Abstract
INTRODUCTION: During the first pandemic wave of the influenza A H1N1 2009 virus, morbidity was particularly high in Brazil. Hospitalizations resulting from severe respiratory disease due to suspected influenza-like illness created an opportunity to identify other respiratory viruses causing lower respiratory infections. OBJECTIVE: The purpose of this study was to assess viral etiologies among samples collected during the first pandemic wave of H1N1 2009 from hospitalized patients with suspected cases in a Brazilian Sentinel Hospital. PATIENTS AND METHODS: Viral etiologies were investigated in samples from 98 children and 61 adults with fever, cough and dyspnea who were admitted to São Paulo Sentinel Hospital with suspected H1N1 infection. RESULTS: From August to November 2009, in 19.5% (31/159) of the samples 2009 H1N1 virus was detected with 23% (14/61) in adults (median age 25 years, range: 14-55 years) and 18.4% (17/92) in children (median age 5 years, range: 4 months - 11 years). Among the negative samples, a wide range of causative etiologic agents was identified. Human rhinovirus was the most frequent virus (23.91%) in children and human metapneumovirus (11.48%) was the second most frequent in adults, following 2009 H1N1 virus (22.95%). CONCLUSION: These data highlight the need to diagnose other viral infections that can co-circulate with influenza and may have been neglected by physicians as causes of severe respiratory diseases.
influenza virus; influenza A H1N1 2009 virus; rhinovirus; human metapneumovirus; human respiratory syncytial virus; respiratory viruses
ORIGINAL ARTICLE
Respiratory virus infections among hospitalized patients with suspected influenza A H1N1 2009 virus during the first pandemic wave in Brazil
Aripuanã Sakurada Aranha WatanabeI; Emerson CarraroII; Luciana MoreiraIII; Clarice CamargoIV; Juliana SinoharaIII; Diane PuerariIII; Sandra GuaturaV; Celso GranatoVI; Nancy BelleiVI
IMSc, PhD student, Clinical Virology Laboratory, Infectious Diseases Unit, Medicine Department, Universidade Federal de São Paulo (Unifesp), São Paulo, Brazil
IIPhD; Laboratory Assistant, Clinical Virology Laboratory, Infectious Diseases Unit, Medicine Department, Unifesp, São Paulo, Brazil
IIIMSc student; Laboratory student, Clinical Virology Laboratory, Infectious Diseases Unit, Medicine Department, Unifesp, São Paulo, Brazil
IVPhD; Laboratory student, Clinical Virology Laboratory, Infectious Diseases Unit, Medicine Department, Unifesp, São Paulo, Brazil
VMSc; Laboratory supervisor, Unifesp, São Paulo, Brazil
VIPhD, Professor; Laboratory Chief, Unifesp, São Paulo, Brazil
Correspondence to Correspondence to: Aripuanã Sakurada Aranha Watanabe Rua Pedro de Toledo, 781, andar 15 - Vila Clementino São Paulo - SP, Brazil 04039-032 almasurfe@yahoo.com.br
ABSTRACT
INTRODUCTION: During the first pandemic wave of the influenza A H1N1 2009 virus, morbidity was particularly high in Brazil. Hospitalizations resulting from severe respiratory disease due to suspected influenza-like illness created an opportunity to identify other respiratory viruses causing lower respiratory infections.
OBJECTIVE: The purpose of this study was to assess viral etiologies among samples collected during the first pandemic wave of H1N1 2009 from hospitalized patients with suspected cases in a Brazilian Sentinel Hospital.
PATIENTS AND METHODS: Viral etiologies were investigated in samples from 98 children and 61 adults with fever, cough and dyspnea who were admitted to São Paulo Sentinel Hospital with suspected H1N1 infection.
RESULTS: From August to November 2009, in 19.5% (31/159) of the samples 2009 H1N1 virus was detected with 23% (14/61) in adults (median age 25 years, range: 14-55 years) and 18.4% (17/92) in children (median age 5 years, range: 4 months - 11 years). Among the negative samples, a wide range of causative etiologic agents was identified. Human rhinovirus was the most frequent virus (23.91%) in children and human metapneumovirus (11.48%) was the second most frequent in adults, following 2009 H1N1 virus (22.95%).
CONCLUSION: These data highlight the need to diagnose other viral infections that can co-circulate with influenza and may have been neglected by physicians as causes of severe respiratory diseases.
Keywords: influenza virus; influenza A H1N1 2009 virus; rhinovirus; human metapneumovirus; human respiratory syncytial virus; respiratory viruses.
INTRODUCTION
In April 2009, a novel variant of influenza virus (2009) H1N1 of swine origin appeared and spread rapidly worldwide. On June 11, the World Health Organization (WHO) raised its pandemic alert to the highest level, phase 6.1 The morbidity and mortality of 2009 H1N1 virus were particularly high in Brazil during the first pandemic wave on the southern hemisphere.2 The pandemic stage resulted in increased surveillance and sampling, school closings and other measures for pandemic control.2,3 In addition, hospitalized cases4 of 2009 H1N1 virus resulted in increased demand for health care. Severe respiratory disease hospitalizations due to suspected influenza-like illness created an opportunity to identify other respiratory viruses causing lower respiratory infections.
OBJECTIVE
The purpose of this study was to assess viral etiologies among the samples collected from hospitalized patients with suspected influenza during the first pandemic wave of influenza A H1N1 2009 virus in a Brazilian Sentinel Hospital.
PATIENTS AND METHODS
Sample collection
From August 19 to November 30, 2009 (epidemiological week 34 to 43), a total of 159 nasal/throat swab samples (one sample per patient) were collected from patients at São Paulo Hospital, São Paulo, Brazil. Patient inclusion criteria followed the National Program Protocol: clinical suspicion of potentially severe influenza A H1N1 2009 virus infection defined as fever, cough and dyspnea.
From August to November 2009, the peak of the first pandemic wave in São Paulo, a single sample from each of 159 hospitalized patients (98 children and 61 adults) was tested in the clinical virology laboratory. This study was approved by the Ethics Committee of Universidade Federal de São Paulo.
Viral total-RNA extraction
Viral RNA and DNA were extracted from 200 µL of each clinical specimen using the RTP DNA/RNA Virus MiniKit (Invitek, Germany) according to the manufacturer's instructions.
Molecular diagnostics
Detection of influenza A seasonal (IAV) and H1N1 2009 viruses was performed according to the Real Time protocol published by the CDC on April 28, 2009.5 After excluding influenza A infection (H1N1 2009 and seasonal influenza), the negative samples were tested for the presence of the following viruses as described previously: seasonal influenza virus B (IBV) with primers targeting the NS gene,6 human rhinovirus (HRV) with primers targeting the 5'NCR and VP4/VP2 genes,7 human metapneumovirus (hMPV) with primers targeting the fusion protein gene,8 adenovirus (AdV) with primers targeting the hexon gene9 and human respiratory syncytial virus (HRSV) with primers targeting the fusion protein gene.10 The eluted RNA was transcribed into cDNA by moloney murine reverse transcriptase (MMLV-RT; Invitrogen, Carlsbad, CA) using random primers (Invitrogen, Carlsbad, CA) for 1 h at 37ºC. The MMLV-RT was subsequently denatured at 70ºC. The cDNA was amplified with virus-specific oligonucleotide primers and Platinum Taq DNA Polymerase (Invitrogen, Carlsbad, CA). All PCR parameters are shown in Table 1.
Seasonal IAV typing
Positive samples identified utilizing the Centers for Disease Control and Prevention (CDC) protocol for seasonal IAV were also tested using an in-house RT-PCR reaction followed by restriction fragment length polymorphism (RFLP) analysis. This method was performed as described previously11 and discriminates seasonal IAV H3N2 from seasonal IAV H1N1.
Statistical analysis
We evaluated viral infections in adult and child populations using the Chi-squared test (SPSS v11.5).
RESULTS
We studied 159 hospitalized patients: 61 adults, median age 32 years (13 - 91 years), and 98 children, median age 2 years (2 days - 11 years); 97/159 (61%) tested positive for at least one virus, and 6/159 (3.77%) were coinfected with multiple viruses. Overall, 64.78% (103/159) of the viral etiologies were characterized. Patient samples were examined according to age group (Figure 1). Children (< 12 years old) had a significantly higher frequency (67/92; 73%) of identified etiology when compared to adults (30/61; 49%) (p = 0.0021).
Influenza: 31/159 (19.5%) samples tested were positive for influenza A H1N1 2009 virus, 14/61 (23%) adults and 17/98 (17.3%) children. Only 10/159 (6.29%) samples were positive for seasonal IAV. None of the samples studied were positive for IBV. The median age of patients with influenza A H1N1 2009 virus was 25 years (14 - 55 years) in adults and 5 years (4 months - 11 years) in children. Among the 10 seasonal IAV positive samples (median age 7 years old, range: 6 months - 51 years), 2 (20%) were characterized as H1N1, and the remaining 8 (80%) were characterized as H3N2.
Other respiratory viruses were detected in 62/118 (52.54%) samples that were negative for influenza A. Of those, 27 (22.88%) tested positive for HRV, 24 (20.34%) for hMPV, 3 (2.54%) for AdV and 2 (1.69%) for HRSV. HRV was the most frequently detected virus in children (23.91%), and hMPV (11.48%) was the second most frequently detected virus in adults, following 2009 H1N1 virus. Six specimens (6/118 - 5.08%) presented with coinfections with 2 viruses (4 HRV/hMPV, 1 Adv/HRSV and 1 AdV/HRV). The median age of coinfected patients was 1.79 years (range: 1 month - 7 years).
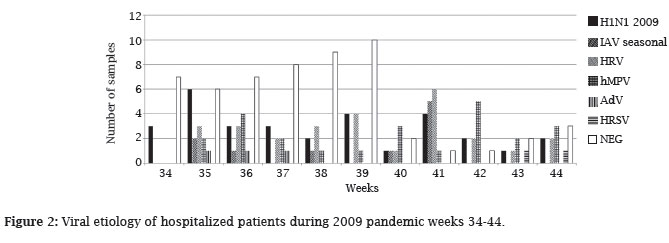
The etiology of viral diagnoses over the course of the study (as a function of time in "weeks") is shown in Figure 2. H1N1 virus was the most frequently detected infection (30% positivity) during the peak of the pandemic in São Paulo City (August 2009: 34 and 35 weeks; Figure 1), coincident with the beginning of the school calendar. HMPV was also observed at this time, and HRV was observed frequently throughout the study period.
DISCUSSION
The emergence of a new pandemic influenza A lineage has led to an increase in surveillance of respiratory infections as well as hospitalizations and detection of other viruses.3
Here, we characterized the viral etiologies in two thirds of the patient samples. Higher detection rates could be achieved if others etiologies were investigated, such as atypical bacterial pneumonias.
In agreement with other reports, our study demonstrated that children had a higher frequency of identifiable etiology when compared to adults, and this was likely influenced by the high viral load present in these patients.12 Additionally, it is likely that the national protocol guidance13 calling for hospitalization of all symptomatic children less than 2 years old with an acute respiratory infection influenced these results.
The HRV was frequently associated with childhood hospitalization in our study. Wheezing is a frequent clinical presentation in the emergency room that is possibly related to HRV infection,14 but we report here that it was misdiagnosed as H1N1 virus infection during the pandemic wave.
In Thailand15 and Egypt,16 hMPV has been reported to be a community-acquired infection in patients presenting with pneumonia. However, the elevated rate of infection among non-elderly hospitalized adults was previously unreported in Brazil. This points to the severity of respiratory disease related to this virus, and indicates that it may be responsible for a greater portion of disease than previously suspected by physicians.15,17
We observed a low rate of coinfection (3.77%) that appeared exclusively in young children (median age 1.79 years old), in agreement with others reports.15
H1N1 2009 virus infected children were older (5 years) than H1N1 negative children (median age 1 year old), in accordance with other studies.2,14,18 Pandemic influenza occurred in young adults (25 years old), as previously described.2,14,18 These data will contribute to the growing body of knowledge regarding influenza clinical management and immunization programs.
H1N1 2009 was the most prevalent virus worldwide during our expected influenza season, from June to September,19,20 and this delayed seasonal H3N2 circulation until October and early November, extending the influenza season.
Other viral etiologies accounting for increased health care demands and hospitalizations during the first wave of H1N1 2009 virus were underdiagnosed by physicians. It is likely that these account for an overestimation in influenza infection rates and may have resulted in inappropriate antibiotic prescriptions in unconfirmed cases.2,14
This study highlights the relevance and impact of both novel and established respiratory viruses that are responsible for lower respiratory tract infections in adults and suggests the need to maintain surveillance during non-pandemic periods.
ACKNOWLEDGMENTS
The authors acknowledge the financial support of Fundação de Amparo a Pesquisa do Estado de São Paulo (FAPESP Project number: 07/01166-9), and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES).
Submitted on: 09/20/2010
Approved on: 01/30/2011
Financial Support: The authors acknowledge the financial support of Fundação de Amparo a Pesquisa do Estado de São Paulo (FAPESP Project number: 07/01166-9), and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES).
We declare no conflict of interest.
- 1. World Health Organization. Pandemic (H1N1) 2009 - update 68. http://www.who.int/csr/don/20091002/en/index.html
- 2. Oliveira W, Carmo E, Penna G et al. Pandemic H1N1 influenza in Brazil: analysis of the first 34,506 notified cases of influenza-like illness with severe acute respiratory infection (SARI). Euro Surveill. 2009; 14(42),pii:19362.
- 3. Ghedin E, Wentworth D, Halpin RA et al. Unseasonal Transmission of H3N2 Influenza A Virus during the Swine-Origin H1N1 Pandemic. J Virol. 2010; 84(11):5715-8.
- 4. Cox A, Guglielmetti P, Coulombier D. Assessing the impact of the 2009 H1N1 influenza pandemic on reporting of other threats through the Early Warning and Response System. Euro Surveill. 2009; 14(45).pii:19397.
- 5. CDC protocol of real time RT-PCR for influenza A (H1N1). www.who.int/csr/resources/publications/swineflu/realtimeptpcr/en/index.html
- 6. Carraro E, Neto DF, Benfica D, SittaPerosa AH, Granato CF, Bellei NC. Applications of a duplex reverse transcription polymerase chain reaction and direct immunofluorescence assay in comparison with virus isolation for detection of influenza A and B. Diagn Microbiol Infect Dis. 2007; 57(1):53-7.
- 7. Savolainen C, Mulders MN, Hovi T. Phylogenetic analysis of rhinovirus isolates collected during successive epidemic seasons. Virus Res. 2002; 85(1):41-6.
- 8. Falsey AR, Erdman D, Anderson LJ, Walsh EE. Human metapneumovirus infections in young and elderly adults. J Infect Dis. 2003; 187(5):785-90.
- 9. Casas I, Avellon A, Mosquera M et al. Molecular identification of adenoviruses in clinical samples by analyzing a partial hexon genomic region. J Clin Microbiol. 2005; 43(12):6176-82.
- 10. Erdman DD, Weinberg GA, Edwards KM et al. GeneScan reverse transcription-PCR assay for detection of six common respiratory viruses in young children hospitalized with acute respiratory illness. J Clin Microbiol. 2003; 41(9):4298-303.
- 11. Carraro E, Watanabe AS, Neto DF, Granato CF, Bellei NC. Influenza detection and subtyping by reverse transcriptase polymerase chain reaction-restriction fragment length polymorphism for laboratory surveillance in Brazil. Diagn Microbiol Infect Dis. 2008; 60(4):445-7.
- 12. Gerna G, Piralla A, Rovida F et al. Correlation of rhinovirus load in the respiratory tract and clinical symptoms in hospitalized immunocompetent and immunocompromised patients. J Med Virol. 2009; 81(8):1498-507.
- 13. National protocol guidance for Influenza A H1N1 2009. portal.saude.gov.br/portal/arquivos/pdf/protocolo_de_manejo_clinico_05_08_2009.pdf
- 14. Follin P, Lindqvist A, Nyström K, Lindh M. A variety of respiratory viruses found in symptomatic travellers returning from countries with ongoing spread of the new influenza A(H1N1)v virus strain. Euro Surveill. 2009 Jun 18; 14(24).pii:19242.
- 15. Olsen SJ, Thamthitiwat S, Chantra S et al. Incidence of respiratory pathogens in persons hospitalized with pneumonia in two provinces in Thailand. Epidemiol Infect. 2010; 138(12):1811-22.
- 16. El Sayed Zaki M, Raafat D, El-Metaal AA, Ismail M. Study of human metapneumovirus-associated lower respiratory tract infections in Egyptian adults. Microbiol Immunol. 2009; 53(11):603-8.
- 17. Wolf DG, Greenberg D, Shemer-Avni Y, Givon-Lavi N, Bar-Ziv J, Dagan R. Association of human metapneumovirus with radiologically diagnosed community-acquired alveolar pneumonia in young children.J Pediatr. 2010; 156(1):115-20.
- 18. Schout D, Hajjar LA, Galas FR et al. Epidemiology of human infection with the novel virus influenza A (H1N1) in the Hospital das Clínicas, São Paulo, Brazil-June-September 2009. Clinics (São Paulo) 2009; 64(10):1025-30.
- 19. Thomazelli LM, Vieira S, Leal AL et al. Surveillance of eight respiratory viruses in clinical samples of pediatric patients in southeast Brazil. J Pediatr (Rio J) 2007; 83(5):422-8.
- 20. Bellei N, Carraro E, Perosa A, Granato C. Patterns of influenza infections among different risk groups in Brazil. Braz J Infect Dis. 2007; 11(4):399-402.
Correspondence to:
Publication Dates
-
Publication in this collection
06 June 2011 -
Date of issue
June 2011
History
-
Received
20 Sept 2010 -
Accepted
30 Jan 2011