Abstract
Microsatellite DNA markers have been used to assess genetic diversity and to study ecological behavioral characteristics in animals. Although these markers are powerful tools, their development is labor intensive and costly. Thus, before new markers are developed it is important to prospect the use of markers from related species. In the present study we investigated the possibility of using microsatellite markers developed for Alligator mississipiensis and Caiman latirostris in South American crocodilians. Our results demonstrate the use of microsatellite markers for Paleosuchus palpebrosus, Caiman crocodilus and Caiman yacare.
SSR; STR; primers; crocodilians; Alligatorinae
SHORT COMMUNICATION
Cross-species microsatellite amplification in South American Caimans (Caiman spp and Paleosuchus palpebrosus)
Rodrigo Barban ZucolotoI, II, IV; Priscilla Marqui Schimidt VillelaII; Luciano Martins VerdadeIII; Luiz Lehmann CoutinhoII
IUniversidade de São Paulo, Centro de Energia Nuclear na Agricultura, Piracicaba, SP, Brazil
IIUniversidade de São Paulo, Escola Superior de Agricultura "Luiz de Queiroz", Laboratório de Biotecnologia, Piracicaba, SP, Brazil
IIIUniversidade de São Paulo, Escola Superior de Agricultura "Luiz de Queiroz", Laboratório de Ecologia Animal, Piracicaba, SP, Brazil
IVCurrent address: Universidade Federal da Bahia, Departamento de Biologia Geral, Laboratório de Biologia Celular e Molecular, Salvador, Ba, Brazil
Send correspondence to Send correspondence to Luiz Lehmann Coutinho Universidade de São Paulo, Escola Superior de Agricultura "Luiz de Queiroz", Laboratório de Biotecnologia, Caixa Postal 9 13418-900 Piracicaba, São Paulo, Brazil E-mail: llcoutin@carpa.ciagri.usp.br
ABSTRACT
Microsatellite DNA markers have been used to assess genetic diversity and to study ecological behavioral characteristics in animals. Although these markers are powerful tools, their development is labor intensive and costly. Thus, before new markers are developed it is important to prospect the use of markers from related species. In the present study we investigated the possibility of using microsatellite markers developed for Alligator mississipiensis and Caiman latirostris in South American crocodilians. Our results demonstrate the use of microsatellite markers for Paleosuchus palpebrosus, Caiman crocodilus and Caiman yacare.
Key words: SSR, STR, primers, crocodilians, Alligatorinae.
Microsatellite DNA markers are simple sequence repeats (Tautz et al., 1986) distributed along the genome (Litt and Luty, 1989) that have been used to assess genetic diversity and to study ecological behavioral characteristics such as mating system and dispersal pattern in reptiles and amphibians (Avise, 1994; Forstner and Forstner, 2002), including the timber rattlesnake Crotalus horridus (Villareal et al., 1995), Alligator mississipiensis (Glenn et al., 1996; Glenn et al., 1998; Davis et al., 2001a), and Crocodylus spp. (Dever et al., 2001; FitzSimmons et al., 2001; Verdade et al., 2002).
Microsatellite markers are powerful research tools but their development is labor intensive and costly. Consequently, researchers have tried to use microsatellite markers developed for one species in another (Moore et al., 1991). Microsatellite markers developed for Alligator mississipiensis have been successfully used in closely related Alligatorinae species (Glenn et al., 1998); however, transference is more effective at the family or subfamily level (Glenn et al., 1998; Zucoloto, 1998).
All South American crocodilians (Caiman spp., Melanosuschus niger and Paleosuchus spp.) belong to the Alligatorinae subfamily (King and Burke, 1989). To date the only Alligatorinae species with specific microsatellite markers currently developed are Alligator mississipiensis and Caiman latirostris (Glenn et al., 1998; Zucoloto, 2002). Thus, transference of microsatellite markers to other Alligatorinae species could help conservation programs, genetic diversity studies as well as mating behavior and ecological studies
The present study tested the ability of microsatellite markers previously developed for Alligator mississipiensis (Glenn et al., 1998) and Caiman latirostris (Zucoloto et al., 2002) to amplify orthologous loci in the related South American Alligatorinae species Caiman crocodilus, Caiman yacare and Paleosuchus palpebrosus.
The blood samples used in this study were from the Brazilian crocodilians P. palpebrosus, C. yacare and C. crocodiles. Samples were obtained from crocodilians maintained at the Department of Zoology, São Paulo State University, Rio Claro, São Paulo (SP), Brazil (UNESP, Rio Claro, SP) and were stored at the Biotechnology laboratory, ESALQ, University of São Paulo, Piracicaba, SP, Brazil.
Blood was collected from three C. crocodilus specimens (Cc1, Cc2 and Cc3), three C. yacare specimens (Cy1, Cy2 and Cy3) and two P. palpebrosus specimens (Pp1 and Pp2) by puncturing the dorsal branch of the superior cava vein, which runs along the interior of the vertebral column of large reptiles (Olson, 1975). After collection, blood was mixed with lysis buffer (100 mM Tris-HCl, pH 8.0; 100 mM EDTA, pH 8.0; 0.5% SDS (w/v); 10 mM NaCl) (Hoelzel, 1992). The DNA from these samples was then purified by CTAB and chloroform extraction followed by isopropyl alcohol precipitation (Sambrook et al., 1989).
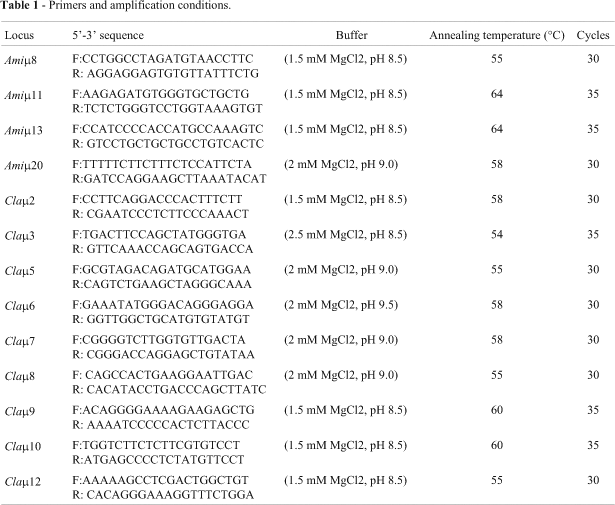
The Amiµ8, Amiµ11, Amiµ13 and Amiµ20 markers developed for Alligator mississipiensis (Glenn et al., 1998) and successfully used in Caiman latirostris (Zucoloto, 1998) and the Claµ2, Claµ3, Claµ5, Claµ6, Claµ7, Claµ8, Claµ9, Claµ10 e Claµ12 markers (Table 1) developed for C. latirostris (Zucoloto et al., 2002) were tested. The PCR conditions were: 60 mM Tris-HCl and 25 mM Ammonium sulfate and different concentrations of Mg2+ and pH (Table 1), 0.2 mM each dNTP, 0.4 µM each primer pair, 1U Taq DNA polymerase and 100 ng DNA in a 25 µl reaction. After 3 min at 94 °C, 30 or 35 cycles (depending on the individual microsatellite) were performed for 1 min at 94 °C, 1 min at the annealing temperature specific for each locus (Table 1), 2 min at 72 °C, and a final extension step of 10 min at 72 °C.
The PCR products were loaded onto 2% agarose gel containing a positive control consisting of the amplification product of the locus analyzed in individuals of C. latirostris under the conditions described in Zucoloto (2002), a negative PCR control, and a fx Hae III DNA size marker to estimate the size of the amplified products. Positive amplifications were loaded in a Megabace 1000 DNA sequencer for genotyping. Allele sizes were obtained using the Genotyper software (GE Healthcare).
Markers developed for A. mississipiensis (Amiµ8, Amiµ11, Amiµ13 and Amiµ20) presented amplification products and polymorphism for all species tested with the exception of the Amiµ8 marker that showed no amplification for P. palpebrosus. The Claµ2, Claµ3, Claµ5, Claµ6, Claµ7, Claµ8, Claµ9, Claµ10 and Claµ12 markers developed for C. latirostris presented amplification products but the Claµ3 and Claµ12 markers showed nonspecific amplification products for C. latirostris (Claµ3) and Palpebrosus (Claµ12). Several loci were monomorphic in at least one species, while the Claµ12 marker was monomorphic in all the species investigated, although it would be premature to assume that these loci are truly monomorphic for the species investigated because only a small number of specimens were used in our study. An exception is the Claµ12 marker, which showed no polymorphism in C. latirostris even when more than 90 individuals were tested (Zucoloto et al., 2002). An interesting observation was that we found that although Claµ3 gave poor amplification results in C. latirostris it worked well in C. crocodilus, C. yacare and P. palpebrosus.
Despite some exceptions, allele sizes for C. crocodilus and C. yacare were in agreement with the size range observed for C. latirostris by Zucoloto et al. (2002) (Table 2). The Amiµ8 marker showed no PCR amplification product for P. palpebrosus and allele sizes for C. crocodilus and C. yacare were out of the range of those observed for C. latirostris (Table 2). Amplification for P. palpebrosus diverged from that observed for the other species, as can be observed in Table 2 for the Amiµ13, Amiµ20, Claµ3, Claµ5, Claµ6, Claµ7, Claµ8 and Claµ10 markers.
The efficiency of heterologous amplification observed in this study was 100% among the caimans and 84.6% between C. latirostris and P. palpebrosus (Table 2). These results were to be expected considering the evolutionary distance between the species (Primmer et al, 1996).
This study supplied the first set of data showing heterologous amplification of microsatellites for C. crocodilus, C. yacare and P. palpebrosus. Future studies with larger sample sizes are necessary to establish if the markers show Mendelian segregation and determine polymorphism information content (PIC) in the 'caiman complex'. Once these markers are fully characterized they may be able to contribute to the evaluation of genetic diversity, conservation efforts and the elucidation of possible genetic flow between C. crocodilus crocodilus and C. crocodilus yacare.
Acknowledgments
We thank Dr. Augusto Shinya Abe (UNESP, Rio Claro) who kindly granted us access to his caiman collection for blood samples. This work was supported by grants from CNPq (grant number 200153/93-5) and FAPESP (grant numbers 00/01495-3 and 99/02605-8).
Received: October 19, 2004; Accepted: May 10, 2005.
Associate Editor: Sergio Furtado dos Reis
- Avise JC (1994) Molecular Markers: Natural History and Evolution. Chapman and Hall, NY, 511 pp.
- Brazaitis P, Madden R, Amato G, Rebelo G, Yamashita C and Watanabe ME (1997) The South American and Central American caiman (Caiman) complex. In: Special Report to the US Fish and Wildlife Service, Arlington, 62 pp.
- Davis LM, Glenn TC, Elsey RM, Brisbin Jr IL, Rhodes WE, Dessauer HC and Sawyer RH (2001a) Genetic structure of six populations of American alligators: A microsatellite analysis. In: Griggs GC, Seebacher F and Franklin CE (eds) Crocodilian Biology and Evolution. Surrey Beatty and Sons, Chipping Norton, pp 38-50.
- Davis LM, Glenn TC, Dessauer HC, Elsey RM and Sawyer RH (2001b) Multiple paternity and mating patterns in the American alligator, Alligator mississipiensis Mol Ecol 10:1011-1024.
- Dever JA, Strauss RE, Rainwater TR, McMurry ST and Densmore LD (2002) Genetic diversity, population subdivision and gene flow in Morelet's crocodile (Crocodylus moreletii) from Belize, Central America. Copeia 2002:1078-1091.
- FitzSimmons NN, Tanksley S, Forstner MR, Louis EE, Daglish R, Gratten J and Davis S (2001) Microsatellite markers for Crocodylus: New genetic tools for population genetics, mating system studies and forensics. In: Griggs GC, Seebacher F and Franklin CE (eds) Crocodilian Biology and Evolution. Surrey Beatty and Sons, Chipping Norton, pp 51-57.
- Forstner M and Forstner JM (2002) Aplicaciones del DNA en la conservación de los crocodilianos. In: Verdade LM and Larriera A (eds) La Conservación y el Manejo de Caimanes y Cocodrilos de América Latina, v. 2. CN Editoria, Piracicaba, pp 99-117.
- Glenn TC, Stephan W, Dessauer HD and Braun MJ (1996) Allelic diversity in alligator microsatellite loci is negatively correlated with GC content of flanking sequences and evolutionary conservation of PCR amplifiability. Mol Biol Evol 13:1151-1154.
- Glenn TC, Dessauer HC and Braun MJ (1998) Characterization of microsatellite DNA loci in American Alligators. Copeia 1998:591-601.
- Hoezel AR (1992) Molecular Genetic Analysis of Populations. IRL Press, Oxford, pp 59-88.
- King FW and Burke RL (1989) Crocodilian, Tuatara, and Turtle Species of the World: A Taxonomic and Geographic Reference. Association of Systematics Collections, Washington, DC, 216 pp.
- Litt M and Luty JA (1989) A hypervariable microsatellite revealed by in vitro amplification of a dinucleotide repeat within the cardiac muscle actin gene. Am J Hum Genet 44:397-401.
- Moore SS, Sargeant LL, King TJ, Mattick JS, Georges M and Hetzel JS (1991) The conservation of dinucleotide microsatellite among mammalian genomes allows the use of heterologous PCR primer pairs in closely related species. Genomics 10:654-660.
- Olson GA, Hessler JR and Faith RE (1975) Techniques for blood collection and intravascular infusion of reptiles. Lab Anim Sci 25:783-786.
- Primmer CR, Moller AP and Ellegren H (1996) A wide-range survey of cross-species microsatellite amplification in birds. Molecular Ecology 5:365-378.
- Sambrook JE, Fritsch EF and Maniatis T (1989) Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratory Press, NY, 1626 pp.
- Tautz D, Trick M and Dover GA (1986) Cryptic simplicity in DNA is a major source of genetic variation. Nature 322:652-656.
- Verdade LM, Zucoloto RB and Coutinho LL (2002) Microgeographic variation in Caiman latirostris J Exp Zool Part B (Mol Dev Evol) 294:387-396.
- Villareal X, Bricker J, Reinert HK, Gelbert L and Bushar LM (1995) Isolation and characterization of microsatellite loci for use in population genetic analysis in the timber rattlesnake, Crotalus horridus J Hered 87:152-155.
- Wang Z, Weber JL, Zhong G and Tanksley SD (1994) Survey of plant short tandem DNA repeats. Theor Appl Genet 88:1-6.
- Zucoloto RB (1998) Avaliação da diversidade genética e teste de maternidade em jacaré-de-papo-amarelo (Caiman latirostris) através do uso de seqüências de DNA microssatélite. Masters Dissertation, Universidade de São Paulo, Piracicaba.
- Zucoloto RB (2002) Desenvolvimento de seqüências de DNA microssatélite para estudo de populações remanescentes de jacaré-de-papo-amarelo (Caiman latirostris) da região central do Estado de São Paulo. PhD Thesis, Universidade de São Paulo, Piracicaba.
- Zucoloto RB, Verdade LM and Coutinho LL (2002) Microsatellite DNA library for Caiman latirostris J Exp Zool Part B (Mol Dev Evol) 294:346-351.
Send correspondence to
Publication Dates
-
Publication in this collection
10 Mar 2006 -
Date of issue
2006
History
-
Received
19 Oct 2004 -
Accepted
10 May 2005