Abstract
Only a limited number of simple sequence repeat (SSR) markers is available for the genome of garlic (Allium sativum L.) despite the fact that SSR markers have become one of the most preferred DNA marker systems. To develop new SSR markers for the garlic genome, garlic expressed sequence tags (ESTs) at the publicly available GarlicEST database were screened for SSR motifs and a total of 132 SSR motifs were identified. Primer pairs were designed for 50 SSR motifs and 24 of these primer pairs were selected as SSR markers based on their consistent amplification patterns and polymorphisms. In addition, two SSR markers were developed from the sequences of garlic cDNA-AFLP fragments. The use of 26 EST-SSR markers for the assessment of genetic relationship was tested using 31 garlic genotypes. Twenty six EST-SSR markers amplified 130 polymorphic DNA fragments and the number of polymorphic alleles per SSR marker ranged from 2 to 13 with an average of 5 alleles. Observed heterozygosity and polymorphism information content (PIC) of the SSR markers were between 0.23 and 0.88, and 0.20 and 0.87, respectively. Twenty one out of the 31 garlic genotypes were analyzed in a previous study using AFLP markers and the garlic genotypes clustered together with AFLP markers were also grouped together with EST-SSR markers demonstrating high concordance between AFLP and EST-SSR marker systems and possible immediate application of EST-SSR markers for fingerprinting of garlic clones. EST-SSR markers could be used in genetic studies such as genetic mapping, association mapping, genetic diversity and comparison of the genomes of Allium species.
expressed sequence tags (EST); simple sequence repeat markers; genetic relationship
Introduction
Garlic (Allium sativum L.) is one of the important cultivated species of the genus Allium and it is classified as belonging to the Alliaceae family. Garlic has a very large genome (33.5 pg/2C, Ranjekar et al., 1978Ranjekar, P. K.; Pallotta, D.; Lafontaine, J. G. 1978. Analysis of plant genomes. V. Comparative study of molecular properties of DNAs of seven Allium species. Biochemical Genetics 16: 957-970.) which is a characteristic of many Allium species, that complicates molecular and genetic studies of garlic compared to other molecularly well-investigated plant species with smaller genome sizes. Most of the molecular studies on garlic have focused on the analysis of genetic diversity among cultivated garlic genotypes.
The genetic diversity between garlic clones has generally been assessed using amplified fragment length polymorphism (AFLP), randomly amplified polymorphic DNA (RAPD) and isozyme markers (Ipek et al., 2003Ipek, M.; Ipek, A.; Simon, P. W. 2003. Comparison of AFLPs, RAPD markers, and isozymes for diversity assessment of garlic and detection of putative duplicates in germplasm collections. Journal of the American Society for Horticultural Science 128: 246-252.; Buso et al., 2008Buso, G. S. C.; Paiva, M. R.; Torres, A. C.; Resende, F. V.; Ferreira, M. A.; Buso, J. A.; Dusi, A. N. 2008. Genetic diversity studies of Brazilian garlic cultivars and quality control of garlic-clover production. Genetic and Molecular Research 7: 534-541.; Ipek et al., 2008aIpek, M.; Ipek, A.; Simon, P. W. 2008a. Molecular characterization of Kastamonu garlic: an economically important garlic clone in Turkey. Scientia Horticulturae 115: 203-208., Paredes et al., 2008Paredes, C. M.; Becerra, V. V.; González, A. M. I. 2008. Low genetic diversity among garlic (Allium sativum L.) accessions detected using random amplified polymorphic DNA (RAPD). Chilean Journal of Agricultural Research 68: 3-12.; Morales et al., 2013Morales, R. G. F.; Resende, J. T. V.; Resende, F. V.; Delatorre, C. A.; Figueiredo, A. S. T.; Da-Silva, P. R. 2013. Genetic divergence among Brazilian garlic cultivars based on morphological characters and AFLP markers. Genetic and Molecular Research 12: 270-281.). In addition, molecular markers have been utilized for the development of a low density genetic map for garlic (Ipek et al., 2005Ipek, M.; Ipek, A.; Almquist, S. G.; Simon, P. W. 2005. Demonstration of linkage and development of the first low-density genetic map of garlic based on AFLP markers. Theoretical and Applied Genetics 110: 228-236.; Zewdie et al., 2005Zewdie, Y.; Havey, M. J.; Prince, J. P.; Jenderek, M. M. 2005. The first genetic linkages among expressed regions of the garlic genome. Journal of the American Society for Horticultural Science 130: 569-574.). In order to increase the density of the genetic maps and anchor dominant markers such as AFLPs and RAPDs, it is essential to develop co-dominant DNA markers like simple sequence repeats (SSRs) and single nucleotide polymorphisms (SNPs) for the garlic genome.
SSR markers have become one of the most preferred marker systems because they are typically co-dominant, reproducible, cross-species transferable and highly polymorphic. Recently, DNA, cDNA and EST (expressed sequence tag) sequence databases of plant species are mined with computer programs to identify SSR motifs. Using this effective approach, large numbers of SSR markers for expressed regions have been successfully developed for many plant species (Lu et al., 2013Lu, J. J.; Kang, J. Y.; Feng, S. G.; Zhao, H. Y.; Liu, J. J.; Wang, H. Z. 2013. Transferability of SSR markers derived from Dendrobium nobile expressed sequence tags (ESTs) and their utilization in Dendrobium phylogeny analysis. Scientia Horticulturae 158: 8-15.; Blair and Hurtado, 2013Blair, M. W.; Hurtado, N. 2013. EST-SSR markers from five sequenced cDNA libraries of common bean (Phaseolus vulgaris L.) comparing three bioinformatic algorithms. Molecular Ecology Resources 13: 688-695.; Mohanty et al., 2013Mohanty, P.; Sahoo, L.; Parida, K.; Das, P. 2013. Development of polymorphic EST-SSR markers in Macrobrachium rosenbergii by data mining. Conservation and Genetic Resources 5: 133-136.).
With the development of next generation DNA sequencing technologies, large nucleotide sequence databases of EST and genomic DNA have been generated for many plant species and are available for screening simple sequence repeats (Yang et al., 2012Yang, T.; Bao, S. Y.; Ford, R.; Jia, T. J.; Guan, J. P.; He, Y. H.; Sun, X. L.; Jiang, J. Y.; Hao, J. J.; Zhang, X. Y.; Zong, X. X. 2012. High-throughput novel microsatellite marker of faba bean via next generation sequencing. BMC Genomics 13: 602.; Zalapa et al., 2012Zalapa, J. E.; Cuevas, H.; Zhu, H.; Steffan, S.; Senalik, D.; Zeldin, E.; Mccown, B.; Harbut, R.; Simon, P. 2012. Using next-generation sequencing approaches to isolate simple sequence repeat (SSR) loci in the plant sciences. American Journal of Botany 99: 193-208.; Wu et al., 2013Wu, H. L.; Chen, D.; Li, J. X.; Yu, B.; Qiao, X. Y.; Huang, H. L.; He, Y. M. 2013. De novo characterization of leaf transcriptome using 454 sequencing and development of EST-SSR markers in Tea (Camellia sinensis). Plant Molecular Biology Reporter 31: 524-538.; Gilmore et al., 2013Gilmore, B.; Bassil, N.; Nyberg, A.; Knaus, B.; Smith, D.; Barney, D. L.; Hummer, K. 2013. Microsatellite marker development in Peony using next generation sequencing. Journal of the American Society for Horticultural Science 138: 64-74.). SSR markers based on cDNA or EST have a number of additional advantages such as being tightly linked to traits of interest, having a high rate of transferability between related species, and providing genetic diversity and mapping analyses for the expressed regions (Varshney et al., 2005Varshney, R. K.; Graner, A.; Sorrells, M. E. 2005. Genic microsatellite markers in plants: features and applications. Trends in Biotechnology 23: 48-55.).
For the garlic genome, however, a limited number of SSR markers have been developed and too few are available to generate high density genetic maps in a crop with such a large genome size (Cunha et al., 2012Cunha, C. P.; Hoogerheide, E.S.; Zucchi, M. I.; Monteiro, M.; Pinheiro, J. B. 2012. New microsatellite markers for garlic, Allium sativum (Alliaceae). American Journal of Botany 99: e17-19.; Ipek et al., 2012Ipek, M.; Ipek, A.; Cansev, A.; Seniz, V.; Simon, P. W. 2012. Development of EST based SSR markers for the garlic genome. Acta Horticulturae 969: 81-83.; Khar et al., 2012Khar, A. 2012. Cross amplification of onion derived microsatellites and mining of garlic EST database for assessment of genetic diversity in garlic. Acta Horticulturae 969: 289-295.). To improve genetic maps and genetic diversity analysis, more SSR markers need to be developed for the garlic genome. This study aimed to develop new SSR markers from the garlic EST sequences and to validate the utility of these markers by evaluating genetic relationships between diverse garlic clones.
Materials and Methods
Thirty one diverse garlic clones obtained in Pullman (46o43’54” N, 117o09’28” W), in the state of Washington, USA were used to find allelic polymorphisms in EST-SSR markers (Table 1). Twenty one of these garlic clones had been previously characterized with AFLP markers (Ipek et al., 2003Ipek, M.; Ipek, A.; Simon, P. W. 2003. Comparison of AFLPs, RAPD markers, and isozymes for diversity assessment of garlic and detection of putative duplicates in germplasm collections. Journal of the American Society for Horticultural Science 128: 246-252.) and they were selected to represent ten phylogenetic groups identified in this AFLP genetic diversity assessment study. Young leaf samples were collected from each clones at the 3-5 leaf stages, freeze-dried for 3-4 days and powdered with a paint shaker.
DNA samples were extracted from 20 mg of lyophilized leaf samples by using DNeasy Plant Mini Kit (Qiagen, Germany). The DNA concentration of each sample was measured using a Qubit Fluorometer and adjusted to 30 ng µL–1 in tris-ethylene diamine tetraacetic acid (TE) buffer.
SSR motifs of 2-6 nucleotide repeats in 7720 non-redundant garlic EST sequences at the publicly available GarlicESTdb (http://garlicdb.kribb.re.kr) (Kim et al., 2009Kim, D. W.; Jung, T. S.; Nam, S. H.; Kwon, H. R.; Kim, A.; Chae, S. H.; Choi, S. H.; Kim, D. W.; Kim, R. N.; Park, H. S. 2009. GarlicESTdb: an online database and mining tool for garlic EST sequences. BMC Plant Biology 9: 61.) and in ESTs derived from cDNA-AFLP fragments (unpublished EST sequences ) were screened using SSR Locator v.1 computer software . Primer pairs were designed from the flanking sequences to SSR repeat motifs using the SSR Locater program. Forward primers were tailed with M13 sequence (GACGTTGTAAAACGACGGCC) (Schuelke, 2000Schuelke, M. 2000. An economic method for the fluorescent labeling of PCR fragments. Nature Biotechnology 18: 233-234.).
Each 20 μL PCR reaction mixture contained 1.0 U Taq DNA polymerase (Fermentas, CA, USA) with supplied reaction buffer at 1 × concentration, 0.25 mM of each dNTP, 1.5 mM MgCl2, 0.10 μM of M13 sequence tailed forward primer, 0.20 μM of reverse primer, 0.20 μM of M13 primer labeled with LI-COR infrared dye either at 700 nm or 800 nm (LI-COR, Lincoln, Neb., USA) and 45 ng of template DNA.
The thermal cycling conditions were as follows: 2 min at 94 °C; 6 cycles of 45 sec at 94 °C, 1 min starting at 5 °C above the annealing temperature of each primer set. The annealing temperature was reduced 1 °C after each cycle, and 1 min and 10 sec at 72 °C for extension; 28 cycles of 45 sec at 94 °C, 1 min at the annealing temperature of each primer set, and 1 min and 10 sec at 72 °C; 7 cycles of 45 sec at 94 °C, 1 min at 54 °C, and 1 min and 10 sec at 72 °C and a final extension step of 15 min at 72 °C.
An Applied Biosystems thermal cycler was used for the PCR reactions. The PCR products, diluted 20 times in formamide loading buffer, were denatured at 94 °C for 4 min and immediately placed on ice. The denatured PCR products were separated on 6 % denaturing polyacrylamide sequencing gels by running at 30 W for 2-3 h with a LI-COR 4300 automated sequencer system. The putative function of EST sequences with polymorphic SSR repeats were determined by “blasting” their putative amino acid sequences to the GenBank database at National Center for Biotechnology Information (NCBI) by using BLASTX program.
The PCR reaction mixtures for analysis of polymorphic SSR markers, thermal cycling conditions and size fractionations of PCR products were performed as described above. Allele sizes of each SSR marker were determined using Saga GT Software (LI-COR) according to the 50-350 bp sizing standard labeled with either IRDye700 or IRDye800 (LI-COR). All alleles were manually identified from each SSR marker and were scored as absent (0) or present (1) to obtain binary data. The EST-SSR data were bootstrapped by resampling 2000 times using TREECON v.1.3b computer software. During the construction of UPGMA dendrogram, Nei and Li (1979)Nei, M.; Li, W. H. 1979. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proceedings of the National Academy of Sciences of the United States of America 76: 5269-5273. distance coefficient was used. Observed heterozygosity and the polymorphism information content (PIC) values for each SSR marker were calculated using PIC Calculator Extra (http://www.genomics.liv.ac.uk/animal/pic.html).
Results and Discussion
Screening 7720 non-redundant garlic EST sequences using the minimum repeat unit criteria of nine for dinucleotides, six for trinucleotides and four for tetranucleotides, pentanucleotides, and hexanucleotides revealed 132 SSR motifs in 125 ESTs (some ESTs contained more than one SSR motif). The frequency of ESTs with SSR motifs in garlic was 2 % which was low compared to the frequency of ESTs with SSR motifs of 8 % in barley (Hordeum vulgare L.) (Thiel et al., 2003Thiel, T.; Michalek, W.; Varshney, R. K.; Graner, A.; 2003. Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theoretical and Applied Genetics 106: 411-422.), 8 % in sesame (Sesamum indicum L.) (Wei et al., 2008Wei, L. B.; Zhang, H. Y.; Zheng, Y. Z.; Guo, W. Z.; Zhang, T. Z. 2008. Developing EST-derived microsatellites in sesame (Sesamum indicum L.). Acta Agronomica Sinica 34: 2077-2084.) and 19 % in opium poppy (Papaver somniferum L.) (Şelale et al., 2013Şelale, H.; Çelik, I.; Gültekin, V.; Allmer, J.; Doğanlar, S.; Frary, A. 2013. Development of EST-SSR markers for diversity and breeding studies in opium poppy. Plant Breeding 132: 344-351.).
Among the 132 repeats, dinucleotide repeats (60 %) were the most abundant repeat motifs and 34 (57 %) of these dinucleotide repeats were (AT/TA)n repeats. The remaining 34 repeats (26 %) were tri-, 16 repeats (12 %) were tetra-, six repeats (5 %) were penta- and 16 repeats (12 %) were hexa-nucleotides. This result supported the findings of a previous study indicating that garlic genome is rich in A/T bases (Kirk et al., 1970Kirk, J. T.; Rees, O. H.; Evans, G. 1970. Base composition of nuclear DNA within the genus Allium. Heredity 25: 507-512.). Lengths of SSR motifs ranged from 16 to 172 bp. SSR motifs with 16 bp were obtained from (tetranucleotide)4 repeat motif and 172 bp were obtained from (GA)86 repeat motif.
Trinucleotide motifs are the most common repeats in plant species (Varshney et al., 2005Varshney, R. K.; Graner, A.; Sorrells, M. E. 2005. Genic microsatellite markers in plants: features and applications. Trends in Biotechnology 23: 48-55.) because simple sequence repeats, other than trinucleotides or hexanucleotides, cause frame shift mutations which result in nonfunctional gene products if they are located in the coding region of a gene (Kalia et al., 2011Kalia, P. J.; Rai, M. K.; Kalia, S.; Singh, R.; Dhawan, A. D. 2011. Microsatellite markers: an overview of the recent progress in plants. Euphytica 177: 309-334.). However, dinucleotide repeats were the most abundant in garlic. Trinucleotide repeats were the second most abundant and (AAG)n repeats were the most frequent trinucleotide repeat motif in garlic, as in other plant species (Li et al., 2004Li, Y. C.; Korol, A. B.; Fahima, T.; Nevo, E. 2004. Microsatellites within genes: structure, function, and evolution. Molecular Biology and Evolution 21: 991-1007.). Hexanucleotide repeats were more abundant than pentanucleotide repeats, which were comparable in number to tetranucleotide repeats.
Primer pairs were synthesized for 50 SSR motifs identified in garlic EST sequences at GarlicEST db by using SSR Locater program. Amplification patterns of these 50 SSR motifs were optimized using eight diverse garlic genotypes. Forty five primer pairs amplified DNA fragments but the remaining five did not produce any amplicon. Twenty four primer pairs with good amplification pattern were selected, based on their consistent amplification patterns among the garlic genotypes and polymorphisms (Table 2 and 3; Figure 1). In addition, two more primer pairs designed from the flanking sequences of the SSR motifs in the sequences of garlic cDNA-AFLP fragments were also used. EST sequences carrying polymorphic SSR repeats were blasted to the GenBank database at the National Center for Biotechnology Information (NCBI) to determine their putative functions. Of these 26 polymorphic ESTs, 17 sequences matched with nucleotide sequences of genes in GenBank while nine (35 %) did not match with any sequences indicating the presence of uncharacterized expressed genes which were unique to the garlic genome (Table 2).
– Primer sequences, repeat motifs, primer annealing temperature (Ta), NCBI BLASTX results and GarlicEST db ID number for each SSR loci.
Genetic relationships between 31 garlic genotypes were assessed using 26 EST-SSR markers developed in the present study. Twenty six primer pairs generated 130 polymorphic DNA fragments among 31 garlic genotypes. Number of polymorphic alleles per primer pair ranged from 2 to 13 with an average of 5 alleles per primer pair. The sizes of alleles of EST-SSR markers ranged from 103 to 341 bp. Observed heterozygosity values of EST-SSR markers were between 0.23 and 0.88 while PIC values ranged from 0.20 to 0.87 (Table 3). Cunha et al. (2012)Cunha, C. P.; Hoogerheide, E.S.; Zucchi, M. I.; Monteiro, M.; Pinheiro, J. B. 2012. New microsatellite markers for garlic, Allium sativum (Alliaceae). American Journal of Botany 99: e17-19. developed ten SSR markers from SSR enriched genomic library of garlic and they found 44 alleles among the 75 garlic accessions with an average of 4.4 alleles per locus, similar to our results. Cunha et al. (2012)Cunha, C. P.; Hoogerheide, E.S.; Zucchi, M. I.; Monteiro, M.; Pinheiro, J. B. 2012. New microsatellite markers for garlic, Allium sativum (Alliaceae). American Journal of Botany 99: e17-19. also found PIC values of these SSR markers ranging from 0.19 to 0.78.
For seven (27 %) EST-SSR markers, there were more than two alleles in the genome of a given garlic clone, indicating that these ESTs are present in more than one copy in the diploid genome of garlic (Table 3). Although amino acid sequences of three of these multiple copy ESTs did not match any of the amino acid sequences in GenBank, amino acid sequences of the remaining four multiple copy ESTs matched with seryl-tRNA synthetase (EMS50639.1), elongation factor 1-alpha (BAN42602.1), heat shock protein (ADF31772.1) and conserved transmembrane protein (BAD61224.1) (Table 2). In previous studies, Ipek et al. (2005Ipek, M.; Ipek, A.; Almquist, S. G.; Simon, P. W. 2005. Demonstration of linkage and development of the first low-density genetic map of garlic based on AFLP markers. Theoretical and Applied Genetics 110: 228-236.; 2008b) identified at least six alleles of alliinase, five alleles of chitinase, three alleles of SST-1 and eight alleles of gaLFY in the diploid genome of garlic indicating that at least three copies of alliinase and chitinase, two copies of SST-1 and four copies of gaLFY genes in the garlic genome. In a mapping study of garlic, Ipek et al. (2005)Ipek, M.; Ipek, A.; Almquist, S. G.; Simon, P. W. 2005. Demonstration of linkage and development of the first low-density genetic map of garlic based on AFLP markers. Theoretical and Applied Genetics 110: 228-236. also found numerous AFLP markers segregated in a 15:1 ratio, typical for duplicated loci. Similarly, King et al. (1998)King, J. J.; Bradeen, J. M.; Bark, O.; McCallum, J. A.; Havey, M. J. 1998. A low-density genetic map of onion reveals a role for tandem duplication in the evolution of an extremely large diploid genome. Theoretical and Applied Genetics 96: 52-62. found that about 21 % of the RFLP loci evaluated were duplicated in another Allium species, onion (A. cepa L.). These results support the hypothesis that duplications are common in the garlic genome (Jones and Rees, 1968Jones, R.; Rees, H. 1968. Nuclear DNA variation in Allium. Heredity 23: 591-605.; King et al., 1998King, J. J.; Bradeen, J. M.; Bark, O.; McCallum, J. A.; Havey, M. J. 1998. A low-density genetic map of onion reveals a role for tandem duplication in the evolution of an extremely large diploid genome. Theoretical and Applied Genetics 96: 52-62.).
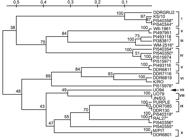
Of 31 diverse garlic genotypes analyzed using these polymorphic EST-SSR markers, 21 had been previously analyzed by Ipek et al. (2003)Ipek, M.; Ipek, A.; Simon, P. W. 2003. Comparison of AFLPs, RAPD markers, and isozymes for diversity assessment of garlic and detection of putative duplicates in germplasm collections. Journal of the American Society for Horticultural Science 128: 246-252. using AFLP, RAPD and isozyme markers and these 21 genotypes represent ten phylogenetic groups determined in that study (Table 1). Genetic relationships between the remaining ten genotypes had not been previously assessed. The UPGMA dendrogram demonstrating genetic relationship between these genotypes was developed using the Nei and Li (1979)Nei, M.; Li, W. H. 1979. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proceedings of the National Academy of Sciences of the United States of America 76: 5269-5273. distance matrix based on the 130 SSR alleles (Figure 2).
– UPGMA dendrogram demonstrating genetic relationships between the 31 garlic genotypes. The dendrogram is based on the Nei and Li (1979) distance matrix calculated using the EST-SSR marker data. Numbers on the branches are the bootstrap values (2000 replications). *Indicates the genotypes which had not been previously analyzed. Roman numerals are the group numbers previously identified with AFLP markers by Ipek et al. (2003).
Ten groups previously determined by Ipek et al. (2003)Ipek, M.; Ipek, A.; Simon, P. W. 2003. Comparison of AFLPs, RAPD markers, and isozymes for diversity assessment of garlic and detection of putative duplicates in germplasm collections. Journal of the American Society for Horticultural Science 128: 246-252. were also identified in the present study and genotypes clustered in these groups were the same in both studies. These results indicate that discrimination of garlic clones using EST-SSR markers agreed with AFLP markers, although EST-SSR markers were developed from expressed regions of garlic genome while AFLP markers were likely derived from throughout the genome.
Accession, U094 was classified as A. longicuspis L. in the Germplasm Resources Information Network (GRIN) of the US Department of Agriculture Agricultural Research Service (USDA-ARS) but the assessment of genetic relationships between U094 and other accessions from A. sativum L. based on AFLP and EST-SSR markers did not suggest that these species are genetically distinct (Figure 2). Accessions from A. sativum and A. longicuspis were clustered together based on their flowering ability but not their geographic origins in the present study. Accessions in groups I, V, VI, VII, VIII, IX and X were flowering, accessions in III and IV were not flowering and accessions in group II were incomplete flowering genotypes (Ipek et al., 2003Ipek, M.; Ipek, A.; Simon, P. W. 2003. Comparison of AFLPs, RAPD markers, and isozymes for diversity assessment of garlic and detection of putative duplicates in germplasm collections. Journal of the American Society for Horticultural Science 128: 246-252.).
Five of ten garlic genotypes which had not been previously analyzed were also placed into ten phylogenetic groups previously identified with AFLP markers, demonstrating the immediate utility of these EST-SSR markers. These five genotypes share common phenotypic characteristics such as flowering ability with garlic clones in the same groups (data not presented). The remaining five newly analyzed genotypes were not placed in any of the ten groups but formed three new groups (Figure 2). PI540319, RAL27 and PI540356 which were clustered together in a new group. Although all of these three genotypes were able to flower, their geographic origins were different (Table 1).
Ipek et al. (2003)Ipek, M.; Ipek, A.; Simon, P. W. 2003. Comparison of AFLPs, RAPD markers, and isozymes for diversity assessment of garlic and detection of putative duplicates in germplasm collections. Journal of the American Society for Horticultural Science 128: 246-252. compared AFLP, RAPD and isozyme markers for their power of discrimination in garlic genotypes. While RAPD and isozyme markers did not find any polymorphism within any group, AFLP markers were able to detect intra group variation. Similarly, in the present genetic diversity analysis based on EST-SSRs, intra group variation was detected in groups I, II, III, IV, V, VI, and IX (Figure 2). EST-SSR markers developed in the present study were effective for determining genetic relationships between garlic genotypes and discriminating closely related garlic clones.
In summary, garlic EST sequences were found to provide numerous SSR motifs and 26 of 52 SSR motifs identified in this study were polymorphic from among the diverse garlic clones tested. Additional polymorphic SSR markers can be developed from garlic EST sequences in a rapid and cost effective way. EST-SSR markers developed in this study can be utilized for diversity and genetic mapping studies for expressed regions of the garlic genome effectively. However, there is still a need to develop many more SSR markers for the generation of high density genetic maps and other genetic studies in the garlic genome.
Acknowledgements
To The Commission of Scientific Research Projects of Uludag University, project number: UAP (Z)-2010/50.
References
- Blair, M. W.; Hurtado, N. 2013. EST-SSR markers from five sequenced cDNA libraries of common bean (Phaseolus vulgaris L.) comparing three bioinformatic algorithms. Molecular Ecology Resources 13: 688-695.
- Buso, G. S. C.; Paiva, M. R.; Torres, A. C.; Resende, F. V.; Ferreira, M. A.; Buso, J. A.; Dusi, A. N. 2008. Genetic diversity studies of Brazilian garlic cultivars and quality control of garlic-clover production. Genetic and Molecular Research 7: 534-541.
- Cunha, C. P.; Hoogerheide, E.S.; Zucchi, M. I.; Monteiro, M.; Pinheiro, J. B. 2012. New microsatellite markers for garlic, Allium sativum (Alliaceae). American Journal of Botany 99: e17-19.
- Gilmore, B.; Bassil, N.; Nyberg, A.; Knaus, B.; Smith, D.; Barney, D. L.; Hummer, K. 2013. Microsatellite marker development in Peony using next generation sequencing. Journal of the American Society for Horticultural Science 138: 64-74.
- Ipek, M.; Ipek, A.; Simon, P. W. 2008a. Molecular characterization of Kastamonu garlic: an economically important garlic clone in Turkey. Scientia Horticulturae 115: 203-208.
- Ipek, M.; Ipek, A.; Almquist, S. G.; Simon, P. W. 2005. Demonstration of linkage and development of the first low-density genetic map of garlic based on AFLP markers. Theoretical and Applied Genetics 110: 228-236.
- Ipek, M.; Ipek, A.; Cansev, A.; Seniz, V.; Simon, P. W. 2012. Development of EST based SSR markers for the garlic genome. Acta Horticulturae 969: 81-83.
- Ipek, M.; Ipek, A.; Simon, P. W. 2003. Comparison of AFLPs, RAPD markers, and isozymes for diversity assessment of garlic and detection of putative duplicates in germplasm collections. Journal of the American Society for Horticultural Science 128: 246-252.
- Ipek, M.; Ipek, A.; Simon, P. W. 2008b. Rapid characterization of garlic clones with locus-specific DNA markers. Turkish Journal of Agriculture and Forestry 32: 357-362.
- Jones, R.; Rees, H. 1968. Nuclear DNA variation in Allium Heredity 23: 591-605.
- Kalia, P. J.; Rai, M. K.; Kalia, S.; Singh, R.; Dhawan, A. D. 2011. Microsatellite markers: an overview of the recent progress in plants. Euphytica 177: 309-334.
- Khar, A. 2012. Cross amplification of onion derived microsatellites and mining of garlic EST database for assessment of genetic diversity in garlic. Acta Horticulturae 969: 289-295.
- Kim, D. W.; Jung, T. S.; Nam, S. H.; Kwon, H. R.; Kim, A.; Chae, S. H.; Choi, S. H.; Kim, D. W.; Kim, R. N.; Park, H. S. 2009. GarlicESTdb: an online database and mining tool for garlic EST sequences. BMC Plant Biology 9: 61.
- King, J. J.; Bradeen, J. M.; Bark, O.; McCallum, J. A.; Havey, M. J. 1998. A low-density genetic map of onion reveals a role for tandem duplication in the evolution of an extremely large diploid genome. Theoretical and Applied Genetics 96: 52-62.
- Kirk, J. T.; Rees, O. H.; Evans, G. 1970. Base composition of nuclear DNA within the genus Allium Heredity 25: 507-512.
- Li, Y. C.; Korol, A. B.; Fahima, T.; Nevo, E. 2004. Microsatellites within genes: structure, function, and evolution. Molecular Biology and Evolution 21: 991-1007.
- Lu, J. J.; Kang, J. Y.; Feng, S. G.; Zhao, H. Y.; Liu, J. J.; Wang, H. Z. 2013. Transferability of SSR markers derived from Dendrobium nobile expressed sequence tags (ESTs) and their utilization in Dendrobium phylogeny analysis. Scientia Horticulturae 158: 8-15.
- Mohanty, P.; Sahoo, L.; Parida, K.; Das, P. 2013. Development of polymorphic EST-SSR markers in Macrobrachium rosenbergii by data mining. Conservation and Genetic Resources 5: 133-136.
- Morales, R. G. F.; Resende, J. T. V.; Resende, F. V.; Delatorre, C. A.; Figueiredo, A. S. T.; Da-Silva, P. R. 2013. Genetic divergence among Brazilian garlic cultivars based on morphological characters and AFLP markers. Genetic and Molecular Research 12: 270-281.
- Nei, M.; Li, W. H. 1979. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proceedings of the National Academy of Sciences of the United States of America 76: 5269-5273.
- Paredes, C. M.; Becerra, V. V.; González, A. M. I. 2008. Low genetic diversity among garlic (Allium sativum L.) accessions detected using random amplified polymorphic DNA (RAPD). Chilean Journal of Agricultural Research 68: 3-12.
- Ranjekar, P. K.; Pallotta, D.; Lafontaine, J. G. 1978. Analysis of plant genomes. V. Comparative study of molecular properties of DNAs of seven Allium species. Biochemical Genetics 16: 957-970.
- Schuelke, M. 2000. An economic method for the fluorescent labeling of PCR fragments. Nature Biotechnology 18: 233-234.
- Şelale, H.; Çelik, I.; Gültekin, V.; Allmer, J.; Doğanlar, S.; Frary, A. 2013. Development of EST-SSR markers for diversity and breeding studies in opium poppy. Plant Breeding 132: 344-351.
- Thiel, T.; Michalek, W.; Varshney, R. K.; Graner, A.; 2003. Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theoretical and Applied Genetics 106: 411-422.
- Varshney, R. K.; Graner, A.; Sorrells, M. E. 2005. Genic microsatellite markers in plants: features and applications. Trends in Biotechnology 23: 48-55.
- Wei, L. B.; Zhang, H. Y.; Zheng, Y. Z.; Guo, W. Z.; Zhang, T. Z. 2008. Developing EST-derived microsatellites in sesame (Sesamum indicum L.). Acta Agronomica Sinica 34: 2077-2084.
- Wu, H. L.; Chen, D.; Li, J. X.; Yu, B.; Qiao, X. Y.; Huang, H. L.; He, Y. M. 2013. De novo characterization of leaf transcriptome using 454 sequencing and development of EST-SSR markers in Tea (Camellia sinensis). Plant Molecular Biology Reporter 31: 524-538.
- Yang, T.; Bao, S. Y.; Ford, R.; Jia, T. J.; Guan, J. P.; He, Y. H.; Sun, X. L.; Jiang, J. Y.; Hao, J. J.; Zhang, X. Y.; Zong, X. X. 2012. High-throughput novel microsatellite marker of faba bean via next generation sequencing. BMC Genomics 13: 602.
- Zalapa, J. E.; Cuevas, H.; Zhu, H.; Steffan, S.; Senalik, D.; Zeldin, E.; Mccown, B.; Harbut, R.; Simon, P. 2012. Using next-generation sequencing approaches to isolate simple sequence repeat (SSR) loci in the plant sciences. American Journal of Botany 99: 193-208.
- Zewdie, Y.; Havey, M. J.; Prince, J. P.; Jenderek, M. M. 2005. The first genetic linkages among expressed regions of the garlic genome. Journal of the American Society for Horticultural Science 130: 569-574.
-
Edited by: Martin Boer
Publication Dates
-
Publication in this collection
Jan-Feb 2015
History
-
Received
11 Apr 2014 -
Accepted
24 June 2014