Abstracts
Crop management may modify soil characteristics, and as a consequence, alter incidence of diseases caused by soilborne pathogens. This study evaluated the suppressiveness to R. solani in 59 soil samples from a microbasin. Soil sampling areas included undisturbed forest, pasture and fallow ground areas, annual crops, perennial crops, and ploughed soil. The soil samples were characterized according to abiotic variables (pH; electrical conductivity; organic matter content; N total; P; K; Ca; Mg; Al; H; S; Na; Fe; Mn; Cu; Zn; B; cation exchange capacity; sum of bases and base saturation) and biotic variables (total microbial activity evaluated by the CO2 evolution and fluorescein diacetate hydrolysis; culturable bacterial, fungal, actinomycetes, protozoa, fluorescent Pseudomonas and Fusarium spp. communities). The contribution and relationships of these variables to suppression to R. solani were assessed by path analysis. When all samples were analyzed together, only abiotic variables correlated with suppression of R. solani, but the entire set of variables explained only 51% of the total variation. However, when samples were grouped and analyzed by vegetation cover, the set of evaluated variables in all cases accounted for more than 90% of the variation in suppression of the pathogen. In highly suppressive soils of forest and pasture/fallow ground areas, several abiotic variables and fluorescein diacetate hydrolysis correlated with suppression of R. solani and the set of variables explained more than 98% of suppressiveness.
soilborne pathogen; soil microorganisms; biological control; path analysis; suppressive soil
As atividades agrícolas podem modificar as características do solo e, como conseqüência, alterar a incidência de patógenos veiculados pelo solo. Este trabalho avaliou a supressividade a R. solani em 59 amostras de solos de uma microbacia. As áreas amostradas foram selecionadas quanto à vegetação, incluindo mata, pasto/pousio, culturas anuais, culturas perenes e solo arado. As amostras de solo foram caracterizadas quanto às propriedades abióticas (pH, condutividade elétrica, teor de matéria orgânica, N total, P, K, Ca, Mg, Al, H, S, Na, Fe, Mn, Cu, Zn, B, capacidade de troca catiônica, soma de bases e saturação de bases) e bióticas (atividade microbiana do solo, avaliada pelo desprendimento de CO2 e hidrólise de diacetato de fluoresceína; comunidades de bactérias, fungos, actinomicetos, protozoários, Pseudomonas fluorescentes e Fusarium spp.). A contribuição e relação dessas variáveis para a supressividade a R. solani foram quantificadas por análise de coeficientes de trilha. Quando se avaliaram todas as amostras em conjunto, somente variáveis abióticas apresentaram correlação com a supressão a R. solani, mas o conjunto das variáveis explicou somente 51% da variação total. Entretanto, quando as amostras foram agrupadas e analisadas considerando o tipo de cobertura vegetal, o conjunto de variáveis explicou mais de 90% da variação da supressividade. Para as áreas de floresta e pasto/pousio, as quais foram classificadas como as mais supressivas, algumas variáveis abióticas e a hidrólise de diacetato de fluoresceína apresentaram correlação com a supressão de R. solani e o conjunto de variáveis explicou mais de 98% da supressividade nesses solos.
patógeno de solo; microrganismos do solo; controle biológico; análise de trilha; solo supressivo
SOILS AND PLANT NUTRITION
Biotic and abiotic factors associated with soil suppressiveness to Rhizoctonia solani
Fatores bióticos e abióticos associados à supressividade de solos a Rhizoctonia solani
Raquel Ghini* * Corresponding author < raquel@cnpma.embrapa.br> ; Marcelo Augusto Boechat Morandi
Embrapa Meio Ambiente, C.P. 69 - 13820-000 - Jaguariúna, SP - Brasil
ABSTRACT
Crop management may modify soil characteristics, and as a consequence, alter incidence of diseases caused by soilborne pathogens. This study evaluated the suppressiveness to R. solani in 59 soil samples from a microbasin. Soil sampling areas included undisturbed forest, pasture and fallow ground areas, annual crops, perennial crops, and ploughed soil. The soil samples were characterized according to abiotic variables (pH; electrical conductivity; organic matter content; N total; P; K; Ca; Mg; Al; H; S; Na; Fe; Mn; Cu; Zn; B; cation exchange capacity; sum of bases and base saturation) and biotic variables (total microbial activity evaluated by the CO2 evolution and fluorescein diacetate hydrolysis; culturable bacterial, fungal, actinomycetes, protozoa, fluorescent Pseudomonas and Fusarium spp. communities). The contribution and relationships of these variables to suppression to R. solani were assessed by path analysis. When all samples were analyzed together, only abiotic variables correlated with suppression of R. solani, but the entire set of variables explained only 51% of the total variation. However, when samples were grouped and analyzed by vegetation cover, the set of evaluated variables in all cases accounted for more than 90% of the variation in suppression of the pathogen. In highly suppressive soils of forest and pasture/fallow ground areas, several abiotic variables and fluorescein diacetate hydrolysis correlated with suppression of R. solani and the set of variables explained more than 98% of suppressiveness.
Key words: soilborne pathogen, soil microorganisms, biological control, path analysis, suppressive soil
RESUMO
As atividades agrícolas podem modificar as características do solo e, como conseqüência, alterar a incidência de patógenos veiculados pelo solo. Este trabalho avaliou a supressividade a R. solani em 59 amostras de solos de uma microbacia. As áreas amostradas foram selecionadas quanto à vegetação, incluindo mata, pasto/pousio, culturas anuais, culturas perenes e solo arado. As amostras de solo foram caracterizadas quanto às propriedades abióticas (pH, condutividade elétrica, teor de matéria orgânica, N total, P, K, Ca, Mg, Al, H, S, Na, Fe, Mn, Cu, Zn, B, capacidade de troca catiônica, soma de bases e saturação de bases) e bióticas (atividade microbiana do solo, avaliada pelo desprendimento de CO2 e hidrólise de diacetato de fluoresceína; comunidades de bactérias, fungos, actinomicetos, protozoários, Pseudomonas fluorescentes e Fusarium spp.). A contribuição e relação dessas variáveis para a supressividade a R. solani foram quantificadas por análise de coeficientes de trilha. Quando se avaliaram todas as amostras em conjunto, somente variáveis abióticas apresentaram correlação com a supressão a R. solani, mas o conjunto das variáveis explicou somente 51% da variação total. Entretanto, quando as amostras foram agrupadas e analisadas considerando o tipo de cobertura vegetal, o conjunto de variáveis explicou mais de 90% da variação da supressividade. Para as áreas de floresta e pasto/pousio, as quais foram classificadas como as mais supressivas, algumas variáveis abióticas e a hidrólise de diacetato de fluoresceína apresentaram correlação com a supressão de R. solani e o conjunto de variáveis explicou mais de 98% da supressividade nesses solos.
Palavras-chave: patógeno de solo, microrganismos do solo, controle biológico, análise de trilha, solo supressivo
INTRODUCTION
In suppressive soils (Baker & Cook, 1974), the development of diseases is suppressed even when pathogens are present and hosts are susceptible. Soil suppressiveness could result from biotic and abiotic factors, in a diverse and complex set of mechanisms. A holistic approach is generally needed to study suppressiveness, since the contributing factors normally work interactively (Schneider, 1982). Once a soil becomes suppressive to a target pathogen, the determination of its main physical, chemical and biological attributes can be useful for understanding the mechanisms of suppressiveness, and to exploit the information in other areas where the same pathogen is a problem (Rodríguez-Kábana & Calvet, 1994).
Rhizoctonia solani has great importance among soilborne plant pathogens, and damages a wide range of hosts worldwide. Ghini & Zaroni (2001) evaluated and grouped statistically by Wardxs method (Everitt, 1981), soils inside a microbasin in categories of suppressiveness against R. solani. The authors defined the microbasin as the physiographic unit for the work based on its climatic homogeneity, convergent water drainage and diversity of vegetation. In general, the pasture and fallow ground areas, followed by undisturbed forest covered areas, were correlated with suppressiveness, while ploughed soils and cultivated soils with annual (corn) or perennial (sugar cane and coffee) crops were most correlated with conducivity.
However, other factors besides vegetation were involved in the suppressiveness to R. solani. The literature describes several factors that could influence the suppressiveness of soils to the pathogen, including soil microbiota (Kobayashi & Ko, 1985), the Trichoderma community (Chet & Baker, 1980; Henis et al., 1979; Liu & Baker, 1980) and pH, moisture and type of soil (Henis et al., 1979; Liu & Baker, 1980).
The objective of the present work was to evaluate the contribution and relationships of abiotic (chemical properties) and biotic (microorganisms) factors to the suppressiveness of soils from the Córrego Taquara Branca microbasin to R. solani.
MATERIAL AND METHODS
Soil sampling
Sampling included 59 areas within the Córrego Taquara Branca microbasin (22º50' to 22º55'S; 47º15'00" to 47º18'45"W), São Paulo state, Brazil. Areas were selected based on vegetation cover, and included undisturbed forest (10), pasture and fallow ground areas (11), annual crops (18), perennial crops (16), and ploughed soil (4). From each area, seven soil sub-samples (80 x 80 x 80 mm) were taken randomly, homogenized and sieved (4 mm). Soil for microbiological analysis was processed within 24 h after sampling.
Biotic soil characteristics
The following biotic variables were evaluated on each soil sample: total microbial activity; culturable bacterial, fungal, actinomycetes and protozoa communities; and, populations of Fusarium spp. and fluorescent Pseudomonas spp. The total microbial activity of the soil was evaluated by measuring the respiratory activity (determination of CO2 evolution) and overall enzymatic activity (hydrolysis of fluorescein diacetate, FDA). To evaluate the CO2 evolution, 200-g soil samples (n = 3) were placed in plastic, hermetic flasks (2.5 L) containing 10 mL of KOH 0.5 N, and incubated at 25ºC in the dark. After seven, 14 and 21 days, the KOH solution was titrated with HCl (Grisi, 1978), for analysis of the total amount of CO2 (µg CO2 g-1 dry soil).
For FDA hydrolysis, the method of Chen et al. (1988) was used. Soil samples (5 g; n = 5) were placed in 250 mL Erlenmeyer flasks and mixed with 20 mL of potassium phosphate buffer (60 mM; pH 7.6). The hydrolysis reaction was triggered by the addition of 0.2 mL of FDA stock solution (2 mg FDA mL-1 acetone) and the flasks were incubated on a rotary shaker (200 rpm) at 25ºC for 20 min. The reaction was halted by the addition of 20 mL of acetone per flask, and the content of each flask was filtered on Whatman nº 1 filter paper. The absorbance (490 nm) of the filtrate was determined spectrophotometrically. To determine the concentration of the hydrolyzed fluorescein (µg hydrolyzed FDA g-1 of dry soil), the absorbance of the samples were compared against a standard curve, obtained by following the same methodology, except that a known concentration of FDA was added to the potassium phosphate buffer and the flasks were kept at 100ºC for 60 min to hydrolyze the FDA, before it was mixed with the soil samples.
All other variables, except protozoa communities, were assessed by counting colonies forming units (CFUs) on selective agar media, as follows: Martin's medium for fungi; agar-nutrients plus nistatine 42 mg L-1 for bacteria; alkalinized water-agar medium (pH 10.5) for actinomycetes; King's medium B for fluorescent Pseudomonas spp.; and a selective medium for Fusarium spp. (Nash & Snyder, 1962). For each soil sample, aliquots of 100 µL from two dilutions (10-1 and 10-2 for general fungi and Fusarium spp.; 10-2 and 10-3 for fluorescent Pseudomonas spp.; and 10-3 and 10-4 for bacteria and actinomycetes) were applied to Petri dishes (n = 3) containing the specific medium, and incubated upside down at 25ºC until colonies were visible (after one day for bacteria; two days for general fungi, Fusarium spp. and fluorescent Pseudomonas; and five days for actinomycetes). Protozoan communities were assessed using the method described by Stotzky et al. (1993).
Abiotic soil characteristics
The following abiotic variables were estimated, for each soil sample: pH; electrical conductivity (EC); organic matter content; N total; P; K; Ca; Mg; Al; H; S; Na; Fe; Mn; Cu; Zn; B; cation exchange capacity (CEC); sum of bases (S value, sum of Ca, Mg, K, and Na); and base saturation (V%) (Silva, 1999).
Suppression of R. solani growth
The suppression of mycelial growth of R. solani AG 4 HG-I, isolated from symptomatic plants of common bean (Phaseolus vulgaris L.), was evaluated on 15-cm Petri dishes containing the soil samples (50 g of soil per dish) (Ko & Ho, 1983). Moisture of soil samples was adjusted to 70% of field capacity. Samples were then placed in the dishes and covered with a thin layer of water agar medium. One disk (7 mm of diameter) of R. solani mycelium grown on PDA was transferred to the center of each dish (n = 3). The dishes were kept at 25ºC in the dark and perpendicular diameter of the R. solani colonies were measured daily using a stereomicroscope (12x). The Area Under Growth Curve (AUGC) of R. solani from the first to the fourth day of incubation for each soil sample was calculated and used for analysis.
Data analysis
The relative importance of individual biotic and abiotic factors on R. solani growth suppression and their interactions were assessed by path coefficient analysis (Wright, 1921). Correlations between variables were divided over direct and indirect influences along different paths in a system. The direct influences, or path coefficients, of a set of variables (causes) upon a certain variable (effect) indicate the degree to which variation of that effect is determined in each particular cause. The assumption of path analysis is that relationships between variables can be causally structured (van Bruggen & Arneson, 1986). If variables are not completely explained by other variables in the system, an extra variable that contains residual influences and experimental error can be added. A general relational diagram showing all the possible relationships between biotic and abiotic, explicative variables measured and the R. solani growth suppression (dependent variable) is given (Figure 1).
Correlation matrices between data for growth suppression of R. solani and the biotic and abiotic variables (independent variables) were generated for each sampled soil, grouped by plant cover (forest, pasture, annual crops, perennial crops, and ploughed soil). Another matrix correlating R. solani growth suppression with all samples, independently of the soil coverage, was also generated. The path coefficient analysis was performed using Software 'GENES' (Quantitative Genetics and Experimental Statistics Analysis, vs. 0.1.0, http://www.ufv.br/dbg/home.htm) (Cruz, 2001), and the studied variables were grouped in a causal diagram incorporating possible pathways of influence on R. solani growth suppression.
The results of the path analysis were interpreted according to directives proposed by Singh & Chaudary (1979): 1. If the correlation coefficient (r) between the causal factor and the effect is almost the same as its direct effect, the correlation can explain the actual relation between them and a direct selection based on this factor will be efficient; and, 2. If the correlation is positive (or negative) but the direct effect is negative, positive or can be ignored, the indirect effects seem to be the cause of the correlation. In such situations, the indirect causal factors should be evaluated simultaneously. To select the major variable(s) and its/their influence on R. solani suppressiveness, the following criteria were used: a) statistical significance on correlation at a = 0.05 by t test; and b) the fulfillment of the directives of Singh & Chaudary (1979).
RESULTS AND DISCUSSION
Ghini & Zaroni (2001) had previously grouped the same areas of the microbasin in categories of suppressiveness, based on vegetation cover. The pasture, fallow ground and forest covered areas were classified as suppressive soils, while ploughed soils and cultivated soils, with annual or perennial crops, as conducive soils. However, in that work it was not possible to determine the factors that contributed to the gradient of suppressiveness of the soils to R. solani.
In this study, biotic and abiotic characteristics of the soils were assessed (Table 1), and the contribution of each variable to the suppression of R. solani growth was determined by path coefficient analysis. This analysis has rarely been used in plant pathology, but van Bruggen & Arneson (1986) indicate the potential use of this technique, such as to assess the relative importance of certain factors contributing to pathogen growth and disease development. Path analysis is not just a multiple linear regression analysis, because it allows several diagrams to be drawn and the establishment by the researcher of a cause-effect model according to one's hypothesis. Furthermore, path analysis can generate more information than multiple regression, and such is the case in this study.
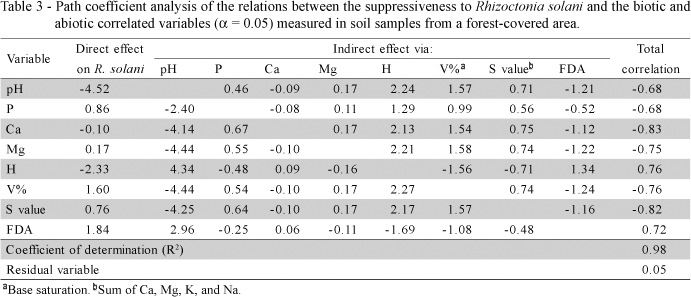
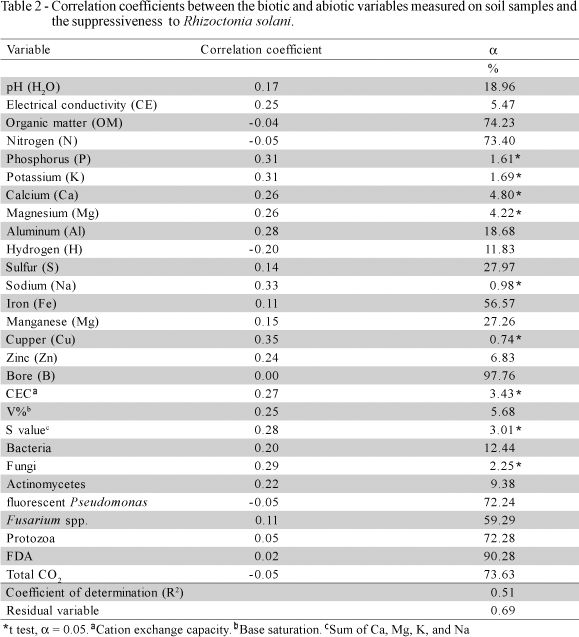
When all samples were analyzed together, independently of the soil type and vegetation, only abiotic variables (P, K, Ca, Mg, Na, Cu, CEC and S value) were correlated (a < 0.05) with R. solani growth suppression, and the entire set of variables explained only 51% of the total variation (R2 = 0.51; residual variable, Rv = 0.69) (Table 2), which is not useful in predicting soil suppressiveness or the mechanism of suppression. However, when samples were grouped and analyzed by vegetation cover, the set of evaluated variables in all instances explained more than 90% of the variation in growth suppression of R. solani. In highly suppressive forest soils and pasture/fallow ground areas, several abiotic variables and FDA hydrolysis correlated with growth suppression of R. solani and the set of variables explained more than 98% of suppressiveness (Tables 3 and 4, respectively). For less suppressive soils, no correlations of suppressiveness with the biotic variables were found. Several abiotic variables were correlated with suppressiveness of soils from annual crops (Table 5), but no correlations were found for ploughed soils and perennial crops. These findings stress the importance of the vegetation cover and soil management on soil supressiveness. To understand the nature of soil suppressiveness, the ecological context within which the major players act must be considered, since the interactions among soil microorganism communities as well as the abiotic variables may influence both the structure and activities of populations, including those of pathogens and their antagonists (Weller et al., 2002).
Suppressive soils are common in ecologically balanced environments with ecosystems in climax, where the physico-chemical and microbiological constituents of the soil are stabilized (Schneider, 1982). In theory, the greater the complexity of the biological community of the soil, the greater is the stability of the soil and, consequently, the level of natural biological control. Many soils possess similarities with regard to microorganisms involved in disease suppression, while other attributes are unique to specific pathogen-suppressive soil systems. The manipulation of microbial communities to induce a disease-suppressive soil environment is a potential tool in the management of soil borne plant diseases (Mazzola, 2002).
Colony counts of bacterial, fungal and actinomycete communities on the selective media and the total number of protozoan cells were not correlated with soil supressiveness to R. solani. Considering that there is no universal culture media, and that only a small percentage (0.9 to 22%) of total microbial communities can be cultivated on available media (Herbert, 1990), the number of CFU obtained per group of microorganisms evaluated was not enough to detect differences among the analyzed soils. The potential of other organisms, such as protozoa, to contribute to the suppressiveness to plant pathogens in soils are poorly explored. One of the few works on this topic was done by Homma & Ishii (1984), who observed perforation on R. solani hyphae by Arachnula impatiens (amoebae). Once the predominantly protozoan groups were not identified, it was not possible to determine their contribution to pathogen suppressiveness.
However, the total microbial activity, inferred by the amount of hydrolyzed FDA, was positively and highly correlated with the pathogen suppression at the pasture/fallow ground and forested areas. The suppressiveness to R. solani was mainly related to biotic soil characteristics, represented by the total microbial activity of the soil (FDA hydrolysis), since this variable was correlated for the most suppressive soils (pasture/fallow ground and forest areas). These results reinforce early reports that the pasture, fallow ground and forest covered areas, which were classified as suppressive soils, became conducive after fumigation (Ghini & Zaroni, 2001). Positive relationships of total microbial activity and suppressiveness of soils to soilborne pathogens are well documented in the literature (Weller et al., 2002).
The two variables used to evaluate the total microbial activity (CO2 evolution and FDA hydrolysis) were correlated (r = 0.71) when all samples were analyzed together, independently of the soil type and vegetation. However, when samples were grouped and analyzed by vegetation cover, the two variables were correlated (r = 0.85) only on the pasture/fallow ground areas. In these soils, both FDA hydrolysis and CO2 evolution were correlated with suppressiveness to R. solani. There was no correlation between the two variables and only the FDA hydrolysis was correlated with suppressiveness in forested areas. The differences in contribution to suppressiveness between FDA hydrolysis and CO2 evolution probably reflect different processes of microbial activity, influenced by different environmental conditions. The FDA is hydrolyzed by several enzymes (lipase, protease and esterase) from live cells (Inbar et al., 1991), while CO2 evolution depends on the total respiratory activity of the living cells.
Besides the FDA hydrolysis, the suppressiveness against R. solani of the two most suppressive soils (pasture/fallow ground and forest areas) was correlated with different groups of abiotic variables. Although only the significantly correlated variables at 5% are shown, several other variables contributed to suppressiveness at a low level of significance. These results reinforce the current knowledge that soil suppressiveness against R. solani could result from the biotic and abiotic factors of the soil, in a complex set of interactions among the chemical, biological and physical properties of the soil (Chet & Baker, 1980; Henis et al., 1979; Kobayashi & Ko, 1985; Liu & Baker, 1980).
Suppressiveness to R. solani was measured only by mycelial growth tests, where extension growth was measured on agar-covered soils. This separates the fungus from the soils and not all of the measured properties are going to influence directly the mycelial growth. Sieved soil and physical orientation of the soil particles placed in Petri dishes also can influence the growth of R. solani. Despite these methodology limitations, it has been used to study soil suppressiveness to several plant pathogens (De Boer et al., 1998; van Elsas et al., 2002). This study evaluated the major factors in the analyzed soil samples that influence the growth of the pathogen, which is an important suppressiveness component but the infection of plants with R. solani was not tested. Suppression of infection may show different results, since new variables of the plant growth should be included in the analysis. Future in-ground experiments in each vegetation type can evaluate more precisely pathogen-suppressive and disease-suppressive soils.
ACKNOWLEDGEMENTS
To Prof. Dr. Paulo Cézar Ceresine from UNESP/FEIS for the classification of the anastomosis group of the R. solani isolates, and to CNPq for research grant to first author.
Received July 22, 2005
Accepted March 08, 2006
- BAKER, R.; COOK, J. Biological control of plant pathogens San Francisco: W.H. Freeman, 1974. 433p.
- CHEN, W.; HOITINK, A.J.; SCHMITTHENNER, A.F.; TUOVINEN, O.H. The role of microbial activity in suppression of damping-off caused by Pythium ultimum Phytopathology, v.78, p.314-322, 1988.
- CHET, H.; BAKER, R. Induction of suppressiveness to Rhizoctonia solani in soil. Phytopathology, v.70, p.994-998, 1980.
- CRUZ, C.D. Software Genes - Windows version - Computational system on Genetics and Statistics. Viçosa: Editora UFV, 2001.
- De BOER, W.; GUNNEWIEK, P.J.A.K.; WOLDENDORP, J.W. Suppression of hyphal growth of soil-borne fungi by dune soils from vigorous and declining stands of Ammophila arenaria New Phytologist, v.138, p.107-116, 1998.
- EVERITT, B. Cluster analysis 2.ed. New York: Wiley, 1981.
- GHINI, R.; ZARONI, M.M.H. Relação entre coberturas vegetais e supressividade de solos a Rhizoctonia solani Fitopatologia Brasileira, v.26, p.10-15, 2001.
- GRISI, B.M. Método químico de medição da respiração edáfica: alguns aspectos técnicos. Ciência e Cultura, v.30, p.82-88, 1978.
- HENIS, Y.; GRAFFAR, A.; BAKER, R. Factors affecting suppressiveness to Rhizoctonia solani in soil. Phytopathology, v.69, p.1164-1169, 1979.
- HERBERT, R.A. Methods for enumerating microorganisms and determining biomass in natural environments. In: GRIGOROVA, R.; NORRIS, J.R. (Ed.) Techniques in microbial ecology San Diego: Academic Press, 1990. p.1-39. (Methods in Microbiology, 22).
- HOMMA, Y.; ISHII, M. Perforation of hyphae and sclerotia of Rhizoctonia solani Kuhn by mycophagous soil amoebae from vegetable field soils in Japan. Annals of the Phytopathological Society of Japan, v.50, p.229-240, 1984.
- INBAR, Y.; BOEHM, M.J.; HOITINK, H.A.J. Hydrolysis of fluorescein diacetate in sphagnum peat container media for predicting suppressiveness to damping-off caused by Pythium ultimum Soil Biology and Biochemistry, v.23, p.479-483, 1991.
- KO, W.; HO, W. Screening soils for suppressiveness to Rhizoctonia solani and Pythium splendens Annals of the Phytopathological Society of Japan, v.49, p.1-9, 1983.
- KOBAYASHI, N.; KO, W.H. Nature of suppression of Rhizoctonia solani in Hawaiian soils. Transactions of the British Mycological Society, v.84, p.691-694, 1985.
- LIU, S.; BAKER, R. Mechanism of biological control in soil suppressive to Rhizoctonia solani Phytopathology, v.70, p.404-412, 1980.
- MAZZOLA, M. Mechanisms of natural soil suppressiveness to soilborne diseases. Antonie van Leeuwenhoek, v.81, p.557-564, 2002.
- NASH, S.M.; SNYDER, W.C. Quantitative estimations by plate counts of propagules of the bean root rot Fusarium in field soils. Phytopathology, v.52, p.567-572, 1962.
- RODRÍGUEZ-KÁBANA, R.; CALVET, C. Capacidad del suelo para controlar enfermidades de origen edafico. Fitopatologia Brasileira, v.19, p.129-138, 1994.
- SCHNEIDER, R.W. Suppressive soils and plant disease St. Paul: APS, 1982. 88p.
- SILVA, F.C. Manual de análises químicas de solos, plantas e fertilizantes Brasília: Embrapa Comunicação para Transferência de Tecnologia, 1999. 370p.
- SINGH, R.K.; CHAUDARY, B.D. Biometrical methods in quantitative genetic analysis New Delhi: Kalyani Publishers, 1979. 304p.
- STOTZKY, G.; BRODER, M.W.; DOYLE, J.D.; JONES, R.A. Selected methods for the detection and assessment of ecological effects resulting from the release of genetically engineered microorganisms to the terrestrial environment San Diego: Academic Press, 1993. 98p. (Advances in Applied Microbiology, 38).
- Van BRUGGEN, A.H.C.; ARNESON, P.A. Path coefficient analysis of effects of Rhizoctonia solani on growth and development of dry beans. Phytopathology, v.76, p.874-878, 1986.
- Van ELSAS, J.D.; GARBEVA, P.; SALLES, J. Effects of agronomical measures on the microbial diversity of soil as related to the suppression of soil-borne plant pathogens. Biodegradation, v.13, p.29-40, 2002.
- WELLER, D.M.; RAAIJMAKERS, J.M.; GARDENER, B.B.M.; THOMASHOW, L.S. Microbial populations responsible for specific soil suppressiveness to plant pathogens. Annual Review of Phytopathology, v.40, p.309-348, 2002.
- WRIGHT, S. Correlation and causation. Journal of Agricultural Research, v.20, p.557-585, 1921.
Publication Dates
-
Publication in this collection
17 Apr 2006 -
Date of issue
Apr 2006
History
-
Accepted
08 Mar 2006 -
Received
22 July 2005