Abstracts
Maize rayado fino virus (MRFV), present in various countries in Latin America, has shown similarities to corn streak virus that occurs in Brazil, regarding pathogenic, serological and histological characteristics. In the current report both virus were molecularly compared to confirm the similarities between them. MRFV was identified by nucleic acid hybridization in samples of maize tissues exhibiting symptoms of "corn stunt" disease, collected from two Brazilian States - São Paulo and Minas Gerais. The coat protein gene and 3'non-translated region of MRFV were amplified from infected tissues by reverse transcription-polymerase chain reaction (RT-PCR) using MRFV-specific primers, and were characterized by nucleotide sequence and phylogenetic analysis of the cloned PCR products. Phylogenetic analysis of the relationships between the Brazilian isolates and isolates obtained from Latin America and the United States reveals a close relationship to isolates from Brazil, Peru and Bolivia. Results support the proposal that the Brazilian corn streak virus be regarded as an isolate of MRFV and provide evidence for the presence of MRFV in "corn stunt' disease in Brazil.
MRF virus; corn viruses; phytoplasma; corn stunt
O vírus do rayado fino do milho (MRFV), presente em vários países da América Latina, tem mostrado semelhança ao vírus da risca do milho que ocorre no Brasil, em relação a características patogênicas, sorológicas e histológicas. No presente trabalho, ambos os vírus foram comparados molecularmente, visando confirmar a similaridade entre os mesmos. O MRFV foi identificado por hibridização de ácido nucléico em amostras de tecido que apresentavam sintomas de enfezamento, coletadas nos estados de São Paulo e Minas Gerais. O gene da capa protéica e a região 3' não traduzida do MRFV foram amplificados, a partir de tecidos infectados, através da transcrição reversa por reação em cadeia da polimerase (RT-PCR), usando-se os iniciadores específicos para o vírus, obtendo-se a seqüência nucleotídica e a análise filogenética dos produtos de PCR clonados. A análise filogenética relacionando isolados brasileiros, da América Latina e dos Estados Unidos revela a estreita relação entre isolados do Brasil, do Peru e da Bolívia. Os resultados demonstraram que o vírus da risca do milho, presente no Brasil, pode ser considerado como um isolado do MRFV e também evidenciaram a presença do MRFV associado à doença conhecida como enfezamento do milho que ocorre no Brasil.
vírus da risca; viroses do miho; fitoplasma; enfezamento do milho
NOTE
Molecular confirmation of Maize rayado fino virus as the Brazilian corn streak virus
Confirmação molecular do 'Maize rayado fino virus' como vírus da estria do milho
Rosemarie Wahnbaeck HammondI; Ivan Paulo BedendoII,* * Corresponding author < ipbedend@esalq.usp.br>
IUnited States Department of Agriculture USDA-BARC West - Beltsville, MD 20705, USA
IIESALQ/USP - Depto. de Entomologia, Fitopatologia e Zoologia Agrícola, C.P. 09 - 13418-900 - Piracicaba, SP - Brasil
ABSTRACT
Maize rayado fino virus (MRFV), present in various countries in Latin America, has shown similarities to corn streak virus that occurs in Brazil, regarding pathogenic, serological and histological characteristics. In the current report both virus were molecularly compared to confirm the similarities between them. MRFV was identified by nucleic acid hybridization in samples of maize tissues exhibiting symptoms of "corn stunt" disease, collected from two Brazilian States - São Paulo and Minas Gerais. The coat protein gene and 3'non-translated region of MRFV were amplified from infected tissues by reverse transcription-polymerase chain reaction (RT-PCR) using MRFV-specific primers, and were characterized by nucleotide sequence and phylogenetic analysis of the cloned PCR products. Phylogenetic analysis of the relationships between the Brazilian isolates and isolates obtained from Latin America and the United States reveals a close relationship to isolates from Brazil, Peru and Bolivia. Results support the proposal that the Brazilian corn streak virus be regarded as an isolate of MRFV and provide evidence for the presence of MRFV in "corn stunt' disease in Brazil.
Key words: MRF virus, corn viruses, phytoplasma, corn stunt
RESUMO
O vírus do rayado fino do milho (MRFV), presente em vários países da América Latina, tem mostrado semelhança ao vírus da risca do milho que ocorre no Brasil, em relação a características patogênicas, sorológicas e histológicas. No presente trabalho, ambos os vírus foram comparados molecularmente, visando confirmar a similaridade entre os mesmos. O MRFV foi identificado por hibridização de ácido nucléico em amostras de tecido que apresentavam sintomas de enfezamento, coletadas nos estados de São Paulo e Minas Gerais. O gene da capa protéica e a região 3' não traduzida do MRFV foram amplificados, a partir de tecidos infectados, através da transcrição reversa por reação em cadeia da polimerase (RT-PCR), usando-se os iniciadores específicos para o vírus, obtendo-se a seqüência nucleotídica e a análise filogenética dos produtos de PCR clonados. A análise filogenética relacionando isolados brasileiros, da América Latina e dos Estados Unidos revela a estreita relação entre isolados do Brasil, do Peru e da Bolívia. Os resultados demonstraram que o vírus da risca do milho, presente no Brasil, pode ser considerado como um isolado do MRFV e também evidenciaram a presença do MRFV associado à doença conhecida como enfezamento do milho que ocorre no Brasil.
Palavras-chave: vírus da risca, viroses do miho, fitoplasma, enfezamento do milho
INTRODUCTION
Corn stunt, a disease identified in maize in the Southern US, and Central and South America, is considered to be a syndrome induced by a complex constituted by a phytoplasma, a spiroplasma, and a virus (Maize rayado fino virus-MRFV), which can all be transmitted simultaneously by their leafhopper vector, mainly Dalbulus maidis De Long & Wolcott (Gamez, 1973; Nault & Bradfute, 1979; Nault et al., 1980). MRFV is the type member of the genus Marafivirus, family Tymoviridae, which includes Oat blue dwarf virus (Gamez & Leon, 1988). The sequence of MRFV has already been reported (Hammond & Ramirez, 2001).
The disease has been detected in all Central American countries, including Costa Rica, El Salvador, Guatemala, Honduras, Nicaragua, Panama, as well as Mexico, Colombia, Peru, Venezuela, Uruguay, the United States, and Brazil (Gamez, 1980; Gamez et al., 1979; Kogel et al., 1996). In Brazil, rayado fino is known as "corn streak" (Costa et al., 1971) and the virus is vectored by D. maidis (Kitajima et al., 1976). In Central America, local maize lines were usually tolerant to MRFV, but the virus caused significant losses in newly introduced varieties. In Brazil, the virus is important in the "safrinha" system, and together with corn stunt spiroplasma and maize bushy stunt phytoplasma, MRFV formed a complex referred to as "corn stunt" causing significant losses (Massola et al., 1999).
The molecular epidemiology of MRFV was previously examined by performing a phylogenetic analysis of the coat protein gene of fourteen isolates collected from several countries in Latin America and the United States (Hammond et al., 1997). The nucleotide sequence was highly conserved among the isolates, yet three phylogenetic clusters were revealed. The current report describes the characterization of the coat protein gene of three virus isolates obtained of samples from the São Paulo and Minas Gerais regions of Brazil. The phylogenetic relationships of the Brazilian isolates to those previously characterized were examined. A brief report appeared earlier (Hammond & Bedendo, 2001) .
MATERIAL AND METHODS
Sampling - A total of 37 maize leaf samples exhibiting symptoms of corn stunt were collected and preserved from four Brazilian States during 1995 and 1996 (Bedendo et al., 1997).
Nucleic acid extractions, hybridizations, and RT-PCR - Total nucleic acid extracts were prepared from dried leaf tissue using a previously described method (Hammond et al., 1997). MRFV was detected in samples spotted onto nylon membranes (Kogel et al., 1996). MRFV-specific cDNA was synthesized by reverse transcription in a reaction containing 2-3 µL total nucleic acid, primer MRFV-10 (5' GCCCACAGGTCTTATGGCCGACCTGCTACC 3', complementary to a region in the 3' NTR of MRFV) in reaction conditions previously described (Hammond & Bedendo, 2001). PCR reactions contained an aliquot of first-strand cDNA and primers MRFV-09 (5' CCTCTTCCTGATCCTCCTCGTGTGCT GGAGACG 3') and MRFV-10 (Hammond et al., 1997). PCR products were analyzed by gel electrophoresis to confirm their size and were cloned directly into the pCR2.1 plasmid vector (Invitrogen).
Nucleotide sequencing and sequence comparisons - The nucleotide sequences of the 633 bp MRFV fragments inserted in the pCR2.1 plasmids were obtained using an automated DNA sequencer (Applied Biosystems International Model 373A). Sequence data were analyzed using Lasergene software by DNASTAR for the Apple Macintosh. Multiple sequence alignments were performed using the Clustal Method Version 5 (Higgins et al., 1992). Cladistic analyses were performed with the computer program phylogenetic analysis using parsimony (PAUP) version 3.1 written by Swofford (Swofford, 1993) on a Power Macintosh Model 8100. Uninformative characters were excluded from the analysis and the phylogenetic trees were constructed by a branch and bound search, using random stepwise addition. Bootstrap analyses (100 replicates) were performed to estimate the support for the inferred phylogenies (Felsenstein, 1987). Gen bank accession numbers of previously sequenced isolates are provided in Hammond et al. (1997). Oat blue dwarf virus (OBDV- Gen bank accession no. U87832) was used as the outgroup (Edwards et al., 1997).
RESULTS AND DISCUSSION
Sixteen out of 37 samples tested for the presence of MRFV by nucleic acid hybridization using an MRFV-specific probe were positive for MRFV (data not shown). Reverse transcription-polymerase chain reaction (RT-PCR) products were obtained from three of the samples (Brazil 17, 26, and 48) using primers MRFV 9 and MRFV 10 and consisted of the expected 633 bp fragment (data not shown). No amplification occurred from healthy plants. The fragment was subsequently cloned into the plasmid vector pCR2.1 for further analysis. Single clones representing each isolate were selected for nucleotide sequencing. The 633 bp fragment encodes nt. 3-533 of the coat protein gene and 99 bp of the 3'NTR (Hammond et al., 1997).
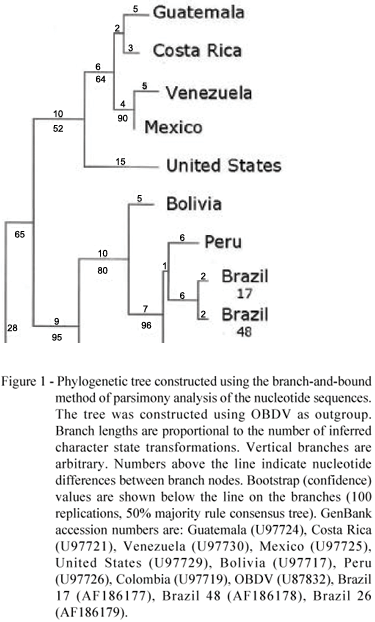
The isolates from Brazil share 98% similarity among themselves and from 90 to 97% similarity to other MRFV isolates. Phylogenetic relationships between the isolates were inferred from the dendrogram (Figure 1). The Brazilian isolates cluster with the isolates from Peru and Bolivia, which are geographically adjoining Brazil.
The results obtained from sequence analysis of RT-PCR products of Brazilian corn streak virus confirm the results obtained previously with serology (Gamez et al., 1979; Kitajima et al., 1975), ie., isolates of the virus obtained from São Paulo and Brasilia were found to be related to, but distinguishable from, isolates from Costa Rica, Mexico, Guatemala, El Salvador, Honduras, Nicaragua, Panama, Peru, Uruguay, and Colombia. This research supports the proposal that Brazilian corn streak virus be regarded as an isolate of MRFV.
The samples analyzed in this study originate from the Ribeirão Preto region of São Paulo and the National Center for Corn Research, Sete Lagoas region of Minas Gerais. Samples 17 and 26 were collected from São Paulo in areas approximately 100 km apart; sample 48 was collected in Minas Gerais at approximately 400 km from samples 17 and 26.
The contribution of MRFV to the corn stunt symptoms exhibited by maize plants is unknown. The usual symptoms of MRFV infection on maize consist of numerous short and long chlorotic stripes, wilting of young plants, general chlorosis, and stunting. Symptoms are most noticeable at the base of leaves, with an increase in severity in the younger leaves (Gamez, 1980; Nault et al., 1980). Symptom severity is not always proportional to viral replication within the host and may be influenced by environmental conditions or germ plasm (Bustamente et al., 1998). In addition, the symptoms of MRFV infection could be masked by the symptoms produced by co-infection with corn stunt agents. Further examination of the occurrence of MRFV in Brazil and its association with corn stunt disease may answer these questions. The use of existing resistance or tolerance germ plasm (Bustamente et al., 1998) or the development of resistant germ plasm through biotechnology will benefit control measures and reduce losses to the diseases caused by MRFV.
Received June 27, 2005
Accepted October 27, 2005
- BEDENDO, I.P.; DAVIS, R.E.; DALLY, E.L. Molecular evidence for the presence of maize bushy stunt phytoplasma in corn in Brazil. Plant Disease, v.81, p.957, 1997.
- BUSTAMENTE, P.I.; HAMMOND, R.; RAMIREZ, P. Evaluation of maize germ plasm for resistance to maize rayado fino virus. Plant Disease, v.82, p.50-56, 1998.
- COSTA, A.S.; KITAJIMA, E.W.; ARRUDA, S.C. Molestias de virus e micoplasmas de milho em São Paulo. Revista da Sociedade Brasileira de Fitopatologia, v.4, p.39-41, 1971.
- EDWARDS, M.C.; ZHANG, Z.; WEILAND, J.J. Oat blue dwarf marafivirus resembles the tymoviruses in sequence, genome organization, and expression strategy. Virology, v.232, p.217-229, 1997.
- FELSENSTEIN, J. Confidence limits on phylogenies: an approach using bootstrap. Evolution, v.39, p.783-791, 1987.
- GAMEZ, R. Transmission of rayado fino virus of maize (Zea mays L.) by Dalbulus maidis Delong & Wolcott. Annals of Applied Biology, v.73, p.285-292, 1973.
- GAMEZ, R. Rayado fino disease of maize in the American tropics. Tropical Pest Management, v.26, p.26-33, 1980.
- GAMEZ, R.; LEON, P. Maize rayado fino and related viruses. In: KOENIG, R.(Ed.) The plant viruses New York: Plenum Press, 1988. p.213-223.
- GAMEZ, R.; KITAJIMA, E.W.; LIN, M.T. The geographical distribution of maize rayado fino virus. Plant Disease Reporter, v.63, p.830-833, 1979.
- HAMMOND, R.W.; BEDENDO, I.P. Role of Maize rayado fino virus in the etiology of "Red Stunt" disease in Brazil. Plant Disease, v.85, p.99, 2001.
- HAMMOND, R.W.; RAMIREZ, P. Molecular characterization of the genome of Maize rayado fino virus, the type member of the genus Marafivirus Virology, v.282, p.338-347, 2001.
- HAMMOND, R.W.; KOGEL, R.; RAMIREZ, P. Variability of geographically distinct isolates of maize rayado fino virus in Latin America. Journal of General Virology, v.78, p.3153-3159, 1997.
- HIGGINS, D.G.; BLEASBY, A.J.; FUCHS, R. Clustal V: improved software for multiple sequence alignment. Computer Applications in the Biosciences, v.8, p.189-191, 1992.
- KITAJIMA, E.W.; GAMEZ, R.; LIN, M.T. A serological and histological comparison of the rayado fino virus from Costa Rica and Brazilian corn streak virus. Proceedings of the American Phytopathology Society, v.2, p.76, 1975.
- KITAJIMA, E.W.; YANO, T.; COSTA, S. Purification and intracellular location of isometric particles associated with the Brazilian corn streak virus infection. Ciência e Cultura, v.28, p.427-430, 1976.
- KOGEL, R.; HAMMOND, R.W.; RAMIREZ, P. Incidence and geographic distribution of maize rayado fino virus (MRFV) in Latin America. Plant Disease, v.80, p.679-683, 1996.
- MASSOLA, N.S.; BEDENDO, I.P.; AMORIM, L.; LOPES, J.R.S. Quantificação de danos causados pelo enfezamento vermelho e enfezamento pálido do milho em condições de campo. Fitopatologia Brasileira, v.24, p.136-142, 1999.
- NAULT, L.R.; BRADFUTE, O.E. Corn stunt: involvement of a complex of leafhopper pathogens. In MARAMOROSCH, K; HARRIS, K. (Ed.) Leafhopper vectors and plant disease agents. New York: Academic Press, 1979. p.561-586.
- NAULT, L.R.; GINGERY, R.E.; GORDON, D.T. Leafhopper transmission and host range of maize rayado fino virus. Phytopathology, v.70, p.709-712, 1980.
- SWOFFORD, D.L. PAUP: Phylogenetic analysis using parsimony Program (version 3.1). Champaign: Illinois Natural History Survey, 1993.
Publication Dates
-
Publication in this collection
12 Dec 2005 -
Date of issue
Dec 2005
History
-
Accepted
27 Oct 2005 -
Received
27 July 2005