ABSTRACT
Some germplasm collections have a high number of accessions, which makes it difficult to explore the genetic variability present in the germplasm bank due to the redundancy and the difficulty of detailed analysis of all conserved accessions. Therefore, our study aimed to analyze the genetic diversity of 153 lima bean (Phaseolus lunatus) accessions for the purpose of constructing a core collection. Eleven SSRs were used for this purpose. The 153 lima bean accessions can be represented by low redundancy using a minimum of 34 accessions, thus representing 22 % of the size of the entire germplasm bank. The core collection had a higher Shannon diversity index and expected heterozygosity (1.906 and 0.811, respectively) than those presented by the entire germplasm bank (1.605 and 0.713, respectively), indicating a higher polymorphism of the representative cultivars in relation to the entire collection. The accessions selected for the core collection may be used in future studies of genome association as well as in genetic crosses in breeding programs aimed at developing improved cultivars with high genetic diversity which can meet current and future market needs.
germplasm; genetic diversity; genetic structure; microsatellite markers
Introduction
The extinction of materials grown in centers of diversity is the main cause of genetic erosion in crops. One of the strategies for solving this problem is the establishment of germplasm banks containing as many genetic components as possible, such as varieties, lineages and clones, with the highest variability possible (Bueno et al., 2001Bueno, L.C.S.; Mendes, A.N.G.; Carvalho, S.P. 2001. Genetic Plant Breeding: Principles and Procedures = Melhoramento Genético de Plantas: Princípios e Procedimentos. UFLA, Lavras, MG, Brazil (in Portuguese).).
Such germplasm banks function as an ex situ strategy of conservation, where a sample of the genetic variability of a particular species is conserved under artificial conditions, outside its habitat, in order to avoid loss of genetic resources and to conserve gene sources for current and future use (Heiden et al., 2007Heiden, G.; Barbieri, R.L.; Couto, M.E.O.; Medeiros, A.R.M.; Sinigaglia, C. 2007. Peppers and chilies from southern Brazil: Creole varieties maintained by Embrapa temperate climate = Pimentas e pimentões do sul do Brasil: variedades crioulas mantidas pela Embrapa clima temperado. Revista Brasileira de Agroecologia 2: 1 (in Portuguese).). The management, evaluation and use of large germplasm banks is expensive and inefficient due to redundancies, duplications and the difficulty of analyzing all conserved accessions in detail (Brown, 1989Brown, A.H.D. 1989. Core collections: a practical approach to genetic resources management. Genome 31: 818-824.; Xu et al., 2016Xu, Y.; Chen, C.; Ji, D.; Xu, K.; Xie, X.; Xie, C. 2016. Developing a core collection of Pyropia haitanensis using simple sequence repeat markers. Aquaculture 452: 351-356.).
In this context, core collections, defined as a subset of accessions that represents as much of the genetic diversity contained in a germplasm bank with as much minimal redundancy as possible (Frankel, 1984Frankel, O.H. 1984. Genetic Perspectives of Germplasm Conservation. Cambridge University Press, Cambridge, UK.; Liu et al., 2017Liu, X.B.; Li, J.; Yang, Z.L . 2017.Genetic diversity and structure of core collection of winter mushroom (Flammulina velutipes) developed by genomic SSR markers. Hereditas 155: 3.), are being developed. To validate the representativeness of the accessions of a core collection, an appropriate evaluation of the genetic diversity of these accessions is required (Wang et al., 2010Wang, Y.K.; Wu, G.L.; Li, D.K.; Ji, A.Q.; Sui, C.L.; Zhao, A.L. 2010. Advances in core collection of fruit germplasm. Journal of Plant Genetic Resources 11: 380-385.). Kim et al. (2007)Kim, K.W.; Chung, H.K.; Cho, G.T.; Ma, H.; Chandrabalan, D. 2007. PowerCore: a program applying the advanced M strategy with a heuristic search for establishing core sets. Bioinformatics 23: 2155-2162. such as a strategy developed by a program known as PowerCore, which can be applied to such an evaluation. According to the method of this program, a core collection is developed based on genetic data by an extension of a heuristic algorithm using an advanced maximization strategy (M). The PowerCore analysis allows for the retention of all alleles in a core collection with the minimum possible number of accessions as possible.
The species Phaseolus lunatus L., known as lima bean, is the second most important legume of its botanical genus (Baudet, 1977Baudet, J.C. 1977. The taxonomic status of the cultivated types of Lima bean (Phaseolus lunatus L.). Trop Grain Legume 7: 29-30.). This crop is important to the northeastern region of Brazil, as it stands out as a subsistence species, being produced under rainfed regime, mainly by smallholder farmers (Oliveira-Silva et al., 2017Oliveira-Silva, R.N.; Burle, M.L.; Pádua, J.G.; Lopes, A.C.A.; Gomes, R.L.F.; Martínez-Castillo, J. 2017. Phenotypic diversity in lima bean landraces cultivated in Brazil, using the Ward-MLM strategy. Chilean Journal of Agricultural Research 77: 35.). The Germplasm Active Bank of P. lunatus of the Federal University of Piauí (BGP-UFPI) has 1,025 accessions, the majority originating in different Brazilian states and the Federal District. To efficiently manage and use the accessions conserved in the BGP-UFPI, it is necessary to develop a representative core collection, with the least number of redundant accessions. In this context, the aim of this study was to establish a core collection for lima bean accessions of the BGP-UFPI, and evaluate their genetic diversity and structuring based on variations as revealed by SSR markers.
Materials and Methods
Plant material and DNA extraction
In the present study, 153 accessions of lima beans were analyzed, constituting a representative sample of the Germplasm Active Bank of P. lunatus L. of the Federal University of Piauí (BGP-UFPI). These accessions were selected based on their origin, in order to maximize the representation of accessions from distinct geographical regions. The majority of the accessions were collected in the main areas of lima bean cultivation in Brazil, comprising eight states and the Federal District. Thus, the sampled accessions consisted of: 40 from Paraíba (one of the greatest lima bean-producing states in the country); 34 from Piauí; 34 from Ceará; 16 from Minas Gerais; 11 from Maranhão; two from São Paulo; two from Espírito Santo; one from Goiás; three from the Federal District; and ten of unknown origin. The seeds of the accessions were sown in pots in a greenhouse, and after 30 days, young leaf tissue samples were collected for genomic DNA extraction, after lyophilization and leaf milling, using the protocol based on CTAB as described by Doyle and Doyle (1990)Doyle, J.J.; Doyle, J.L. 1990. Isolation of plant DNA from fresh tissue. Focus 1: 13-15.. To assess the quality and concentration of the extracted DNA, aliquots of DNA obtained from each accession were quantified on Sybr-safe 1 % (w/v) agarose gels and compared with standard phage lambda DNA. Next, the DNA of each sample was diluted to a final concentration of 10 ng μL1.
SSR genotyping
For the study of genetic diversity 11 microsatellite markers (Gaitán-Solís et al., 2002Gaitán-Solís, E.; Duque, M.C.; Edwards, K.J.; Tohme, J. 2002. Microsatellite repeats in common bean (Phaseolus vulgaris L.): isolation, characterization, and cross-species amplification in Phaseolus ssp. Crop Science 42: 2128-2136.) isolated and optimized for Phaseolus vulgaris L. were used in the lima bean accessions (Table 1). Amplification reactions for the 11 loci were induced with 20 ng of DNA, 1 U of Taq DNA polymerase, 2.0 mM of magnesium chloride (MgCl2), 0.2 mM of each dNTP, 0.1 μM of each primer (“forward” and “reverse”) and 1 × PCR reaction buffer in a final volume of 20 μL. The amplification conditions used were as follows: 1) 94 °C for 2 min; 2) 94 °C for 15 s; 3) annealing temperature (Ta specific for each SSR) for 15 s; 4) 72 °C for 15 s; 5) repetition of steps 2-4 30 times; and 6) 72 °C for 10 min.
Genotyping was performed on a semi-automated sequencer LI-COR 4300 DNA Analyzer, which uses infrared fluorescence technology. The data were read using the SAGA GT analysis software for the assignment of alleles in each sample.
Development of the core collection for P. lunatus
The development of the core collection was established based on the genotyping data using the advanced maximization strategy (M), implemented by modifying the heuristic algorithm in the PowerCore software program as described by Kim et al. (2007)Kim, K.W.; Chung, H.K.; Cho, G.T.; Ma, H.; Chandrabalan, D. 2007. PowerCore: a program applying the advanced M strategy with a heuristic search for establishing core sets. Bioinformatics 23: 2155-2162.. The advanced maximization strategy M selects the most diverse accessions to represent the total variability of the entire collection. The PowerCore software program minimizes allele loss and therefore effectively selects the most diverse accessions, reducing the number of redundant accessions as described by Kim et al. (2007)Kim, K.W.; Chung, H.K.; Cho, G.T.; Ma, H.; Chandrabalan, D. 2007. PowerCore: a program applying the advanced M strategy with a heuristic search for establishing core sets. Bioinformatics 23: 2155-2162..
Genetic diversity analysis
Polymorphic information content (PIC), as described by Botstein et al. (1980)Botstein, D.; White R.L.; Skolnick, M.; Davis, R.W. 1980. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. American Journal of Human Genetics 32: 314-331., was calculated as a function of the number of alleles detected and their distribution and frequency in the groups of accessions. The values of the PIC per locus were determined by
, where pi and pj are the frequencies of the alleles i and j in the accession groups. The calculation is based on the number of alleles detected per given locus and the relative frequency of each allele in the total set of hits.The estimates of genetic diversity (Na = mean number of alleles per locus, I = Shannon diversity index, Ho = observed heterozygosity, He = expected heterozygosity), inbreeding coefficients, the analysis of molecular variance (AMOVA) and principal coordinate analysis (PCoA) were obtained using GENALEX 6.5 software analysis (Peakall and Smouse, 2012Peakall, R.; Smouse, P.E. 2012. GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28: 2537-2539.).
A cluster analysis was performed using the unweighted pair group method with arithmetic means (UPGMA), based on Rogers’ distances (1972), to determine the genetic relationships between accessions using the Populations software package (Langella, 2002Langella, O. 2002. Populations v.1.2.32. Available at: http://bioinformatics.org/~tryphon/populations/index.html#pubdas [Accessed Aug 31, 2017]
http://bioinformatics.org/~tryphon/popul...
). The stability of the clusters was tested using the bootstrap procedure based on 1,000 resamples, and the final tree was formatted using the MEGA software tool, version 6.0 (Tamura et al., 2013Tamura, K.; Stecher, G.; Peterson, D.; Fillipski, A.; Kumar, S. 2013. MEGA6 Molecular evolutionary genetics analyses version 6.0. Molecular Biology and Evolution 30: 2725-2729.).
The genetic structure of the core collection was analyzed by the STRUCTURE 2.3.3 software package (Pritchard et al., 2000Pritchard, J.K.; Stephens, M.; Donnelly, P. 2000. Inference of population structure using multilocus genotype data. Genetics 155: 945-959.) based on the admixture model with correlated allelic frequencies. The models were tested for K values ranging from 1 to 12, with 10 independent runs per K value. For each simulation, the initial period (burnin) was 100,000 steps followed by 500,000 Monte Carlo Markov chain (MCMC) iterations. To determine the most likely value of K, the ΔK method (Evanno et al., 2005Evanno, G.; Regnaut, S.; Goudet, J. 2005. Detecting the number of clusters of individuals using the software structure: a simulation study. Molecular Ecology 14: 2611-2620.) was used as implemented in the Harvester Structure program (Earl and von Holdt, 2012Earl, D.A.; von Holdt, B.M. 2012. Structure harvester: a website and program for visualizing structure output and implementing the Evanno method. Conservation Genetics Resources 4: 359-361.).
Results
Genetic diversity of the BGP-UFPI
The 11 SSR markers analyzed in 153 accessions from the BGP-UFPI generated a total of 113 alleles with a mean of 10.27 alleles per locus, ranging from 6 (BM 146 and BM 160) to 16 alleles (BM 141 and GATS 91) per locus. The mean PIC was 0.675, and values ranged from 0.472 (BM 212) to 0.903 (BM 140). The set of 153 lima bean accessions presented a value of 1.605 for the Shannon diversity index, the observed heterozygosity (Ho) was 0.105, and the expected heterozygosity (He) value was 0.713. The coefficient of inbreeding (0.859) indicates that these lima bean accessions present a large heterozygote deficit.
Development of the core collection and its genetic diversity
Based on the results of the PowerCore software package, the 113 alleles obtained from the 11 SSR loci for the 153 lima bean accessions can be represented using a minimum of 34 accessions. Therefore, these accessions may form a lima bean core collection, represented by 22 % of the BGP-UFPI accessions and with low redundancy. The core collection had a Shannon diversity index and expected heterozygosity (1.0906 and 0.811, respectively) higher than those presented by the BGP-UFPI, indicating a higher genetic diversity at the individual level of the selected accessions (Table 2). Heterozygosity is used to estimate the degree of genetic variation in a population.
– Estimates of genetic diversity for 11 microsatellite loci in BGP-UFPI and in the core collection of lima bean (Phaseoulus lunatus). Shannon diversity index (I), expected heterozygosity (He) and polymorphic information content (PIC).
Similarly, the mean PIC for the BGP-UFPI was 0.675, while it was 0.773 for the nuclear collection (Table 2). The PIC provides an estimate of the discriminatory power of the locus and is related to the number and frequency of alleles. According to the classification of Botstein et al. (1980)Botstein, D.; White R.L.; Skolnick, M.; Davis, R.W. 1980. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. American Journal of Human Genetics 32: 314-331., markers with values of PIC greater than 0.5 are considered very informative, values between 0.25 and 0.50 are moderately informative and values lower than 0.25 are minimally informative. As in the case of the Shannon diversity indexes, PIC values were higher in all loci for accessions of the core collection than in the BGP-UFPI accessions (Table 2).
The estimates of genetic diversity for the core collection and for the BGP-UFPI are summarized in Table 3. The t-tests comparing the mean genetic diversity estimates (Na, He, Ho and I) of the nuclear collection and the BGP-UFPI were not significant (p > 0.01), which shows that the genetic diversity of the core collection has no significant difference in relation to the complete collection. The estimates of genetic diversity, heterozygosity and PIC values of the core collection were higher than the values obtained for the BGP-UFPI, an expected result, since genetic diversity increases with the elimination of genetically similar (or redundant) accessions (Agrama et al., 2009Agrama, H.A.; Yan, W.G.; Lee, F.; Fjellstrom, R.; Chen, M-H.; Jia, M.; McClung, A. 2009. Genetic assessment of a minicore subset developed from the USDA rice genebank. Crop Science 49: 1336-1346.).
– Estimates of genetic diversity for the BGP-UFPI and the core collection of the lima bean (Phaseoulus lunatus), based on 11 SSR markers. Number of accessions (N), mean number of alleles (Na), Shannon diversity index (I), observed (Ho) and expected (He) heterozygosity and inbreeding coefficient (F).
The representativeness of the core collection was also validated by principal coordinate analysis (PCoA), showing the distribution of the BGP-UFPI and core collection accessions along the first two coordinates (Figure 1). The variation explained by the first two coordinates was 22 %, which gave rise to the observation of a general overlap between accessions selected for the core collection and the accessions from the BGP-UFPI. This result suggests that the core collection consists of an appropriate representation of the total genetic diversity in the BGP-UFPI lima bean.
– Graph of the principal coordinates analysis, showing the dispersion of the accessions from the core collection (N = 34, black diamonds) and from the lima bean (Phaseolus lunatus) BGP-UFPI (N = 153, white squares).
In addition to allelic variation, other information of great importance is the genetic distance, which allows for the evaluation of the redundancy of germplasm collections. From the analysis of the molecular data in this study, a genetic distance matrix was generated using Rogers’ distances (1972), which was used to group individuals by constructing a dendrogram with UPGMA (Figure 2). The dendrogram indicates that the accessions were not grouped according to their geographical origins. Considering the arbitrary threshold of 0.40 for the Rogers’ distance, the dendrogram can be divided into 3 major groups (I-III). Group I has only one accession, and group II has two accessions, all from the state of Paraíba, which show greater genetic dissimilarity in relation to accessions coming from the other locations. Group III consists of the remaining 150 accessions (98 % of the BGP-UFPI). The lima bean accessions in group III include samples from the nine sites: Piauí, Paraíba, Ceará, Maranhão, Minas Gerais, Espírito Santo, São Paulo, the Federal District and Goiás.
– UPGMA dendrogram built with Rogers’ distances (1972), illustrating the grouping of 153 accessions from the lima bean (Phaseolus lunatus) BGP-UFPI, based on variation of eleven microsatellite loci. Black diamonds indicate the 34 accessions selected by PowerCore for the development of a core collection.
The 34 lima bean accessions selected by PowerCore for the development of the core collection include all samples of groups I and II, plus 31 accessions from the group III. The core collection accessions are from the states of Piauí, Maranhão, Minas Gerais, Federal District, Ceará and Paraíba.
Core collection genetic structure
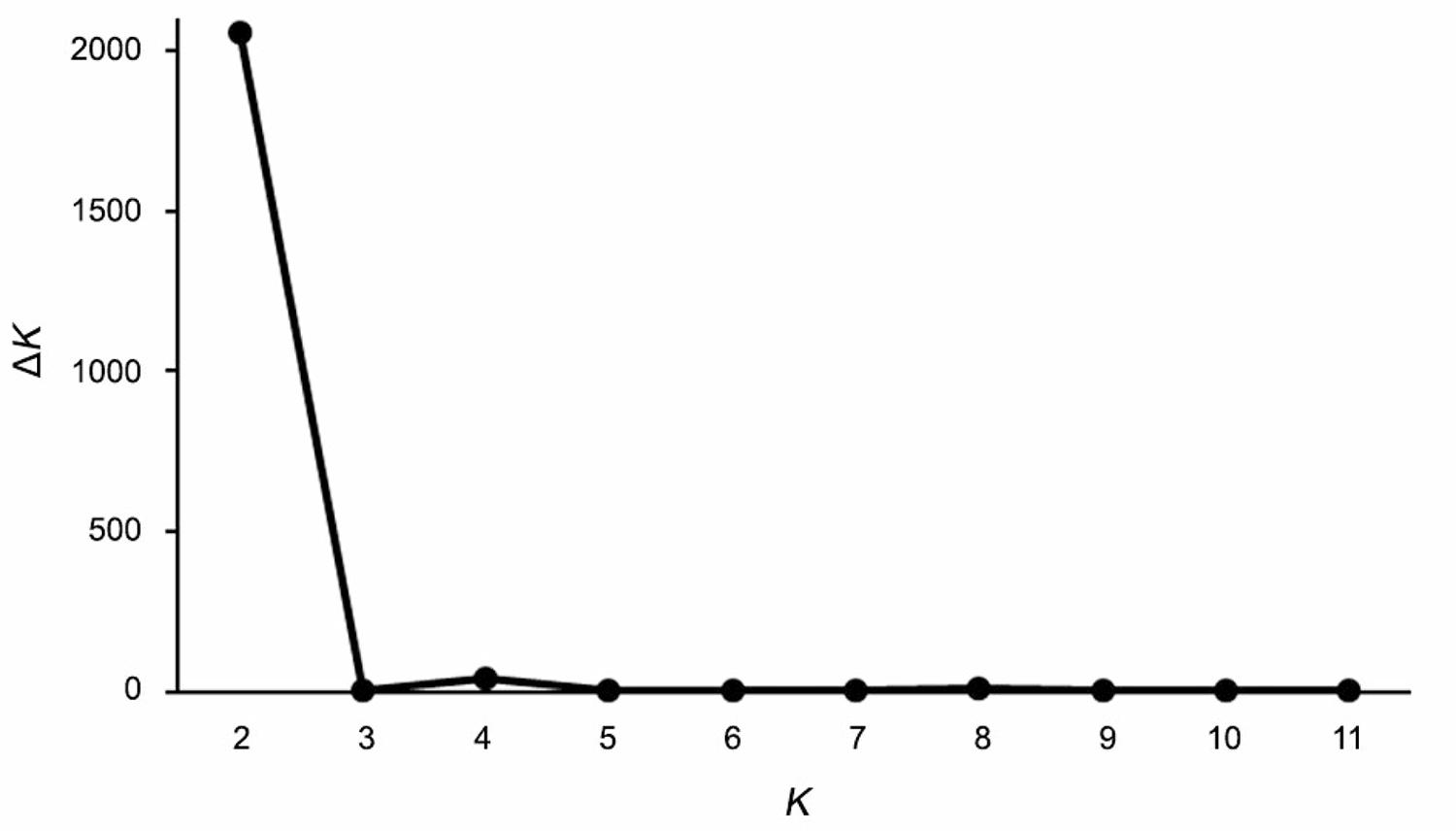
The population structure of the lima bean accessions was evaluated using the Bayesian analysis of the STRUCTURE software program. The ΔK method (Evanno et al., 2005Evanno, G.; Regnaut, S.; Goudet, J. 2005. Detecting the number of clusters of individuals using the software structure: a simulation study. Molecular Ecology 14: 2611-2620.) suggested K = 2 as the most probable number of genetic clusters (Figure 3), indicating the presence of two main groups. Group 1 comprised 22 accessions of the core collection, from the states of Piauí, Paraíba, Ceará, Minas Gerais and the Federal District. Group 2 comprised 12 accessions of the core collection, from the states of Piauí, Paraíba, Maranhão and the Federal District (Figure 4). Although a number of different accessions from the same state were assigned to distinct groups, the analysis of the genetic structure showed little evidence of admixture among lima bean accessions. This result suggests genetic structuring among lima bean varieties grown in different states (Figure 4).
– Graphical representation of the optimal number of groups (K) in the STRUCTURE inferred using the ΔK criterion of Evanno et al. (2005)Evanno, G.; Regnaut, S.; Goudet, J. 2005. Detecting the number of clusters of individuals using the software structure: a simulation study. Molecular Ecology 14: 2611-2620.. The analysis was based on data obtained from the 11 SSR loci used to evaluate the genetic diversity of 153 accessions from the lima bean (Phaseolus lunatus) BGP-UFPI.
– Bar plot representation at K = 2 of the Bayesian analysis of STRUCTURE, showing membership coefficients for 153 accessions from the lima bean (Phaseolus lunatus) BGP-UFPI. Accessions are ordered according to their geographical origins (ND = non-determined). Different colors represent distinct genetic groups.
The AMOVA considering groups of accessions from the BGP-UFPI and from the core collection indicated that only 3 % of the variation is between these groups and 97 % of the variation is within each group (Table 4). The small proportion of variation between groups indicates that most of the genetic variability of BGP-UFPI was maintained in the core collection. This result also suggests that the method used to establish the lima bean core collection was effective in representing the genetic diversity present in the BGP-UFPI.
– Analysis of molecular variance (AMOVA) considering the entire BGP-UFPI and the core collection. Degrees of freedom (d.f.).
Discussion
The genetic profile of large germplasm collections of crop species is essential to the identification of small and diverse sets of accessions for efficient use in breeding programs (Choudhury et al., 2014Choudhury, D.R.; Singh, N.; Singh, A.K.; Kumar, S.K.; Srinivasan, K.; Tyagi, R.K.; Ahmand, A.; Singh, N.K.; Singh, R. 2014. Analysis of genetic diversity and population structure of rice germplasm from north-eastern region of India and development of a core germplasm set. Plos One 9: e113094.). The present study is the first effort to create a core collection of lima bean accessions. We successfully developed a basic core collection of lima bean from the BGP-UFPI based on 11 SSR markers, which were efficient and selective markers in the genome analysis of lima beans.
Ideally, the diversity of a core collection should be at least 70 % of that found in the entire collection (Brown and Spillane, 1999Brown, A.H.D.; Spillane, C. 1999. Implementing core collections: principles, procedures, progress, problems and promise. p. 1-9. In: Johnson, R.C.; Hodgkin, T., eds. Core collections for today and tomorrow. IPGRI, Rome, Italy.). The 34 accessions selected for the core collection represent all the allelic variability present in the initial germplasm collection. In addition, the greater genetic diversity estimates of the core collection in comparison to the BGP-UFPI lima bean demonstrates a desirable characteristic of our proposed core collection (Odong et al., 2013Odong, T.L.; Jansen, J.; van Eeuwijk, F.A.; van Hintum, T.J.L. 2013. Quality of core collections for effective utilisation of genetic resources review, discussion and interpretation. Theoretical and Applied Genetics 126: 289-305.). The results demonstrated that the advanced M strategy of PowerCore is efficient in capturing allelic diversity in a basic collection. We propose a core collection representing 22 % of the accessions currently maintained at the BGP-UFPI. The percentage of accessions to form the core collection of our study is higher than the percentage (approximately 10 %) recommended in the literature (Odong et al., 2013Odong, T.L.; Jansen, J.; van Eeuwijk, F.A.; van Hintum, T.J.L. 2013. Quality of core collections for effective utilisation of genetic resources review, discussion and interpretation. Theoretical and Applied Genetics 126: 289-305.). However, the PowerCore method has been used successfully in the establishment of core collections of many economically important crops, such as Oryza sativa (Tiwari et al., 2015Tiwari, K.K.; Singh, A.; Pattnaik, S.; Sandhu, M.; Kaur, S.; Jain, S.; Tiwari, S.; Mehrotra, S.; Anumalla, M.; Samal, R.; Bhardwaj, J.; Dubey, N.; Sahu, V.; Kharshing, G.A.; Zeliang, P.K.; Sreenivasan, K.; Kumar, P.; Parida, S.K.; Mithra, S.V.A.; Rai, V.; Tyagi, W.; Agrawal, P.K.; Rao, A.R.; Pattanayak, A.; Chandel, G.; Singh, A.K.; Bisht I.S.; Bhat, K.V.; Rao, G.J.N.; Khurana, J.P.; Singh, N.K.; Mohapatra, T. 2015. Identification of a diverse mini-core panel of Indian rice germplasm based on genotyping using microsatellite markers. Plant Breeding 134: 164-171.), Glycine max (Kaga et al., 2012Kaga, A.; Shimizu, T.; Watanabe, S.; Tsubokura, Y.; Katayose, Y.; Harada, K.; Vaughan, P.A.; Tomookan, N. 2012. Evaluation of soybean germplasm conserved in NIAS genebank and development of mini core collections. Breeding Science 61: 566-592.; Gireesh et al., 2017Gireesh, C.; Husain, S.M.; Shivakumar.; Satpute, G.K.; Kumawat, G.; Arya, M.; Agarwal, D.K.; Bhatia, V.S. 2017. Integrating principal component score strategy with power core method for development of core collection in Indian soybean germplasm. Plant Genetic Resources 15: 230-238.), Cucumis melo (Hu et al., 2015), Vicia faba (Göl et al., 2017Göl, Ş.; Doğanlar, S.; Frary, A. 2017. Relationship between geographical origin, seed size and genetic diversity in faba bean (Vicia faba L.) as revealed by SSR markers. Molecular Genetics and Genomics 5: 991-999.) and Capsicum spp. (Mongkolporn et al., 2015Mongkolporn, O.; Hanyong, S.; Chunwongse, J.; Wasee, S. 2015. Establishment of a core collection of chilli germplasm using microsatellite analysis. Plant Genetic Resources 13: 104-110.; Lee et al., 2016Lee, H.Y.; Ro, N.Y.; Jeong, H.J.; Kwon, J.K.; Jo, J.; Há, Y.; Jung, A.; Han, J.W.; Venkatesh, J.; Kang, B.C. 2016. Genetic diversity and population structure analysis to construct a core collection from a large Capsicum germplasm. BMC Genetics 17: 142.). Higher estimates of genetic diversity for the core collection in relation to the lima bean BGP-UFPI may be explained simply by the PowerCore method, which results in a better sampling of allelic diversity using a smaller number of individuals when compared to the entire collection. In addition, the inclusion of accessions from various states maximizes the representation of the existing allelic diversity in the crop. As plant domestication proceeds, distinct characteristics may be selected in different locations in response to local preference, which in turn, may result in high levels of genetic diversity (Ladizinsky, 1998Ladizinsky, G. 1998. Plant Evolution under Domestication. Springer, Dordrecht, The Netherlands.).
Through the analysis of genetic structuring we investigated the genetic divergence between the accessions from the core collection and the BGP-UFPI lima bean. Two methods of analysis, STRUCTURE and PCoA, were used to interpret the genetic structure of lima bean accessions. PCoA showed a greater dispersion of the core collection accessions (black diamonds), confirming that they retain much of the genetic diversity of the entire collection (Figure 1). The Bayesian analysis performed with STRUCTURE, as well as the dispersion of the accessions observed in PCoA suggest that there is little or no admixture, demonstrating that there is also a great level of genetic divergence among lima bean accessions of the BGP-UFPI. Bayesian analysis shows that accessions form two distinct genetic groups.
The efficiency of the development of the core collection was further demonstrated by the results of Bayesian analysis in STRUCTURE, and cluster analysis with UPGMA. The accessions selected for the core collection are present in both genetic groups detected by STRUCTURE. Although the UPGMA dendrogram had limited agreement with the geographical origin of the accessions, the accessions of the nuclear collection were distributed across its three major groups. Once again, these results suggest that the genetic variability of BGP-UFPI was well represented in the core collection.
Analysis of the genetic structure of populations is an effective method for studying the genetic relationships of plant germplasm collections (Haouane et al., 2011Haouane, H.; Bakkali, A.E.; Moukhli, A.; Tollon, C.; Santoni, S.; Oukabli, A.; Modafar, C.E.; Khadari, B. 2011. Genetic structure and core collection of the world olive germplasm bank of Marrakech: towards the optimized management and use of Mediterranean olive genetic resources. Genetica 139: 1083-1094.; Emanuelli et al., 2013Emanuelli, F.; Lorenzi, S.; Grzeskowiak, L.; Catalano, V.; Stefanini, M.; Troggio, M.; Myles, S.; Martinez-Zapater, J.M.; Zyprian, E.; Moreira, F.M.; Grando, M.S. 2013. Genetic diversity and population structure assessed by SSR and SNP markers in a large germplasm collection of grape. BMC Plant Biology 13: 39. DOI:10.1186/1471-2229-13-39; Cristo-Araújo et al., 2015Cristo-Araújo, M.; Rodrigues, M.; Astolfi-Filho, S.; Clement, C.R. 2015. Peach palm core collection in Brazilian Amazonia. Crop Breeding and Applied Biotechnology 15: 18-25.). Previous studies have shown that the analysis of the genetic structure combined with cluster analysis are able to classify precisely the groups of complex populations without obvious genetic differentiation or with large gene flow among the different groups (Wu et al., 2010Wu, C.L.; Zhang, Q.Q.; Dong, B.X.; Li, S.F.; Zhang, C.Q. 2010. Analysis of genetic structure and genetic relationships of partial maize inbred lines in China. Acta Agronomica Sinica 36: 1820-1831.; Song et al., 2014Song, Y.; Fan, L.; Chen, H.; Zhang, M.; Ma, Q.; Zhang, S.; Wu, J. 2014. Identifying genetic diversity and a preliminary core collection of Pyrus pyrifolia cultivars by a genome-wide set of SSR markers. Scientia Horticulturae 167: 5-16.).
For the proper use of the genetic resources of a germplasm bank, it is essential to know the genetic diversity among the available accessions which allows for the appropriate selection of genotypes according to the objectives of the breeding program (Perseguini et al., 2011Perseguini, J.M.K.C.; Chiorato, A.F.; Zucchi, M.I.; Colombo, C.A.; Carbonell, S.A.M.; Mondego, J.M.C.; Gazaffi, R.; Garcia, A.A.F.; Campos, T.; Souza, A.P.; Rubiano, L.B. 2011. Genetic diversity in cultivated carioca common bean based on molecular marker analysis. Genetics and Molecular Biology 34: 88-102.). The low levels of genetic diversity and the great deficit of heterozygotes found for the lima bean accessions may be explained by the reproductive characteristics of this crop and by the conservation of these accessions in the form of a germplasm bank. According to Penha et al. (2017)Penha, J.S.; Lopes, A.C.A.; Gomes, R.L.F.; Pinheiro, J.B.; Assunção-Filho, J.R.; Silvestres, E.A.; Viana, J.P.G.; Castilho-Martinez, J. 2017. Estimation of natural outcrossing rate and genetic diversity in Lima bean (Phaseolus lunatus L. var. lunatus) from Brazil using SSR markers: implications for conservation and breeding. Genetic Resources and Crop Evolution 64: 1355-1364.P. lunatus had a mixed system with a predominance of self-fertilization, with only 38 % of natural crossing. High frequency of selfing results in the increasing of homozygosity (Hartl and Clarck, 2007Hartl, D.L.; Clarck, A.G. 2007. Principles of Population Genetics. Sinauer, Sunderland, MA, USA.), which may have possibly contributed to increased levels of inbreeding in lima bean accessions. In addition, the limited amount of genetic diversity conserved due to sampling strategies is an intrinsic difficulty of establishing a germplasm collection (Thormann et al., 2006Thormann, I.; Dullo, M.; Engels, J. 2006. Techniques for ex situ plant conservation. p. 7-36. In: Henry, R.J., ed. Plant conservation genetics. Food Products Press, Binghamton, NY, USA). Therefore, it may be expected that low levels of genetic diversity are represented in collections of naturally low-diversity species. Nevertheless, all of our results demonstrated that the methodology used to establish the nuclear collection was appropriate because it maintained the genetic diversity present in the base collection.
Understanding the genetics of lima bean accessions conserved in Brazilian germplasm banks is extremely important for the use of this crop in breeding programs. Despite their importance, improved lima bean varieties have not yet been produced in Brazil, and there is still little information about their breeding systems and genetic diversity (Penha et al., 2017Penha, J.S.; Lopes, A.C.A.; Gomes, R.L.F.; Pinheiro, J.B.; Assunção-Filho, J.R.; Silvestres, E.A.; Viana, J.P.G.; Castilho-Martinez, J. 2017. Estimation of natural outcrossing rate and genetic diversity in Lima bean (Phaseolus lunatus L. var. lunatus) from Brazil using SSR markers: implications for conservation and breeding. Genetic Resources and Crop Evolution 64: 1355-1364.). In this context, this study is of extreme importance, since it is one of the pioneers in proposing the development of a core collection for this species. It is likely that the development of core collections for lima beans will significantly reduce the task of germplasm bank curators since they will be able to multiply and maintain fewer accessions in the medium term as active collections to meet the needs of breeding programs (Bryan, 2006Bryan, G. 2006. Techniques for ex situ plant conservation. p. 131-148. In: Henry, R.J., ed. Plant conservation genetics. Food Products Press Binghamton, NY, USA.). In this way, this core collection of lima bean germplasms may be managed more efficiently in terms of time and cost.
Conclusion
Overall, our study indicates a high deficit of heterozygotes among the 153 accessions of the BGP-UFPI lima bean, probably due to the reproductive biology of the crop. The maintenance of these accessions as discrete units within a larger germplasm collection also contributes to the absence of recombination among them, which difficult the reduction of heterozygote deficit observed for this collection. Even though the levels of genetic diversity were low, cluster and Bayesian analysis evidenced the existence of very divergent accessions. If these accessions show contrasting desirable agronomic characteristics it may be possible to use them in the creation of desired heterotic groups. Based on genetic data the proposed core collection represents very well the variability currently stored in the BGP-UFPI lima bean. It is important to note that the 34 selected accessions represent an efficient approach in the development of a proper core collection of P. lunatus, since it allows for the conservation of maximum genetic diversity in a limited number of accessions, thus providing more opportunities to capture the desirable variation involving complex traits of agronomic interest. The proposed core collection will facilitate further genetic characterization, and the study of the morphological variation in the characteristics of agronomic interest of lima beans as well as being important for conducting field evaluations and genomic association studies. Nonetheless, the proposed nuclear collection should be periodically updated, including additional lima bean germplasms in the base collection; moreover, it should be examined in detail for resistance to diseases, such as anthracnose and drought. This information can then be used together with the genetic diversity data to develop improved varieties of lima beans that present superior performance and a broad genetic base.
Acknowledgments
The authors thank the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for funding the research with Phaseolus lunatus L., the Coordenação de Aperfeiçoamento Pessoal de Nível Superior (CAPES) for the granting of the scholarships, and the Laboratório de Diversidade Genética e Melhoramento at the Escola Superior de Agricultura Luiz de Queiroz (ESALQ), for the accomplishment of the study, and AlessandroAlves Pereira thanks CAPES for a post-doctoral scholarship.
References
- Agrama, H.A.; Yan, W.G.; Lee, F.; Fjellstrom, R.; Chen, M-H.; Jia, M.; McClung, A. 2009. Genetic assessment of a minicore subset developed from the USDA rice genebank. Crop Science 49: 1336-1346.
- Baudet, J.C. 1977. The taxonomic status of the cultivated types of Lima bean (Phaseolus lunatus L.). Trop Grain Legume 7: 29-30.
- Botstein, D.; White R.L.; Skolnick, M.; Davis, R.W. 1980. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. American Journal of Human Genetics 32: 314-331.
- Brown, A.H.D. 1989. Core collections: a practical approach to genetic resources management. Genome 31: 818-824.
- Brown, A.H.D.; Spillane, C. 1999. Implementing core collections: principles, procedures, progress, problems and promise. p. 1-9. In: Johnson, R.C.; Hodgkin, T., eds. Core collections for today and tomorrow. IPGRI, Rome, Italy.
- Bryan, G. 2006. Techniques for ex situ plant conservation. p. 131-148. In: Henry, R.J., ed. Plant conservation genetics. Food Products Press Binghamton, NY, USA.
- Bueno, L.C.S.; Mendes, A.N.G.; Carvalho, S.P. 2001. Genetic Plant Breeding: Principles and Procedures = Melhoramento Genético de Plantas: Princípios e Procedimentos. UFLA, Lavras, MG, Brazil (in Portuguese).
- Choudhury, D.R.; Singh, N.; Singh, A.K.; Kumar, S.K.; Srinivasan, K.; Tyagi, R.K.; Ahmand, A.; Singh, N.K.; Singh, R. 2014. Analysis of genetic diversity and population structure of rice germplasm from north-eastern region of India and development of a core germplasm set. Plos One 9: e113094.
- Cristo-Araújo, M.; Rodrigues, M.; Astolfi-Filho, S.; Clement, C.R. 2015. Peach palm core collection in Brazilian Amazonia. Crop Breeding and Applied Biotechnology 15: 18-25.
- Doyle, J.J.; Doyle, J.L. 1990. Isolation of plant DNA from fresh tissue. Focus 1: 13-15.
- Earl, D.A.; von Holdt, B.M. 2012. Structure harvester: a website and program for visualizing structure output and implementing the Evanno method. Conservation Genetics Resources 4: 359-361.
- Emanuelli, F.; Lorenzi, S.; Grzeskowiak, L.; Catalano, V.; Stefanini, M.; Troggio, M.; Myles, S.; Martinez-Zapater, J.M.; Zyprian, E.; Moreira, F.M.; Grando, M.S. 2013. Genetic diversity and population structure assessed by SSR and SNP markers in a large germplasm collection of grape. BMC Plant Biology 13: 39. DOI:10.1186/1471-2229-13-39
- Evanno, G.; Regnaut, S.; Goudet, J. 2005. Detecting the number of clusters of individuals using the software structure: a simulation study. Molecular Ecology 14: 2611-2620.
- Frankel, O.H. 1984. Genetic Perspectives of Germplasm Conservation. Cambridge University Press, Cambridge, UK.
- Gaitán-Solís, E.; Duque, M.C.; Edwards, K.J.; Tohme, J. 2002. Microsatellite repeats in common bean (Phaseolus vulgaris L.): isolation, characterization, and cross-species amplification in Phaseolus ssp. Crop Science 42: 2128-2136.
- Gireesh, C.; Husain, S.M.; Shivakumar.; Satpute, G.K.; Kumawat, G.; Arya, M.; Agarwal, D.K.; Bhatia, V.S. 2017. Integrating principal component score strategy with power core method for development of core collection in Indian soybean germplasm. Plant Genetic Resources 15: 230-238.
- Göl, Ş.; Doğanlar, S.; Frary, A. 2017. Relationship between geographical origin, seed size and genetic diversity in faba bean (Vicia faba L.) as revealed by SSR markers. Molecular Genetics and Genomics 5: 991-999.
- Haouane, H.; Bakkali, A.E.; Moukhli, A.; Tollon, C.; Santoni, S.; Oukabli, A.; Modafar, C.E.; Khadari, B. 2011. Genetic structure and core collection of the world olive germplasm bank of Marrakech: towards the optimized management and use of Mediterranean olive genetic resources. Genetica 139: 1083-1094.
- Hartl, D.L.; Clarck, A.G. 2007. Principles of Population Genetics. Sinauer, Sunderland, MA, USA.
- Heiden, G.; Barbieri, R.L.; Couto, M.E.O.; Medeiros, A.R.M.; Sinigaglia, C. 2007. Peppers and chilies from southern Brazil: Creole varieties maintained by Embrapa temperate climate = Pimentas e pimentões do sul do Brasil: variedades crioulas mantidas pela Embrapa clima temperado. Revista Brasileira de Agroecologia 2: 1 (in Portuguese).
- Hu. J.; Wang, P.; Su, Y.; Wang, R.; Li, Q.; Sun, K. 2015. Microsatellite diversity, population structure, and core collection formation in melon germplasm. Plant Molecular Biology Reporter 33: 439-447.
- Kaga, A.; Shimizu, T.; Watanabe, S.; Tsubokura, Y.; Katayose, Y.; Harada, K.; Vaughan, P.A.; Tomookan, N. 2012. Evaluation of soybean germplasm conserved in NIAS genebank and development of mini core collections. Breeding Science 61: 566-592.
- Kim, K.W.; Chung, H.K.; Cho, G.T.; Ma, H.; Chandrabalan, D. 2007. PowerCore: a program applying the advanced M strategy with a heuristic search for establishing core sets. Bioinformatics 23: 2155-2162.
- Ladizinsky, G. 1998. Plant Evolution under Domestication. Springer, Dordrecht, The Netherlands.
- Langella, O. 2002. Populations v.1.2.32. Available at: http://bioinformatics.org/~tryphon/populations/index.html#pubdas [Accessed Aug 31, 2017]
» http://bioinformatics.org/~tryphon/populations/index.html#pubdas - Lee, H.Y.; Ro, N.Y.; Jeong, H.J.; Kwon, J.K.; Jo, J.; Há, Y.; Jung, A.; Han, J.W.; Venkatesh, J.; Kang, B.C. 2016. Genetic diversity and population structure analysis to construct a core collection from a large Capsicum germplasm. BMC Genetics 17: 142.
- Liu, X.B.; Li, J.; Yang, Z.L . 2017.Genetic diversity and structure of core collection of winter mushroom (Flammulina velutipes) developed by genomic SSR markers. Hereditas 155: 3.
- Mongkolporn, O.; Hanyong, S.; Chunwongse, J.; Wasee, S. 2015. Establishment of a core collection of chilli germplasm using microsatellite analysis. Plant Genetic Resources 13: 104-110.
- Odong, T.L.; Jansen, J.; van Eeuwijk, F.A.; van Hintum, T.J.L. 2013. Quality of core collections for effective utilisation of genetic resources review, discussion and interpretation. Theoretical and Applied Genetics 126: 289-305.
- Oliveira-Silva, R.N.; Burle, M.L.; Pádua, J.G.; Lopes, A.C.A.; Gomes, R.L.F.; Martínez-Castillo, J. 2017. Phenotypic diversity in lima bean landraces cultivated in Brazil, using the Ward-MLM strategy. Chilean Journal of Agricultural Research 77: 35.
- Peakall, R.; Smouse, P.E. 2012. GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28: 2537-2539.
- Penha, J.S.; Lopes, A.C.A.; Gomes, R.L.F.; Pinheiro, J.B.; Assunção-Filho, J.R.; Silvestres, E.A.; Viana, J.P.G.; Castilho-Martinez, J. 2017. Estimation of natural outcrossing rate and genetic diversity in Lima bean (Phaseolus lunatus L. var. lunatus) from Brazil using SSR markers: implications for conservation and breeding. Genetic Resources and Crop Evolution 64: 1355-1364.
- Perseguini, J.M.K.C.; Chiorato, A.F.; Zucchi, M.I.; Colombo, C.A.; Carbonell, S.A.M.; Mondego, J.M.C.; Gazaffi, R.; Garcia, A.A.F.; Campos, T.; Souza, A.P.; Rubiano, L.B. 2011. Genetic diversity in cultivated carioca common bean based on molecular marker analysis. Genetics and Molecular Biology 34: 88-102.
- Pritchard, J.K.; Stephens, M.; Donnelly, P. 2000. Inference of population structure using multilocus genotype data. Genetics 155: 945-959.
- Rogers, J.S. 1972. Measures of genetic similarity and genetic distance. p. 145-153. In: Studies in genetics VII. University of Texas, Austin, TX, USA.
- Song, Y.; Fan, L.; Chen, H.; Zhang, M.; Ma, Q.; Zhang, S.; Wu, J. 2014. Identifying genetic diversity and a preliminary core collection of Pyrus pyrifolia cultivars by a genome-wide set of SSR markers. Scientia Horticulturae 167: 5-16.
- Tamura, K.; Stecher, G.; Peterson, D.; Fillipski, A.; Kumar, S. 2013. MEGA6 Molecular evolutionary genetics analyses version 6.0. Molecular Biology and Evolution 30: 2725-2729.
- Thormann, I.; Dullo, M.; Engels, J. 2006. Techniques for ex situ plant conservation. p. 7-36. In: Henry, R.J., ed. Plant conservation genetics. Food Products Press, Binghamton, NY, USA
- Tiwari, K.K.; Singh, A.; Pattnaik, S.; Sandhu, M.; Kaur, S.; Jain, S.; Tiwari, S.; Mehrotra, S.; Anumalla, M.; Samal, R.; Bhardwaj, J.; Dubey, N.; Sahu, V.; Kharshing, G.A.; Zeliang, P.K.; Sreenivasan, K.; Kumar, P.; Parida, S.K.; Mithra, S.V.A.; Rai, V.; Tyagi, W.; Agrawal, P.K.; Rao, A.R.; Pattanayak, A.; Chandel, G.; Singh, A.K.; Bisht I.S.; Bhat, K.V.; Rao, G.J.N.; Khurana, J.P.; Singh, N.K.; Mohapatra, T. 2015. Identification of a diverse mini-core panel of Indian rice germplasm based on genotyping using microsatellite markers. Plant Breeding 134: 164-171.
- Wang, Y.K.; Wu, G.L.; Li, D.K.; Ji, A.Q.; Sui, C.L.; Zhao, A.L. 2010. Advances in core collection of fruit germplasm. Journal of Plant Genetic Resources 11: 380-385.
- Wu, C.L.; Zhang, Q.Q.; Dong, B.X.; Li, S.F.; Zhang, C.Q. 2010. Analysis of genetic structure and genetic relationships of partial maize inbred lines in China. Acta Agronomica Sinica 36: 1820-1831.
- Xu, Y.; Chen, C.; Ji, D.; Xu, K.; Xie, X.; Xie, C. 2016. Developing a core collection of Pyropia haitanensis using simple sequence repeat markers. Aquaculture 452: 351-356.
Edited by
Publication Dates
-
Publication in this collection
02 Sept 2019 -
Date of issue
2020
History
-
Received
26 Apr 2018 -
Accepted
13 Aug 2018