ABSTRACT:
Genetic redundancy in cassava (Manihot esculenta Crantz) presents a challenge to efficient management of genetic resources. This study aimed to identify and define the genetic structure of duplicates in cassava germplasm from various Embrapa research units, using single-nucleotide polymorphism (SNP) markers. We evaluated 2,371 accessions with 20,712 SNPs. The identification of duplicates was performed based on multilocus genotypes (MLG), adopting a maximum genetic distance threshold of 0.05. The population structure was defined based on discriminant analysis of principal components (DAPC). A total of 1,757 unique and 614 duplicate accessions were identified. The redundancy of the collections ranged from 17 % (Belém, PA – Brazil) to 39 % (Petrolina, PE – Brazil), with an average of 21 %. This redundancy between different research units is probably due to the historical sharing of accessions, as well as collections carried out in the same region, or even to the intense germplasm exchange between farmers with different genotype names. In terms of genetic structure, the 250 principal components explained 88 % of the genetic variation of the SNP markers and defined the hierarchical structure of the duplicate cassava germplasm in 12 groups. Since heterotic groups have not yet been identified for cassava, crosses between accessions of the 12 DAPC groups may be promising. All MLGs were allocated within the same DAPC group, corroborating duplicate analyses yet still revealing high variability between groups that were quite distinct based on the first two discriminant functions. Our results contribute to optimizing the conservation of genetic resources, together with understanding diversity and its use in crop improvement.
Keywords:
Manihot esculenta Crantz; genotyping-by-sequencing; multilocus; genetic resources
Introduction
Cassava (Manihot esculenta Crantz) is native to South America, where Brazil is its probable center of origin and diversity (Olsen, 2004Olsen, K.M. 2004. SNPs SSRs and inferences on cassava's origin. Plant Molecular Biology 56: 517-526.). The Cassava Germplasm Bank (CGB) conserves a wide variety of accessions of this species, with great relevance to future use in several genetic studies and breeding programs. However, the maintenance costs of the CGB are relatively high because the plants are kept in the field and there is still a need for in vitro backup. According to Shands (2010Shands, H.; Hawtin, G.; MacNeil, G. 2010. The Cost to the CGIAR Centres of Maintaining and Distributing Germplasm. CGIAR, Montpellier, France. (CGIAR Consortium Study Paper, 10).), the annual cost for maintenance and distribution of cassava (capital and variable costs) of CIAT (International Center for Tropical Agriculture) is approximately US$ 92.30 per accession/year. Therefore, to reduce the number of copies, measures aimed at the rationalization of germplasm collections are of paramount importance to optimizing the physical storage space of the accessions in both laboratory and field, as well as to reducing the maintenance cost of the collections (Treuren and Hintum, 2003Treuren, R.; Hintum, T.J.L. 2003. Marker-assisted reduction of redundancy in germplasm collections: genetic and economic aspects. Acta Horticulturae 623: 139-149.).
With advances in the molecular biology of cassava, numerous opportunities have arisen to apply this information to increase efficiency in the use of genetic resources and conventional breeding. At the moment, it is necessary to use all this information to integrate the current knowledge on cassava genomics so as to consolidate the use of advanced techniques, genetics and molecular biology strategies in the daily activities of the genetic cassava breeding program. Although several types of molecular markers have been used in cassava, currently the SNP (single-nucleotide polymorphism) markers are those most commonly used in several molecular studies focused on genomic selection (Oliveira et al., 2012Oliveira, E.J.; Resende, M.D.V.; Santos, V.S.; Fereira, C.F.; Oliveira, G.A.F.; Silva, M.S.; Oliveira, L.A.; Aguilar-Vildoso, C.I. 2012. Genome-wide selection in cassava. Euphytica 187: 263-276.), linkage and mapping of quantitative trait loci (QTL) (Nzuki et al., 2017Nzuki, I.; Katari, M.S.; Bredeson, J.V.; Masumba, E.; Kapinga, F.; Salum, K.; Mkamilo, G.S.; Shah, T.; Lyons, J.B.; Rokhsar, D.S.; Rounsley, S.; Myburg, A.A.; Ferguson, M.E. 2017. QTL mapping for pest and disease resistance in cassava and coincidence of some QTL with introgression regions derived from Manihot glaziovii. Frontiers in Plant Science. doi: https://doi.org/10.3389/fpls.2017.01168
https://doi.org/10.3389/fpls.2017.01168...
), and identification of duplicate accessions (Rabbi et al., 2015Rabbi, I.Y.; Kulakow, P.A.; Manu-Aduening, J.A.; Dankyi, A.A.; Asibuo, J.Y.; Parkes, E.Y.; Abdoulaye, T.; Girma, G.; Gedil, M.A.; Ramu, P.; Reyes, B.; Maredia, M.K. 2015. Tracking crop varieties using genotyping-by-sequencing markers: a case study using cassava (Manihot esculenta Crantz). BMC Genetics. doi: https://doi.org/10.1186/s12863-015-0273-1
https://doi.org/10.1186/s12863-015-0273-...
). SNP markers have also been used as an auxiliary tool in the analysis of duplicates in germplasm banks as they have abundant polymorphisms and can be automated, resulting in a high analytical yield (Mammadov et al., 2012Mammadov, J.; Aggarwal, R.; Buyyarapu, R.; Kumpatla, S. 2012. SNP markers and their impact on plant breeding. International Journal of Plant Genomics 2012: 1-12.). Along with the phenotypic and passport data, the SNP markers can greatly contribute to a clear and efficient distinction of cassava germplasm.
The cost of identifying the duplicate cassava accession using molecular characterization is 12 times lower than the cost of conserving and using the material as a different accession in the germplasm collection (Horna et al., 2010Horna, D.; Debouck, D.; Dumet, D.; Hanson, J.; Payne, T.; Sackville-Hamilton, R.; Sanchez, I.; Upadhyaya, H.D.; Van Den Houwe, I. 2010. Evaluating Cost-Effectiveness of Collection Management: Ex-situ Conservation of Plant Genetic Resources in the CG System. CGIAR, Montpellier, France.). Therefore, this molecular information should be used as ancillary information to maximize the conservation efficiency of cassava germplasm so that curators can precisely identify accessions with redundant genetic backgrounds. Thus, the objective of this study was to identify the presence of duplicates of accessions in the cassava germplasm bank from Brazil, as well as to evaluate the genetic structure of the accession duplicates using SNP markers.
Materials and Methods
Plant material
A total of 2,371 accessions belonging to the Cassava Germplasm Bank (CGB) conserved at Embrapa were analyzed, of which 1,553 accessions belong to Cruz das Almas, BAa (CNPMF, 224 m altitude, 12°40’ S latitude and 39°06’ W longitude); 356 accessions to Belém, PA (CPATU, 12 m altitude, 1°28’ S latitude and 48°27’ W longitude); 327 accessions to Planaltina, Federal District (CPAC, 1,007 m altitude, 15°35’ S latitude and 47°42’ W longitude); and 135 accessions to Petrolina, PE (CPATSA, 370 m altitude, 09°09’ S latitude and 40°22’ W longitude), Brazil.
DNA extraction
DNA was extracted from young leaves according to the CTAB protocol (cetyltrimethylammonium bromide) as described by Doyle and Doyle (1987Doyle, J.J.; Doyle, J.L. 1987. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin 19: 11-15.), with minor modifications, such as the addition of polyvinylpyrrolidone (PVP) and an increase in the 2-mercaptoethanol concentration to 0.4 %. The DNA quality was assessed through 1 % (w/v) agarose gel quantification stained with ethidium bromide (1.0 mg L−1) in 0.5x TBE buffer (45mM Tris-borate, 1mM EDTA and distilled water qsp), visualized in UV light, and recorded by a high-performance system for capturing images. The quantity of DNA was estimated by comparing the fluorescent yield of the samples with a series of lambda (λ) DNA standards at varying known concentrations. The DNA was diluted in TE buffer (10 mM Tris-HCl and 1 mM EDTA) to a final concentration of 60 ng μL−1, and the quality was checked through the digestion of 250 ng of the genomic DNA from 10 random samples with the restriction enzyme EcoRI at 65 °C for two hours and thereafter visualized on agarose gel.
Genotyping by sequencing (GBS)
The basic protocol of GBS has been described by Elshire et al. (2011Elshire, R.J.; Glaubitz, J.C.; Sun, Q.; Poland, J.A.; Kawamoto, K.; Buckler, E.S.; Mitchell, S.E. 2011. A robust simple genotyping-by-sequencing (GBS) approach for high diversity species. Plos One 6: 1-10.), wherein the DNA is digested by the enzyme ApeKI, a type II restriction endonuclease that recognizes a 5-base degenerate sequence (GCWGC, where W is A or T), in lengths of 100 bp, as recommended by Hamblin and Rabbi (2014Hamblin, M.T.; Rabbi, I.Y. 2014. The effects of restriction-enzyme choice on properties of genotyping-by-sequencing libraries: a study in cassava (Manihot esculenta). Crop Science 54: 2603-2608.). The binding between ApeKI cleavage fragments and the adapter was performed after the digestion of samples and a 192-plex sequencing run. In order to analyze the sequences and quality filters, the software Tassel package, version 5.2.37 (Bradbury et al., 2007Bradbury, P.J.; Zhang, Z.; Kroon, D.E.; Casstevens, T.M.; Ramdoss, Y.; Buckler, E.S. 2007. TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics Applications Note 23: 2633-2635.) was used to remove loci with a minimum frequency (MAF) of less than 5 %, keep variants that had been successfully genotyped at least in 50 % of individuals, and SNPs with more than 20 % of missing data, resulting in 20,712 SNPs of high quality.
Identification of duplicate accessions
Hamming distance was calculated by the bitwise.dist function from the poppr package version 2.3.0 of the R program version 3.3.4 (R Development Core Team, 2017) and is defined as the number of allelic differences between two accessions, forcing the missing data to not match with any other allele, including other missing data. The identification of duplicates was carried out based on the detection of multilocus genotypes (MLGs) using the function mlg.filter, which utilizes the Hamming distance for collapsing multilocus genotypes that are under a specific distance threshold. The threshold of 0.05 was arbitrarily defined as the minimum distance for considering two genotypes unique, and any cassava accessions below that will be clustered into the same MLG. Moreover, underlying the mlg.filter function, the algorithm “nearest neighbor” was used for deciding what accessions go together to the same MLG.
This method was specifically developed for analysing clonal populations, and was implemented with the objective of visualizing relationships between unknown MLGs (Kamvar et al., 2014Kamvar, Z.N.; Tabima, J.F.; Grunwald, N.J. 2014. Poppr: an R package for genetic analysis of populations with clonal partially clonal and/or sexual reproduction. Peer J. doi: https://doi.org/10.7717/peerj.281
https://doi.org/10.7717/peerj.281...
). However, studies in clonal populations, based only on the identification and comparison of MLGs between individuals, may not be accurate, as the existence of somatic mutations and possible errors in the identification of genetic markers should be taken into account. With the generation of large amounts of data via next-generation sequencing (NGS), the genetic resolution has increased greatly, although the possibility of genotyping errors and high numbers of missing data are also frequent (Elshire et al., 2011Elshire, R.J.; Glaubitz, J.C.; Sun, Q.; Poland, J.A.; Kawamoto, K.; Buckler, E.S.; Mitchell, S.E. 2011. A robust simple genotyping-by-sequencing (GBS) approach for high diversity species. Plos One 6: 1-10.). However, the mlg.filter function considers these possible biases and allows users to choose the genetic distance and the approach for clustering common MLGs that are more biologically relevant to the population studied considering the ploidy level and the nature of the DNA markers used (Kamvar et al., 2015Kamvar, Z.N.; Brooks, J.C.; Grunwald, N.J. 2015. Novel R tools for analysis of genome-wide population genetic data with emphasis on clonality. Frontiers in Genetics 6: 1-10.).
Clusters of duplicate accessions
The discriminant analysis of principal components (DAPC) available in the adegenet package version 2.0.1 of the R program version 3.3.4 (R Development Core Team, 2017) was used to define the clusters of duplicate cassava accessions, as this technique does not require an a priori definition of the genetic clusters (Jombart et al., 2010Jombart, T.; Devillard, S.; Balloux, F. 2010. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics. doi: https://doi.org/10.1186/1471-2156-11-94
https://doi.org/10.1186/1471-2156-11-94...
). DAPC is based on the preliminary transformation of the data, using principal component analysis (PCA) as a step prior to discriminant analysis (DA) and their number is lower than the individuals analyzed, without necessarily implying loss of genetic information. This transformation allows the DA to be applied to any genetic data, while the analysis minimizes the differences between individuals within groups and maximizes them between groups, in which accessions are better discriminated in clusters (Jombart et al., 2010Jombart, T.; Devillard, S.; Balloux, F. 2010. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics. doi: https://doi.org/10.1186/1471-2156-11-94
https://doi.org/10.1186/1471-2156-11-94...
).
Successive clustering analysis with the K-means and the Bayesian information criterion (BIC) was used to define the ideal number of clusters, in which the lowest BIC value is assigned to represent the most probable number of clusters of the data set under analysis. In the presence of a hierarchical structure, BIC values can be reduced after identification of the true value of K. Therefore, the reduction in BIC values was empirically analyzed to identify the K value at which the BIC value is slightly reduced (Jombart et al., 2010Jombart, T.; Devillard, S.; Balloux, F. 2010. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics. doi: https://doi.org/10.1186/1471-2156-11-94
https://doi.org/10.1186/1471-2156-11-94...
). The number of clusters was assessed using the function find.clusters, covering a range of possible clusters from 1 to 80. After defining the number of clusters, we kept 250 PCs, the number required to explain more than 88 % of the genetic variance.
Results
Identification of duplicate accessions based on multilocus analysis
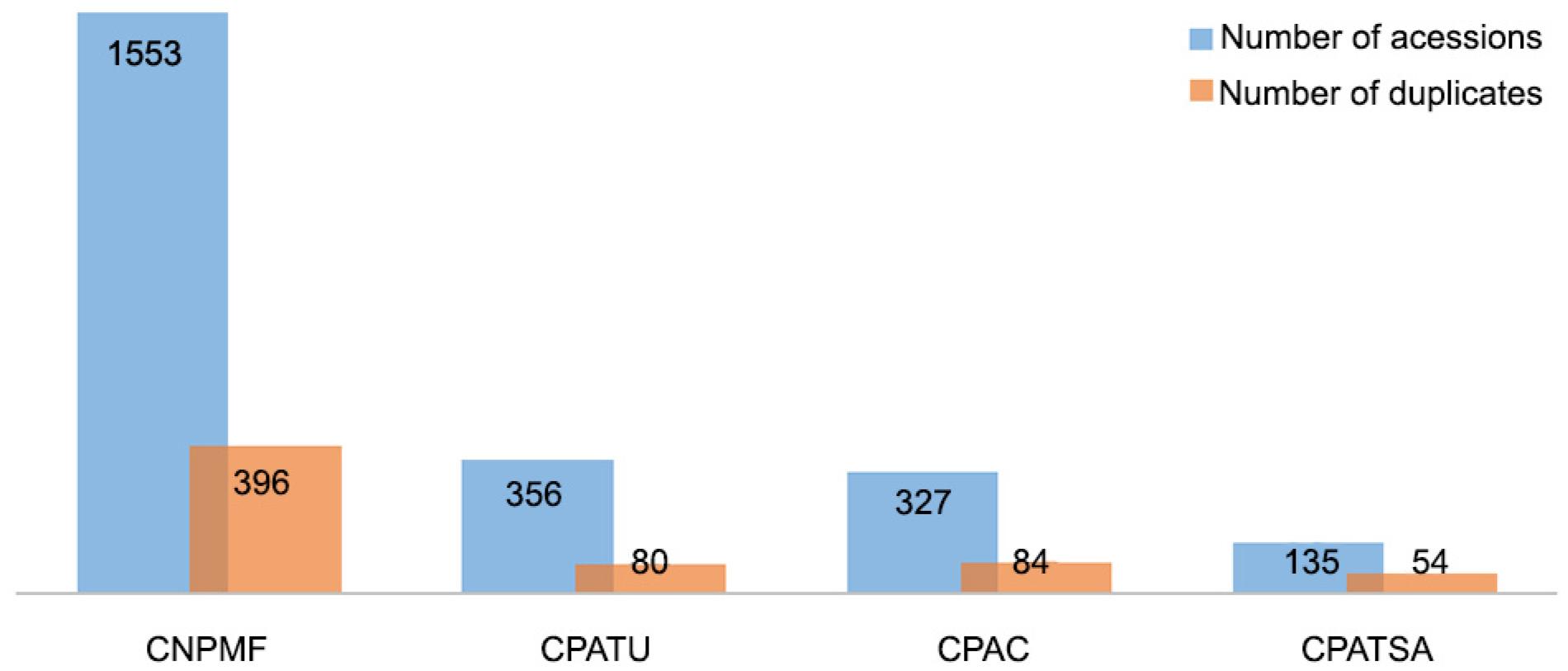
Of the 2,371 cassava accessions, 1,757 samples had unique genotypes (single MLGs) and were thus not duplicate clones, while the other 614 accessions presented a nonexclusive SNP profile, assumed to be duplicates as each MLG corresponded to a single genotype (Arnaud-Haond et al., 2007Arnaud-Haond, S.C.M.; Alberto, F.; Serrão, E.A. 2007. Standardizing methods to address clonality in population studies. Molecular Ecology 16: 5115-5139.). The numbers of duplicate accessions were 54, 80, 84, and 396 on the germplasm banks from CPATSA, CPATU, CPAC, and CNPMF, respectively (Figure 1).
Relationship between the number of accessions conserved in cassava germplasm banks from Brazil and the number of accessions with similar SNP profiles, being considered as duplicates.
It was possible to identify accessions with similar genetic profiles among and within Embrapa's cassava collections. Specifically, in the case of CPATSA, all 135 accessions were duplicated from the CNPMF germplasm because they presented the same MLGs (Table 1). Possibly this occurred due to the historical accession exchange between these institutions, as well as the fact that many germplasm collections in the semiarid region of Brazil have been carried out in the areas covered by CPATSA's mandate. The CNPMF also has 87 and 82 accessions maintained by CPAC and CPATU, respectively, while CPATU and CPAC presented 31 duplicate accessions maintained by both units. CPAC and CPATSA presented only ten duplicate accessions in common. However, between CPATU and CPATSA, there were no duplicates maintained by these research units (Table 1). Similar to CPATSA, the history of germplasm exchange in Brazil is well known, although in many situations the exchange of names, or even the lack of passport data standardization, can lead to doubts about the origin of the accessions collected. In this case, molecular analysis can solve doubts about the genetic identity of the accessions with a high degree of reliability.
Number of duplicate and unique cassava accessions based on the multilocus genotype (MLG) analysis within and among Embrapa's cassava collections, based on 20,712 SNP markers.
Based on the analysis of duplicates, 124 unique genotypes (MLGs) were identified within Embrapa's CGB (86, 19, 17, and two MLGs formed by accessions belonging to CNPMF, CPATU, CPAC, and CPATSA, respectively) (Table 1). In addition, there were accessions that shared MLGs between different CGBs, of which 33 were between CNPMF and CPATSA, 20 between CNPMF and CPAC, 19 between CNPMF and CPATU, seven between CPAC and CPATSA, and two between CPAC and CPATSA. Therefore, the greatest number of unique genotypes among and within the CGBs was observed in CNPMF, which has been a repository of cassava accessions for other germplasm banks. However, there is no germplasm exchange between CPATU and CPATSA. This is probably due to the greater climatic difference between these regions, considering that CPATU is located in the northern region of Brazil (Belém-PA), whose main climatic characteristics are 26.7 °C annual average temperature, 84 % relative humidity, and 3,000 mm of annual rainfall, while CPATSA is located in the northeastern region of Brazil (Petrolina, PE) with 26.1 °C annual average temperature, 60 % relative humidity, and 530 mm of annual rainfall (INMET, 2016Instituto Nacional de Meteorologia [INMET]. 2016. BDMAP: historical data = BDMAP: dados históricos. Available at: http://www.inmet.gov.br/portal/index.php?r=home2/index [Accessed Jan 21, 2017) (in Portuguese).
http://www.inmet.gov.br/portal/index.php...
). Thus, annual rainfall is one of the main differentiating factors between these regions, and, as a result, the cassava accessions from the north present lower adaptability to the Brazilian northeast and vice versa.
In total, the 614 cassava accessions (26 %) that expressed a nonexclusive SNP profile represented 124 different unique genotypes, so there were 1,881 different SNP profiles on the CGB, representing 21 % of duplicate accessions. Within Embrapa's units, the CPATSA had the highest percentage of duplicate accessions (39 %) with similar genetic profiles based on the SNP. In the other Embrapa units, the percentage of duplicates was 17 %, 20 % and 20 % for CPATU, CNPMF, and CPAC, respectively.
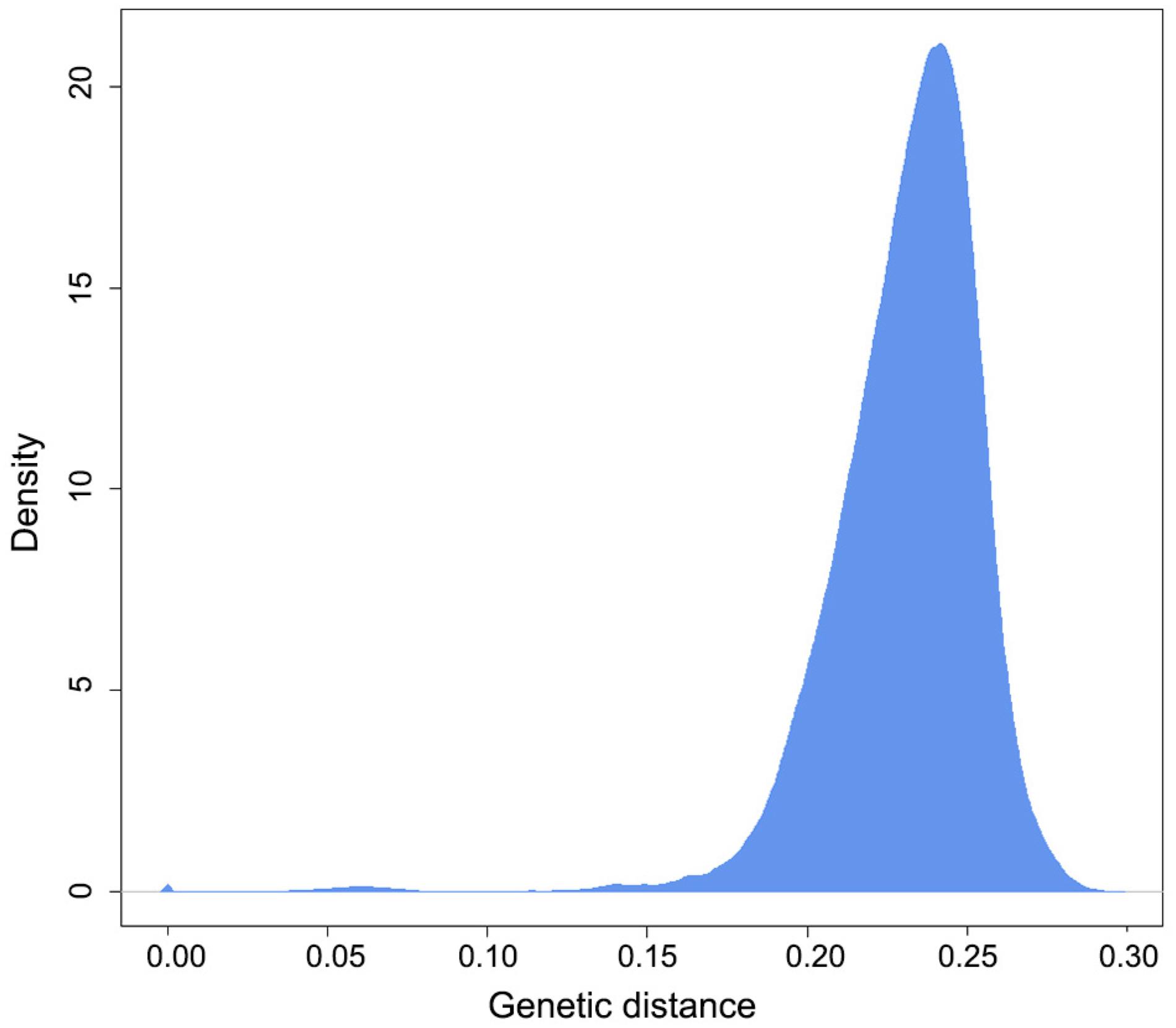
The genetic diversity of the 2,371 cassava accessions, based on the Hamming genetic distance, ranged from 0.014 to 0.297, with a mean of 0.231 (Figure 2). Most of the accessions diverged at a distance greater than 0.20, indicating high genetic diversity stored in Brazilian cassava germplasm. Considering the maximum genetic distance threshold (0.05) to define the different MLGs, it was observed that accessions BGM2004 and BGM1635 had the lowest genetic distance (0.014), while in the other duplicate accessions this distance ranged from 0.021 to 0.050, with an average of 0.041.
Genetic distance among 2,371 cassava accessions based on 20,712 single-nucleotide polymorphism (SNP) markers.
Population structure of duplicate accessions
DAPC was used to investigate the pattern of genetic diversity of accessions considered duplicates by multilocus analysis. The maintenance of 250 principal components (PCs) in the preliminary step of data transformation allowed the DAPC to explain more than (88 %) of the total genetic variation observed by the SNP markers. The definition of the number of PCs to be retained in the analysis is a point of discussion about the power of dimensionality reduction of the data. In general, the use of PCs that retain more than 80 % of the genetic variance is recommended (Jombart et al., 2010Jombart, T.; Devillard, S.; Balloux, F. 2010. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics. doi: https://doi.org/10.1186/1471-2156-11-94
https://doi.org/10.1186/1471-2156-11-94...
). In the DAPC, it is necessary to establish a balance between the power of cluster discrimination and the stability of genotype assignment in each cluster. Therefore, the analysis with 250 PCs guaranteed high statistical power for evaluating the genetic structure of the duplicate accessions conserved in the four main CGBs in Brazil.
Figure 3 indicates the presence of a hierarchical structure in duplicate cassava germplasm and illustrates the definition of the ideal number of clusters. Based on the K-means grouping, the first elbow curve of BIC values as a function of k occurred with K = 12, this being the number of clusters selected to represent the diversity of the duplicate cassava germplasm and also to analyze if the MLGs were clustered together.
Distribution of the Bayesian information criterion (BIC) according to the number of clusters, based on the analysis of 614 duplicate cassava accessions with 20,712 single-nucleotide polymorphisms (SNPs). The red arrow indicates the number of clusters chosen for grouping analysis.
The probability of assignment of individuals of each accession in a given cluster by DAPC was 100 % for all 614 duplicate accessions. Therefore, the clustering of duplicate cassava accessions based on the first and two discriminant functions revealed a clear separation of the 12 diversity groups (Figure 4). The number of accessions per group was 12 (Group 6) to 129 (Group 11), with an average of 51 accessions per cluster. In contrast, the number of MLGs ranged from 4 (Group 4) to 51 (Group 11), with an average of 16 MLGs per DAPC cluster (Table 2). Therefore, the cassava accessions from Group 11 still store a wide genetic diversity within the cluster.
Discriminant analysis of principal components (DAPC) of 614 cassava accessions based on the linear discriminant functions 1 and 2 (LD1 and LD2, respectively), obtained from the analysis of 20,712 single-nucleotide polymorphism (SNP) markers. Clusters are represented by colors. The graph at top right represents the contribution of the eigenvalues of the principal components selected, while the graph at bottom right indicates the variance explained by the eigenvalues of the two discriminant functions of the scatterplot.
Number of duplicate and unique cassava accessions based on the multilocus genotype (MLG) analysis belonging to the 12 genetic diversity groups identified by discriminant analysis of principal components (DAPC) based on the analysis from 20,712 single-nucleotide polymorphism (SNP) markers.
Due to the great germplasm exchange of the same accessions in the same region by different Embrapa units, the DAPC clustering was not formed by accessions belonging exclusively to a single Embrapa unit (Table 2). In general, DAPC clusters were composed of accessions belonging to two (Groups 3 and 6), three (Groups 2, 4, 5, 7, 8, 10, and 11), or four Embrapa units (Groups 1, 9, and 12). All MLGs were allocated within the same DAPC cluster, which demonstrates the agreement on the cassava clustering based on two completely different genetic approaches (i.e., MLG and DAPC analysis). This information reflects the high degree of accuracy in defining the similarity of duplicate accessions identified in the cassava germplasm.
The mean distance within the DAPC clusters ranged from 0.037 to 0.045 in Groups 1 and 12, respectively. On the other hand, the greatest variation in the genetic distance occurred in groups 6 (0.021 to 0.050) and 10 (0.014 to 0.050) (Table 2). This indicates that the maximum genetic variability was effectively divided among the DAPC clusters, considering that the principle of discriminant analysis presents as differential a better cluster separation and visualization while seeking to summarize all genetic differentiation by minimizing the genetic differences between individuals and maximizing the differences between clusters.
Discussion
Identification of duplicate accessions based on the multilocus approach
Historically, the movement of germplasm between the Amazon and the central region of Brazil allowed for domestication of the species, and the formation of several landraces during the migrations of native peoples allowed hybridization of the cultivars’ and wild relatives’ genotypes, mainly in the central region of Brazil (Nassar, 2002Nassar, N.M. 2002. Cassava Manihot esculenta Crantz genetic resources: origin of the crop, its evolution and relationships with wild relatives. Genetics and Molecular Research 1: 298-305.). Although hybridization still occurs in nature and in a controlled way in breeding programs, cassava germplasm is almost exclusively maintained by vegetative propagation under field and in vitro conditions. In addition, there is still an intense exchange of propagation material between different regions. This often presents problems as accession registration and labeling are not always standardized; therefore, new accessions present limited information about their correct identity and origin (Kilian and Graner, 2012Kilian, B.; Graner, A. 2012. NGS technologies for analyzing germplasm diversity in genebanks. Briefings in Functional Genomics 11: 38-50.). In addition, farmers tend to adopt different names for the exchanged accessions: in the formation of cassava germplasm banks, different accessions with the same name and the same accessions with different names were certainly collected. This is a reality that affects most germplasm banks of plant species that needs to be rectified.
Considering Brazil as the center of origin and diversity of cassava (Olsen, 2004Olsen, K.M. 2004. SNPs SSRs and inferences on cassava's origin. Plant Molecular Biology 56: 517-526.), Embrapa has sought to maintain a diverse collection of genetic resources of M. esculenta Crantz for immediate and future use. The effective maintenance of this cassava genetic diversity is particularly important in the context of modern agriculture, which aims to guarantee high levels of productivity in a competitive way through the search for better varieties with different characteristics from those used by farmers. Therefore, the introduction of additional genetic diversity by cross-pollination with landraces and even wild relatives should be perpetrated using the germplasm of the species, which is an important source of new variation. However, the cost of accession conservation under both field and in vitro conditions is extremely high and imposes limitations on the maintenance and expansion of germplasm banks. Accordingly, it is fundamental that the incorporation of new accessions should be done in a strategic way to capture the additional genetic diversity, which has potential value for end users.
The presence of duplicates of accession certainly contributes to increasing the cost of preserving cassava germplasm. As a consequence, curators need to use different tools to identify and reduce duplicate accessions. The molecular profile analysis of the 2,371 cassava accessions belonging to Embrapa's different collections revealed the presence of more than 20 % of them as duplicate accessions that were priority collected over more than 40 years in Brazil. The variation in the percentage of duplicates between the different CGBs (17 % for CPATU and 39 % for CPATSA) is probably due to the practice of germplasm exchange between the national cassava germplasm banks as well as to the collection of accessions in the same agroecological areas by different Embrapa units. While these redundancies need to be reduced among different cassava collections, it is necessary to encourage the preservation of cassava accessions in agroecological zones that are quite diverse, such as those located in the north, northeast, and south of Brazil, especially to guarantee the survival of accessions with specific adaptations to these regions.
In other cassava studies, isoenzymatic, microsatellite, and AFLP markers along with seven morphological descriptors were used to identify duplicates in a core collection of 521 cassava accessions, in which the results showed that there were at least 1 % of duplicates, even after using several tools to avoid duplicates (Chavarriaga-Aguirre et al., 1999Chavarriaga-Aguirre, P.; Maya, M.M.; Tohme, J.; Duque, M.C.; Iglesias, C.; Bonierbale, M.W.; Kresovich, S.; Kochert, G. 1999. Using microsatellites isozymes and AFLPs to evaluate genetic diversity and redundancy in the cassava core collection and to assess the usefulness of DNA-based markers to maintain germplasm collections. Molecular Breeding 5: 263-273.).
Although there are a number of differences between the germplasm stored in the CIAT and the Brazilian germplasm, as well as differences in strategies and techniques to detect duplicates, similar values of redundant accessions were verified (around 20 %). Regardless of approach, reduction in duplicate accessions in germplasm banks and the comparison of new and existing entries in the collection to avoid redundancy are strategies that must be implemented in cassava germplasm banks.
High precision identification of genetically identical samples requires that the selected markers be sufficiently variable and take into account the reality of the population structure and relationship between individuals (Gross et al., 2012Gross, B.L.; Volk, G.M.; Richards, C.M.; Forsline, P.L.; Fazio, G.; Chao, C.T. 2012. Identification of “duplicate” accessions within the USDA-ARS National Plant Germplasm System Malus Collection. Journal of the American Society for Horticultural Science 137: 333-342.). In addition, Arnaud-Haond et al. (2007Arnaud-Haond, S.C.M.; Alberto, F.; Serrão, E.A. 2007. Standardizing methods to address clonality in population studies. Molecular Ecology 16: 5115-5139.) used the Monte Carlo procedure to ensure that a set of microsatellite markers had sufficient discriminatory power to identify all MLGs in 220 genotypes of the marine grass, Cymodocea nodosa. They reported that only seven microsatellite loci were sufficient to determine the number of MLGs present in the samples with high reliability. In addition to the number of markers, one of the most common problems affecting estimates of diversity and the relationship between individuals is the limited polymorphism of the markers. This prevents the precise discrimination of different accessions that could be considered identical, leading to the overestimation of the occurrence of duplicates. Therefore, in numerical and polymorphism terms, it is expected that 20,712 SNPs used in the present study are well suited to determining the different MLGs based on the genetic profiles of the 2,371 cassava accessions.
In the process of identifying duplicates, it is also necessary to consider the overestimation of the number of different genotypes due to the presence of multiple MLGs belonging to the same genotype, whose occurrence is due to the presence of somatic mutations or genotyping errors (Douhovnikoff and Dodd, 2003Douhovnikoff, V.; Dodd, R.S. 2003. Intra-clonal variation and a similarity threshold for identification of clones: application to Salix exigua using AFLP molecular markers. Theoretical and Applied Genetics 106: 1307-1315.). According to Arnaud-Haond et al. (2007Arnaud-Haond, S.C.M.; Alberto, F.; Serrão, E.A. 2007. Standardizing methods to address clonality in population studies. Molecular Ecology 16: 5115-5139.), the adoption of a genetic distance threshold can be defined under the hypothesis that distinct MLGs belonging to the same genotype should not be rejected. However, certain authors have mentioned that the amount of genetic diversity acceptable between genetically similar accessions is still not well defined (Lund et al., 2003Lund, B.; Ortiz, R.; Skovgaard, I.M.; Waugh, R.; Andersen, S.B. 2003. Analysis of potential duplicates in barley gene bank collections using re-sampling of microsatellite data. Theoretical and Applied Genetics 106: 1129-1138.). Indeed, Fu et al. (2006Fu, Y-B.; Richards, K.W.; Peterson, G.W. 2006. Genetic variability in multiple accessions of two Canadian heritage crop cultivars as revealed by AFLP markers. Communications in Biometry and Crop Science 1: 1-10.) studied the genetic variation between six accessions of Marquis-type wheat and six accessions of Canadian Thorpe-type barley using AFLP markers to develop a limit value for declaring accession duplicates. These authors observed the true duplicates of these two species presenting a variation of up to 5 % in the AFLP fragments. Therefore, the definition of MLGs in cassava germplasm based on the maximum genetic distance of 0.05 between accessions seems to be quite realistic considering that the automation of SNP genotyping by GBS may further reduce the error rate in the definition of the genetic profile of each accession. Indeed, according to Zhou et al. (2015Zhou, L.; Matsumoto, T.; Tan, H-W.; Meinhardt, L.W.; Mischke, S.; Wang, B.; Zhang, D. 2015. Developing single nucleotide polymorphism markers for the identification of pineapple (Ananas comosus) germplasm. Horticulture Research. doi: 10.1038/hortres.2015.56
https://doi.org/10.1038/hortres.2015.56...
), the biallelic nature of the SNPs may facilitate a reduction in the error rate in the genotyping of accessions, in addition to promoting the compatibility of results among laboratories. In addition, the possibility of identifying duplicates with high reliability, accuracy, and automation with minimum lost data and genotyping errors is an important advantage of the analysis.
Population structure of duplicate cassava accessions
The use of molecular markers with high genomic coverage such as SNPs is relatively recent in cassava breeding programs. In addition, SNPs for identification of accession duplicates have been little explored, so this is one of the first studies seeking to understand the population structure in cassava accessions with high genetic redundancy. Based on the DAPC in the 124 MLGs, 12 diversity groups were identified. There was strong correlation between the DAPC clustering and the MLGs, since the DAPC grouped the 124 MLGs into 12 different groups, where all the individuals allocated in the same MLGs also remained in the same DAPC clusters.
The DAPC revealed clear separations between the 12 different groups of duplicate accessions, as a consequence of the sensitivity of this technique for detecting substructure in hierarchical models (Jombart et al., 2010Jombart, T.; Devillard, S.; Balloux, F. 2010. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics. doi: https://doi.org/10.1186/1471-2156-11-94
https://doi.org/10.1186/1471-2156-11-94...
). Population structure may be influenced by the presence of unique alleles of certain groups that have disproportionate influence on the DAPC. Therefore, most of the clusters are clearly separated from others using only the first two discriminant functions (Figure 4). Exceptions occurred only in Groups 1 and 5, which presented a slight overlap of a number of accessions.
According to Jombart et al. (2010Jombart, T.; Devillard, S.; Balloux, F. 2010. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics. doi: https://doi.org/10.1186/1471-2156-11-94
https://doi.org/10.1186/1471-2156-11-94...
), DAPC may be usefully applied to a wide variety of organisms, regardless of their ploidy and rate of genetic recombination, as the methodology is independent of any population genetic model and is therefore free of assumptions about Hardy-Weinberg or linkage disequilibrium. In addition, this analysis can be applied to large data sets and requires lower computational demand than Bayesian clustering algorithms based on pre-defined population genetics models such as the STRUCTURE or BAPS softwares (Jombart et al., 2010Jombart, T.; Devillard, S.; Balloux, F. 2010. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics. doi: https://doi.org/10.1186/1471-2156-11-94
https://doi.org/10.1186/1471-2156-11-94...
).
The presence of high levels of diversity is an important factor in the interpretation of population differentiation and the definition of genetic contrasts within certain duplicate groups. Therefore, these 12 groups formed by the DAPC can not only guide conservation strategies for cassava germplasm, but also contribute to the definition of genetic groups that can be crossed to generate segregating populations within breeding programs.
The ability to identify breeding populations with high agronomic performance is an objective in the development of new varieties of cassava (Oliveira et al., 2015Oliveira, E.J.; Santana, F.A.; Oliveira, L.A.; Santos, V.S. 2015. Genotypic variation of traits related to quality of cassava roots using affinity propagation algorithm. Scientia Agricola 72: 53-61.). In addition, according to Semagn et al. (2012Semagn, K.; Magorokosho, C.; Vivek, B.S.; Makumbi, D.; Beyene, Y.; Mugo, S.; Prasanna, B.M.; Warburton, M.L. 2012. Molecular characterization of diverse CIMMYT maize inbred lines from eastern and southern Africa using single-nucleotide polymorphic markers. BMC Genomics. doi: https://doi.org/10.1186/1471-2164-13-113
https://doi.org/10.1186/1471-2164-13-113...
), it is recognized that close groups of diversity tend to increase redundancy in several breeding programs; therefore, their use in crossbreeding may result in a waste of time and resources since crossbreeding low genetic complementarity parents can produce progenies of low performance (Dias et al., 2003Dias, L.A.S.; Marita, J.; Cruz, C.D.; Barros, E.G.; Salomão, T.M.F. 2003. Genetic distance and its association with heterosis in cacao. Brazilian Archives of Biology and Technology 46: 339-347.; Benin et al., 2012Benin, G.; Matei, G.; Costa de Oliveira, A.; Silva, G.O.; Hagemann, T.R.; Lemes da Silva, C.; Pagliosa, E.S.; Beche, E. 2012. Relationships between four measures of genetic distance and breeding behavior in spring wheat. Genetics and Molecular Research 16: 2390-2400.). On the other hand, crosses of genetically divergent parents may result in high phenotypic variation in progenies. Thus, the population structure and classification of cassava accessions in different groups based on the SNP markers may encourage breeders to better plan their crosses to maximize the chances of obtaining high phenotypic contrast progenies for various agronomic traits.
Perspectives for optimization and use of cassava germplasm
According to the FAO (2010Food and Agriculture Organization [FAO]. 2010. The state of ex situ conservation. p. 55-90. In: The second report on the state of the world's plant genetic resources for food and agriculture. FAO, Rome, Italy.), of the 7.4 million accessions stored in 1,700 germplasm banks, it is estimated that between 70 and 75 % are duplicates based on genotypic and passport data. In addition, the maintenance of cassava germplasm in the field has a high cost, about US$ 92.30 per accession/year according to Shands (2010Shands, H.; Hawtin, G.; MacNeil, G. 2010. The Cost to the CGIAR Centres of Maintaining and Distributing Germplasm. CGIAR, Montpellier, France. (CGIAR Consortium Study Paper, 10).). Therefore, it is necessary to invest in adequate methods of unequivocal identification of duplicates to ensure the continued maintenance of the highest priority genetic resources using the limited financial resources available, especially in the public sector.
In general, the goal of germplasm conservation is to maintain the greatest possible genetic diversity for a given plant species. Until some time ago, the lack of efficient strategies and methods for identifying duplicates in germplasm banks compelled the curators/breeders to keep all accessions in the collection, in face of evidence of the presence of duplicates based on morphoagronomic descriptors. However, now that it is possible to analyze differences directly at the DNA level and several computational tools are available to associate phenotypic and genotypic information, the disposal of accessions could be done in a safer and more reliable way. Nonetheless, the identification of duplicate samples using a set of common markers, as well as problems in the reproducibility of some molecular marker data among different laboratories has caused some difficulty in advancing studies of duplicates and comparing different germplasm collections. On the other hand, standardized NGS approaches, such as GBS, can identify useful SNPs to overcome these problems and thus represent an ideal information platform to reduce germplasm redundancy (Kilian and Graner, 2012Kilian, B.; Graner, A. 2012. NGS technologies for analyzing germplasm diversity in genebanks. Briefings in Functional Genomics 11: 38-50.). It is very probable that these modern approaches to conserving genetic resources will generate important inputs for curators/breeders in the prioritization of the maintenance of unique genotypes.
Once duplicates are identified, curators can evaluate the maintenance or disposal of these accessions to optimize germplasm conservation and also introduce new samples. However, accessions identified as identical, based on SNP markers, must still be characterized at the phenotypic level to determine whether they have similar enough characteristics to be considered synonymy in the collection. This information will certainly provide greater reliability for discarding redundant accessions, considering that somaclonal variation (mutations) in the cassava germplasm with the same SNP profile can occur throughout the cultivation and domestication process. Indeed, observations of this nature have been reported in other species of clonal propagation, such as pineapple, whose SNP analysis resulted in the identification of 64 unique accessions out of 170 evaluated (Zhou et al., 2015Zhou, L.; Matsumoto, T.; Tan, H-W.; Meinhardt, L.W.; Mischke, S.; Wang, B.; Zhang, D. 2015. Developing single nucleotide polymorphism markers for the identification of pineapple (Ananas comosus) germplasm. Horticulture Research. doi: 10.1038/hortres.2015.56
https://doi.org/10.1038/hortres.2015.56...
). In addition, these authors reported that certain pineapple accessions within the same groups of duplicates presented apparent morphological differences, despite having unique SNP profiles, such as the Cayenne 7898 QC accession whose pulp color is yellow, while the Cayenne 7898 4N has white pulp.
The characterization of cassava accessions belonging to the same MLG based on phenotypic descriptors is still essential for complementing the SNP molecular profile to precisely define duplicates’ accession for final disposal purposes. Therefore, in addition to improving the efficiency of the cassava germplasm banks’ routine activities, the information generated in this study can be useful for verifying and controlling the origin of varieties, directing the introduction of new germplasm, and even helping to minimize registration problems of synonymy varieties.
In summary, this study provides a comprehensive view of the redundancy of the M. esculenta germplasm in Brazil, the country considered the center of origin and diversity of the species (Olsen, 2004Olsen, K.M. 2004. SNPs SSRs and inferences on cassava's origin. Plant Molecular Biology 56: 517-526.). The use of more than 20,000 SNPs allowed for the identification of more than 20 % duplicates in this germplasm and some redundancy within Embrapa's collections that resulted from institutional germplasm exchange and collection of common accessions in different geographic regions. In the latter case, the intense exchange of cassava propagation material among farmers in Brazil favors the dissemination of genotypes of interest throughout the national territory. Generally, this germplasm exchange is accompanied by name changes, which certainly raises some doubts about origin. In general, a curator will decide to keep an accession in the collection until some kind of characterization can define its correct genetic background.
Despite the high reliability of SNP in defining accession duplicates, the occurrence of spontaneous mutations in vegetative propagation commonly mentioned in the literature means that a single gene alteration could generate important variations for the crop and, therefore, must be preserved. Thus, the final decision to discard an accession will be made after full phenotypic characterization and evaluation for agronomic parameters, resistance to diseases and abiotic stresses (water deficit and postharvest physiological deterioration), as well as root and starch quality.
The clear clustering of cassava accessions based on DAPC analysis was strongly correlated with the different MLGs such that all 124 MLGs were maintained in the same DAPC clusters. This demonstrates high reliability in the definition of accession duplicates, considering that both methods use different approaches for clustering. In addition, high genetic diversity was verified in the duplicate accessions considering that only the first two discriminant functions were able to distinguish the cluster with a high probability of assignment in the cassava accessions. All this information will be useful in improving efficiency in the management of cassava germplasm and will benefit the users of these genetic resources.
Acknowledgments
The authors thank the Fundação de Amparo à Pesquisa do Estado da Bahia (FAPESB), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for their financial assistance and scholarship support.
References
- Arnaud-Haond, S.C.M.; Alberto, F.; Serrão, E.A. 2007. Standardizing methods to address clonality in population studies. Molecular Ecology 16: 5115-5139.
- Benin, G.; Matei, G.; Costa de Oliveira, A.; Silva, G.O.; Hagemann, T.R.; Lemes da Silva, C.; Pagliosa, E.S.; Beche, E. 2012. Relationships between four measures of genetic distance and breeding behavior in spring wheat. Genetics and Molecular Research 16: 2390-2400.
- Bradbury, P.J.; Zhang, Z.; Kroon, D.E.; Casstevens, T.M.; Ramdoss, Y.; Buckler, E.S. 2007. TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics Applications Note 23: 2633-2635.
- Chavarriaga-Aguirre, P.; Maya, M.M.; Tohme, J.; Duque, M.C.; Iglesias, C.; Bonierbale, M.W.; Kresovich, S.; Kochert, G. 1999. Using microsatellites isozymes and AFLPs to evaluate genetic diversity and redundancy in the cassava core collection and to assess the usefulness of DNA-based markers to maintain germplasm collections. Molecular Breeding 5: 263-273.
- Dias, L.A.S.; Marita, J.; Cruz, C.D.; Barros, E.G.; Salomão, T.M.F. 2003. Genetic distance and its association with heterosis in cacao. Brazilian Archives of Biology and Technology 46: 339-347.
- Douhovnikoff, V.; Dodd, R.S. 2003. Intra-clonal variation and a similarity threshold for identification of clones: application to Salix exigua using AFLP molecular markers. Theoretical and Applied Genetics 106: 1307-1315.
- Doyle, J.J.; Doyle, J.L. 1987. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin 19: 11-15.
- Elshire, R.J.; Glaubitz, J.C.; Sun, Q.; Poland, J.A.; Kawamoto, K.; Buckler, E.S.; Mitchell, S.E. 2011. A robust simple genotyping-by-sequencing (GBS) approach for high diversity species. Plos One 6: 1-10.
- Food and Agriculture Organization [FAO]. 2010. The state of ex situ conservation. p. 55-90. In: The second report on the state of the world's plant genetic resources for food and agriculture. FAO, Rome, Italy.
- Fu, Y-B.; Richards, K.W.; Peterson, G.W. 2006. Genetic variability in multiple accessions of two Canadian heritage crop cultivars as revealed by AFLP markers. Communications in Biometry and Crop Science 1: 1-10.
- Gross, B.L.; Volk, G.M.; Richards, C.M.; Forsline, P.L.; Fazio, G.; Chao, C.T. 2012. Identification of “duplicate” accessions within the USDA-ARS National Plant Germplasm System Malus Collection. Journal of the American Society for Horticultural Science 137: 333-342.
- Hamblin, M.T.; Rabbi, I.Y. 2014. The effects of restriction-enzyme choice on properties of genotyping-by-sequencing libraries: a study in cassava (Manihot esculenta). Crop Science 54: 2603-2608.
- Horna, D.; Debouck, D.; Dumet, D.; Hanson, J.; Payne, T.; Sackville-Hamilton, R.; Sanchez, I.; Upadhyaya, H.D.; Van Den Houwe, I. 2010. Evaluating Cost-Effectiveness of Collection Management: Ex-situ Conservation of Plant Genetic Resources in the CG System. CGIAR, Montpellier, France.
- Instituto Nacional de Meteorologia [INMET]. 2016. BDMAP: historical data = BDMAP: dados históricos. Available at: http://www.inmet.gov.br/portal/index.php?r=home2/index [Accessed Jan 21, 2017) (in Portuguese).
» http://www.inmet.gov.br/portal/index.php?r=home2/index - Jombart, T.; Devillard, S.; Balloux, F. 2010. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genetics. doi: https://doi.org/10.1186/1471-2156-11-94
» https://doi.org/10.1186/1471-2156-11-94 - Kamvar, Z.N.; Brooks, J.C.; Grunwald, N.J. 2015. Novel R tools for analysis of genome-wide population genetic data with emphasis on clonality. Frontiers in Genetics 6: 1-10.
- Kamvar, Z.N.; Tabima, J.F.; Grunwald, N.J. 2014. Poppr: an R package for genetic analysis of populations with clonal partially clonal and/or sexual reproduction. Peer J. doi: https://doi.org/10.7717/peerj.281
» https://doi.org/10.7717/peerj.281 - Kilian, B.; Graner, A. 2012. NGS technologies for analyzing germplasm diversity in genebanks. Briefings in Functional Genomics 11: 38-50.
- Lund, B.; Ortiz, R.; Skovgaard, I.M.; Waugh, R.; Andersen, S.B. 2003. Analysis of potential duplicates in barley gene bank collections using re-sampling of microsatellite data. Theoretical and Applied Genetics 106: 1129-1138.
- Mammadov, J.; Aggarwal, R.; Buyyarapu, R.; Kumpatla, S. 2012. SNP markers and their impact on plant breeding. International Journal of Plant Genomics 2012: 1-12.
- Nassar, N.M. 2002. Cassava Manihot esculenta Crantz genetic resources: origin of the crop, its evolution and relationships with wild relatives. Genetics and Molecular Research 1: 298-305.
- Nzuki, I.; Katari, M.S.; Bredeson, J.V.; Masumba, E.; Kapinga, F.; Salum, K.; Mkamilo, G.S.; Shah, T.; Lyons, J.B.; Rokhsar, D.S.; Rounsley, S.; Myburg, A.A.; Ferguson, M.E. 2017. QTL mapping for pest and disease resistance in cassava and coincidence of some QTL with introgression regions derived from Manihot glaziovii Frontiers in Plant Science. doi: https://doi.org/10.3389/fpls.2017.01168
» https://doi.org/10.3389/fpls.2017.01168 - Oliveira, E.J.; Resende, M.D.V.; Santos, V.S.; Fereira, C.F.; Oliveira, G.A.F.; Silva, M.S.; Oliveira, L.A.; Aguilar-Vildoso, C.I. 2012. Genome-wide selection in cassava. Euphytica 187: 263-276.
- Oliveira, E.J.; Santana, F.A.; Oliveira, L.A.; Santos, V.S. 2015. Genotypic variation of traits related to quality of cassava roots using affinity propagation algorithm. Scientia Agricola 72: 53-61.
- Olsen, K.M. 2004. SNPs SSRs and inferences on cassava's origin. Plant Molecular Biology 56: 517-526.
- Rabbi, I.Y.; Kulakow, P.A.; Manu-Aduening, J.A.; Dankyi, A.A.; Asibuo, J.Y.; Parkes, E.Y.; Abdoulaye, T.; Girma, G.; Gedil, M.A.; Ramu, P.; Reyes, B.; Maredia, M.K. 2015. Tracking crop varieties using genotyping-by-sequencing markers: a case study using cassava (Manihot esculenta Crantz). BMC Genetics. doi: https://doi.org/10.1186/s12863-015-0273-1
» https://doi.org/10.1186/s12863-015-0273-1 - Semagn, K.; Magorokosho, C.; Vivek, B.S.; Makumbi, D.; Beyene, Y.; Mugo, S.; Prasanna, B.M.; Warburton, M.L. 2012. Molecular characterization of diverse CIMMYT maize inbred lines from eastern and southern Africa using single-nucleotide polymorphic markers. BMC Genomics. doi: https://doi.org/10.1186/1471-2164-13-113
» https://doi.org/10.1186/1471-2164-13-113 - Shands, H.; Hawtin, G.; MacNeil, G. 2010. The Cost to the CGIAR Centres of Maintaining and Distributing Germplasm. CGIAR, Montpellier, France. (CGIAR Consortium Study Paper, 10).
- Treuren, R.; Hintum, T.J.L. 2003. Marker-assisted reduction of redundancy in germplasm collections: genetic and economic aspects. Acta Horticulturae 623: 139-149.
- Zhou, L.; Matsumoto, T.; Tan, H-W.; Meinhardt, L.W.; Mischke, S.; Wang, B.; Zhang, D. 2015. Developing single nucleotide polymorphism markers for the identification of pineapple (Ananas comosus) germplasm. Horticulture Research. doi: 10.1038/hortres.2015.56
» https://doi.org/10.1038/hortres.2015.56
Edited by
Publication Dates
-
Publication in this collection
Jul-Aug 2019
History
-
Received
12 Nov 2017 -
Accepted
02 Mar 2018