ABSTRACT:
Despite important biotic stresses to common bean, Anthracnose (ANT) and Angular Leaf Spot (ALS) can cause losses of up to 80 % and occur in more than 60 countries around the world. Genetic resistance is the most sustainable strategy to manage these diseases. Thus, we aimed to (1) identify new SNP markers associated with ALS and ANT resistance loci in elite common bean lines, and (2) provide a functional characterization of the DNA sequences containing the identified SNP markers. We evaluated 60 inbred lines, under field conditions, which represent the elite germplasm developed by the Embrapa common bean breeding program across 22 years, in terms of severity of the ALS and ANT. The lines were genotyped with 5,398 SNPs. Then, a Mixed Linear Model was run to determine the SNP-trait associations. We observed two-significant marker-trait associations reacting to ANT, both located on chromosome Pv-02. These markers explained 25 % of the phenotypic variation. For ALS, only one significant marker-trait association was observed, which is located in chromosome Pv-10 and explained 19 % of the phenotypic variation. These markers, along with others already used, will be useful to add or keep ANT and ALS resistance loci identified in this work in the new carioca and black seeded cultivars.
Keywords:
Colletotrichum lindemuthianum; Pseudocercospora griseola; disease resistance; elite lines; marker-assisted selection
Introduction
Anthracnose (ANT), caused by hemibiotrophic fungus Colletotrichum lindemuthianum (Sacc. and Magnus) Briosi and Cavara, is one important fungal pathogen affecting the sustainability of bean production worldwide. Mild temperatures and high humidity favor disease development that can lead to a complete crop failure (Trabanco et al., 2015Trabanco, N.; Campa, A.; Ferreira, J.J. 2015. Identification of a new chromosomal region involved in the genetic control of resistance to anthracnose in common bean. Plant Genome 8: 79.). The pathogen can attack all aerial parts of bean plants and damage can reach up to 100 % yield loss in extreme cases (Singh and Schwartz, 2010Singh, S.P.; Schwartz, H.F. 2010. Breeding common bean for resistance to diseases: a review. Crop Science 50: 2199-2223.).
Another important disease affecting common bean crop is the Angular Leaf Spot (ALS), caused by the hemibiotrophic fungus Pseudocercospora griseola (Sacc.) Crous and Braun. ALS can cause losses as high as 80 % and is found in more than 60 countries around the world (Oblessuc et al., 2015Oblessuc, P.R.; Matiolli, C.C.; Chiorato, A.F.; Camargo, L.E.A.; Benchimol-Reis, L.L.; Melotto, M. 2015. Common bean reaction to angular leaf spot comprises transcriptional modulation of genes in the ALS10.1 QTL. Frontiers in Plant Science 6: 152.). Infection and sporulation occur within a broad temperature range, from 10 to 33 °C (Allorent and Savary, 2005Allorent, D.; Savary, S. 2005. Epidemiological characteristics of angular leaf spot of bean: a systems analysis. European Journal of Plant Pathology 113: 329-341.).
Genetic resistance is the most cost-effective, easy-to-use and ecologically sustainable strategy to manage ALS and ANT diseases (Burt et al., 2015Burt, A.J.; William, H.M.; Perry, G.; Khanal, R.; Pauls, K.P.; Kelly, J.D.; Navabi, A. 2015. Candidate gene identification with SNP marker-based fine mapping of anthracnose resistance gene Co-4 in common bean. Plos One 10: 10.; Keller et al., 2015Keller, B.; Manzanares, C.; Jara, C.; Lobaton, J.D.; Studer, B.; Raatz, B. 2015. Fine-mapping of a major QTL controlling angular leaf spot resistance in common bean (Phaseolus vulgaris L.). Theoretical and Applied Genetics 128: 813-826.). The high virulence diversity of their causal agents and recurring emergence of new strains of these pathogens challenge the development of cultivars with effective resistance (Souza et al., 2016Souza, T.L.P.O.; Gonçalves-Vidigal, M.C.; Raatz, B.M.C.; Mukankusi, Â.F.B.; Melo, L.C.; Pastor-Corrales, M.A. 2016. Major loci controlling resistance to the angular leaf spot of common bean. Annual Report Bean Improvement Cooperative 59: 15-18.; Abadio et al., 2012Abadio, A.K.R.; Lima, S.S.; Santana, M.F.; Salomao, T.M.F.; Sartorato, A.; Mizubuti, E.S.G.; Araujo, E.F.; Queiroz, M.V. 2012. Genetic diversity analysis of isolates of the fungal bean pathogen Pseudocercospora griseola from central and southern Brazil. Genetics Molecular Research 11: 1272-1279.).
Therefore, molecular markers offer significant potential to breeding programs for their resistance to these diseases, as markers tightly linked to Quantitative Trait Loci (QTL) are available. These markers can be used to deploy multiple resistance alleles to cultivars, compromising their resistance due to the emergence of new races considerably less probable (Burt et al., 2015Burt, A.J.; William, H.M.; Perry, G.; Khanal, R.; Pauls, K.P.; Kelly, J.D.; Navabi, A. 2015. Candidate gene identification with SNP marker-based fine mapping of anthracnose resistance gene Co-4 in common bean. Plos One 10: 10.). In this sense, genome-wide association studies (GWAS) is one of the most efficient methods, in terms of time, cost, and precision, to identify candidate genes or genome regions associated with the genetic control of agriculturally important traits (Tang et al., 2016Tang, Y.; Liu, X.L.; Wang, J.B.; Li, M.; Wang, Q.S.; Tian, F.; Su, Z.B.; Pan, Y.C.; Liu, D.; Lipka, A.E.; Buckler, E.S.; Zhang, Z.W. 2016. GAPIT version 2: an enhanced integrated tool for genomic association and prediction. Plant Genome 9: 120.). Furthermore, one of the main objectives of molecular genetics is to identify and characterize genes that govern important traits (Oblessuc et al., 2015Oblessuc, P.R.; Matiolli, C.C.; Chiorato, A.F.; Camargo, L.E.A.; Benchimol-Reis, L.L.; Melotto, M. 2015. Common bean reaction to angular leaf spot comprises transcriptional modulation of genes in the ALS10.1 QTL. Frontiers in Plant Science 6: 152.), helping plant breeders to improve the selection process through the identification of new markers or pathways related to diseases resistance.
In this study, we aimed to identify SNP markers associated with ALS and ANT resistance loci using elite inbred lines that represent the Brazilian germplasm of common bean from the two main market classes (around 90 % of market share). In addition, we provided a functional characterization of the DNA sequences containing the SNP markers that are associated with common bean resistance to ALS and ANT.
Materials and Methods
Field experiments
For this study, 33 breeding lines of common bean (elite inbred lines) from the “black market” class and 27 from the “carioca market” class were evaluated under field conditions (Faria et al., 2013Faria, L.C.; Melo, P.G.S.; Pereira, H.S.; Peloso, M.J.; Bras, A.; Moreira, J.A.A.; Carvalho, H.W.L.; Melo, L.C. 2013. Genetic progress during 22 years of improvement of carioca-type common bean in Brazil. Field Crops Research 142: 68-74.; Faria et al., 2014Faria, L.C.; Melo, P.G.S.; Pereira, H.S.; Wendland, A.; Borges, S.F.; Pereira, I.A.; Diaz, J.L.C.; Calgaro, M.; Melo, L.C. 2014. Genetic progress during 22 years of black bean improvement. Euphytica 199: 261-272.). These lines formed a panel that represents the elite germplasm developed by Embrapa (The Brazilian Agricultural Research Corporation) breeding program of common bean across 22 years (1985 to 2006).
Seeds of these breeding lines were multiplied during the winter crop season of 2008 (planted in May) in Santo Antônio de Goiás (16°30’17” S; 49°17’01” W; 766 m), Goiás State, Brazil, to standardize germination vigor. Field experiments were performed in three common bean-producing areas of Brazil (South, Central-West, and Northeast) on samples of plants sown during the rainy and fall-winter crop seasons in nine environments to evaluate ALS and in four environments to evaluate ANT. Table 1 lists the locations and research sites where the evaluations were performed as well as the crop seasons/years and the corresponding sowing dates.
Environments in the main Brazilian common bean-producing areas where field trials were conducted (in 2008-2010) for severity evaluation of the angular leaf spot (ALS) resistance and anthracnose (ANT) resistance in common bean.
The experiments were performed in a randomized complete block design, with two replicates, in plots with two four-meter rows, with 0.5 m spacing between rows, and 15 seeds sown per meter in the rows. Fertilizers were applied according to the results of soil analyses to ensure ideal conditions for development and production. Insect pests, invasive plants and weeds, as well as irrigation were controlled as needed according to official recommendations for the common bean crop. Disease control was not carried out in these experiments.
Severity of ALS and ANT corresponded to the percentage of infection in leaves and organs from infected plants. The descriptive scale used corresponded to 1 (absence of symptoms), 3 (up to 5 % infection), 5 (up to 20 % infection), 7 (up to 60 % infection) and 9 (up to 100 % infection, i.e., most of the plant was leafless or dead). In all cases, two evaluators scored the disease severity.
Deviance analysis and genotypic values prediction
Using the ALS and ANT severity scores, we carried out the Deviance analysis (ANADEV) according to Gilmour et al. (2009Gilmour, A.R.; Gogel, B.J.; Cullis, B.R.; Thompson, R. 2009. ASReml User Guide Release 3.0. VSN International, Hemel Hempstead, UK.). Additionally, the variance components and genotypic values of the lines were estimated in each index by a Restricted Maximum Likelihood/Best Linear Unbiased Predictor (REML/BLUP) as follows:
where: y is the vector of the traits (ALS or ANT); r is the replicate within the environment effect vector, considered fixed; l is the environment effect vector, considered fixed; g is the line effect vector and was considered random where g ~ N (0, I); i is the Genotype by Environment (GE) effect vector that was considered as random, where i ~ N (0, I).; e is the experimental error, where e ~ N (0,). X, T, Z and W are incidence matrices that relate the independent vector effects from each matrix with the dependent y vector.
The broad-sense heritability was estimated based on the mean of environments:
where: is the genotypic variance, is the genetic by environment variance, is the residual variance, r is the number of replicates per environment and l the number of locations.
Genotyping and data quality control
Samples of leaf tissue from each line were used for DNA extraction and genotyping. DNA concentrations were measured using a Nanodrop spectrum- photometer and its quality was checked on an agarose gel. Then, the lines were genotyped using an Illumina BARCBean6K_3 BeadChip with 5,398 SNPs (Song et al., 2015Song, Q.J.; Jia, G.F.; Hyten, D.L.; Jenkins, J.; Hwang, E.Y.; Schroeder, S.G.; Osorno, J.M.; Schmutz, J.; Jackson, S.A.; McClean, P.E.; Cregan, P.B. 2015. SNP assay development for linkage map construction, anchoring whole-genome sequence, and other genetic and genomic applications in common bean. G3 5: 2285-2290.).
The SNP genotype matrix was converted to numeric digits, with 0 for reference allele homozygotes, 1 for heterozygotes and 2 for alternative allele homozygotes. The quality control of the genomic data was performed with the snpReady package [https://github.com/italo-granato/SnpReady/tree/master/R] and required a minimum Call Rate of 85 % and Minor Allele Frequency (MAF) of 3 %. After filtering for low quality and markers without known positions on the genome, 1,490 SNPs were retained for further analysis.
The population structure was determined using the principal component analysis (PCA). The kinship matrix was developed using (VanRaden, 2008VanRaden, P.M. 2008. Efficient methods to compute genomic predictions. Journal of Dairy Science 91: 4414-4423.) method. Both were obtained by GAPIT 2 (Tang et al., 2016Tang, Y.; Liu, X.L.; Wang, J.B.; Li, M.; Wang, Q.S.; Tian, F.; Su, Z.B.; Pan, Y.C.; Liu, D.; Lipka, A.E.; Buckler, E.S.; Zhang, Z.W. 2016. GAPIT version 2: an enhanced integrated tool for genomic association and prediction. Plant Genome 9: 120.) and, when necessary, included into the association analysis to correct for population structure and cryptic relatedness.
GWAS
A Mixed Linear Model (MLM) was run based on Endelman (2011Endelman, J.B. 2011. Ridge regression and other kernels for genomic selection with R package rrBLUP. Plant Genome 4: 250-255.) to determine the SNP-trait associations. The MLM equation used in the analysis was as follows:
where: y is the BLUP of the genotypes for ANT or ALS; a is the vector of fixed effects of the SNP; b is the vector of fixed effect of the population structure (first principal components used, depending on the trait); n is the random effect of the relative kinship, where ; ε is the error term, where . S, P and K are incidence matrices that relate the independent vector effects from each matrix with the dependent y vector.
To determine the p-value threshold, we used a resampling method. Therefore, first, the phenotypic values are shuffled, breaking their association with markers and then the random association between all markers to the phenotype is estimated and the corresponding best marker score (minimum p-value obtained among all markers) is recorded. This procedure was repeated 200 times for each trait and the 95 % quantile from the 200 best scores was defined as the threshold to declare a significant association (Churchill and Doerge, 1994Churchill, G.A.; Doerge, R.W. 1994. Empirical threshold values for quantitative trait mapping. Genetics 138: 963-971.).
Functional annotation
Using BLASTN with E-value ≤ 1.0E−10, the flanking sequences (60 bp upstream and downstream) of the SNPs associated with ALS and ANT resistance were aligned against the genome of P. vulgaris from the Andean origin (Schmutz et al., 2014Schmutz, J.; McClean, P.E.; Mamidi, S.; Wu, G.A.; Cannon, S.B.; Grimwood, J.; Jenkins, J.; Shu, S.Q.; Song, Q.J.; Chavarro, C. 2014. A reference genome for common bean and genome-wide analysis of dual domestications. Nature Genetics 46: 707-713.) [http://www.phytozome.net/]. Functional annotations of the genes around the associated SNPs were analyzed.
Results
ANADEV and population parameters
Considering the joint analysis of deviance for ALS and ANT, the sources of variation Environment and Line were significant for both diseases. The Line × Environment interaction was significant only for ANT (Table 2). Regarding the genetic parameters, heritability was 0.64 and 0.93 for ANT and ALS, respectively (Table 2), which are considered high, indicating high reliability in field phenotypes. The correlation between the genotypic values of the lines for their reaction against these two diseases was low (0.26) and not significant.
Heritability, means, and genotypic correlation, the Wald test of fixed effects, and likelihood radio test (LRT) of random effects angular leaf spot (ALS) resistance and anthracnose (ANT) resistance in common bean.
Population structure and genomic data
After quality control of the marker data, 1,490 informative SNPs were selected for further analyses. The SNPs were split into 135, 143, 93, 118, 182, 93, 67, 189, 111, 164, and 195 in the first to the eleventh chromosome, respectively. The average linkage disequilibrium (LD, r2) along the genome was 0.04, with a standard deviation of 0.09. Furthermore, the estimated LD decay over the whole genome was 0.23 on 9K bp.
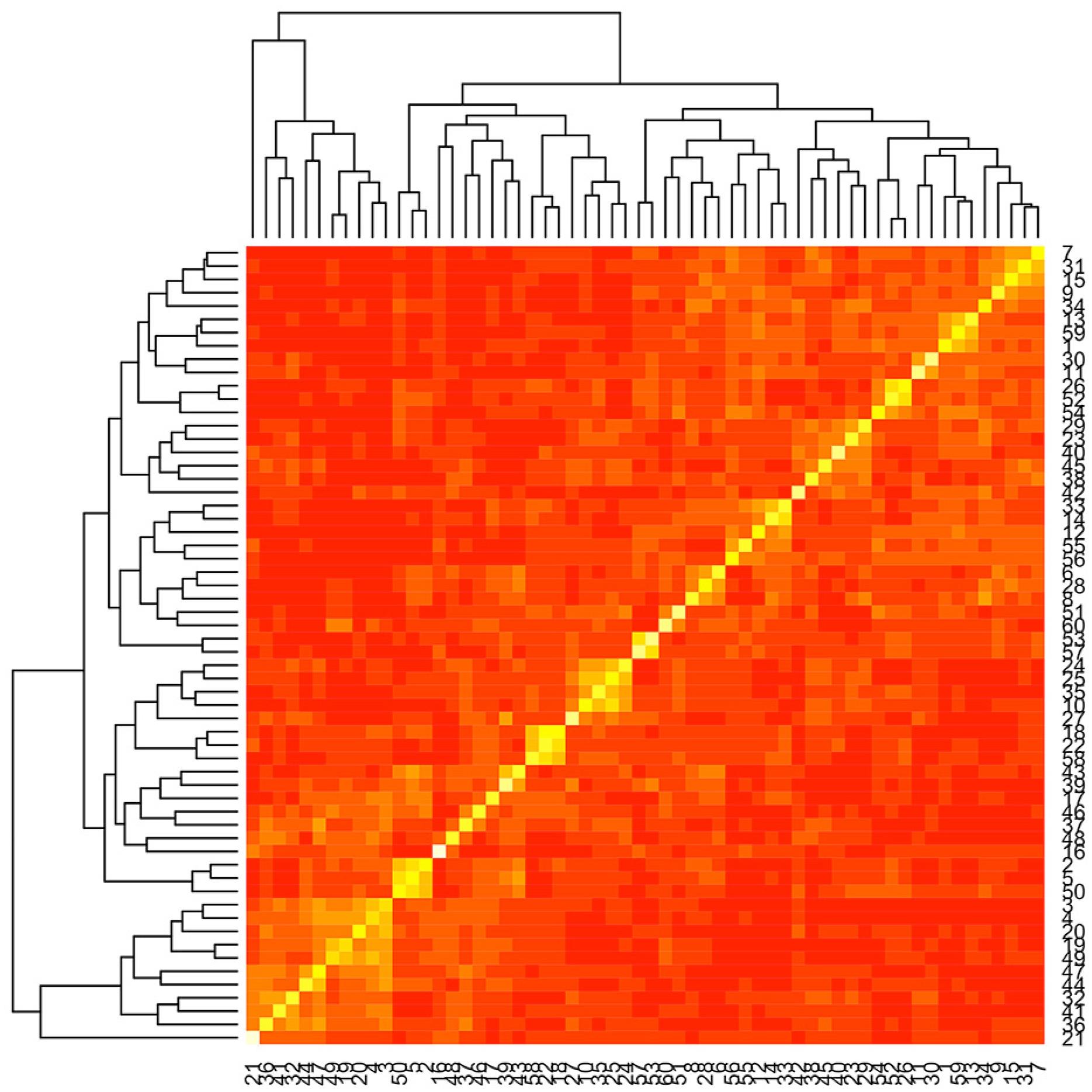
The population structure was represented graphically in two ways. In the first method, the heatmap from the genomic kinship matrix (K) (Figure 1) was used to identify the degree of individual relationship and possible subgroups in the panel. Based on that, it was verified that the lines are not closely related with no strong population structure. The second method was based on the two first principal components of matrix K (Figure 2). The first component (PC1) explained about 12 % of the total variance, while the second (PC2) accounted for 7 %. Moreover, there was no clear separation of lineages either by grain type or by any other source of variation (Figure 2).
Associations of SNPs with phenotypic characteristics
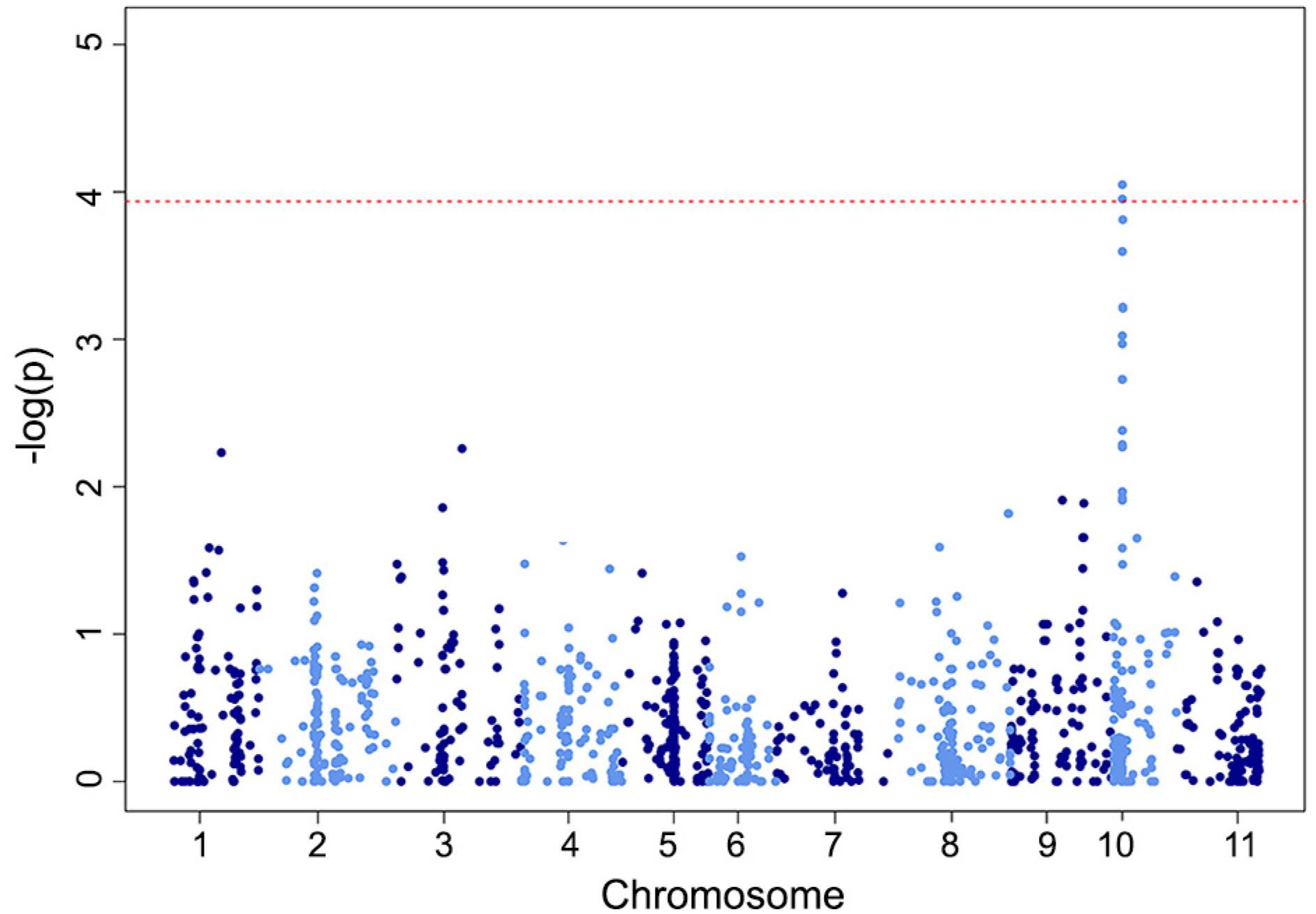
Two-significant marker-trait associations were observed reacting to ANT (p-value < 1.6 × 10−4), both located in chromosome Pv-02 (Figure 3). These markers explained about 25 % of the phenotypic variation regarding the reaction of lines to this disease. On the other hand, for ALS, only one significant marker-trait association was observed (p-value < 1.2 × 10−4), in chromosome Pv-10 (Figure 4) and explained about 19 % of the phenotypic variation of the trait.
Manhattan plot showing candidate SNP markers and their -log10 (p-values) from GWAS for anthracnose (ANT) resistance. The dotted red line is the significance threshold.
Manhattan plot showing candidate SNP markers and their -log10 (p-values) from GWAS for angular leaf spot (ALS) resistance. The dotted red line is the significance threshold.
For both SNPs associated with reaction to ANT, genotypes with two copies of the alternative allele (genotype of class 2) tended to be more resistant (negative values) and with less phenotypic variation compared to genotype class 0 (Figure 5). Heterozygotes tend to be more susceptible than homozygous individuals are. However, for the ALS variable, no homozygous individuals were observed for the unfavorable allele (Figure 5). The dominance deviation effects were estimated for these three significant SNPs. Even though dominance estimates were pronounced, their p-values (3.8 × 10−3, 2.5 × 10−4 and p < 2.2 × 10−2, respectively) were higher than the threshold established for the significance test.
Boxplots relating the genotypic values of the common bean lines to the SNP significantly associated with anthracnose (ANT) resistance and angular leaf spot (ALS) resistance.
Annotation of the associated loci
A BlastN of the SNP probes against the Andean genome (Schmutz et al., 2014Schmutz, J.; McClean, P.E.; Mamidi, S.; Wu, G.A.; Cannon, S.B.; Grimwood, J.; Jenkins, J.; Shu, S.Q.; Song, Q.J.; Chavarro, C. 2014. A reference genome for common bean and genome-wide analysis of dual domestications. Nature Genetics 46: 707-713.) positioned the two SNPs associated with ANT within 100 Kb apart at chromosome Pv-02. The first SNP associated with ANT reaction (BARCPV_1.0_Chr02_23542475_A_G) was mapped within the last intron of gene Phvul.002G115900 that encode a protein similar with Interleukin-1 Receptor-Associated Kinase 4 (IRAK4). This protein is an essential component of the signal transduction cascade that occurs after stimulation through Interleukin-1 Receptor, often associated with disease resistance genes (Table 3).
Chromosome (Chr), position, proportion explained by the model (R2) with and without the SNP, minor allele frequency (MAF), and annotation for the SNP significantly associated with angular leaf spot (ALS) resistance and anthracnose (ANT) resistance in common bean.
The second SNP associated with ANT reaction (BARCPV_1.0_Chr02_23644618_G_A) is in an intergenic region, ~5 Kb downstream of gene Phvul.002G116500 (with no annotation) and ~15 Kb upstream of gene Phvul.002G116400. This last gene is annotated as a RAB escort protein, which is a small GTPases that serves a regulatory role in vesicular membrane traffic (Table 3). Furthermore, the region within these two SNPs associated with ANT seems to contain a cluster of six genes annotated as involved in disease resistance or as protein kinases.
For ALS, the only SNP related to its reaction (BARCPV_1.0_Chr10_20935383_C_T) was mapped in an intergenic region on chromosome 10. Its location is ~5 Kb downstream of gene Phvul.010G072700. The protein encoded by this gene is annotated as a “Scarecrow-like protein,” which contains a GRAS domain involved gibberellin signaling, and is necessary for plant development. This gene is the only one within 70 Kb window from the SNP cited above (Table 3).
These SNPs may be located in regions with extensive linkage disequilibrium. Therefore, based on the proportion explained, possibly causal polymorphisms were not detected.
Discussion
Phenotypes and marker effects
A significant Line factor in the ANADEV analysis indicates the existence of genetic variability in the panel. This result was expected given the composition of the population in this study that aims to represent the black and carioca bean germplasms over the last 22 years of Embrapa breeding program (Faria et al., 2014Faria, L.C.; Melo, P.G.S.; Pereira, H.S.; Wendland, A.; Borges, S.F.; Pereira, I.A.; Diaz, J.L.C.; Calgaro, M.; Melo, L.C. 2014. Genetic progress during 22 years of black bean improvement. Euphytica 199: 261-272.; Faria et al., 2013Faria, L.C.; Melo, P.G.S.; Pereira, H.S.; Peloso, M.J.; Bras, A.; Moreira, J.A.A.; Carvalho, H.W.L.; Melo, L.C. 2013. Genetic progress during 22 years of improvement of carioca-type common bean in Brazil. Field Crops Research 142: 68-74.). Regarding the environmental factor, the results indicate that the sites and years used for the evaluation had distinct edaphoclimatic characteristics, which modified the reaction to the diseases of the common bean lines studied. However, only for ANT reaction, these environmental differences caused differential performance of the lines across the environments (Table 2). As the aim of the program was to develop lines with stable resistance across environments, the GE interaction was disregarded (minimized) in the subsequent analyses.
The 1,490 markers identified after quality control are well distributed throughout all 11 common bean chromosomes. Moreover, the average and standard deviation LD along the genome indicates that, although saturation was not so high, the markers represented well the possible haplotypes and segregate practically independently in the panel (Korte and Farlow, 2013Korte, A.; Farlow, A. 2013. The advantages and limitations of trait analysis with GWAS: a review. Plant Methods 9: 29.).
The significant marker-trait associations explain a satisfactory proportion of the reaction to the diseases, especially considering the low density of markers and small genotyping panel (Korte and Farlow, 2013Korte, A.; Farlow, A. 2013. The advantages and limitations of trait analysis with GWAS: a review. Plant Methods 9: 29.). The small population size may reduce power in detecting significant signals, especially for small effect QTLs. In addition, field resistance to ANT and ALS evaluated by natural incidence of the causal pathogens allowed the identification of QTLs stable across the different environments, that is, those targeted by the breeding programs. However, QQ-plots (Figures 6 and 7) show little inflation of p-values in lower tails, supporting that the modeling of population structure was appropriate (Yu et al., 2006Yu, J.M.; Pressoir, G.; Briggs, W.H.; Bi, I.V.; Yamasaki, M.; Doebley, J.F.; McMullen, M.D.; Gaut, B.S.; Nielsen, D.M.; Holland, J.B.; Kresovich, S.; Buckler, E.S. 2006. A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nature Genetics 38: 203-208.). Moreover, deviation from the uniform distribution (null hypothesis) in higher tails shows the presence of a significant signal in the data, which was clearly located in the Manhattan plots (Figures 3 and 4), with signal peaks above the significance threshold. The lowest SNP p-values for both ANT and ALS have neighboring SNPs with decreasingly low p-values, forming peaks, indicating the presence of some SNPs in LD around the peak. The presence of these peaks also means a good genomic coverage around the association loci. A higher population size might have allowed to detect a larger number of significantly associated SNPs, but those markers would explain a smaller fraction of the genotypic variation for these traits, concerning the breeding population under study.
QQ-plot showing the relationship between the expected and obtained –log10 (p-values) from GWAS for anthracnose (ANT) resistance.
QQ-plot showing the relationship between the expected and obtained –log10 (p-values) from GWAS for angular leaf spot (ALS) resistance.
In a GWAS study in common beans for the same diseases, Perseguini et al. (2016Perseguini, J.M.K.C.; Oblessuc, P.R.; Rosa, J.R.B.F.; Gomes, K.A.; Chiorato, A.F.; Carbonel, S.A.M.; Garcia, A.A.F.; Vianello, R.P.; Benchimol-Reis, L.L. 2016. Genome-wide association studies of anthracnose and angular leaf spot resistance in common bean (Phaseolus vulgaris L.). PLoS One 11: e0150506.) evaluated a diversity panel with more individuals, but genotyped with fewer SNP markers (369) than we used in this work. Nevertheless, the authors found 21 and 17 markers associated with ANT and ALS, respectively. Among them, the most important ones were located in chromosomes 7 and 4, explaining about 11 % and 17 % of the variance for ANT and ALS, respectively. On the other hand, as described above, we found different regions associated with the same characteristics, explaining 25 % of the variation. Thus, our findings reveal new relevant genomic regions in common beans, which could be used together with those found by Perseguini et al. (2016Perseguini, J.M.K.C.; Oblessuc, P.R.; Rosa, J.R.B.F.; Gomes, K.A.; Chiorato, A.F.; Carbonel, S.A.M.; Garcia, A.A.F.; Vianello, R.P.; Benchimol-Reis, L.L. 2016. Genome-wide association studies of anthracnose and angular leaf spot resistance in common bean (Phaseolus vulgaris L.). PLoS One 11: e0150506.) to perform marker-assisted selection in breeding programs.
The performance of the lines for the reaction to ALS and ANT was observed to be independent, as indicated by the non-significant correlation between the line genotypic values (Table 2). Furthermore, the SNPs associated to the main QTL for the diseases are located in different chromosomes, which may partially explain the low correlation between the traits. This is important, because it means that we can select for resistant genotypes for both diseases at the same time.
The heterozygote lines for the SNPs associated with ANT reaction tended to be more susceptible than the homozygous individuals did. This might indicate a disruptive selection due to a negative dominance deviation (Rueffler et al., 2006Rueffler, C.; Van Dooren, T.J.M.; Leimar, O.; Abrams, P.A. 2006. Disruptive selection and then what? Trends in Ecology Evolution 21: 238-245.). On other the hand, for reaction to ALS, no homozygous individuals were seen for the susceptible allele. It may be attributed to negative selection, making susceptible individuals more likely to be discarded by breeders. However, since there were few lines, this trend could not be confirmed based on this dataset. Thus, a hypothesis is raised for a future work. Nevertheless, in a common bean breeding program, which works mainly through the Single Seed Descendent method (SSD) (Brim, 1966Brim, C.A. 1966. A modified pedigree method of selection in soybeans. Crop Science 6: 220.), negative dominance deviations tend not to affect the selective process, even in this scenario. Notwithstanding, it would be interesting to identify from F2 and advance only the progenies with two copies of the superior allele for the markers in question to maximize the gains with the selection, eliminating the lower individuals as early as possible.
Annotation and further use of this results
The two SNP markers identified as associated with ANT reaction (BARCPV_1.0_Chr02_23542475_A_G and BARCPV_1.0_Chr02_23644618_G_A) are mapped in a genomic region that seems to contain a cluster of six genes annotated as involved in disease resistance or as protein kinases. This result is coherent with the phenotype observed and indicates that these SNPs should be selected as important candidate markers for marker-assisted selection (MAS). Resistance loci to ANT have already been reported in chromosome Pv02, such as major genes Co-u (BAT 93) (Geffroy et al., 2008Geffroy, V.; Sevignac, M.; Billant, P.; Dron, M.; Langin, T. 2008. Resistance to Colletotrichum lindemuthianum in Phaseolus vulgaris: a case study for mapping two independent genes. Theoretical and Applied Genetics 116: 407-415.) and CoPv02 (Xana) (Campa et al., 2014), and QTL AN2.1 (Oblessuc et al., 2014Oblessuc, P.R.; Baroni, R.M.; Pereira, G.S.; Chiorato, A.F.; Carbonel, S.A.M.; Briñes, B.; Silva, L.C.; Garcia, A.A.F.; Camargo, L.E.A.; Kelly, J.D.; Benchimol, L.L. 2014. Quantitative analysis of race-specific resistance to Colletotrichum lindemuthianum in common bean. Molecular Breeding 34: 1313-1329.), all from the Mesoamerican gene pool. However, these loci were physically positioned apart from the SNP sequences identified as associated to ANT in this work, based on in silico analysis in the sequenced bean genotype G19833 (www.phytozome.net). To date, these loci have not been targeted for MAS by the Embrapa common bean breeding program.
The SNP associated with ALS reaction (BARCPV_1.0_Chr10_20935383_C_T) contains a GRAS domain involved in gibberellin signaling, which is important for plant development. A QTL for ALS resistance from the Andean gene pool was already reported on Pv10 (ALS10.1) (Oblessuc et al., 2012Oblessuc, P.R.; Baroni, R.M.; Garcia, A.A.F.; Chiorato, A.F.; Carbonel, S.A.M.; Camargo, L.E.A.; Benchimol, L.L. 2012. Mapping of angular leaf spot resistance QTL in common bean (Phaseolus vulgaris L.) under different environments. BMC Genetics 13: 50.), which was recently renamed as the locus Phg-5 (Souza et al., 2016Souza, T.L.P.O.; Gonçalves-Vidigal, M.C.; Raatz, B.M.C.; Mukankusi, Â.F.B.; Melo, L.C.; Pastor-Corrales, M.A. 2016. Major loci controlling resistance to the angular leaf spot of common bean. Annual Report Bean Improvement Cooperative 59: 15-18.). However, all genotypes used in this work are from the Mesoamerican gene pool. For this reason, the SNP marker identified here may be linked to an ALS resistance gene different from Phg-5. In addition, the in silico analysis (www.phytozome.net) also showed that the SNP sequence identified as associated with ALS resistance in this work is physically positioned apart from Phg-5.
After validation with additional experiments, probably using specific biparental segregating populations (F2:3 or RILs), the SNP markers identified in this work will be converted into TaqMan markers and incorporated into the MAS routine of breeding programs of common bean at Embrapa. These markers, along with others already used, will be useful for all the programs around the world to add or keep ANT and ALS resistance loci identified here in the development of new carioca and black seeded cultivars.
References
- Abadio, A.K.R.; Lima, S.S.; Santana, M.F.; Salomao, T.M.F.; Sartorato, A.; Mizubuti, E.S.G.; Araujo, E.F.; Queiroz, M.V. 2012. Genetic diversity analysis of isolates of the fungal bean pathogen Pseudocercospora griseola from central and southern Brazil. Genetics Molecular Research 11: 1272-1279.
- Allorent, D.; Savary, S. 2005. Epidemiological characteristics of angular leaf spot of bean: a systems analysis. European Journal of Plant Pathology 113: 329-341.
- Brim, C.A. 1966. A modified pedigree method of selection in soybeans. Crop Science 6: 220.
- Burt, A.J.; William, H.M.; Perry, G.; Khanal, R.; Pauls, K.P.; Kelly, J.D.; Navabi, A. 2015. Candidate gene identification with SNP marker-based fine mapping of anthracnose resistance gene Co-4 in common bean. Plos One 10: 10.
- Churchill, G.A.; Doerge, R.W. 1994. Empirical threshold values for quantitative trait mapping. Genetics 138: 963-971.
- Endelman, J.B. 2011. Ridge regression and other kernels for genomic selection with R package rrBLUP. Plant Genome 4: 250-255.
- Faria, L.C.; Melo, P.G.S.; Pereira, H.S.; Peloso, M.J.; Bras, A.; Moreira, J.A.A.; Carvalho, H.W.L.; Melo, L.C. 2013. Genetic progress during 22 years of improvement of carioca-type common bean in Brazil. Field Crops Research 142: 68-74.
- Faria, L.C.; Melo, P.G.S.; Pereira, H.S.; Wendland, A.; Borges, S.F.; Pereira, I.A.; Diaz, J.L.C.; Calgaro, M.; Melo, L.C. 2014. Genetic progress during 22 years of black bean improvement. Euphytica 199: 261-272.
- Geffroy, V.; Sevignac, M.; Billant, P.; Dron, M.; Langin, T. 2008. Resistance to Colletotrichum lindemuthianum in Phaseolus vulgaris: a case study for mapping two independent genes. Theoretical and Applied Genetics 116: 407-415.
- Gilmour, A.R.; Gogel, B.J.; Cullis, B.R.; Thompson, R. 2009. ASReml User Guide Release 3.0. VSN International, Hemel Hempstead, UK.
- Keller, B.; Manzanares, C.; Jara, C.; Lobaton, J.D.; Studer, B.; Raatz, B. 2015. Fine-mapping of a major QTL controlling angular leaf spot resistance in common bean (Phaseolus vulgaris L.). Theoretical and Applied Genetics 128: 813-826.
- Korte, A.; Farlow, A. 2013. The advantages and limitations of trait analysis with GWAS: a review. Plant Methods 9: 29.
- Oblessuc, P.R.; Baroni, R.M.; Garcia, A.A.F.; Chiorato, A.F.; Carbonel, S.A.M.; Camargo, L.E.A.; Benchimol, L.L. 2012. Mapping of angular leaf spot resistance QTL in common bean (Phaseolus vulgaris L.) under different environments. BMC Genetics 13: 50.
- Oblessuc, P.R.; Baroni, R.M.; Pereira, G.S.; Chiorato, A.F.; Carbonel, S.A.M.; Briñes, B.; Silva, L.C.; Garcia, A.A.F.; Camargo, L.E.A.; Kelly, J.D.; Benchimol, L.L. 2014. Quantitative analysis of race-specific resistance to Colletotrichum lindemuthianum in common bean. Molecular Breeding 34: 1313-1329.
- Oblessuc, P.R.; Matiolli, C.C.; Chiorato, A.F.; Camargo, L.E.A.; Benchimol-Reis, L.L.; Melotto, M. 2015. Common bean reaction to angular leaf spot comprises transcriptional modulation of genes in the ALS10.1 QTL. Frontiers in Plant Science 6: 152.
- Perseguini, J.M.K.C.; Oblessuc, P.R.; Rosa, J.R.B.F.; Gomes, K.A.; Chiorato, A.F.; Carbonel, S.A.M.; Garcia, A.A.F.; Vianello, R.P.; Benchimol-Reis, L.L. 2016. Genome-wide association studies of anthracnose and angular leaf spot resistance in common bean (Phaseolus vulgaris L.). PLoS One 11: e0150506.
- Rueffler, C.; Van Dooren, T.J.M.; Leimar, O.; Abrams, P.A. 2006. Disruptive selection and then what? Trends in Ecology Evolution 21: 238-245.
- Schmutz, J.; McClean, P.E.; Mamidi, S.; Wu, G.A.; Cannon, S.B.; Grimwood, J.; Jenkins, J.; Shu, S.Q.; Song, Q.J.; Chavarro, C. 2014. A reference genome for common bean and genome-wide analysis of dual domestications. Nature Genetics 46: 707-713.
- Singh, S.P.; Schwartz, H.F. 2010. Breeding common bean for resistance to diseases: a review. Crop Science 50: 2199-2223.
- Song, Q.J.; Jia, G.F.; Hyten, D.L.; Jenkins, J.; Hwang, E.Y.; Schroeder, S.G.; Osorno, J.M.; Schmutz, J.; Jackson, S.A.; McClean, P.E.; Cregan, P.B. 2015. SNP assay development for linkage map construction, anchoring whole-genome sequence, and other genetic and genomic applications in common bean. G3 5: 2285-2290.
- Souza, T.L.P.O.; Gonçalves-Vidigal, M.C.; Raatz, B.M.C.; Mukankusi, Â.F.B.; Melo, L.C.; Pastor-Corrales, M.A. 2016. Major loci controlling resistance to the angular leaf spot of common bean. Annual Report Bean Improvement Cooperative 59: 15-18.
- Tang, Y.; Liu, X.L.; Wang, J.B.; Li, M.; Wang, Q.S.; Tian, F.; Su, Z.B.; Pan, Y.C.; Liu, D.; Lipka, A.E.; Buckler, E.S.; Zhang, Z.W. 2016. GAPIT version 2: an enhanced integrated tool for genomic association and prediction. Plant Genome 9: 120.
- Trabanco, N.; Campa, A.; Ferreira, J.J. 2015. Identification of a new chromosomal region involved in the genetic control of resistance to anthracnose in common bean. Plant Genome 8: 79.
- VanRaden, P.M. 2008. Efficient methods to compute genomic predictions. Journal of Dairy Science 91: 4414-4423.
- Yu, J.M.; Pressoir, G.; Briggs, W.H.; Bi, I.V.; Yamasaki, M.; Doebley, J.F.; McMullen, M.D.; Gaut, B.S.; Nielsen, D.M.; Holland, J.B.; Kresovich, S.; Buckler, E.S. 2006. A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nature Genetics 38: 203-208.
Edited by
Publication Dates
-
Publication in this collection
Jul-Aug 2019
History
-
Received
28 Aug 2017 -
Accepted
23 Feb 2018