Abstract
Cytogenetic studies were carried out in three populations of Bryconamericus aff. iheringii from two hydrographic systems of the Paranapanema and Ivaí Rivers, separated by a watershed, both belonging to the upper Paraná River basin. Specimens had a constant diploid number 2n = 52 chromosomes. However, three karyotype formulae were identified in the three populations: B. aff. iheringii from the Maringá stream had 12M+18SM+8ST+14A (FN = 90); specimens from Keller River showed 8M+28SM+6ST+10A (FN = 94) and specimens from the Tatupeba stream had 8M+20SM+8ST+16A (FN = 88). Nucleolar organizer regions (NORs) were identified by silver nitrate staining and fluorescent in situ hybridization (FISH) with an 18S rDNA probe. Specimens from Tatupeba stream had a simple NOR system located in a terminal position of the short arm of a pair of large submetacentric chromosomes. Ag-NOR and FISH methodologies revealed multiple NORs in specimens of the Maringá stream and Keller River. Differences in chromosome structure and in NOR patterns in the three populations of B. aff. iheringii revealed fixed evolutionary chromosome divergence. Aspects related to karyotypic variations and to geographic isolation of these populations are discussed.
Bryconamericus; Characid fish; chromosome divergence; fluorescent in situ hybridization; NOR polymorphism
FISH CYTOGENETICS
RESEARCH ARTICLE
Chromosome divergence and NOR polymorphism in Bryconamericus aff. iheringii (Teleostei, Characidae) in the hydrographic systems of the Paranapanema and Ivaí Rivers, Paraná, Brazil
Thiago Gomes CapistanoI; Ana Luiza de Brito Portela CastroII; Horácio Ferreira JulioJuniorII
IDepartamento de Biologia Celular e Genética, Universidade Estadual de Maringá, Maringá, PR, Brazil
IIDepartamento de Biologia Celular e Genética, Universidade Estadual de Maringá, Nupélia, Maringá, PR, Brazil
Send correspondence to Send correspondence to: Ana Luiza de Brito Portela Castro Departamento de Biologia Celular e Genética Universidade Estadual de Maringá Av. Colombo 5790 87020900 Maringá, PR, Brazil Email: albpcastro@nupelia.uem.br
ABSTRACT
Cytogenetic studies were carried out in three populations of Bryconamericus aff. iheringii from two hydrographic systems of the Paranapanema and Ivaí Rivers, separated by a watershed, both belonging to the upper Paraná River basin. Specimens had a constant diploid number 2n = 52 chromosomes. However, three karyotype formulae were identified in the three populations: B. aff. iheringii from the Maringá stream had 12M+18SM+8ST+14A (FN = 90); specimens from Keller River showed 8M+28SM+6ST+10A (FN = 94) and specimens from the Tatupeba stream had 8M+20SM+8ST+16A (FN = 88). Nucleolar organizer regions (NORs) were identified by silver nitrate staining and fluorescent in situ hybridization (FISH) with an 18S rDNA probe. Specimens from Tatupeba stream had a simple NOR system located in a terminal position of the short arm of a pair of large submetacentric chromosomes. AgNOR and FISH methodologies revealed multiple NORs in specimens of the Maringá stream and Keller River. Differences in chromosome structure and in NOR patterns in the three populations of B. aff. iheringii revealed fixed evolutionary chromosome divergence. Aspects related to karyotypic variations and to geographic isolation of these populations are discussed.
Key words:Bryconamericus, Characid fish, chromosome divergence, fluorescent in situ hybridization, NOR polymorphism.
Introduction
An important characteristic of nucleolar organizer regions (NORs) in fish is its inter and intraspecies polymorphism. NOR characterization can be a cytogenetic marker of for cytotaxonomic studies and can even aid in constructing phylogenetic hypotheses (cytosystematics) for several fish groups (Amemyia and Gold, 1988; Galetti Jr, 1998; AlmeidaToledo, 2000). Some fish groups present a simple NOR system characterized by ribosomal cistrons on only one chromosome pair, whereas others have a multiple NOR system composed of cistrons dispersed over several chromosomes (Galetti Jr., 1998).
Small characids are characterized by extensive heterogeneity with regard to NOR patterns. A simple NOR system was identified in the species Gymnocorymbus ternetzi (Alberdi and Fenocchio, 1997), Tetragonopterus argenteus (Alberdi and Fenocchio, 1997), Moenkhausia intermedia (Portela et al., 1988; PortelaCastro and Julio Júnior, 2002) and Moenkhausia costae (Portela et al., 1988). In others characids, occurrence of a multiple NOR system is common, as has been observed in the genus Astyanax, chiefly in the Astyanax scabripinnis complex (e.g., Maistro et al., 1998; Mizoguchi and MartinsSantos, 1998; Mantovani et al., 2000; MarcoFerro et al., 2001; Souza et al., 2001; Kavalco and MoreiraFilho, 2003; Mantovani et al., 2005). Both conditions have been reported in some cases: specimens of Moenkhausia sanctaefilomenae from the Tietê River (SP, Brazil) analyzed by Foresti et al. (1989) had a multiple NOR system, whereas Moenkhausia sanctaefilomenae from the Paraná River (PR, Brazil) analyzed by PortelaCastro and Júlio Jr. (2002) exhibited a simple NOR pattern. Differentiation in the localization of 18S and 5S ribosomal sites was detected between two Hyphessobrycon anisitsi populations which presented similar karyotype in number and formulae (Centofante et al., 2003).
Bryconamericus is one of the 88 genera listed as "Incertae sedis" group of the Characidae family with 51 valid species (Lima et al., 2003). Actual diversity in the genus Bryconamericus is unknown and its systematics is unresolved. The genus comprises smallsized species, not more than 10 cm in length, that are distributed throughout several continental aquatic ecosystems in South and Central America (Vari and Siebert, 1990).
Chromosome analyses of the genus Bryconamericus are rare and the diploid number of the species studied up to the moment is restricted to 2n = 52 (Portela et al., 1988; Wasko et al., 1996; Wasko and Galetti Jr., 1998; PaintnerMarques et al., 2002a, 2003), although great diversity in chromosome structure has been revealed.
In the current study three populations of Bryconamericus aff. iheringii from two hydrographic systems of the Paranapanema and Ivaí River basins, separated by a watershed were studied. Karyotypes were analyzed with emphasis on the identification of NORs by silver nitrate staining (AgNO3) and fluorescent in situ hybridization (FISH) with 18S rDNA probes.
Material and Methods
Cytogenetic studies were carried out in three populations of Bryconamericus aff. iheringii from two hydrographic system of the upper Paraná River basin in the state of Paraná, Brazil (Figure 1): the Maringá stream belongs to the Paranapanema River basin; Keller River and the Tatupeba stream belong to the Ivaí River basin. Amongst the 54 specimens analyzed, 21 (8 males and 13 females) were collected from the Tatupeba stream; 16 specimens (5 males and 11 females) were collected from Maringá stream and 17 specimens (10 males and 7 females) were collected from Keller River. Mitotic metaphases were obtained from kidney cells by airdrying, as described by Bertollo et al. (1978). NORs were identified by silver nitrate (AgNO3) following the Howell and Black (1980) method, and by fluorescent in situ hybridization (FISH) method with 18S rDNA probes. Two types of probes were used to detect 18S rDNA segments in FISH analysis: (1) genomic DNA of Astyanax scabripinnis amplified by PCR using the primers NS1 (5'GTAGTCATATGCTTGTCTC3') and NS8 (5'TCCGCAGGTTCACCTACGGA3'), recommended by White et al. (1990); (2) amplified and cloned fragments of Oreochromis niloticus (kindly provided by Dr. Cesar Martins of the Universidade Estadual Paulista, Botucatu, SP, Brazil). The probes were labeled with biotin 14dATP by nick translation (Bio Nick Labeling System Gibco, BRL). The FISH protocol followed the methods of HeslopHarrison et al. (1991) and Cuadrado and Jouve (1994).
Results
Bryconamericus aff. iheringii specimens had a constant diploid number of 2n = 52 chromosomes, however, three karyotype formulae were identified. No chromosome differences were found between the sexes.
Bryconamericus aff. iheringii population of the Maringá stream (Cytotype I)
This karyotype is composed of 12M+18SM+8ST+ 14A with fundamental number (FN) of 90, (Figure 2a). Silver nitratestained metaphases (AgNOR) showed terminal labeling on the short arm of two to four chromosomes, with inter and intraindividual variation in signal numbers (Figure 3a). By FISH we detected six fluorescent signals of which four corresponded to AgNOR chromosomes (Figure 3b). Four fluorescent signals on the telomeres of the short arm of the submetacentric pairs 7 and 10 showed greater intensity, whereas a low intensity fluorescent spot was detected on the telomere of the short arm of submetacentric pair 14. A size heteromorphism occurred on pair 7.
Population of Bryconamericus aff. iheringii from Keller River (Cytotype II)
This karyotype is composed of 8M+28SM+6ST+ 10A and FN = 94, (Figure 2b). Silver nitrate staining revealed terminal labels in the short arm of two to four chromosomes (Figure 3c), with inter and intraindividual numerical variations. AgNORs sites were eventually detected in the telomere of the long arm of one of the homologs of the large subtelocentric pair. FISH revealed ten ribosomal sites that included AgNOR markings (Figure 3d). Intense fluorescent signals were detected in the short arm of submetacentric telomeres (pairs 7 and 8). In addition to the above chromosomes, smaller signs were visible in six more chromosomes which included the AgNOR region in a subtelocentric chromosome. Size heteromorphism was detected in chromosome pair 7.
Population of Bryconamericus aff. iheringii from Tatupeba stream (Cytotype III)
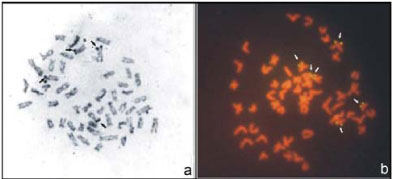
The karyotype structure of these specimens consists of 8M+20SM+8ST+16A, FN = 88 (Figure 2c). Silver nitratestained metaphases showed only one pair of NORbearing submetacentric chromosomes (n. 7) with labeling on the short arm, coinciding with a secondary constriction (Figure 3e). Fluorescent signals confirmed the AgNOR pair by FISH. This chromosome pair also had a NORsize heteromorphism (Figure 3f).
Discussion
Although the diploid number of 2n = 52 chromosomes is the most frequent one in the genus Bryconamericus, karyotype formulae are variable even at the intraspecific level. Structural chromosome diversity is corroborated by results of the current analysis. PaintnerMarques et al. (2003) reported 2n = 52 chromosomes in B. aff. iheringii specimens from the Água da Floresta River (Tibagi River basin, Paraná) distributed as 8M+22SM+ 10ST+12A (FN = 92). The three karyotype formulae obtained in the present study and the karyotype reported by PaintnerMarques et al. (2003) suggest the occurrence of fixed extensive evolutionary chromosome diversification in this species. The fundamental number in this genus ranges from 84 to 102 (Portela et al., 1988; Wasko et al., 1996; Wasko and Galetti Jr., 1998; PaintnerMarques et al., 2002a, 2003). Differences in karyotype structure have been evidenced mainly as divergence in acrocentric chromosomes number. Variation in the fundamental number (FN) may be the result of chromosome rearrangements of the pericentric inversion type which are considered to be the main mechanism of karyotype evolution in this fish group (Wasko and Galetti Jr., 1998).
Chromosome banding in Bryconamericus species has contributed towards a better understanding of the structural chromosome diversity of the group. Bryconamericus showed extensive variability of NORs (Wasko and Galetti Jr., 1999; PaintnerMarques et al., 2002b), and each species could be characterized by a specific Cbanding pattern (Wasko and Galletti Jr., 1998).
A multiple NOR system is the most frequent condition in the genus Bryconamericus. Although the two populations of B. aff. iheringii from the Maringá stream and Keller River have multiple ribosomal sites, they differ in the number of NORbearing chromosomes, as revealed by the FISH technique. Chromosome rearrangements, such as transposition and/or translocations resulting in dispersion of ribosomal genes, seem to occur in several fish species (Galetti Jr et al., 1995; Castro et al., 1996; Mantovani et al., 2000). These mechanisms may explain NOR variation in each isolated population of B. aff. iheringii. Silver nitratestained chromosomes (AgNOR) in populations of B. aff. iheringii with multiple NORs (Maringá stream and Keller River) are less than the number of FISHidentified ribosomal sites. This variation shows that not all ribosomal sites (AgNOR) are active. A similar condition has been reported in Bryconamericus aff. exodon, where 2 to 5 AgNOR sites and eight 18S rDNA sites were detected (PaintnerMarques et al., 2002b). The FISH approach using an 18S rDNA probe was extremely important for the distinction between cytotypes I and II with regard to numbers of structural NORs. Additionally, both methodologies revealed a simple NOR pattern in the Tatupeba stream B. aff. iheringii population. Although a single nucleolar pair is an uncommon condition in the genus, results are in accordance with the simple NOR system reported in B. aff. iheringii of the Tibagi River basin analyzed by PaintnerMarques et al. (2003).
The three populations analyzed have a common NOR phenotype, or rather a pair of large submetacentric chromosomes is always labeled. This submetacentric pair in the three studied populations also showed a more intense fluorescent signal, suggesting that it contains a larger number of rDNA gene copies. These chromosomes may be considered as the main nucleolus organizer and probably contain a cytogenetic marker preserved in this species. Wasko and Galetti Jr. (1999) detected up to 9 AgNOR phenotypes in four Bryconamericus species and registered the occurrence of a NOR phenotype (NOR in the short arm of a medium acrocentric chromosome) in three species, indicating that this pair may be an ancient NOR feature among these fishes.
The karyotype divergence and NOR polymorphism detected between the populations of B. aff. iheringii suggests that their geographic isolation could favor the fixation of chromosomal rearrangements that probably occurred during karyotype evolution of the genus Bryconamericus. Differential selective pressures in each environment may have been decisive for karyotype differentiation and may have produced the detected chromosome diversification. The above hypothesis is based on the biological characteristics of small characids, which is a group that comprises species with high levels of endemism and fast speciation rates (Böhlke et al., 1978).
According to Silva (2004), Bryconamericus is a polyphyletic genus with many groups of species. Based mainly on the position and shape of maxillary teeth, this author recognized three groups of Bryconamericus species in South America; the groups exodon, microcephalus and iheringii. The latter includes all species found in the southern region of South America. The occurrence of different B. aff. iheringii cytotypes requires a detailed taxonomic evaluation of this species, and cytogenetic data can be important tools in their identification.
Acknowledgments
We would like to thank Dra. Carla S. Pavanelli for taxonomic identification and CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico) for financial support.
Received: August 22, 2006; Accepted: May 11, 2007.
Associate Editor: Fausto Foresti
License information: This is an openaccess article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
- Alberdi AJ and Fenocchio AS (1997) Karyotypes of five Tetragonopterinae species (Pisces, Characidae) from Argentina. Cytologia 62:171176.
- AlmeidaToledo LF (2000) Karyotypic evolution in Neotropical freshwater fish. Chrom Today 13:169182.
- Amemiya CT and Gold JR (1988) Chromosomal NORs taxonomic and systematic characters in North American cyprinid fishes. Genetica 76:8190.
- Bertollo LAC, Takahashi CS and MoreiraFilho O (1978) Cytotaxonomic consideration on Hoplias lacerdae (Pisces, Erythrinidae). Rev Bras Genet 1:103120.
- Bölke JE, Weitzman SH and Menezes NA (1978) Estado atual da sistemática dos peixes de água doce da América do Sul. Acta Amazônica 8:657677.
- Castro J, Viñas A, Sánchez L and Martínez P (1996) Characterization of an atypical NOR site polymorphism in brown trout (Salmo trutta) with Ag and CMA3 staining, and fluorescent in situ hybridization. Cytogenet Cell Genet 75:234239.
- Centofante L, Bertollo LAC, Miyazawa CS and MoreiraFilho O (2003) Chromosomal differentiation among allopatric populations of Hyphessobrycon anisitsi (Pisces, Tetragonopterinae). Cytologia 68:283288.
- Cuadrado A and Jouve N (1994) Mapping and organization of highlyrepeated DNA sequences by means of simultaneous and sequential FISH and Cbanding in 6xtriticale. Chrom Res 2:331338.
- Foresti F, AlmeidaToledo LF and Toledo SA (1989) Supernumerary chromosome system, Cbanding pattern characterization and multiple nucleolus organizer regions in Moenkhausia sanctaefilomenae (Pisces, Characidae). Genetica 79:107114.
- Galetti Jr PM, Mestriner CA, Monaco PJ and Rasch EM (1995) Postzygotic modifications and intra andinter individual nucleolar organizing regions variations in fish: Report of a case involving Leporinus friderici Chrom Res 3:285290.
- Galetti Jr PM (1998) Chromosome diversity in neotropical fish. NOR studies. Ital J Zool 65(Suppl):5356.
- HeslopHarrison JS, Schwarzacher T, AnamthawJónsson K, Leitch AR, Shi M and Leitch IJ (1991) In situ hybridization with automated chromosome denaturation. Technique J Meth Cell Mol Biol 3:109116.
- Howell WM and Black DA (1980) Controlled silver staining of organizer regions with a protective colloidal developer: A 1step method. Experientia 36:10141015.
- Kavalco KF and MoreiraFilho O (2003) Cytogenetics analyses in four species of the genus Astyanax (Pisces, Characidae) from Paraíba do Sul River Basin. Caryologia 56:453461.
- Lima FCT, Malabarba LR, Buchup PA, Silva JFP, Vari RP, Harold A, Benine R, Oyakawa OT, Pavanelli CS, Menezes NA, et al. (2003) Characidae genera Incertae sedis in Characidae. In: Reis RE, Kullander SO and Ferraris Jr CJ (eds) Check List of the freshwater fishes of the South and Central America. EDIPUCRS, Porto Alegre, pp 106169.
- Maistro EL, Oliveira C and Foresti F (1998) Comparative cytogenetic and morphological of Astyanax scabripinnis paranae (Pisces, Characidae, Tetragonopterinae) Genet Mol Biol 21:201206.
- Mantovani M, Abel LDS, Mestriner CA and MoreiraFilho O (2000) Accentuated polymorphism of heterochromatin and nucleolar regions in Astyanax scabripinnis (Pisces, Characidae): Tools for understanding karyotypic evolution. Genetica 109:161168.
- Mantovani M, Abel LDS and MoreiraFilho O (2005) Conserved 5S and variable 45S rDNA chromosomal localization revealed by FISH in Astyanax scabripinnis (Pisces, Characidae). Genetica 123:211216.
- MarcoFerro DA, Neo DM, MoreiraFilho O and Bertollo LAC (2001) Nucleolar organizing regions, 18S and 5S rDNA in Astyanax scabripinnis (Pisces, Characidae): Populations distribution and functional diversity. Genetica 110:5562.
- Mizoguchi SMHN and MartinsSantos IC (1998) Activation patterns of the nucleolar organizer region in Astyanax scabripinnis populations (Pisces, Characidae) Cytologia 63:259265.
- PaintnerMarques TR, GiulianoCaetano L and Dias AL (2002a) Karyotypic diversity in a Bryconamericus aff. exodon population (Characidae, Tetragonopterinae). Cytologia 67:397402.
- PaintnerMarques TR, GiulianoCaetano L and Dias AL (2002b) Multiple NORs in Bryconamericus aff. exodon (Osteichthyes, Characidae, Tetragonopterinae). Hereditas 137:107112.
- PaintnerMarques TR, GiulianoCaetano L and Dias AL (2003) Cytogenetic characterization of a population of Bryconamericus aff. iheringii (Characidae, Tetragonopterinae). Genet Mol Biol 26:145149.
- Portela ALBS, Galetti Jr PM and Bertollo LAC (1988). Considerations on the chromosome evolution of Tetragonopterinae (Pisces, Characidae). Rev Bras Genet 11:307316.
- PortelaCastro ALB and JúlioJúnior HF (2002) Karyotype relationships among species of the subfamily Tetragonopterinae (Pisces, Characidae): Cytotaxonomic and evolution aspects. Cytologia 67:329336.
- Silva JFP (2004) Two new species of Bryconamericus Eigenmann (Characiformes, Characidae) from Southern Brazil. Neotrop Ichthyol 2:5560.
- Souza IL, Galían J, De La Rua P, Bertollo LAC and MoreiraFilho O (2001) Nonrandom distribution and nucleolar rDNA sites on Astyanax scabripinnis chromosomes. Cytologia 66:8591.
- Vari RP and Siebert DJ (1990) A new unusually sexually dimorphic species of Bryconamericus (Pisces, Ostariophysi, Characidae) from the Peruvian Amazon. Proc Biol Soc Wash 103:516524.
- Wasko AP, Vênere PC and Galetti Jr PM (1996) Chromosome divergence between two sympatric characid fishes of the genus Bryconamericus. Braz J Genet 19:225230.
- Wasko AP and Galetti Jr PM (1998) Karyotype diversity in the neotropical fish Bryconamericus (Characidae, Tetragonopterinae). Cytobios 94:185193.
- Wasko AP and Galetti Jr PM (1999) Extensive NOR variability in fishes of the genus Bryconamericus (Characidae). Cytologia 64:6367.
- White TJ, Bruns T, Lee S and Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Shinsky JJ and White TJ (eds) PCR Protocols: A Guide to Methods and Applications. Academic Press, New York, pp 315322.
Send correspondence to:
Publication Dates
-
Publication in this collection
03 June 2008 -
Date of issue
2008
History
-
Received
22 Aug 2006 -
Accepted
11 May 2007