Abstract
Halo blight, caused by Pseudomonas savastanoi pv. phaseolicola (Psp), is considered to be an important bacterial disease on common bean (Phaseolus vulgaris L.) in Serbia. Use of pathogen-free seeds is one of the most effective control measures against this disease. The aim of this study was to evaluate a detection method for Psp on untreated common bean seeds (23 genotypes) from commercial crops grown within Serbia. Detection of this pathogen was made by plating onto the modified sucrose peptone (MSP) and Milk Tween (MT) semi-selective mediums from soaked whole common bean seed. Colonies growing on the MSP medium were light yellow, convex and shiny, whereas on the MT medium, they were creamy white, flat and circular. The pathogenicity of the obtained strains was confirmed by the inoculation of germinated bean seed. The isolates recovered from the seed assay were further confirmed to be Psp by using both Enzyme-linked immunosorbent assay (ELISA) and polymerase chain reaction (Nested-PCR) detection methodologies. The International Seed Testing Association (ISTA) method selected for this work was found to be effective in detecting the presence of Psp in common bean seed. The bacterium Psp was detected in only two of the 23 seed samples analyzed by this method, which shows that the bacterium is not widespread in Serbia.
MSP; MT; ELISA; Nested-PCR; halo blight; pathogenicity
NOTE
Application of semi-selective mediums in routine diagnostic testing of Pseudomonas savastanoi pv. phaseolicola on common bean seeds
Tatjana PopovicI; Predrag MilovanovicII; Goran AleksicI; Veljko GavrilovicI; Mira StarovicI; Mirjana VasicIII; Jelica BalažIV
IInstitute for Plant Protection and Environment, Teodora Drajzera, 9, PO Box 33-79, 11040, Belgrade, Republic of Serbia,
E-mail:tanjaizbis@gmail.com
IIGalenika/Fitofarmacija, Batajniki drum b.b., 11080, Belgrade, Republic of Serbia
IIIInstitute for Field and Vegetable Crops, Maksima Gorkog 30, 21000, Novi Sad, Republic of Serbia
IVUniversity of Novi Sad/Faculty of Agriculture, Dositeja Obradovia 8, 21000, Novi Sad, Republic of Serbia
ABSTRACT
Halo blight, caused by Pseudomonas savastanoi pv. phaseolicola (Psp), is considered to be an important bacterial disease on common bean (Phaseolus vulgaris L.) in Serbia. Use of pathogen-free seeds is one of the most effective control measures against this disease. The aim of this study was to evaluate a detection method for Psp on untreated common bean seeds (23 genotypes) from commercial crops grown within Serbia. Detection of this pathogen was made by plating onto the modified sucrose peptone (MSP) and Milk Tween (MT) semi-selective mediums from soaked whole common bean seed. Colonies growing on the MSP medium were light yellow, convex and shiny, whereas on the MT medium, they were creamy white, flat and circular. The pathogenicity of the obtained strains was confirmed by the inoculation of germinated bean seed. The isolates recovered from the seed assay were further confirmed to be Psp by using both Enzyme-linked immunosorbent assay (ELISA) and polymerase chain reaction (Nested-PCR) detection methodologies. The International Seed Testing Association (ISTA) method selected for this work was found to be effective in detecting the presence of Psp in common bean seed. The bacterium Psp was detected in only two of the 23 seed samples analyzed by this method, which shows that the bacterium is not widespread in Serbia.
Keywords: MSP, MT, ELISA, Nested-PCR, halo blight, pathogenicity
Introduction
Pseudomonas savastanoi pv. phaseolicola (Burkholder) Gardan et al. (Psp) which causes halo blight of bean (Phaseolus vulgaris L.) is a pathogen present in all countries where beans are cultivated. The disease is induced by cool temperatures and humidity and is widespread in Europe and North America. Many states and countries have strict quarantine laws regarding Psp (Grogan and Kimble, 1967; Taylor, 1970; Taylor et al., 1979; Webster et al., 1983). Halo blight was the main problem in bean production in Serbia during the 1970-80s (Balaž, 1990). Earlier cases have been reported where snap bean crops have completely collapsed in years where the spring weather was cool and rainy. However, in recent years, Psp has not been a major problem, which can probably be contributed to the weather conditions (the emergence and first phase of bean development generally follow warmer and drier weather conditions) and the use of resistant bean cultivars.
Pseudomonas savastanoi pv. phaseolicola is a seed-borne pathogen (Fahy and Lloyd, 1983). The detection of bacterium on seeds is essential for effective control of the disease. Several methods have been described for testing the presence of Psp in bean seed: a seed soak-plant inoculation technique (Lahman and Schaad, 1985; Webster et al., 1983), immunological methods (Guthrie et al., 1965; Van Vuurde et al., 1983), plating on King B agar (Van Vuurde and Van den Bovenkamp, 1987) and semiselective modified sucrose peptone (MSP) agar (Mohan and Schaad, 1987). Various PCR assays have been proposed for the detection of Psp (Prosen et al., 1993; Schaad et al., 1995, 2001; Mosqueda and Herrera, 1997). All of them rely on the detection of DNA sequences involved in the biosynthesis of phaseolotoxin (Prosen et al., 1993; Schaad et al., 1995; Audy et al., 1996; Güven et al., 2004).
Considering the necessity for the development of a practical and effective method for the routine analysis of bean seed in Serbia, this study was carried out using Psp isolation in semi-selective mediums developed for this purpose. Pathogenicity testing and rapid ELISA and PCR methods were used to confirm identification of obtained strains.
Materials and Methods
Untreated common bean seeds from 23 genotypes grown in Serbia (the Novi Sad area 45º15' N and 19°50' E) were analyzed for the presence of Psp. There were 12 samples of Serbian dry bean cultivars (Balkan, Belko, Dvadesetica, Galeb, Maksa, Medijana, Oplenac, Panonski gradistanac, Panonski tetovac, Slavonski zutozeleni, Sremac, and Zlatko) and 11 samples of foreign common bean genotypes from collection (Oreol, Dobrudzanski rani, Dobrudzanski rani 7, KB 100 and KB 101 from Bulgaria; C 20, Naya Nayahit, HR 45, Xan 159, Xan 208 and Xan 273 from USA). All samples came from untreated commercial crops. Five 1000-seed sub-samples were gathered for each cultivar, with a total of 115 sub-samples. Pure culture of Psp reference strain (Ps12, Faculty of Agriculture, Novi Sad) was used in all assays.
Extraction of bacteria from the seed
Each 1000-seed sub-sample was first washed in running water and then placed on sterile laboratory paper to dry. After that, sub-samples were weighed and soaked in seed extraction solution (0.85 % saline with Tween 20) in the proportion of 1:2 (1 g of seed in 2 mL of solution) for 24 h in the fridge (5 ºC). After soaking, the liquid was sampled. Extraction of the bacterium from whole bean seeds was done according to Jansing and Rudolph (1990), ISF (2006) and Kurowski and Remeeus (2007, 2008) methods.
Isolation on semi-selective media
After incubating, 0.1 mL of undiluted extracts obtained from the seeds and their 10-fold dilution series (to 10-5) were plated onto the surface of two semi-selective media, Modified Sucrose Peptone Agar (MSP) and Milk Tween Agar (MT), described by Mohan and Schaad (1987) and Goszczynska and Serfontein (1998), respectively. The series of dilution were prepared with the aim to get single colonies of Psp which could be morfologically differentiated from other non-targeted organisms. Two Petri dishes were plated for each concentration. A pure culture of Psp reference strain (Ps12) was also plated on both semi-selective media.
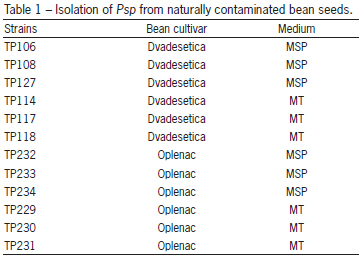
The Petri dishes were incubated at 28-30 ºC for five days. After that, the sample plates were visually assessed for the presence of colonies with typical Psp morphology by comparison with the reference strain. Suspected colonies, as well as the reference strain, were transferred onto a King's B agar (King et al., 1954) and incubated for three-four days at 27 °C. The transferred colonies were again compared with the reference strain. Six typical strains from each positive sample (12 representative strains) were selected for identification (Table 1).
Pathogenicity test
Suspected Psp bacterial colonies were evaluated using pathogenicity testing on the susceptible bean cultivar Oplenac, by the inoculation of germinated seed (Fenwick and Guthrie, 1969; Van Vuurde and Van den Bovenkamp, 1987, 1989; ISF, 2006). Susceptibility of the cultivar Oplenac to the Psp strains from Serbia had been confirmed and described by Balaž (1990). The seeds were incubated in rolled germination paper for four days at 25 °C in the darkness. To provide better conditions for infection, seedlings were sprayed with water two hours before inoculation. Inoculation was made with a sterile toothpick, which was dipped into the bacterial culture of obtained strains growing on the King B medium for two days. The toothpick was pushed through the cotyledon. After that, the toothpick was turned slightly while being pulled out in order to release the bacteria. Four seedlings were inoculated per every representative strain. The positive (reference culture) and negative (sterile water) controls were prepared in a similar manner. The seedlings were placed in a phytotron at 20 °C with 80 % relative humidity and light adequate for plant growth (light:dark 12:12).
The symptoms on the inoculated seedlings were recorded after five and 10 days and compared with the positive and negative controls. Pathogenicity was evaluated by the inspection of the flat inner sides of the cotyledons to determine the presence of typical "greasy" spots at the point of inoculation.
Serological and molecular identification
All strains were identified as Psp by the rapid tests, ELISA (Enzyme-linked immunosorbent assay) and PCR (polymerase chain reaction), regardless of their pathogenicity.
ELISA - Double-antibody sandwich (DAS)-ELISAs and Plate Trapped Antigen (PTA)-ELISAs were performed with commercial kits by Loewe Biochemica GmbH, Germany and ADGEN Phytodiagnostics, Neogen Europe Ltd., Scotland, U.K., respectively. The assays followed the manufacturer's instructions. Bacterial suspensions were prepared in sterile water with pure bacterial cultures grown on the King B for 48 h at 27 °C. A reference strain of Psp was used as a positive control, while a reference strain of Erwinia amylovora (NCPPB 595) was used as the negative control.
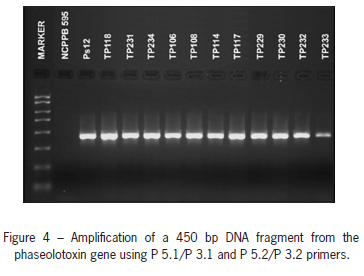
PCR - Nested PCR was conducted with DNA extracted from pure bacterial cultures (Schaad et al., 2001). Cultures were grown on the King B medium at 27 °C for 48 h, and cells from 0.5 mL of water suspension (3 ' 108 CFU mL-1) were used for DNA extraction. The method of Schaad et al. (1995) for the detection of phaseolotoxin genes was modified by Güven et al. (2004). 0.5 mL of cell suspension were boiled for 15 min. The cell debris was removed by centrifugation for 10 min at 11.000 rpm. 2 µL of supernatant were used for amplification. For the first PCR primers P 5.1: 5'-AGC TTC TCC TCA AAA CAC CTG C- 3' and P 3.1: 5'-TGT TCG CCA GAG GCA GTC ATG-3' were used, as suggested by Schaad et al. (1995). Primers P 5.1 and P 3.1 directed the amplification of the 500-bp DNA fragment. For the second PCR, primers P 5.2: 5'-TCG AAC ATC AAT CTG CCA GCC A-3' and P 3.2: 5'-GGC TTT TAT TAT TGC CGT GGG C-3' were used, as suggested by Schaad et al. (2001). Primers P 5.2 and P 3.2 directed the amplification of the 450-bp DNA fragment. A reference strain of Psp (Ps12) was used as a positive control and a reference strain of E. amylovora (NCPPB 595) was used as the negative control.
The PCR amplification assay was performed in a 25 µL reaction mixture containing Taq DNA polymerase 1.25 U, 50 mM KCl, 30 mM Tris-HCl, 1.5 mM Mg2+, 0.1 % igepal-CA630, 200 µM dNTP, 0.4 µM of primers and 1 µL of DNA.A Mastercycler ep gradient S (Eppendorf, Germany) was used for PCR with the following profile amplifications: an initial 3 min incubation at 94 °C, a manual "hot start" step at 80 °C, 25 cycles (1 min at 94 °C, 1 min at 58 °C and 1 min at 72 °C), and a final extension step of 10 min at 72 °C. After the first PCR, products were diluted 10x and 2 µL were used for the second PCR. The amplified DNA fragments were electrophoresed in 2.0 % agarose gels in 1xTBE buffer and visualized with ultraviolet light after ethidium bromide staining.
Results
The presence of Psp on the common bean seeds was observed using the method of isolation on semi-selective media following the ISTA method. This method was supplemented with ELISA and Nested-PCR methods in order to confirm the identity of obtained strains. The method was effective in isolating Psp colonies from infected common bean seeds. All the obtained strains were identified as Psp by the pathogenicity and rapid tests ELISA and PCR. The analysis of 23 common bean seed samples from the Novi Sad locality in Serbia revealed that only two of them (Dvadesetica and Oplenac) were infected with Psp (Table 1). Not a single suspected bacterium colony formed on the 21 bean genotypes (Balkan, Belko, C 20, Dobrudzanski rani, Dobrudzanski rani 7, Galeb, HR 45, KB 100, KB 101, Maksa, Medijana, Naya Nayahit, Oreol, Panonski gradistanac, Panonski tetovac, Slavonski zutozeleni, Sremac, Xan 159, Xan 208, Xan 273 and Zlatko).
After four days of incubation on the MSP medium colonies of Psp formed in 10-1-10-2 dilutions. They were light yellow, convex, shiny, and 2-3 mm in diameter (Figure 1). The medium around the colony turned light yellow.
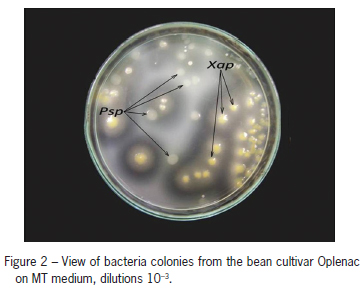
On MT, colonies of Psp were formed in 10-1-10-3 dilutions after four days of incubation. They were creamy white, flat, circular, and 3-5 mm in diameter (Figure 2). Furthermore, in both samples of common bean seeds (cultivar Dvadesetica and Oplenac) on the same medium, colonies of the bacterium Xanthomonas axonopodis pv. phaseoli (Smith) Vauterin et al. (Xap) also formed. Colonies were yellow, convex and mucoid and were surrounded by two zones of hydrolysis, a large clear zone of casein hydrolysis and a smaller milky zone of Tween 80 hydrolysis (Popovi et al., 2010), indicating that the seeds were infected with both bacteria (Figure 2). MT medium can also be used to develop colonies of the bacterium Pseudomonas syringae pv. syringae. This bacterium can be distinguished from Psp using a differential tests described by Lelliott and Stead (1987). The selected colonies grown on King B medium were creamy white and formed a green fluorescent pigment.
The presence of the suspect strains (Table 1) was confirmed by pathogenicity testing. On the inoculated bean cotyledons characteristic dark green, greasy spots formed at the point of inoculation after five days (Figure 3a). After 10 days, the whole cotyledon was affected with small greasy spots (Figure 3b).
Obtained strains of Psp were tested by DAS- and PTA-ELISA. The polyclonal antibody used reacted as expected with all the strains, producing clear positive reactions which indicate that strains of Psp can be detected using commercial ELISA tests. All bacterial strains submitted to the Nested-PCR test were identified as Psp. The used P 5.1/P 3.1 and P 5.2/P 3.2 primers pair directed the amplification of the 450-bp target DNA fragment (Figure 4).
Discussion
The bacterium Psp is a seed-borne pathogen (Taylor, 1970). Seeds contaminated internally or externally are the primary source of infection (Schwartz, 1989). Very low levels of inoculum, such as a single contaminated seed in 2000 or 5000, can result in an epidemic in the field under favorable environmental conditions (Webster et al., 1983). Use of healthy, pathogen-free seed is one of the most effective control measures against halo blight disease.
In Serbia, commercial bean seed is used for planting, from native varieties predominate in the production. Certified seeds are seldom used, and even in the case of the new domestic cultivars for which seed production has been organized, it is used only in the first production year (Todorovi et al., 2008; Popov et al., 2010). The overall objective of this study was to evaluate the detection method and determine the occurrence of Psp on common bean seed collected from different genotypes and populations in Serbia. A number of plating assays and semi-selective media have been developed and improved to detect Psp in bean seed (Mohan and Schaad, 1987; Gozczynska and Serfontein, 1998). We conducted a detection method by sowing the extract obtained from the soaked whole bean seed onto the semi-selective media MSP and MT. This method was previously reported on the artificial infected bean seed by Balaž et al. (2008). The results of our study showed that the direct plating method can be routinely used for the detection of the Psp on bean seed, because of its reliability.
The Psp colonies that formed on the semi-selective medium MSP after four days of incubation were light yellow, convex and shiny and medium around the colony became turned light yellow (Mohan and Schaad, 1987; Jansing and Rudolph, 1990, 1996; Kurowski and Remeeus, 2008). The colonies formed on the semi-selective medium MT are creamy white, flat and circular after four-five days of incubation (Goszczynska and Serfontein, 1998; ISF, 2006; Kurowski and Remeeus, 2008). Many colonies of Psp on MT can form a pale-blue fluorescent pigment when using UV light (ISF, 2006). Goszczynska and Serfontein (1998) described the use of the semi-selective medium MT for routine seed testing in the laboratory and for the detection and distinction of all bacterial pathogens screened for in dry bean seed and plant material. This study confirmed the advantage of the MT medium, which allowed simultaneous detection of Xap and Psp on the tested bean cultivar Oplenac, where seeds were infected with both of the bacteria. Other media such as King B (Taylor, 1970), KBC (Mohan and Schaad, 1987; NSHS, 2002) and LPGA can also be used for the isolation of Psp from bean seeds.
Pathogenicity tests were carried out to establish if the obtained strains were typically pathogenic. Psp formed dark green, greasy spots on inoculated bean cotyledons. The rapid confirmation to identify the obtained strains in this assay as Psp was achieved by using ELISA and PCR tests. When using ELISA test in identifying Psp 104 bacteria in 1 mL can be detected (Barzic and Trigalet, 1982), while the use of the other serological method - immunefluorescence (IF) - makes it possible to detect 102 cells in 1 mL (Bazzi and Calzolari, 1982). Van Vuurde and Van den Bovenkamp (1987, 1989) reported that IF is a fast, simple and cheap method for the detection of Psp on bean seed, but they also noted that the reliability of the results depends on the specific and appropriate solutions of antisera. Van Vuurde et al. (1983) reported the application of IF and ELISA tests as potential routine tests for the detection of Psp and Xap in bean seed. Serological methods were considered to be an important diagnostic test in the detection of bacteria causing blight of bean. Certainly, there are numerous restrictions and warnings that must be done before a method is standardized and accepted for routine seed health testing (Weaver and Guthrie, 1978; Sheppard et al., 1986).
The nested-PCR assay with the specific primers pair P5.1/P3.1 and P5.2/P3.2, which directed the amplification of the 500-bp and 450-bp DNA fragments, respectively (Schaad et al., 2001), was included to confirm the identity of all the obtained strains from this study. Audy et al. (1996) reported the possibility of using the PCR method for simultaneous detection of Xap and Psp in bean seed. A combination of specific primers was used to detect two bacteria specifically, but the other tested pathogenic bacteria did not give positive results. The method is sensitive, can detect as few as 1 infected seed in 10.000 seeds, and has a great potential for the detection of these two bacteria in commercial seeds (Audy et al., 1996).
The presence of Psp was recorded on only two out of 23 common bean seed samples analyzed from the Novi Sad area in Serbia. The lower number of seed samples infected by Psp indicated that the bacterium is not widespread, which is attributed to the warmer and drier weather conditions in Serbia in recent years as well as to the use of halo blight resistant bean cultivars. Presented results provide a good basis for a widely routine use, because the detection method is applicable, fast, reliable and cheap and therefore recommended for the use by the phytosanitary service in Serbia.
References
Audy, P.; Braat, C.E.; Saindon, G.; Huang, H.C.; Laroche, A. 1996. A rapid and sensitive PCR-based assay for concurrent detection of bacteria causing common and halo blights in bean seed. Phytopathology 86: 361-366.
Balaž, J. 1990. Contribution to the study of string-bean and bean susceptibility to Pseudomonas syringae pv. phaseolicola (Burkholder) Young, Dye et Wilkie. Zatita bilja 41: 431-453 (in Serbian, with abstract in English).
Balaž, J.; Popovi, T.; Vasi, M.; Nikoli, Z. 2008. Elaboration of methods for detection of Pseudomonas savastanoi pv. phaseolicola on bean seeds. Pesticidi i Fitomedicina 23: 81-88 (in Serbian, with abstract in English).
Barzic, M.R.; Trigalet, A. 1982. Detection de Pseudomonas phaseolicola (Burkh.) Dowson par la technique ELISA. Agronomie 2: 389-398.
Bazzi, C.; Calzolari, A. 1982. Detection of Pseudomonas phaseolicola (Burkh.) Dowson in bean seed with immunofluorescence staining (indirect method). Informatore Fitopatologico 32: 55-58 (in Italian).
Fahy, P.C.; Lloyd, A.B. 1983. Pseudomonas: The Flourescent Pseudomonads. p. 141-188. In: Fahy, P.C.; Persley, G.J., eds. Plant bacterial diseases: a diagnostic guide. Academic Press, Sydney, Australia.
Fenwick, H.S.; Guthrie, J.W. 1969. An improved Pseudomonas phaseolicola pathogenicity test. Phytopathology 59: 11.
Goszczynska, T.; Serfontein, J.J. 1998. Milk tween agar, a semiselective medium for isolation and differentiation of Pseudomonas syringae pv. syringae, Pseudomonas syringae pv. phaseolicola and Xanthomonas axonopodis pv. phaseoli. Journal of Microbiological Methods 32: 65-72.
Guthrie, J.W.; Huber, D.M.; Fenwick, H.S. 1965. Serological detection of halo blight. Plant Disease Reporter 49: 297-299.
Grogan, R.G.; Kimble, K.A. 1967. The role of seed contamination in the transmission of Pseudomonas phaseolicola in Phaseolis vulgaris. Phytopathology 57: 28-31.
Güven, K.; Jones, J.B.; Momol, M.T.; Dickstein, E.R. 2004. Phenotypic and genetic diversity among Pseudomonas syringae pv. phaseolicola. Journal of Phytopathology 152: 658-666.
International Seed Testing (ISF). 2006. Method for the Detection of Pseudomonas savastanoi pv. phaseolicola on Bean Seed. ISHI-Veg, Nyon, Switzerland. Available at: http://www.worldseed.org/cms/medias/file/TradeIssues/PhytosanitaryMatters/SeedHealthTesting/ISHI-Veg [Accessed Aug. 2, 2011]
Jansing, H.; Rudolph, K. 1990. A sensitive and quick test for determination of bean seed infestation by Pseudomonas syringae pv. phaseolicola. Journal Zeitschrift für Pflanzenkrankheiten und Pflanzenschutz 97: 42-55.
Jansing, H.; Rudolph, K. 1996. Pseudomonas syringae pv. phaseolicola: ISTA Handbook on Seed Health Testing. International Seed Testing Association, Zurich, Switzerland. (Working Sheet, 66).
King, E.O.; Ward, M.K.; Raney, D.E. 1954. Two simple media for the demonstration of pyocyanin and fluorescin. Journal of Laboratory and Clinical Medicine 44: 301-307.
Kurowski, C.; Remeeus, P.M. 2007. Proposal for a new method for detecting Pseudomonas savastanoi pv. phaseolicola on bean seeds. ISTA Method Validation Reports 4: 1-12. Available at: http://www.seedtest.org/upload/cms/user/MVR2007.pdf [Accessed Jan. 1, 2007]
Kurowski, C.; Remeeus, P.M. 2008. Detection of Pseudomonas savastanoi pv. phaseolicola on Phaseolus vulgaris. International Seed Testing Association, Bassersdorf, Switzerland. (International Rules for Seed Testing, 7-023). Available at: http://www.seedtest.org/upload/cms/user/7-023.pdf. [Accessed Jan. 1, 2008]
Lahman, L.K.; Schaad, N.W. 1985. Evaluation of the 'dome test' as a reliable assay for seedborne bacterial blight pathogens of beans. Plant Disease 69: 680-683.
Lelliott, R.A.; Stead, D.E. 1987. Methods for the Diagnosis of Bacterial Diseases of Plants. Blackwell, Oxford, UK.
Mohan, S.K.; Schaad, N.W. 1987. An improved agar plating assay for detecting Pseudomonas syringae pv. syringae and P. s. pv. phaseolicola in contaminated bean seed. Phytopathology 77: 1390-1395.
Mosqueda, C.G.; Herrera, E.L. 1997. A simple and efficient PCR method for the specific detection of Pseudomonas syringae pv. phaseolicola in bean seeds. World Journal of Microbiology & Biotechnology 13: 463-467.
National Seed Health System [NSHS]. 2002. Reference manual B (RM-B): seed health testing and phytosanitary field inspection methods manual. Available at: http://www.seedhealth.org/files/pdf/RMB0402.pdf. [Accessed Sep. 18, 2002]
Popovi, T.; Balaž, J.; Nikoli, Z.; Starovi, M.; Gavrilovi, V.; Aleksi, G.; Vasi, M.; ivkovi, S. 2010. Detection and identification of Xanthomonas axonopodis pv. phaseoli on bean seed collected in Serbia. African Journal of Agricultural Research 5: 2730-2736.
Prosen, D.; Hatziloukas, E.; Schaad, N.W.; Panopoulos, N.J. 1993. Specific detection of Pseudomonas syringae pv. phaseolicola DNA in bean seed by polymerase chain reaction-based amplification of a phaseolotoxin gene region. Phytopathology 83: 965-970.
Schaad, N.W.; Cheong, S.S.; Tamaki, S.; Hatziloukas, E.; Panopoulos, N.J. 1995. A combined biological and enzymatic amplification (BIO-PCR) technique to detect Pseudomonas syringae pv. phaseolicola in bean seed extracts. Phytopathology 85: 243-248.
Schaad, N.W.; Jones, J.B.; Chun, W. 2001. Laboratory Guide for Identification of Plant Pathogenic Bacteria. APS Press, St. Paul, MN, USA.
Schwartz, H.F. 1989. Halo blight. In: Schwartz, H.F.; Pastor-Coralles, M.A., eds. Bean production problems in the tropics. Centro Internacional de Agricultura Tropical, Cali, Colombia.
Sheppard, J.W.; Wright, P.F.; DeSavingy, D.H. 1986. Methods for the evaluation of EIA for use in the detection of seed-borne diseases. Seed Science and Technology 14: 49-59.
Taylor, J.D. 1970. The quantitative estimation of the infection of bean seed with Pseudomonas phaseolicola (Burkh.) Dowson. Annals of Applied Biology 66: 29-36.
Taylor, J.D.; Dudley, C.L.; Presly, L. 1979. Studies of halo-blight seed infection and disease transmission in dwarf beans. Annals of Applied Biology 93: 267-277.
Todorovi, J.; Vasi, M.; Todorovi, J. 2008. Pasulj i Boranija. Institute for Vegetable and Field Crops. Novi Sad, Serbia. .
Van Vuurde, J.W.L.; Van den Bovenkamp, G.W. 1987. Pseudomonas syringae pv. phaseolicola:. ISTA Handbook on Seed Health Testing. International Seed Testing Association. Zurich, Switzerland. (Working Sheet, 65).
Van Vuurde, J.W.L.; Van den Bovenkamp, G.W. 1989. Detection of Pseudomonas syringae pv. phaseolicola in bean. p. 30-40. In: Saettler, A.W.; Schaad, N.W.; Roth, D.A., eds. Detection of bacteria in seed and other planting material, The American Phytopathological Society Press, St. Paul, USA.
Van Vuurde, J.W.L.; Van den Bovenkamp, G.W.; Birnbaum, Y. 1983. Immunofluorescence microscopy and enzymelinked immunosorbent assay as potential routine tests for the detection of Pseudomonas syringae pv. phaseolicola and Xanthomonas campestris pv. phaseoli in bean seed. Seed Science and Technology 11: 547-559.
Weaver, W.M.; Guthrie, J.W. 1978. Enzyme linked immunospecific assay: application to the detection of seed-borne bacteria. Phytopathology News 12: 156.
Webster, D.M.; Atkin, J.D.; Cross, J.E. 1983. Bacterial blights of snap beans and their control. Plant Disease 67: 935-940.
(Received March 03, 2011)
(Accepted November 22, 2011)
Edited by: Cláudio Marcelo Gonçalves de Oliveira
- Audy, P.; Braat, C.E.; Saindon, G.; Huang, H.C.; Laroche, A. 1996. A rapid and sensitive PCR-based assay for concurrent detection of bacteria causing common and halo blights in bean seed. Phytopathology 86: 361-366.
- Balaž, J. 1990. Contribution to the study of string-bean and bean susceptibility to Pseudomonas syringae pv. phaseolicola (Burkholder) Young, Dye et Wilkie. Zatita bilja 41: 431-453 (in Serbian, with abstract in English).
- Balaž, J.; Popovi, T.; Vasi, M.; Nikoli, Z. 2008. Elaboration of methods for detection of Pseudomonas savastanoi pv. phaseolicola on bean seeds. Pesticidi i Fitomedicina 23: 81-88 (in Serbian, with abstract in English).
- Barzic, M.R.; Trigalet, A. 1982. Detection de Pseudomonas phaseolicola (Burkh.) Dowson par la technique ELISA. Agronomie 2: 389-398.
- Bazzi, C.; Calzolari, A. 1982. Detection of Pseudomonas phaseolicola (Burkh.) Dowson in bean seed with immunofluorescence staining (indirect method). Informatore Fitopatologico 32: 55-58 (in Italian).
- Fahy, P.C.; Lloyd, A.B. 1983. Pseudomonas: The Flourescent Pseudomonads. p. 141-188. In: Fahy, P.C.; Persley, G.J., eds. Plant bacterial diseases: a diagnostic guide. Academic Press, Sydney, Australia.
- Fenwick, H.S.; Guthrie, J.W. 1969. An improved Pseudomonas phaseolicola pathogenicity test. Phytopathology 59: 11.
- Goszczynska, T.; Serfontein, J.J. 1998. Milk tween agar, a semiselective medium for isolation and differentiation of Pseudomonas syringae pv. syringae, Pseudomonas syringae pv. phaseolicola and Xanthomonas axonopodis pv. phaseoli Journal of Microbiological Methods 32: 65-72.
- Guthrie, J.W.; Huber, D.M.; Fenwick, H.S. 1965. Serological detection of halo blight. Plant Disease Reporter 49: 297-299.
- Grogan, R.G.; Kimble, K.A. 1967. The role of seed contamination in the transmission of Pseudomonas phaseolicola in Phaseolis vulgaris Phytopathology 57: 28-31.
- Güven, K.; Jones, J.B.; Momol, M.T.; Dickstein, E.R. 2004. Phenotypic and genetic diversity among Pseudomonas syringae pv. phaseolicola. Journal of Phytopathology 152: 658-666.
- International Seed Testing (ISF). 2006. Method for the Detection of Pseudomonas savastanoi pv. phaseolicola on Bean Seed. ISHI-Veg, Nyon, Switzerland. Available at: http://www.worldseed.org/cms/medias/file/TradeIssues/PhytosanitaryMatters/SeedHealthTesting/ISHI-Veg [Accessed Aug. 2, 2011]
- Jansing, H.; Rudolph, K. 1990. A sensitive and quick test for determination of bean seed infestation by Pseudomonas syringae pv. phaseolicola Journal Zeitschrift für Pflanzenkrankheiten und Pflanzenschutz 97: 42-55.
- Jansing, H.; Rudolph, K. 1996. Pseudomonas syringae pv. phaseolicola: ISTA Handbook on Seed Health Testing. International Seed Testing Association, Zurich, Switzerland. (Working Sheet, 66).
- King, E.O.; Ward, M.K.; Raney, D.E. 1954. Two simple media for the demonstration of pyocyanin and fluorescin. Journal of Laboratory and Clinical Medicine 44: 301-307.
- Kurowski, C.; Remeeus, P.M. 2007. Proposal for a new method for detecting Pseudomonas savastanoi pv. phaseolicola on bean seeds. ISTA Method Validation Reports 4: 1-12. Available at: http://www.seedtest.org/upload/cms/user/MVR2007.pdf [Accessed Jan. 1, 2007]
- Kurowski, C.; Remeeus, P.M. 2008. Detection of Pseudomonas savastanoi pv. phaseolicola on Phaseolus vulgaris International Seed Testing Association, Bassersdorf, Switzerland. (International Rules for Seed Testing, 7-023). Available at: http://www.seedtest.org/upload/cms/user/7-023.pdf. [Accessed Jan. 1, 2008]
- Lahman, L.K.; Schaad, N.W. 1985. Evaluation of the 'dome test' as a reliable assay for seedborne bacterial blight pathogens of beans. Plant Disease 69: 680-683.
- Lelliott, R.A.; Stead, D.E. 1987. Methods for the Diagnosis of Bacterial Diseases of Plants. Blackwell, Oxford, UK.
- Mohan, S.K.; Schaad, N.W. 1987. An improved agar plating assay for detecting Pseudomonas syringae pv. syringae and P. s. pv. phaseolicola in contaminated bean seed. Phytopathology 77: 1390-1395.
- Mosqueda, C.G.; Herrera, E.L. 1997. A simple and efficient PCR method for the specific detection of Pseudomonas syringae pv. phaseolicola in bean seeds. World Journal of Microbiology & Biotechnology 13: 463-467.
- Popovi, T.; Balaž, J.; Nikoli, Z.; Starovi, M.; Gavrilovi, V.; Aleksi, G.; Vasi, M.; ivkovi, S. 2010. Detection and identification of Xanthomonas axonopodis pv. phaseoli on bean seed collected in Serbia. African Journal of Agricultural Research 5: 2730-2736.
- Prosen, D.; Hatziloukas, E.; Schaad, N.W.; Panopoulos, N.J. 1993. Specific detection of Pseudomonas syringae pv. phaseolicola DNA in bean seed by polymerase chain reaction-based amplification of a phaseolotoxin gene region. Phytopathology 83: 965-970.
- Schaad, N.W.; Cheong, S.S.; Tamaki, S.; Hatziloukas, E.; Panopoulos, N.J. 1995. A combined biological and enzymatic amplification (BIO-PCR) technique to detect Pseudomonas syringae pv. phaseolicola in bean seed extracts. Phytopathology 85: 243-248.
- Schaad, N.W.; Jones, J.B.; Chun, W. 2001. Laboratory Guide for Identification of Plant Pathogenic Bacteria. APS Press, St. Paul, MN, USA.
- Schwartz, H.F. 1989. Halo blight. In: Schwartz, H.F.; Pastor-Coralles, M.A., eds. Bean production problems in the tropics. Centro Internacional de Agricultura Tropical, Cali, Colombia.
- Sheppard, J.W.; Wright, P.F.; DeSavingy, D.H. 1986. Methods for the evaluation of EIA for use in the detection of seed-borne diseases. Seed Science and Technology 14: 49-59.
- Taylor, J.D. 1970. The quantitative estimation of the infection of bean seed with Pseudomonas phaseolicola (Burkh.) Dowson. Annals of Applied Biology 66: 29-36.
- Taylor, J.D.; Dudley, C.L.; Presly, L. 1979. Studies of halo-blight seed infection and disease transmission in dwarf beans. Annals of Applied Biology 93: 267-277.
- Todorovi, J.; Vasi, M.; Todorovi, J. 2008. Pasulj i Boranija. Institute for Vegetable and Field Crops. Novi Sad, Serbia.
- Van Vuurde, J.W.L.; Van den Bovenkamp, G.W. 1987. Pseudomonas syringae pv. phaseolicola:. ISTA Handbook on Seed Health Testing. International Seed Testing Association. Zurich, Switzerland. (Working Sheet, 65).
- Van Vuurde, J.W.L.; Van den Bovenkamp, G.W. 1989. Detection of Pseudomonas syringae pv. phaseolicola in bean. p. 30-40. In: Saettler, A.W.; Schaad, N.W.; Roth, D.A., eds. Detection of bacteria in seed and other planting material, The American Phytopathological Society Press, St. Paul, USA.
- Van Vuurde, J.W.L.; Van den Bovenkamp, G.W.; Birnbaum, Y. 1983. Immunofluorescence microscopy and enzymelinked immunosorbent assay as potential routine tests for the detection of Pseudomonas syringae pv. phaseolicola and Xanthomonas campestris pv. phaseoli in bean seed. Seed Science and Technology 11: 547-559.
- Weaver, W.M.; Guthrie, J.W. 1978. Enzyme linked immunospecific assay: application to the detection of seed-borne bacteria. Phytopathology News 12: 156.
- Webster, D.M.; Atkin, J.D.; Cross, J.E. 1983. Bacterial blights of snap beans and their control. Plant Disease 67: 935-940.
Publication Dates
-
Publication in this collection
28 Mar 2012 -
Date of issue
2012
History
-
Received
03 Mar 2011 -
Accepted
22 Nov 2011