Abstract
We report the characterization and optimization of 45 heterologous microsatellite loci, and the development of a new set of molecular sex markers for the conservation and management of the Neotropical harpy eagle (Harpia harpyja L. 1758). Of the 45 microsatellites tested, 24 were polymorphic, six monomorphic, 10 uncharacterizable due to multiple bands and five did not amplify. The observed gene diversity of the analyzed sample of H. harpyja was low and similar to that of other threatened Falconiformes. While a high proportion of the microsatellite markers were highly variable, individuals of H. harpyja could be differentiated by a joint analysis of just three (p = 2.79 x 10-4) or four markers (p = 2.89 x 10-5). Paternity could be rejected with 95.23% and 97.83% probabilities using the same three and four markers, respectively. The sex determination markers easily and consistently differentiated males from females even with highly degraded DNA extracted from naturally shed feathers. The markers reported in this study potentially provide an excellent set of molecular tools for the conservation and management of wild and captive H. harpyja and they may also prove useful for the enigmatic Neotropical crested eagle (Morphnus guianensis Daudin 1800).
conservation genetics; Harpia harpyja; microsatellites; raptors; sex markers
EVOLUTIONARY GENETICS
RESEARCH ARTICLE
Genomic resources for the conservation and management of the harpy eagle (Harpia harpyja, Falconiformes, Accipitridae)
Aureo BanhosI; Tomas HrbekI, II; Waleska GravenaI; Tânia SanaiottiIII; Izeni P. FariasI
ILaboratório de Evolução e Genética Animal, Instituto de Ciências Biológicas, Universidade Federal do Amazonas, Manaus, AM, Brazil
IIDepartment of Biology, University of Puerto Rico, Río Piedras, San Juan, Puerto Rico
IIIDepartamento de Ecologia, Instituto Nacional de Pesquisas da Amazônia, Manaus, AM, Brazil
Send correspondence to Send correspondence to: Izeni Pires Farias Laboratório de Evolução e Genética Animal, Instituto de Ciências Biológicas Universidade Federal do Amazonas 69077-000 Manaus, AM, Brazil E-mail: izeni_farias@ufam.edu.br
ABSTRACT
We report the characterization and optimization of 45 heterologous microsatellite loci, and the development of a new set of molecular sex markers for the conservation and management of the Neotropical harpy eagle (Harpia harpyja L. 1758). Of the 45 microsatellites tested, 24 were polymorphic, six monomorphic, 10 uncharacterizable due to multiple bands and five did not amplify. The observed gene diversity of the analyzed sample of H. harpyja was low and similar to that of other threatened Falconiformes. While a high proportion of the microsatellite markers were highly variable, individuals of H. harpyja could be differentiated by a joint analysis of just three (p = 2.79 x 10-4) or four markers (p = 2.89 x 10-5). Paternity could be rejected with 95.23% and 97.83% probabilities using the same three and four markers, respectively. The sex determination markers easily and consistently differentiated males from females even with highly degraded DNA extracted from naturally shed feathers. The markers reported in this study potentially provide an excellent set of molecular tools for the conservation and management of wild and captive H. harpyja and they may also prove useful for the enigmatic Neotropical crested eagle (Morphnus guianensis Daudin 1800).
Key words: conservation genetics, Harpia harpyja, microsatellites, raptors, sex markers.
Introduction
The Neotropical harpy eagle (Harpia harpyja L. 1758, Falconiformes, Accipitridae) is the largest eagle in the Americas and is considered the most powerful bird of prey in the world (Collar, 1989; Sick, 1997). This species inhabits the upper stratum of New World forests from southern Mexico to northeastern Argentina but is comparatively rare throughout its distribution. The main threats to the conservation of H. harpyja is habitat fragmentation, hunting and trade in live birds (Vargas G et al., 2006). The slow reproductive rate and low population densities of H. harpyja make these threats significant throughout its distribution. Harpia harpyja is classified as near threatened by the International Union for the Conservation of Nature (IUCN) and is cited in Appendix I of the Convention on the International Trade of Threatened Species of Fauna and Flora (CITES). While active conservation programs exist in several countries where this eagle occurs, implementation of conservation programs is challenging and conservation success is difficult to assess due to the difficulty of obtaining ecological data. Molecular markers often allow indirect estimates of many ecologically important parameters, and if available would greatly facilitate the conservation and management of H. harpyja.
The knowledge of biology of threatened species is of indispensable interest for conservation. However, as in the case for many threatened raptors, H. harpyja is difficult to study because adults are very difficult to capture and mark, individual birds may move over great distances and while slight size dimorphism exists, there is no sexual dimorphism in plumage making males and females difficult-to-distinguish. However, molecular tools may overcome many of these challenges (e.g. Frankham et al., 2002; Allendorf and Luikart, 2006), especially since such methods can use non-invasive sampling techniques based on DNA extraction from feathers (e.g. Pearce et al., 1997; Segelbacher, 2002; Horváth et al., 2005; Rudnick et al., 2005). Understanding the genetic characteristics of a species is also extremely important for the success of in situ and ex situ conservation programs because this information allows definition of management units needed to minimize the loss of genetic diversity while at the same time preserving the existing genetic structure of the species (Haig, 1998; Hedrick, 2001).
Of the various types of molecular markers used today, microsatellites have many positive attributes, including hypervariability, co-dominance, abundance and tolerance to variation in DNA quality and quantity (Selkoe and Toonen, 2006). Additionally, due to our reasonably good understanding of molecular evolution and the development of robust computational methods, microsatellites are well-suited to answer questions related to effective population size, population structure, migration and colonization rates, and reproductive system, thus providing essential data for conservation. The need to characterize species-specific loci by expensive and laborious isolation and characterization procedures is the primary limitation to the more wide-spread use of microsatellites. Although mutations in the flanking regions of microsatellite loci may prevent amplification, many studies have shown that microsatellites isolated from one species can amplify homologous products in related species (e.g. Martínez-Cruz et al., 2002; Busch et al., 2005), a characteristic known as transferability or cross-species amplification (Selkoe and Toonen, 2006).
Other important molecular markers for conservation are those for sex determination, mainly for species that do not possess apparent sexual dimorphism, whether at the juvenile or adult stage, as is the case of for H. harpyja. In all neognath birds, the female is the heterogametic (WZ) and the male homogametic (ZZ) sex. Molecular methods of sex determination are based on the amplification of paralogous copies of the Chromo helicase DNA binding protein gene (CHD1) present on chromosomes W and Z using mismatch primers, i.e. while the forward primer amplifies CHD1 copies on both W and Z chromosomes, the reverse primers are designed to anneal to either W or Z chromosome and amplify different sized products. Molecular sex markers developed for birds (Griffiths et al., 1998) have limited applicability in Falconiformes due to the fact that the differentiation of amplified CHD1W and CHD1Z fragments is unreliable because both fragments are large and similar in size. Recent publication by Ito et al. (2003) presents a solution that appears to be applicable to all Falconiformes by increasing the size difference of the CHD1W and CHD1Z amplified fragments. However, the relatively large size of these fragments makes PCR amplification from forensic samples difficult.
In this study we present the results of an amplification test of 45 microsatellite loci, isolated and characterized by other authors in various other accipitrid raptors, the characterization and optimization of 30 of these microsatellites and the development of a new set of molecular sex markers for H. harpyja. In addition, we also present a preliminary characterization of genetic diversity of H. harpyja and comment on the usefulness of a subset of these microsatellite markers for assessing the joint probability of the identity of any two samples and of paternity exclusion. We conclude that a carefully chosen subset of microsatellite markers optimized for multiplexing and the newly developed molecular sex markers provide highly valuable and simple-to-use set of molecular tools to assist in the formulation of conservation and management strategies for this threatened raptor.
Material and Methods
Specimens
Molted feathers were collected from individual harpy eagle (Harpia harpyja L. 1758 Falconiformes, Accipitridae) specimens from three main Brazilian biomes: the Amazonian rainforest, the Atlantic rainforest and the Pantanal wetland. Some feather samples were collected from nests in the wild while other samples came from zoos and museums; however, in all cases samples of feather originated from wild-born individuals (Table 1).
Microsatellite loci
There are approximately 100 microsatellite loci isolated and characterized for Falconiformes, some of these being published after we commenced our study. For testing in H. harpyja we chose 45 based in the following criteria: 1) loci were isolated from related falconiform species, 2) loci had at least five alleles in the species for which they were developed, and had preferentially a perfect repeat motif, and 3) when cross-species amplification tests were made, the loci were polymorphic in phylogenetically distantly related taxa. The tested microsatellite loci were taken from six microsatellite panels described by the following authors: Nesje and Røed (2000); Martínez-Cruz et al. (2002); Busch et al. (2005); Hailer et al. (2005); Johnson et al. (2005); Mira et al. (2005); a complete list of loci is presented in Table 2. Preliminary screening was done using two captive H. harpyja specimens for which sufficient quantities of good quality DNA could be extracted. For this preliminary screening and for characterization of specimens in Table 1, total genomic DNA was extracted from a blood clot in the superior umbilicus (a small opening at the proximal tip of the calamus or quill) as recommended by Horváth et al., (2005) using the Qiagen® DNA extraction kit (Valencia, CA, USA). Polymerase chain reactions (PCR) were carried out a total volume of 10 mL consisting of 1 mL of sample DNA (~10 ng), 1 mL each of forward and reverse primer (2 mM), 1 mL of 10X Buffer (200 mM Tris-KCl, pH 8.5), 0.7 mL of MgCl2 (25 mM), 0.8 mL of dNTP (10 mM), 0.2 mL Taq DNA polymerase (5 units/mL; Biotools, Spain) and 4.3 mL of water. All primers were purchased from IDT, Coralville, IA, USA (www.idtdna.com), and dNTPs were purchased from Fermentas, Glen Burnie, MD, USA (www.fermentas.com). Amplification consisted of 35 cycles of denaturation at 93 °C for 15 s, annealing between 50 °C and 55 °C for 30 s and extension at 68 °C for 30 s, and a final seven minute extension at 68 °C was added after the last cycle. For the microsatellite loci that amplified we re-synthesized the forward primer by adding an M13 tail to its 5' end to allow for dynamic fluorescent labeling with FAM-6 labeled M13 primer following the protocol described by Schuelke (2000). Genotyping PCR was performed in a total volume of 10 mL containing 1 µL of reverse primer (0.2 µM), 0.5 mL of M13 labeled forward primer (0.2 µM), 0.5 µL of FAM-6 labeled M13 primer (0.2 µM) and the other reagents described above. Amplification was carried out in a Hybaid PCR thermocycler (Thermo Scientific, USA) and had two main cycling steps (modified from Schuelke, 2000), consisting of an initial denaturation step of 1 min at 93 °C followed by 30 cycles of 30 s at 93 °C, 30 s at 55 °C and 30 s at 68 °C then 20 cycles of 30 s at 93 °C, 30 s at 50 °C, and 30 s at 68 °C. The reaction was completed by a final extension for 30 min at 68 °C to minimize stutter due to non-specific incorporation of adenine (Brownstein et al., 1996). The PCR product was visualized using a MegaBACE1000 (GE Healthcare, United Kingdom) and analyzed with the software Fragment Profiler v1.2 (GE Healthcare, United Kingdom) following the manufacturer's recommendations. For each microsatellite marker we genotyped 10 to 17 (average 15) specimens of H. harpyja originating from all three main Brazilian biomes (Table 1). The variable number of specimens analyzed per microsatellite locus was due to failures in genotyping and the limited quantity of DNA available for repeat analyses, a common problem with forensic samples such as naturally shed feathers (Segelbacher, 2002).
The characterization of each microsatellite locus was based on number of alleles and gene diversity (Nei, 1978), expected (HE) and observed (HO) heterozygosity (Weir, 1996), deviation from Hardy-Weinberg equilibrium (HWE) and linkage disequilibrium between all pairs of loci. All the analyses were performed using the program Arlequin v3.1 (Excoffier et al., 2005), with significance levels for multiple tests being adjusted using the sequential Bonferroni correction (Rice, 1989). To evaluate the potential use of the microsatellite loci for relatedness analyses, we also estimated the probability of paternity exclusion at an individual locus (Q or Pei), and the joint probability of paternity exclusion at all loci (QC or Pet) following Weir (1996). Additionally, we estimated the probability of genetic identity at an individual locus (I) and the joint probability of genetic identity at all loci (IC) according to Paetkau et al. (1995).
Sex determination markers in H. harpyja
For the characterization of sex markers we used the primers developed by Ito et al. (2003). Tests were performed on 10 specimens of H. harpyja collected from museums, zoos and nature (Table 3), of which two samples were of know sex while the other samples were of unknown sex. The PCR reactions for the sex markers were carried out in a total volume of 25 µL containing 1 µL of DNA (~10 ng), 2.5 µL of NP primer (2 µM) 1.3 µL of MP primer (2 µM), 1.3 µL of P2 primer (2 µM), 2.5 µL of 10X Buffer (200 mM Tris-KCl, pH 8.5), 2.5 µL of MgCl2 (25 mM), 2.0 µL of dNTP (10 mM), 0.2 µL Taq DNA polymerase (5 units/mL; Biotools, Spain) and 11.8 µL of deionized water. All primers were purchased from IDT, Coralville, IA, USA (www.idtdna.com), and dNTPs were purchased from Fermentas, Glen Burnie, MD, USA (www.fermentas.com). The thermocycling profile consisted of 1 min denaturation at 93 °C, followed by 35 cycles of denaturation at 93 °C for 10 s, annealing at 52 °C for 35 s and extension at 68 °C for 30 s. The reaction was completed by a final extension for seven minutes at 68 °C. The PCR products were separated on a 3% (w/v) agarose gel. To assess the consistency of the results we repeated each PCR three times for each specimen. The CDH1Z and CDH1W gene fragments of a number of the specimens did not amplify, most likely due to a high degree of DNA degradation; therefore we designed two primers, CHD1Wr (5'-GCTGATCTGGTTTCAGAT TAA-3') and CHD1Zr (5'-AGTCACTATCAGATCCAG AG-3') as substitutes for primers MP (Ito et al., 2003) and P2 (Griffiths et al., 1998) respectively (Table 4). Our new primer set reduced the size of amplicons by nearly 100 bp and using this strategy we were able to sex the remaining specimens of unknown sex.
Results and Discussion
Transferability and characterization of the microsatellites in H. harpyja
Of the 45 microsatellite loci tested in our sample of H. harpyja, 40 amplified successfully but only 30 could be genotyped unambiguously. All 30 loci amplified at 55 °C and produced unambiguous genotypes, thus all PCR reactions were standardized to this annealing temperature. A total of 24 microsatellites loci were polymorphic and the number of alleles per locus ranged from 2 to 11 (Table 4). After sequential Bonferroni correction for multiple comparisons (Rice, 1989), a significant departure from HWE was observed only in the locus BBU46. This and six additional loci that showed HWE deviations before Bonforroni correction were re-scored from original electrophoregrams to eliminate potential scoring biases. The observed and expected gene diversity (Nei, 1978) over all loci was 0.50580 and 0.47242, respectively. We found less than 5% of pairs with significant linkage disequilibrium across all pairs of loci. Indexes of joint probability of paternity exclusion (p = 0.99596) and genetic identity (p = 1.04221 x 10-8) were highly robust.
The high rate of cross-species amplification (40 out of 45 loci) and characterizability (30 out of 45 loci) was attributable to our initial choice of loci. Five of the six microsatellite panels were developed for other Accipitrid species, the same family as the H. harpyja. The transferability of microsatellite primers between species is directly related to the genetic divergence of the species concerned; the greater the genetic divergence, the greater the probability of mutations at priming sites, and thus lower the probability of successful annealing of primers. Although we chose only polymorphic loci, the rates of polymorphism characterized for H. harpyja did not reflect the polymorphism observed in the original studies (reg. R = 0.035, p = 0.402). Levels of polymorphism depend on the sample analyzed, and there is also no expectation of transferability of the degree of polymorphism (Ellegren et al., 1995).
The 24 polymorphic loci appeared to present an excellent panel for populational analyses of H. harpyja. They were also robust markers for estimating kinship and paternity relations (Weir, 1996) and to identify individuals (Paetkau and Strobeck, 1995). However, statistically significant levels of paternity exclusion and genetic identity can be obtained with a panel of only three or four loci. Paternity can be excluded at the p = 0.95233 and genetic identity rejected at the p = 0.00028 levels using just the loci IEAAAG15, IEAAAG04 and Aa43. The addition of the NVHfr206 locus would increase these probabilities to p = 0.97834 and p = 0.00003 levels, respectively. The allelic classes produced by the loci IEAAAG15, IEAAAG04 and Aa43 are non-overlapping, and thus conducive to multiplexing even with one dye set and dynamic fluorescent labeling of alleles (Schuelke, 2000). Inclusion of the locus NVHfr206 would require the use of a second fluorescent dye since its allele sizes overlap with those of the locus IEAAAG15. Dynamic multiplexing with the first three loci would result in a statistically significant estimate of paternity exclusion and/or genetic identity at less than US$ 1 per sample analyzed.
Genetic diversity of H. harpyja
For the IUCN near-threatened H. harpyja the average HO calculated by us was 0.506, similar to that for the recently surveyed accipitrid species Aquila adalberti (the Spanish imperial eagle; HO = 0.516) and Aquila heliaca (the eastern imperial eagle; HO = 0.563) listed as vulnerable by IUCN (Martinez-Cruz et al., 2004). Genetic diversity is necessary for populations and species to adapt to environmental change and reflects their evolutionary potential (Frankham et al., 2002), low genetic diversity therefore being viewed as an indirect measure of extinction threat. It may also be that H. harpyja presents the signature of a genetic bottleneck. Garza and Williamson (2001) have demonstrated that for a population sample of microsatellite loci the mean ratio of the number of alleles to the range in allele size, the M parameter, can be used to detect reductions in population size. The average value of M for the 24 microsatellite loci was 0.84, a value significantly lower than that obtained under simulation of a pre-bottleneck population size (p = 0.026 using the genetic parameter q of 2.24). q is a summary statistic representing four times the product of the effective population size and the mutation rate (Hartl and Clark, 1997). We derived q from estimated census sizes of 104 to 105 harpy eagle individuals (Ferguson-Lee, 2001; Vargas G et al., 2006) assuming that the effective number of individuals is equivalent to 1/10 the census size (Frankham et al., 2002), and that microsatellite mutation rate (m) estimates range from 2.5 x 10-3 to 5.6 x 10-4 (e.g. Dallas, 1992; Weber and Wong, 1993; Brinkmann et al., 1998; Sajantila et al., 1999; Kayser and Sajantila, 2001; Hrbek et al., 2006). Using the most conservative parameter estimates (q = 2.24, µ = 5.6 x 10-4) a value of M = 0.84 reflects a significant reduction in population size (p = 0.026). When the parameter q was estimated directly from the microsatellite data (q = 1.50), the M value was not significant (p = 0.101). However, the q calculated from the data itself is necessarily a lower bound estimate if H. harpyja shows any population structure. Although there is a possibility that H. harpyja has experienced a genetic bottleneck, a more definitive inference can only be made with more extensive sampling, and the determination of any existing population structuring.
Although we cannot extend the perceived threat to the entire distribution of H. harpyja, it seems reasonable to extrapolate low genetic diversity and associated threat potential to other Neotropical regions which are often anthropogenically impacted and near the periphery of the natural distribution of this raptor. Within Brazil, H. harpyja possesses its core and broadest area of distribution and, at least within the Amazon basin, it appears to have suffered limited anthropogenic impact. Yet, the genetic diversity of H. harpyja is lower than that of other accipitrid raptors listed by IUCN in categories which indicate a greater risk of extinction. Furthermore, H. harpyja is not even on the official list of threatened species of the Brazilian Environmental Agency (Instituto Brasileiro do Meio Ambiente e dos Recursos Naturais Renováveis - IBAMA). Although our preliminary data clearly bring into question the current classification status by IUCN and IBAMA, in order to facilitate adequate management and conservation policies, a denser sampling throughout the distribution of H. harpyja is necessary to find out how genetic diversity is distributed over geographic landscape, how genetically diverse is H. harpyja throughout the areas of its distribution, and if it has suffered a genetic bottleneck.
Molecular sex markers
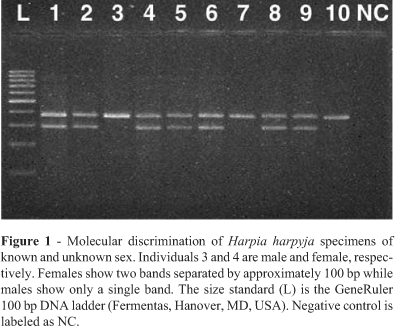
Using the primers from Ito et al. (2003) we were able to confirm the sex of the two H. harpyja specimens of known sex, a male (specimen 3) and a female (specimen 4) (Figure 1). The PCR pattern of males is characterized by a single band and that of females by two bands, with 100 base pairs difference between the two bands. The other eight H. harpyja specimens of differing DNA qualities and concentrations were characterized as two males (specimens 7 and 10) and six females (specimens 1, 2, 5, 6, 8 and 9). Molecular sex determination was repeated three times, each time resulting in the same pattern. The primer set of Ito et al. (2003) minimizes false positive identifications since the female-specific CHD1W gene product is the smaller product. Theoretically even in the case when highly degraded DNA is used and only the smaller CHD1W gene fragment is amplified, this fragment will 100 bp smaller than the CHD1Z gene fragment and therefore this specimen will be easily identifiable as a female. However, some of our specimens showed no amplification of the molecular sex markers, most probably due to the very high levels of DNA degradation common in feathers, which apparently do not allow amplification of the ~300 to 400 bp fragments generated by the markers from Ito et al. (2003). Therefore we designed primers CHD1Wr and CHD1Zr to substitutes for primers MP (Ito et al., 2003) and P2 (Griffiths et al., 1998), respectively in our PCR reactions. The new primer combination NP/CHD1Wr produced a 250 bp fragment while NP/CHD1Zr produced a 300 bp fragment. Primers to obtain even shorter products could not be designed, since the regions amplified span a size variable intron lacking suitably conserved regions. With these new primer combinations we were able to sex our remaining specimens.
Molecular tools for the conservation of the harpy eagle
We have characterized a set of molecular tools useful for in situ and ex situ conservation and management of H. harpyja. The loci IEAAAG15, IEAAAG04 and Aa43 together with sex markers provide powerful and cost effective tools for identifying best potential mates in captive breeding programs. The correct identification of the sex of individual birds in conservation programs, currently a difficult invasive procedure, is clearly fundamental for the success of any breeding program. If the goal of the breeding program is to minimize pedigree inbreeding and maximize genetic diversity, microsatellite markers in addition to the three presented above will need to be used. Captive breeding decisions must be made in light of any potential natural population genetic structuring which, although at present unknown, will be easily determinable with the presented set of 24 polymorphic microsatellite markers once sufficient sampling data are obtained. The existence, or absence, of population structure is also critical for ex situ management and reintroduction programs. With few exceptions, H. harpyja is effectively extinct in the Brazilian Atlantic rainforest and IBAMA has approved a plan presented by the CRAX Society (Sociedade de Pesquisa do Manejo e Reprodução da Fauna Silvestre, MG, Brazil) to reintroduce H. harpyja into the Atlantic Rainforest from a captive population maintained and bred by the CRAX Society. The captive population consists of birds from various biomes, confiscated animals of unknown origin and the hybrid offspring of these animals (Nemésio et al., 2000). The program is currently stalled as specific areas of introduction and financial sponsors have yet to be identified. Even more critically, no data currently exist on whether H. harpyja from the Amazon rainforest and the Atlantic rainforest form one large population and are genetically and demographically interchangeable, or if they represent two differentiated populations. The introduction of inappropriate birds could have serious negative conservation consequences, potentially even leading to the extinction of the remnant Atlantic rainforest populations of H. harpyja (Frankham et al., 2002; Hedrick, 2005; Allendorf and Luikart, 2006). If, however, introductions are scientifically justified, they would be of great benefit in helping to rescue the highly threatened Atlantic rainforest population. Therefore a molecular study of representative specimens from the Amazon and Atlantic rainforests is urgently needed, and the markers reported in this study will greatly facilitate these conservation efforts. Last, but not least, if H. harpyja shows a signal of strong population structuring, these microsatellites could further be used to identify the origin of seized specimens from illegal animal traders and clandestine breeding units. In Brazil, for instance, confiscated specimens of H. harpyja are generally destined for zoos and legalized breeding units (Efe et al., 2006), the principal reason for this bureaucratic decision being the lack of knowledge of the region in which the individuals were clandestinely captured. The assignment of confiscated specimens to their regions of origin could be accomplished with the use of molecular methods, and the confiscated birds could be repatriated to their areas of origin. However, in the case that H. harpyja comprises a panmitic population there will be no restrictions on seized specimens being released in any region within the distribution of H. harpyja and these specimens may be used to augment the severely depleted Atlantic rainforest population. The caveat of these inferences is that they are based on neutral genetic markers. It is possible that H. harpyja from different areas of its distribution may show adaptive differences even if differentiation among regions is not observed at the level of neutral genetic markers, and therefore management and conservation strategies should not solely rely on conclusions drawn from putatively neutral microsatellite markers. In spite of this cautionary note, we believe that the markers reported in this study will prove to be excellent tools for the conservation and management of H. harpyja throughout its geographic distribution, and we also presume that they can be utilized for studies of other raptor species such as the enigmatic species of the genus Morphnus.
Acknowledgments
We thank all the people and institutions that provided feather samples for this study: IBAMA Costa Marques, Foz Tropicana Parque das Aves, Crocodilo Safari Zoo, UFMT Zoo, MPEG Parque Zoobotanico, Bosque Municipal de São José do Rio Preto, Criadouro Sítio Tibagi, Museu de História Natural Capão da Imbuia, Refúgio Biológico Bela Vista - Itaipu Binacional, Estação Vera Cruz - Veracel, Zoológico do Centro Integrado de Guerra na Selva (Zoológico CIGS), and Gerência do Parque Nacional Serra da Bodoquena. We also thank members of the Laboratório de Evolução e Genética Animal (LEGAL UFAM), Projeto Gavião-real of INPA, Coordenação do Curso de Genética, Evolução e Biologia Evolutiva (INPA) and Coordenação de Ecologia (INPA) for help in the laboratory, the field and for discussion. We thank two anonymous reviewers for improving the manuscript. IBAMA (#120/2005 - CGFU/LIC) and CGEN (#088/2005) provided collection and genetic assessment permits, respectively. Financial support for this study was provided by grants from Fundação O Boticário de Proteção à Natureza, Cleveland Zoological Society, Programa Bolsa de Estudos para Conservação da Amazônia (Programa BECA), and Fundação de Amparo a Pesquisa da Amazônia (FAPEAM). This study forms a portion of a Ph.D. dissertation of Aureo Banhos who has a scholarship from the Brazilian Coordenação de Aperfeiçoamento Pessoal de Nível Superior.
Received: March 16, 2007; Accepted: June 11, 2007.
Associate Editor: João S. Morgante
- Allendorf FW and Luikart G (2006) Conservation and the Genetics of Populations. Blackwell Publishing, New York, 644 pp.
- Brinkmann B, Klintschar M, Neuhuber F, Hühne J and Rolf B (1998) Mutation rate in human microsatellites: Influence of the structure and length of the tandem repeat. J Human Genet 62:1408-1415.
- Brownstein MJ, Carpten JD and Smith JR (1996) Modulation of non-templated nucleotide addition by Taq DNA polymerase: Primer modifications that facilitate genotyping. Biotechniques 20:1004-1010.
- Busch JD, Katzner TE, Bragin E and Keim P (2005) Tetranucleotide microsatellites for aquila and haliaeetus eagles. Mol Ecol Notes 5:39-41.
- Collar NJ (1989) Harpy eagle. World Birdswatch 11:5.
- Dallas JF (1992) Estimation of microsatellite mutation rates in recombinant inbred strains of mouse. Mam Genome 3:452-456.
- Efe MA, Martins-Ferreira C, Olmos F, Mohr LV and Silveira LF (2006) Diretrizes da Sociedade Brasileira de Ornitologia para a destinação de aves provenientes do tráfico e cativeiro. Rev Brasil Ornitol 14:67-72.
- Ellegren H, Primmer CR and Sheldon B (1995) Microsatellite evolution: Directionality or bias in locus selection? Nat Genet 11:60-62.
- Excoffier L, Laval G and Schneider S (2005) Arlequin ver. 3.1: An integrated software package for population genetics data analysis. Evol Bioinf Online 1:47-50.
- Ferguson-Lee J (2001) Raptors of the World. Houghton Mifflin, Boston, 872 pp.
- Frankham R, Ballou JR and Briscoe DA (2002) Introduction to Conservation Genetics. Cambridge University Press, Cambridge, 640 pp.
- Garza JC and Williamson EG (2001) Detection of reduction in population size using data from microsatellite loci. Mol Ecol 10:305-318.
- Griffiths RC, Double MC, Orr K and Dawson RJG (1998) A DNA test to sex most birds. Mol Ecol 7:1071-1075.
- Haig SM (1998) Molecular contributions to conservation. Ecology 79:413-425.
- Hailer F, Gautschi B and Helander B (2005) Development and multiplex PCR amplification of novel microsatellite markers in the White-tailed Sea Eagle, Haliaeetus albicilla (Aves, Falconiformes, Accipitridae). Mol Ecol Notes 5:938-940.
- Hartl DL and Clark AG (1997) Principles of Population Genetics. 3rd ed. Sinauer Associates, Sunderland, 542 pp.
- Hedrick PW (2001) Conservation genetics: Where are we now? Trends Ecol Evol 16:629-636.
- Hedrick PW (2005) 'Genetic restoration': A more comprehensive perspective than 'genetic rescue'. Trends Ecol Evol 20:109.
- Horváth MB, Martínez-Cruz B, Negro JJ, Kalmár L and Godoy JA (2005) An overlooked DNA source for non-invasive genetic analysis in birds. J Avian Biol 36:84-88.
- Hrbek T, de Brito RA, Wang B, Pletscher LS and Cheverud JM (2006) Genetic characterization of a new set of recombinant inbred lines (LGXSM) formed from the intercross of SM/J and LG/J inbred mouse strains. Mam Genome 17:417-429.
- Ito H, Sudo-Yamaji A, Abe M, Murase T and Tsubota T (2003) Sex identification by alternative polymerase chain reaction methods in falconiformes. Zool Sci 20:339-344.
- Johnson PCD, Fowlie MK and Amos W (2005) Isolation of microsatellite loci from the common buzzard, Buteo buteo (Aves, Accipitridae). Mol Ecol Notes 5:208-211.
- Kayser M and Sajantila A (2001) Mutations at Y-STR loci: Implications for paternity testing and forensic analysis. Forensic Sci Int 118:116-121.
- Martínez-Cruz B, David VA, Godoy JA, Negro JJ, O'Brien SJ and Johnson WE (2002) Eighteen polymorphic microsatellite markers for the highly endangered Spanish imperial eagle (Aquila adalberti) and related species. Mol Ecol Notes 2:323-326.
- Martinez-Cruz B, Godoy JA and Negro JJ (2004) Population genetics after fragmentation: The case of the endangered Spanish imperial eagle (Aquila adalberti). Mol Ecol 13:2243-2255.
- Mira S, Wolff K and Cancela ML (2005) Isolation and characterization of microsatellite markers in Bonelli's eagle (Hieraaetus fasciatus). Mol Ecol Notes 5:493-495.
- Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583-590.
- Nemésio A, Machado AR and Andrade MA (2000) Reprodução da hápia (Harpia hapyja) em cativeiro. Aves 1:34-35.
- Nesje M and Røed KH (2000) Microsatellite DNA markers from the gyrfalcon (Falco rusticolus) and their use in other raptor species. Mol Ecol 9:1438-1440.
- Paetkau D, Calvert W, Stirling I and Strobeck C (1995) Microsatellite analysis of population structure in Canadian polar bears. Mol Ecol 4:347-354.
- Paetkau D and Strobeck C (1995) The molecular basis and evolutionary history of a microsatellite null allele in bears. Mol Ecol 4:519-520.
- Pearce JM, Fields RL and Scribner KT (1997) Nest materials as a source of genetic data for avian ecological studies. J Field Ornithol 68:471-481.
- Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223-225.
- Rudnick JA, Katzner TE, Bragin EA, Rhodes Jr. OE and DeWoody JA (2005) Using naturally shed feathers for individual identification, genetic parentage analyses, and population monitoring in an endangered Eastern imperial eagle (Aquila heliaca) population from Kazakhstan. Mol Ecol 14:2959-2967.
- Sajantila A, Lukka M and Syvänen A-C (1999) Experimentally observed germline mutations at human micro- and minisatellite loci. Eur J Hum Genet 7:263-266.
- Schuelke M (2000) An economic method for the fluorescent labeling of PCR fragments. Nat Biotech 18:233-234.
- Segelbacher G (2002) Noninvasive genetic analysis in birds: Testing reliability of feather samples. Mol Ecol Notes 2:367-369.
- Selkoe KA and Toonen RJ (2006) Microsatellites for ecologists: A practical guide to using and evaluating microsatellite markers. Ecol Lett 9:615-629.
- Sick H (1997) Ornitologia Brasileira. Editora Nova Fronteira, Rio de Janeiro, 912 pp.
- Vargas GJJ, Whitacre D, Mosquera R, Albuquerque J, Piana R, Thiollay J-M, Márquez C, Sánchez JE, López ML, Midence S et al. (2006) Estado y distribución actual del Águila Arpía (Harpia harpyja) en Centro y Sur América. Ornitol Neotrop 17:39-55.
- Weber JL and Wong CC (1993) Mutation of human short tandem repeats. Hum Mol Genet 2:1123-1128.
- Weir BS (1996) Genetic Data Analysis II: Methods for Discrete Population Genetic Data. Sinauer Associates, Sunderland, 445 pp.
Publication Dates
-
Publication in this collection
28 Feb 2008 -
Date of issue
2008
History
-
Accepted
11 June 2007 -
Received
16 Mar 2007