick, P.; Franks, N. P. Biochim. Biophys. Acta 1999, 1441, 131.
- 12. Ivanov, A. I.; Christodoulou, J.; Parkinson, J. A.; Barnham, K. J.; Tucker, A.; Woodrow, J.; Sadler, P. J. Biol. Chem. 1998, 273, 14721.
- 13. Shaw III, C. F. Chem. Rev. 1999, 99, 2589.
- 14. Pneumatikakis, G.; Psaroulis, P. Inorg. Chim. Acta 1980, 46, 97.
- 15. Espósito, B. P.; Faljoni-Alário, A.; Menezes, J. F. S.; Brito, H. F.; Najjar, R. J. Inorg. Biochem. 1999, 75, 55.
- 16. Dennis, A. M.; Howard, R. A.; Bear, J. L. Inorg. Chim. Acta 1982, 66, L31.
- 17. Trynda, L.; Pruchnik, F. P. J. Inorg. Biochem. 1995, 58, 69.
- 18. Trynda-Lemiesz, L.; Pruchnik, F. P. J. Inorg. Biochem. 1997, 66, 187.
- 19. Vilchez, F. G.; Vilaplana, R.; Blasco, G.; Messori, L. J. Inorg. Biochem. 1998, 71, 45.
- 20. Messori, L.; Orioli, P.; Vullo, D.; Alessio, E.; Iengo, E. Eur. J. Biochem. 2000, 267, 1206.
- 21. Howard, R. A.; Sherwood, E.; Erck, A.; Kimball, A. P.; Bear, J. L. J. Med. Chem. 1977, 20, 943.
- 22. Harris, E. L. V.; Angal, S. Protein Purification Methods - A Practical Approach, IRL Press, Oxford, 11, 1989.
- 23. Kragh-Hansen, U. Dan. Med. Bull. 1990, 37, 57.
- 24. Nothenberg, M. S.; Takeda, G. K. F.; Najjar, R. J. Inorg. Biochem. 1991, 42, 217.
- 25. Dennis, A. M.; Howard, R. A.; Lançon, D.; Kadish, K. M.; Bear, J. L. J. Chem. Soc., Chem. Commun. 1982, 399.
- 26. Lowry, O. H.; Rosebrough, N. J.; Farr, A. L.; Randall, R. J. J. Biol. Chem. 1951, 193, 265.
- 27. Stephanos, J. J. J. Inorg. Biochem. 1996, 62, 155.
- 28. Neault, J. F.; Tajmir-Riahi, H. A. Biochim. Biophys. Acta 1998, 1384, 153.
- 29. Espósito, B. P.; Zyngier, S. B.; Najjar, R.; Paes, R. P.; Ueda, S. M. Y.; Barros, J. C. A. Metal-Based Drugs 1999, 6, 17.
- 30. Matthews, D. E.; Farewell, V. T. Using and Understanding Medical Statistics, 2nd ed, Karger, Basel, 79-87, 1988.
- 31. Das, K.; Simmons, E. L.; Bear, J. L. Inorg. Chem. 1977, 16, 1268.
- 32. Best, S. P.; Chandley, P.; Clark, R. J. H.; McCarthy, S.; Hursthouse, M. B.; Bates, P. A. J. Chem. Soc. Dalton Trans. 1989, 581.
- 33. Benet, L. Z.; Mitchell, J. R.; Sheiner, L. B. Pharmacokinetics: the Dynamics of Drug Absorption, Distribution and Elimination, In Goodman and Gilman's pharmacological basis of therapeutics, 8th ed, Gilman, A. G.; Rall, T. W.; Nies, A. S.; Taylor, P., eds, Pergamon, New York, 12, 1990.
- 34. Espósito, B. P.; Zyngier, S. B.; Souza, A. R.; Najjar, R. Metal-Based Drugs 1997, 4, 133.
Publication Dates
-
Publication in this collection
15 Jan 2001 -
Date of issue
Oct 2000
History
-
Received
13 Dec 1999 -
Accepted
31 Aug 2000