Abstracts
This paper provides an evaluation of the population structure, phenotype and genetic trends of registered Gyr herd cattle in northeast Brazil. The study provides important baseline information for the management, conservation and potential population expansion of this economically and culturally important cattle breed. Pedigree data were analyzed for individuals born between 1964 and 2006. Body weight values were adjusted to 205 and 365 days of age for animals born between 1978 and 2006. Phenotypic change of zebu Gyr in northeast Brazil is solely due to environmental improvement. However, there is potential for artificial selection for weight gain in young cattle. Effective population size decreased during the 1990s and the average inbreeding coefficient increased during the studied period. An increase of the effective population size of Gyr in northeast Brazil is strongly recommended, along with an increase in the management of the mating process to prevent inbreeding and to maintain the genetic variability of the breed.
effective size of the population; generation interval; genetic trend; inbreeding
Com o intuito de fornecer subsídios para programas de conservação, seleção e expansão da raça Gir no Nordeste do Brasil, objetivou-se avaliar o histórico do rebanho Gir registrado no nordeste brasileiro, com base na sua estrutura populacional e no progresso genético e fenotípico de características de desenvolvimento ponderal. Foram utilizadas informações de pedigree de animais nascidos no período de 1964 a 2006 e dados dos pesos ajustados aos 205 e 365 dias de idade de bovinos nascidos de 1978 a 2006. O progresso genético para a raça no Nordeste foi ocasionado exclusivamente pelo melhoramento ambiental. O tamanho efetivo da população reduziu a partir da década de 90, e o coeficiente de endogamia médio aumentou durante o período estudado. É imprescindível que o tamanho efetivo da raça Gir do nordeste seja ampliado e que haja maior controle de acasalamentos entre indivíduos aparentados, para prevenção da endogamia e conservação da variabilidade genética e viabilidade da raça.
endogamia; intervalo de geração; tamanho efetivo; tendência genética
PAPERS
ANIMAL REPRODUCTION
History of registered Gyr breed in Brazilian Northeast: population structure and genetic improvement of growth traits
Histórico da raça Gir registrada no Nordeste Brasileiro: melhoramento genético em características de crescimento e estrutura populacional
Carlos Henrique Mendes MalhadoI,1 1 Autor para correspondência. ; Paulo Luiz Souza CarneiroI; Ana Claudia Mendes MalhadoII; Jorge André Matias MartinsIII; Raimundo Martins FilhoIV; Riccardo BozziV
IDepartamento de Ciências Biológicas, Universidade Estadual do Sudoeste da Bahia (UESB), 45200-000, Jequié, BA, Brasil. E-mail: carlosmalhado@gmail.com
IIDepartamento de Engenharia Agrícola e Meio Ambiente, Universidade Federal de Viçosa (UFV), Viçosa,MG, Brasil
IIIDepartamento de Zootecnia, Universidade Federal do Ceará (UFC), Fortaleza, CE, Brasil
IVProfessor Visitante Sênior Nacional, UFC, Cariri, CE, Brasil
VDepartment of Agricultural Biotechnology, Università di Firenze, Firenze, Itália
ABSTRACT
This paper provides an evaluation of the population structure, phenotype and genetic trends of registered Gyr herd cattle in northeast Brazil. The study provides important baseline information for the management, conservation and potential population expansion of this economically and culturally important cattle breed. Pedigree data were analyzed for individuals born between 1964 and 2006. Body weight values were adjusted to 205 and 365 days of age for animals born between 1978 and 2006. Phenotypic change of zebu Gyr in northeast Brazil is solely due to environmental improvement. However, there is potential for artificial selection for weight gain in young cattle. Effective population size decreased during the 1990s and the average inbreeding coefficient increased during the studied period. An increase of the effective population size of Gyr in northeast Brazil is strongly recommended, along with an increase in the management of the mating process to prevent inbreeding and to maintain the genetic variability of the breed.
Key words: effective size of the population, generation interval, genetic trend, inbreeding.
RESUMO
Com o intuito de fornecer subsídios para programas de conservação, seleção e expansão da raça Gir no Nordeste do Brasil, objetivou-se avaliar o histórico do rebanho Gir registrado no nordeste brasileiro, com base na sua estrutura populacional e no progresso genético e fenotípico de características de desenvolvimento ponderal. Foram utilizadas informações de pedigree de animais nascidos no período de 1964 a 2006 e dados dos pesos ajustados aos 205 e 365 dias de idade de bovinos nascidos de 1978 a 2006. O progresso genético para a raça no Nordeste foi ocasionado exclusivamente pelo melhoramento ambiental. O tamanho efetivo da população reduziu a partir da década de 90, e o coeficiente de endogamia médio aumentou durante o período estudado. É imprescindível que o tamanho efetivo da raça Gir do nordeste seja ampliado e que haja maior controle de acasalamentos entre indivíduos aparentados, para prevenção da endogamia e conservação da variabilidade genética e viabilidade da raça.
Palavras-chave: endogamia, intervalo de geração, tamanho efetivo, tendência genética.
INTRODUCTION
Research and management of domestic animal populations have been recently benefited by the application of molecular genetics techniques (PAIVA et al., 2005; ALBUQUERQUE et al., 2006). These methods have provided significant insights into the genetic structure of economically important species. Despite these advances there is still a need for information about pedigree in order to identify temporal trends and to evaluate inbreeding, confirmation of pedigree, effective size, generation interval, genetic diversity and several other important population parameters (FARIA et al., 2004; CARNEIRO et al. 2009; MALHADO et al., 2008a,b; MARTÍNEZ et al., 2008). Studies utilizing historical pedigree records can identify and uncover important circumstances that have influenced the genetic history of a population (VALERA et al., 2005), moreover, some population parameters that are strongly dependent on management and mating systems have significant impacts on genetic variability.
In addition to productivity, the major concern of cattle breeders is the monitoring of genetic population health. Knowledge of genetic changes in a population is of crucial importance, not only as a guide for genetic interventions, but also to evaluate the results of the particular selection program adopted. However, it should also be noted that the success of genetic improvement programs is critically dependent on the knowledge of several factors that potentially interfere on selection, such as effective size of the population, generation interval and genetic variability (MALHADO et al. 2008a). Moreover, knowledge of population structure combined with information on genetic change in the population can guide future management actions, allowing the development of strategies that promote the genetic improvement and adaptation of a breed in a particular region (MALHADO et al. 2008b).
The objective of this study was to evaluate the history of registered population Gyr breed cattle in northeast Brazil. Specifically, there was analyzed population structure and changes in a range of body development parameters with the aim of providing information to support artificial selection, conservation and possible population expansion of Gyr breed in northeast Brazil.
MATERIAL AND METHODS
Growth data of Gyr was obtained from the Brazilian Association of Zebu Breeders. Pedigree information was available for animals born between 1964 and 2006, sons of 586 sires, from Brazilian Northeast. Data on 3,190 and 2,160 weight values adjusted to 205 (W205) and 365 (W365) days of age, respectively, were analyzed for animals born between 1978 and 2006.
The derivate free maximum likelihood methodology, using animal models, was applied to obtain the variances and estimate the genetic values (BOLDMAN et al., 1995). For W205, a model was applied which included the additive genetic random effects (direct and maternal), maternal permanent environment, the co-variable age of cow at parity (linear and quadratic effects) and the fixed effects of contemporary groups, admitting the co-variance among direct and maternal genetic effects as null. For W365 the same fixed effects and co-variables were considered, with only the direct additive genetic effects as random.
Estimates of genetic and phenotypic trends were obtained by weighing average linear regression of dependent variables (genetic values and observed weights) against year of birth, using SAS (SAS Institute, 2001). Non-parametric regression through articulated polynomials ("spline") was used to describe the observed trends in the plots.
Contemporary groups were composed of animals of the same sex, time (season and year of birth), farm and feed management regime. Groups containing less than three animals were excluded from analysis.
Pedigree analysis and parameter estimates based on gene origin probability, inbreeding, and generation interval were performed using the ENDOG program (GUTIÉRREZ & GOYACHE, 2005).
The effective size of the population (Ne) was estimated, according to the following equation: , where: Nm is the number of males and Nf is the number of females that reproduce.
, where: Nm is the number of males and Nf is the number of females that reproduce.
The effective number of founders represents the number of animals which, under random mating, would produce the same genetic variability as observed in the studied population. The effective number of ancestors represents the minimum number of animals (founders or not) that are necessary to explain the total genetic diversity of the study population (GUTIÉRREZ & GOYACHE, 2005).
The Wright's F statistics (1938) - Fit, Fst and Fis - were calculated by ENDOG program (GUTIÉRREZ & GOYACHE, 2005), where: Fit is the average inbreeding coefficient of population; Fst represents the expected average inbreeding coefficient, estimated in hypothetic populations produced by parents random mating, within each period, and; Fis expresses the deviation generated from actual mating.
RESULTS AND DISCUSSION
Means and standard deviations for the adjusted weights at 205 and 365 days of age were 138.7±35.6 and 197.0±44.5kg respectively. The heritability coefficients (direct effect) were 0.12±0.05 and 0.27±0.07 for W205 and W365, respectively (Table 1). The maternal heritability coefficient for W205 was low and equal to 0.06±0.07. These results are similar to those reported in the literature: LÔBO et al. (2000) and LIRA et al. (2008) in revisions papers which respectively reported average heritability (direct effects) of 0.30 and 0.28 for W205 and 0.37 and 0.31 for W365.
The values of heritability (direct effect) for W365 indicate high direct genetic variability and suggest that genetic gain is possible through selection. Conversely, the maternal and direct heritability coefficients for W205 were low implying that selection for these effects would be problematic.
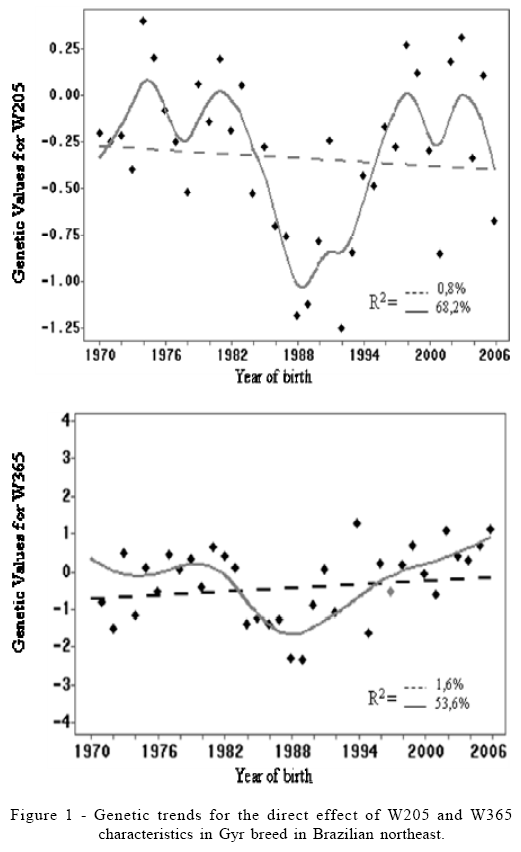
No significant genetic trends, estimated by linear regression, were apparent for the characteristics of W205 and W365. The articulated polynomial regressions identify the 1980's as the period during which the animals presented the lowest genetic values for both characteristics (Figure 1). The absence of genetic gain in growth characteristics could be explained by the fact that, in recent decades, most of the breeders became convinced of Gyr potential to produce milk and they performed artificial selection with the aim to improve this characteristic in their herds.
EUCLIDES FILHO et al. (2000), in their study of weight gain in Gyr cattle throughout Brazil during the period between 1962 to 1994, reported that the genetic gains of the breed were low, particularly, for the post-weaning weight gain.
The maternal genetic trend for W205 was not significant (P>0.05). According to EUCLIDES FILHO et al. (2000), genetic trends close to zero and even negative are not uncommon in literature, mainly where studies have used data from commercial herds where selection criteria are not well defined or uniform.
BIFFANI et al. (1999) suggest that in the rearing conditions typical of the northeast of Brazil, the breeders primarily select animals based on their anatomical and racial characteristics rather than on productivity. This fact, in addition to the low utilization of genetically evaluated animals and the widespread practice of selecting for milk yield, may explain the absence of a genetic trend for both evaluated characteristics. Other zebu breeds in Brazilian northeast showed genetic progress in recent years. Specifically, MALHADO et al. (2008b) report genetic gain in body development characteristics of a zebu population since 1994.
Phenotypic trends (linear regression) for both ages were significant (P<0.05) and equal to 0.62 and 1.05kg per year for W205 and W365, respectively (Figure 2). These results show that these characteristics are improving over time, implying that breeders have obtained significant annual phenotypic gains up to one year of age. Moreover, it should be noted that this progress is exclusively due to environmental improvement since the genetic contribution was null.
The estimates for generation interval (and its standard deviation) for the four breeding pathways were 8.6±0.8 (sires of sires), 8.2±0.2 (sires of dam), 7.8±0.4 (dam of sires) and 7.5±0.2 (dam of dam). The average generation interval was 7.9±0.1 years. The values for the four breeding pathways are slightly lower to those reported by MALHADO et al. (2008b) and higher than the intervals observed by VERCESI FILHO et al. (2002) and FARIA et al. (2002) for registered Nelore, Tabapuã and Nelore, respectively.
The optimization of average generation interval is of fundamental importance for genetic improvement programs since a reduction in the rate of genetic gain it is the main consequence of a long generation interval. Generation interval can be decreased by the use of evaluated young bulls, fast sire substitution with a decrease of the age to the first parity of dams.
Lower generation intervals may be reachable, as demonstrated by the experimental herd at the Animal Science Experimental Station of Sertãozinho- SP in Brazil, where the father-son intervals were 3.66 and 3.74 years for Nelore and Guzerat breeds, respectively (RAZOOK et al., 1993).
The major number of births occurred in the 1980's, with a maximum of 368 in 1986 (Figure 3). In this same decade there was a big rise of effective population size and of the number of sires in reproduction, reaching the maximum in 1991, with 81 reproducing bulls and an effective population size of 250. Moreover, the 1980's was the period with the lowest average genetic value. This trend was possibly driven by a higher utilization of bulls with low genetic value for both characteristics.
FRANKHAM (1995) suggests that an effective population size of 500 animals is needed to prevent the loss of genetic diversity through genetic drift and to maintain an evolutionarily flexible population. However, only effective size of at least 50 are required to prevent inbreeding depression.
The average inbreeding coefficient and the percentage of inbred animals in the population increased across generations (Table 2). The mating of relatives results in inbreeding depression, decrease of individual's performance, mainly in terms of reproduction (WALL et al., 2005) and general health (MIGLIOR et al., 1995).
The reference population, in which both parents are known, contained 5,945 individuals, while 2,952 animals had one or both parents unknown. The effective number of founders (fe) was 260, and the expected increase of inbreeding caused by the founders' unbalanced contribution was 0.09%. The genetic contribution of the 10, 30 and 171 most influential ancestors (founders or not) explained 14.9, 26.8 and 50% of the genetic variability of the whole population respectively. REIS FILHO (2006) studying the dairy Gyr breed in Brazil reported an effective number of founders of 146 and an expected increase of inbreeding of 0.34%. Furthermore he related that only 28 founders were responsible for the origin of 50% of population's genes. This implies that the milk yield-selected population in Brazil represents only part of the population, i.e. several ancestors and founders of the Gyr breed did not contribute to the formation of the "dairy" Gyr cattle.
Wright's F statistics, for the overall population, represented by Fis, Fst and Fit, were -0.0289, 0.0028 and 0.0008, respectively. Fis, which measures the deviation from random mating, was negative and close to zero implying a small amount of mating between subpopulations, which helped to minimize inbreeding. According to MALHADO et al. (2008b) a negative Fis value is probably caused by the use of artificial insemination and sires exchange - practices used for genetic improvement of the breed and connectivity among herds. A low value of Fst can also indicate a lack of population structure, due to the fact that 100% of studied herds use external sires or artificial insemination.
CONCLUSION
Phenotypic change of zebu Gyr in northeast Brazil is solely due to environmental improvement. However, there is a potential for artificial selection for weight gain in young cattle. Effective population size decreased during the 1990s and the average inbreeding coefficient increased during the studied period. An increase of the effective population size of Gyr in northeast Brazil is strongly recommended, along with an increase in the management of the mating process to prevent inbreeding and to maintain the genetic variability of e breed.
ACKNOWLEDGEMENTS
We would like to thank the CNPQ and CAPES for supporting the work of CHMM and RMF, respectively, and the Brazilian Association of Zebu Farmers for access to their data. We also thank Dr Richard Ladle (Oxford University) for useful comments and proof-reading.
Received 05.07.09
Approved 05.06.10
Returned by the author 06.17.10
CR- 1967
- ALBUQUERQUE, M.S.M. et al. Variabilidade genética em búfalos estimada por marcadores RAPD. Pesquisa Agropecuária Brasileira, Brasília, v.41, p.623-628, 2006. Available from: <http://www.scielo.br/scielo.php?pid=S0100-204X2006000400011&script=sci_arttext>. Acessed: May 5, 2010. doi: 10.1590/S0100-204X2006000400011.
- BIFFANI, S. et al. Fatores ambientais e genéticos sobre o crescimento ao ano e ao sobreano de bovinos Nelore, criados no Nordeste do Brasil. Revista Brasileira de Zootecnia, v.28, p.468-473, 1999.
- BOLDMAN, K.G. et al. A manual for use of MTDFREML: a set of programs to obtain estimates of variances and covariances [DRAFT]. Lincoln: Agricultural Research Service, 1995. 120p.
- CARNEIRO, P.L.S. et al. A raça Indubrasil no Nordeste brasileiro. Melhoramento e estrutura populacional. Revista Brasileira de Zootecnia, v.38, n.12, p.2327-2334, 2009. Available from: <http://www.scielo.br/scielo.php?pid=S1516-35982009001200006&script=sci_arttext>. Acessed: May 5, 2010. doi: 10.1590/S1516-35982009001200006.
- EUCLIDES FILHO, K. et al. Tendência genética na raça Gir. Pesquisa Agropecuária Brasileira, v.35, p.787-791, 2000. Available from: <http://www.scielo.br/scielo.php?script=sci_arttext&pid=S0100-204X2000000400016>. Acessed: May 5, 2010. doi: 10.1590/S0100-204X2000000400016.
- FARIA, F.J.C. et al. Estrutura populacional da raça Nelore Mocho. Arquivo Brasileiro de Medicina Veterinária e Zootecnia, v.54, p.501-509, 2002. Available from: <http://www.scielo.br/scielo.php?script=sci_arttext&pid=S0102-09352002000500008&lng=en&nrm=iso&tlng=pt>. Acessed: May 5, 2010. doi: 10.1590/S0102-09352002000500008.
- FRANKHAM, R. Conservation genetics. Annual Review of Genetics, v.29, p.305-327, 1995.
- GUTIÉRREZ, J.P.; GOYACHE, F. A note on ENDOG: a computer program for analysing pedigree information. Journal of Animal Breeding and Genetics, v.122, p.172-176, 2005. Available from: <http://www.ucm.es/info/prodanim/html/JP_Web_archivos/EN_Us_G_.pdf>. Acessed: May 5, 2010. doi: 10.1111/j.1439-0388.2005.00512.x.
- LIRA, T. de. et al. Parâmetros genéticos de características produtivas e reprodutivas em zebuínos de corte (revisão). Ciência Animal Brasileira, v.9, p.1-22, 2008. Available from: <http://www.revistas.ufg.br/index.php/vet/article/viewArticle/3655>. Acessed: May 5, 2010.
- LÔBO, R.N.B. et al. Average estimates of genetic parameters for beef and dairy cattle in tropical regions. Animal Breeding Abstracts, v.68, p.433-462, 2000.
- MALHADO, C.H.M. et al. Melhoramento e estrutura populacional de bubalinos da raça Mediterrâneo no Brasil. Pesquisa Agropecuária Brasileira, v.43, p.215-220, 2008a. Available from: <http://www.scielo.br/scielo.php?script=sci_arttext&pid=S0100-204X2008000200009&lng=end_0100-6916&nrm=iso&tlng=end_0100-6916>. Acessed: May 5, 2010. doi: 10.1590/S0100-204X2008000200009
- MALHADO, C.H.M. et al. Progresso genético e estrutura populacional do rebanho Nelore no Estado da Bahia. Pesquisa Agropecuária Brasileira, v.43, p.1169-1169, 2008b. Avaliable from: <http://www.scielo.br/scielo.php?pid=S0100-204X2009000700010&script=sci_arttext>. Acessed: May 5, 2010. doi: 10.1590/S0100-204X2009000700010.
- MARTÍNEZ, R.A. et al. Genetic variability in Colombian Creole cattle populations estimated by pedigree information. Journal Animal Science, v.86, p.545-552, 2008. Avaliable from: <http://jas.fass.org/cgi/content/short/86/3/545>. Acessed: May 5, 2010. doi: :10.2527/jas.2007-0175.
- MIGLIOR, F. et al. Nonadditive genetic effects and inbreeding depression for somatic cell counts of Holstein cattle. Journal Dairy Science v.78, p.168-1173, 1995. Avaliable from: <http://www.ncbi.nlm.nih.gov/pubmed/7622727>. Acessed: May 5, 2010.
- PAIVA, S.R. et al. Genetic variability of the main Brazilian hair sheep breeds. Pesquisa Agropecuária Brasileira, v.40, p.887-893, 2005. Avaliable from: <http://www.scielo.br/scielo.php?pid=S0100-204X2005000900008&script=sci_abstract>. Acessed: May 5, 2010. doi: 10.1590/S0100-204X2005000900008.
- RAZOOK, A.G. et al. Intensidades de seleção e repostas diretas e correlacionadas em 10 anos de progênies de bovinos das raças Nelore e Guzerá selecionadas para peso pós desmame. Boletim Industria. Animal, v.50, p.147-163, 1993.
- REIS FILHO, J.C. Endogamia na raça Gir 2006. 50f. Dissertação (Mestrado em Genética e Melhoramento Animal) - Universidade Federal de Viçosa, Viçosa, MG.
- SAS INSTITUTE. SAS: user guide: versão 8.2. Cary, 2001. 956p.
- VALERA, M. et al. Pedigree analysis in the Andalusian horse: population structure, genetic variability and influence of the Carthusian strain. Livestock Production Science, v.95, p.57-66, 2005. Avaliable from: <http://www.ucm.es/info/prodanim/html/JP_Web_archivos/Valera.pdf>. Acessed: May 5, 2010. doi:10.1016/j.livprodsci.2004.12.004.
- VERCESI FILHO, A.E. et al. Estrutura populacional do rebanho Tabapuã registrado no Brasil. Arquivo Brasileiro de Medicina Veterinária e Zootecnia, v.54, p.609-617, 2002. Avaliable from: <http://www.scielo.br/scielo.php?pid=S0102-09352002000600009&script=sci_arttext>. Acessed: May 5, 2010. doi: 10.1590/S0102-09352002000600009.
- WALL, E. et al. Impact of nonadditive genetic effects in the estimation of breeding values for fertility and correlated traits. Journal Dairy Science, v.88, p.376-385, 2005. Avaliable from: <http://www.journalofdairyscience.org/article/S0022-0302(05)72697-7/abstract>. Acessed: May 5, 2010. doi:10.3168/jds.S0022-0302(05)72697-7.
Publication Dates
-
Publication in this collection
23 July 2010 -
Date of issue
June 2010
History
-
Received
07 May 2009 -
Accepted
06 May 2010