Abstract
The aim was to characterize the karyotype of rodents of the genus Proechimys from three localities in the central Brazilian Amazon, in the search for new markers that might shed light on our understanding of the taxonomy and evolutionary history of this taxon. Two karyotypes were found, viz., 2n = 28, FN = 46 in individuals from the NRSP (Cuieiras River) and REMAN (Manaus), and 2n = 46, FN = 50 in individuals from the Balbina Hydroelectric Plant. While individuals with the karyotype with 2n = 28 chromosomes were morphologically associated with Proechimys cuvieri, their karyotype shared similarities with those of the same diploid number in two other regions. Although three karyotypes are described for Proechimys cuvieri, no geographic distribution pattern that defined a cline could be identified. Based on the morphological examination of voucher specimens and additional results from molecular analysis, the karyotype with 2n = 46 and FN = 50 could be associated with P. guyannensis.
spiny rats; C-banding; Amazon region; karyotypes; FISH
Karyological analysis of Proechimys cuvieri and Proechimys guyannensis (Rodentia, Echimyidae) from central Amazon
Carlos Eduardo Faresin e SilvaI,III; Eduardo Schmidt ElerI,III; Maria Nazareth F. da SilvaII; Eliana FeldbergIII
IPós-Graduação em Genética, Conservação e Biologia Evolutiva, Instituto Nacional de Pesquisas da Amazônia, Manaus, AM, Brazil
IIColeção de Mamíferos, Instituto Nacional de Pesquisas da Amazônia, Manaus, AM, Brazil
IIILaboratório de Genética Animal, Instituto Nacional de Pesquisas da Amazônia, Manaus, AM, Brazil
Send correspondence to Send correspondence to: Eliana Feldberg Instituto Nacional de Pesquisas da Amazônia, Laboratório de Genética Animal Av. André Araújo 2936, Petrópolis 69011-970 Manaus, AM, Brazil E-mail: feldberg@inpa.gov.br
ABSTRACT
The aim was to characterize the karyotype of rodents of the genus Proechimys from three localities in the central Brazilian Amazon, in the search for new markers that might shed light on our understanding of the taxonomy and evolutionary history of this taxon. Two karyotypes were found, viz., 2n = 28, FN = 46 in individuals from the NRSP (Cuieiras River) and REMAN (Manaus), and 2n = 46, FN = 50 in individuals from the Balbina Hydroelectric Plant. While individuals with the karyotype with 2n = 28 chromosomes were morphologically associated with Proechimys cuvieri, their karyotype shared similarities with those of the same diploid number in two other regions. Although three karyotypes are described for Proechimys cuvieri, no geographic distribution pattern that defined a cline could be identified. Based on the morphological examination of voucher specimens and additional results from molecular analysis, the karyotype with 2n = 46 and FN = 50 could be associated with P. guyannensis.
Key words: spiny rats, C-banding, Amazon region, karyotypes, FISH.
Spiny rats of the genus Proechimys are among the most abundant terrestrial mammals in the Amazon forests (Malcolm et al., 2005), the genus consisting of 25 valid species (Wilson and Reeder, 2005), and 57 karyotype forms (da Silva, 1998; Weksler et al., 2001; Bonvicino et al., 2005; Machado et al., 2005). Patton and Gardner (1972) had previously proposed that chromosome data could be used as efficient markers in the separation of Proechimys species, this having proven to be especially true for locations in which there is sympatry of as yet unknown species of the genus (see da Silva, 1998; Patton et al., 2000).
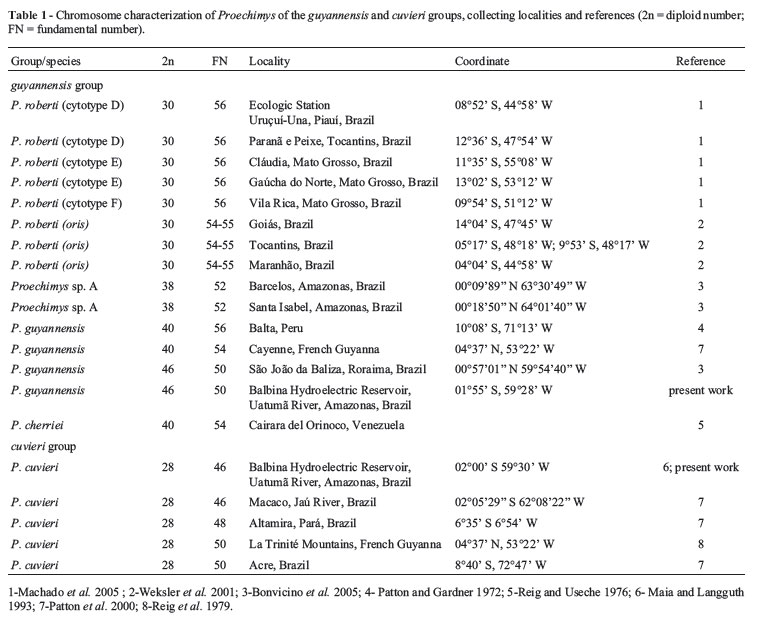
The cuvieri-group comprises only one nominal form, Proechimys cuvieri Petter 1978 (Patton, 1987), with a diploid number of 28 chromosomes. Nevertheless, three karyotype forms have been found in which the fundamental number (FN) is variable (Table 1). Revelation of the existence of clades with divergent mitochondrial DNA has placed in, evidence that P. cuvieri sensu Patton (1987) is composite (da Silva, 1998; Patton et al., 2000). The guyannensis group includes two valid species, viz., P. roberti and P. guyannensis (Weksler et al., 2001; Wilson and Reeder, 2005), although eight karyotype forms have already been described (Table 1).
Herein, the cytogenetic data of specimens recognized as Proechimys cuvieri and Proechimys guyannensis, from three sites in the central Brazilian Amazon, were described, thereby contributing to an understanding of the geographic variation and taxonomy of these taxa.
Eleven individuals of the genus Proechimys were collected for analysis from three localities in the central Brazilian Amazon (Figure 1). The specimens were prepared and deposited in the Mammal Collection of the National Institute of Amazonian Research (INPA, Manaus). Voucher specimens were identified, according to Patton (1987).
Mitotic chromosomes were prepared from femur bone marrow, in accordance with the Ford and Hamerton (1956) protocol. C-banding, G-banding and the detection of nucleolus organizer regions (NORs), were according to protocols described by Sumner (1972), Seabright (1971), and Howell and Black (1980), respectively.
18S rDNA units were amplified according to Gross et al. (2010) from total genomic DNA extracted from Caluromys philander liver tissues (heterologous probe), "all human telomeric probes" were obtained according Ijdo et al. (1991). Probe labeling was with biotin-14-dATP by nick translation (BioNick Labeling System, Invitrogen). In situ fluorescent hybridization was based on protocols described by Pinkel et al. (1986) and Martins and Galetti Jr (2001), with modifications.
Hybridized chromosomes were analyzed using an Olympus BX 51 microscope and the images were captured with a digital camera (Olympus DP70), using the Image-Pro MC 6.0 software. Mitotic metaphases were processed on the Adobe Photoshop CS4 program, and chromosomes measured by means of the Image J public domain program. Chromosomes were classified according to Levan et al. (1964). The fundamental number was based on the number of autosomal arms (FN), as described by Gardner and Patton (1976).
Proechimys cuvieri individuals collected from the NRSP (two males) and the REMAN (two males and one female) showed 2n = 28 chromosomes and FN = 46. The autosomes consisted of seven metacentric, two submetacentric, one subtelocentric and three acrocentric, pairs. The sexual X chromosome was a medium sized acrocentric and the Y was a puntiform chromosome. The largest chromosomes in the karyotype consisted of one metacentric pair (pair 1), one subtelocentric pair (pair 2) and one acrocentric pair (pair 3) (Figure 2a).
Constitutive heterochromatin was encountered in the centromeric region of seven autosomal pairs, viz., three small metacentric pairs, submetacentric pair 5 and all the acrocentric pairs. Nevertheless, heteromorphism was also observed in the heterochromatic blocks in pair 9. Among the sex chromosomes, only the X chromosome had a weakly stained, proximal heterochromatic block in the long arm (Figure 2b). The G-banding pattern was useful in recognizing chromosome pairs (Figure 2c). As visualized by conventional coloration in some metaphases, the nucleolus organizer region (NOR) and 18 S DNAr sequences were all located interstitially in the long arms of submetacentric pair 5, coinciding with secondary constriction (Figure 2d). Telomeric probe hybridization occurred in the telomeric regions of both arms of all the chromosomes (data not shown).
The Proechimys guyannensis individuals collected at the Balbina Hydroelectric Plant (four males and two females) presented a diploid number of 46 chromosomes and FN = 50. Autosomes comprised two metacentric, one submetacentric and 19 acrocentric pairs (Figure 2e). X and Y chromosomes were acrocentric, with the Y chromosomes approximately half the size of the X.
The positive C band, observed in the centromeric region, was distributed among 10 chromosome pairs, one metacentric and one submetacentric pair, the acrocentric pairs and the sex chromosomes. Whereas in X chromosomes, heterochromatin was located only in the centromeric region, Y chromosomes were completely heterochromatic (Figure 2f). The G-banding pattern was useful in recognizing chromosome pairs (Figure 2g). NOR and the 18S rDNA sequences, located interstitially in the long arm of a submetacentric pair (pair 11), coincided with secondary constrictions visualized in a few metaphases of conventional coloration (Figure 2h). Telomeric probe hybridization was observed in the telomeric regions of both arms of all the chromosomes (data not shown).
A diploid number of 2n = 28 was encountered in four species of Proechimys belonging to three species groups (sensu Patton, 1987), viz., in P. quadruplicatus from the goeldii group (Gardner and Emmons, 1984; Patton et al., 2000; Bonvicino et al., 2005), P. longicaudatus (Machado et al., 2005) and P. brevicauda from the longicaudatus group (Patton and Gardner, 1972; Gardner and Emmons, 1984; Patton, 1987; Patton et al., 2000), and P. cuvieri from the cuvieri group (Maia and Langguth, 1993, Patton et al., 2000, and references therein). Considering only those species with 2n = 28, 15 karyotype forms have been found, with fundamental number ranging from 44 to 50 (Weksler et al., 2001).
Proechimys cuvieri is distributed throughout the Amazon basin, from southeastern Peru and the state of Acre (Brazil) in the west, to the Guyanas and the state of Pará (Brazil), in the East. There is partial overlapping of geographic distribution with the goeldii, longicaudatus, simonsi and guyannensis species groups (sensu Patton, 1987). Among all, the only one bearing karyotypes similar to that described here (2n = 28 and FN = 46) is the longicaudatus group. Nevertheless, differences in the size and position of the centromere in the sex chromosomes in P. cuvieri from our material and from the longicaudatus group (Patton et al., 2000; Weksler et al., 2001; Machado et al., 2005) were noted. Based on sex-chromosome morphology and the distribution of constitutive heterochromatin of P. cuvieri specimens from REMAN and the NRSP, their karyotype could be related with those previously described for individuals from the Jaú River (Patton et al., 2000), the Balbina Hydroelectric Plant in the Uatumã River (Maia and Langguth, 1993), and the Jari Valley (Eler et al. in press), all assigned to the cuvieri group.
Currently, three karyotype forms have been recorded for the cuvieri group, all with 2n = 28 chromosomes, but with different fundamental numbers. FN = 46 prevailed in individuals collected from four localities in the central Amazon, namely the Balbina Hydroelectric Plant in the Uatumã River (Maia and Langguth, 1993), the Jaú River (Patton et al., 2000), REMAN and NRSP (present study). FN = 48 occurred in the eastern Amazon, south of the Amazon River in Altamira, and the state of Pará (Brazil) (Patton et al., 2000), and FN = 50 in localities as far distant as the upper Juruá River (state of Acre, Brazil) (Patton et al., 2000) and Cayenne in French Guyana (Reig et al., 1979).
On comparing our data with the conventional coloration of karyotypes from the Jaú River (Patton et al., 2000), it is evident that these karyotypes are very similar to one another, but differ from those from the Balbina Hydroelectric Plant (Maia and Langguth, 1993), as to morphology of the sex chromosomes. In terms of C-band patterns, seven autosomal pairs and a tenuous band in the long arms of sexual chromosome X were present in individuals from REMAN and NRSP, whereas, in individuals from the Balbina Hydroelectric Plant (Maia and Langguth 1993), large heterochromatic blocks were noted in 11 autosomal pairs, and X and Y chromosomes were fully heterochromatic. C-band data for individuals from the Jaú River are inexistent.
The geographic distribution of P. cuvieri cytotypes does not presuppose a cline. Patton et al. (2000) reported FN = 50 in locales situated at both extremes of its distribution, in western (Juruá River, Acre, Brazil) and northeastern (Cayenne, French Guyana) Amazon, FN = 48 in the southeastern section of the Solimões-Amazonas axis, and FN = 46 in central Amazon and the northern part of the Solimões-Amazonas axis. Thus, despite the proposed uniformity in morphological characters for the cuvieri group (Patton, 1987), the presence of chromosome rearrangements appears to indicate that this group is composite. Molecular analysis of the cytochrome b mitochondrial gene (da Silva, 1998; Patton et al., 2000; da Silva et al., unpublished data) certainly indicated differentiated regional units within the cuvieri group presenting degrees of divergence similar to those among other species of Proechimys. These results underscore the need for associating a more ample geographic sampling of P. cuvieri throughout its entire range, with genetic and morphological studies, in order to clarify species composition of this taxon.
In the guyannensis group, four diploid numbers have been described (30, 38, 40 and 46), as well as a variation in the fundamental number from 50 to 56 (Table 1). Weksler et al. (2001) also associated the diploid number of 40 found in P. pattoni and P. gardneri (da Silva, 1998) with the same group, although without presenting an explanation for an association which is not supported by the limited molecular data currently available (see Figure 13 in da Silva, 1998). Weksler et al. (2001) also included specimens from Balta (Peru) with diploid number 40 in the guyannensis group, possibly since these animals, originally referred to as P. guyannensis by Gardner and Patton (1976), were considered as closely related to this species by Gardner and Emmons (1984). Patton (1987), on listing the specimens from Balta as Proechimys sp., placed them provisionally in the group cuvieri. Later on, da Silva (1998) included these specimens in P. pattoni.
The karyotype with 46 chromosomes, as described in the present study, was found in animals from the Balbina Hydroelectric Plant, situated within the areas of geographic distribution of the goeldii, guyannensis and cuvieri species groups (Patton, 1987). This diploid number, identified in P. guairae from the trinitatis group, presents a variation in FN = 68, 70 and 72 (George and Weir, 1973; Reig and Useche, 1976; Aguilera and Corti, 1994). As to diploid (2n = 46) and fundamental (FN = 50) numbers, the form found here is identical to that described by Bonvicino et al. (2005) for specimens from the Jatapu River in the state of Roraima (Brazil), listed as Proechimys sp. (B) and associated with the guyannensis group.
Based on the morphological examination of our voucher specimens, and due to the karyotypic similarities observed between individuals from the Balbina site and those described by Bonvicino et al. (2005), the karyotypic form 2n = 46, FN = 50 from the Balbina Hydroelectric Plant was also associated with the guyannensis group.
Many similarities were found, on comparing this form with that described by Machado et al. (2005) for the region to the north of Manaus, with diploid number 2n = 44 and FN = 52. The apparent differences between the two are the presence of a pair of large submetacentric chromosomes in 2n = 44, and the morphology of the sex chromosomes. In the 2n = 46 karyotype, the X and Y chromosomes are acrocentric, whereas in the 2n = 44, the X chromosome is subtelocentric. Despite the similarities between the two, and the relatively short distance separating the two sampling areas, individuals with 2n = 44 were not assigned by the to any of the Proechimys species groups. We also refrain from doing so here, due to the apparent inexistence of voucher specimens associated to this karyotype. However, when concidering the statement (Machado et al., 2005) that the karyotype 2n = 44, FN = 52 belongs to a different taxonomic entity other than P. cuvieri and P. guyannensis, both currently the only two taxa of Proechimys recognized for central Amazon, it is not clear why they make a claim which certainly lacks support from Malcolm (1994). Thus, it remains unclear whether the diploid number with 44 chromosomes represents a case of chromosome polymorphism in P. guyannensis or, as suggested, a third taxon, as yet unknown in the region. In fact, molecular evidence implies that P. cuvieri and P. guyannensis may be composite.
The distribution of constitutive heterochromatin also differs between the two karyotypes. In the karyotype with 2n = 44, constitutive heterochromatin is found in the centromeric region of all the autosomes and in the Y chromosome, except in pair 1. The X chromosome presented only a pale mark in the proximal region of the short arm. As to 2n = 46, its presence was noted only in autosomal pairs (11, 12, 4, 5, 6, 16, 20, 21, 22), whereas in the X chromosome, centromeric labeling, as well as more tenuous labeling in the interstitial region, were perceptible.
Regarding the C-band pattern, the 2n = 46 karyotype is very similar to the 2n = 38 (guyannensis group) even as regards morphology of the sex chromosomes.
Comparative analysis between the G-band patterns of P. cuvieri and P. guyannensis was inconclusive, as to the identification of homeologies, possibly due to the high number of chromosomal rearrangements within the two. Indeed, molecular evidence presupposes high levels of genetic divergence among Proechimys species (da Silva, 1998).
As Proechimys sp. A (2n = 38), Proechimys sp. B (2n = 46) and Proechimys guyannensis (2n = 40) form a well supported monophyletic clade (Bonvicino et al., 2005), and without discarding inversion re-arrangements, plausibly centric fission/fusion events were involved in differentiation of the diploid number groupwise.
As regards nucleolus organizer regions, Yonenaga-Yassuda et al. (1985) proposed that the nucleolar pair is homeologous in all the species of the Echimyidae family. However, there is considerable variation in the position of this pair in the karyotypes of several Proechimys species, due to non-Robertsonian re-arrangements in other autosomes. The two species under analysis are no exception. This region was located in the 5th pair in P. cuvieri, and in the 11th in P. guyannensis (2n = 46, FN = 50).
Interstitial telomeric signals (ITS) were not detected in P. cuvieri and P. guyannensis, when using telomeric probes, and neither in P. gr. goeldii, 2n = 15 (Machado et al., 2005) and P. guairae guairae, 2n = 48 (Garagna et al., 1997). To date, data on repetitive DNA mapping on Proechimys is very scanty. The lack of chromosomebanding information leaves many gaps, and hinders a more precise analysis of homologies between karyotypes. Consequently, it is, as yet, impossible to define either evolutionary trends, or which karyotype could be considered as the most basal for the genus Proechimys, to thereby trace main chromosomal evolutionary trends, as has been done for other groups of Amazonian organisms, such as bats (Silva et al., 2005), and various families of fish (Feldberg et al., 2003; Artoni and Bertollo, 2001).
The lack of a robust phylogenetic hypothesis for this genus is a primary problem (Lara et al., 1996; da Silva, 1998; Leite and Patton, 2002; Galewski et al., 2005). Obviously, supplementary studies of phylogenetics, which include chromosome markers and a larger number of representative species, are crucial in furthering understanding of chromosomal evolution and genome organization in Proechimys.
In fact, this lack of knowledge is representative of practically all the taxa of the Amazonian mammalian fauna, and underscores the need for biodiversity surveys and associated collection-based data on most of the forest organisms throughout the region.
Acknowledgments
The authors wish to thank R.A. Andrade and M.A.A. Schetino for help with field work, and C.H. Schneider, M.C. Gross and N.D.M. Carvalho for laboratory help and tips. This work was supported by the Instituto Nacional de Pesquisas da Amazônia (INPA) through the Research Institutional Projects (PRJ.05-73), Petrobrás S/A, Instituto de Pesquisas Ecológicas (IPÊ), Programa de apoio a Núcleos de Excelência em Ciência e Tecnologia (PRONEX). C.E.F. e Silva received a granted from the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq). The English was revised by Glenn Sheperd.
Received: July 28, 2011; Accepted: November 4, 2011.
Associate Editor: Yatiyo Yonenaga-Yassuda
License information: This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
- Aguilera M and Corti M (1994) Craniometric differentiation and chromosomal speciation of the genus Proechimys (Rodentia, Echimyidae). Mamm Biol 59:366-377.
- Artoni RF and Bertollo LAC (2001) Trends in the karyotype evolution of Loricariidae fish (Siluriformes). Hereditas 134:201-210.
- Bonvicino CR, Otazú IB and Vilela JF (2005) Karyologic and molecular analysis of Proechimys Allen, 1899 (Rodentia, Echimyidae) from the Amazonian region. Arq Mus Nac 63:191-200.
- da Silva MNF (1998) Four new species of spiny rats of the genus Proechimys (Rodentia, Echimyidae) from the western Amazon of Brazil. Proc Biol Soc Wash 111:436-471.
- Feldberg E, Porto JIR and Bertollo LAC (2003) Chromosomal changes and adaptation of cichlid fishes during evolution. In: Val AL and Kapoor BG (eds) Fish Adaptation. Science Publishers, New Hampshire, pp 285-308.
- Ford C and Hamerton J (1956) A colchicines hypotonic citrate squash sequence for mammalian chromosomes. Stain Technol 31:247-251.
- Galewski T, Mauffrey J, Leite YLR, Patton JL and Douzery EJP (2005) Ecomorphological diversification among South American spiny rats (Rodentia, Echimyidae): A phylogenetic and chrornological approach. Mol Phylogenet Evol 34:601-615.
- Garagna S, Pérez-Zapata A, Zuccotti M, Mascheretti S, Marziliano N, Redi CA, Aguilera M and Capanna E (1997) Genome composition in Venezuelan spiny-rats of the genus Proechimys (Rodentia,Echimyidae). I. Genome size, Cheterochromatin and repetitive DNAs in situ hybridization patterns. Cytogenet Cell Res 78:36-43.
- Gardner AL and Patton JL (1976) Karyotypic variation in Oryzomine rodents (Cricetidae) with comments on chromosomal evolution in the Neotropical cricetinae complex. Occ Pap Mus Zool Univ Mich 49:1-48.
- Gardner AL and Emmons LH (1984) Species group in Proechimys (Rodentia, Echimyidae) as indicated by karyology and bullar morphology. J Mammal 65:10-25.
- George W and Weir BJ (1973) A note on the karyotype of Proechimys guairae (Rodentia, Hystricomorpha). Mammalia 37:330-332.
- Gross MC, Schneider CH, Valente GT, Martins C and Feldberg E (2010) Variability of 18S rDNA locus among Symphysodon fishes: Chromosomal rearrangements. J Fish Biol 76:1117-1127.
- Howell WM and Black DA (1980) Controlled silver-staining of nucleolus organizer region with a protective colloidal developer: A 1-step method. Experientia 36:1014-1015.
- Ijdo JW, Wells RA, Baldini A and Reeders ST (1991) Improved telomere detection using a telomere repeat probe (TTAGGG)n generated by PCR. Nucleic Acids Res 17:4780.
- Lara MC, Patton JL and da Silva MNF (1996) The simultaneous diversification of South American Echimyid rodents (Hystricognathi) based on complete cytochrome b sequences. Mol Phylogenet Evol 5:403-413.
- Leite YLR and Patton JL (2002) Evolution of South American spiny rats (Rodentia, Echimyidae): The star-phylogeny hypotesis revisited. Mol Phylogenet Evol 25:455-464.
- Levan A, Fredga K and Sandberg AA (1964) Nomenclature for centromeric position on chromosomes. Hereditas 52:201-220.
- Machado T, Silva MJJ, Leal-Mesquita ER, Carmignotto AP and Yonenaga-Yassuda Y (2005) Nine karyomorphs for spiny rats of the genus Proechimys (Echimyidae, Rodentia) from North and Central Brazil. Genet Mol Biol 28:682-692.
- Maia V and Langguth A (1993) Constitutive heterochromatin polymorphism and NORs in Proechimys cuvieri Petter, 1978. Rev Bras Genet 16:145-154.
- Malcolm JR (1994) Edge effects in central Amazonian forest fragments. Ecology 75:2438-2445.
- Malcolm JR, Patton JL and da Silva MNF (2005) Small mammal communities in upland and floodplain forests along an Amazonian white water river. In: Lacey EI and Myers P (eds) Mammalian Diversification: From Chromosomes to Phylogeography (A Celebration of the Career of James L. Patton). UC Publications in Zoology Paper, Berkley, 133 pp.
- Martins C and Galetti-Jr PM (2001) Two 5S rDNA arrays in Neotropical fish species: Is it a general rule for fishes? Genetica 111:439-446.
- Patton JL and Gardner AL (1972) Notes on the systematics of Proechimys (Rodentia, Echimyidae), with emphasis on Peruvian forms. Occ Pap Mus Zool LSU 44:1-30.
- Patton JL (1987) Species group of spiny rats, genus Proechimys (Rodentia, Echimyidae). Fieldiana Life Earth Sci 39:305-345.
- Patton JL, da Silva MNF and Malcolm JR (2000) Mammals of the Rio Juruá and the evolutionary and ecological diversification of Amazonia. Bull Am Mus Nat Hist 244:1-306.
- Pinkel D, Straume T and Gray JW (1986) Cytogenetic analysis using quantitative, high-sensitivity, fluorescence hybridization. Proc Natl Acad Sci USA 83:2934-2938.
- Reig AO and Useche M (1976) Diversidad cariotípica y sistemática en poblaciones venezolanas de Proechimys (Rodentia, Echimyidae), con datos adicionales sobre poblaciones de Perú y Colombia. Acta Cient Venez 27:132-140.
- Reig O, Trainer M and Barros MA (1979) Sur l'ídentification chromosomique de Proechimys guyannensis (E. Geoffroy, 1803) et de Proechimys cuvieri Petter, 1978 (Rodentia, Echimyidae). Mammalia 43:501-505.
- Seabright M (1971) A rapid banding technique for human chromosomes. Lancet 2:971-972.
- Silva AM, Marques-Aguiar SA, Barros RMS, Nagamachi CY and Pieczarka JC (2005) Comparative cytogenetic analysis in the species Uroderma magnirostrum and U. bilobatum (cytotype 2n = 42) (Phyllostomidae, Stenodermatinae) in the Brazilian Amazon. Genet Mol Biol 28:248-253.
- Sumner AT (1972) A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res 75:304-306.
- Weksler M, Bonvicino CR, Otazu IB and Junior JSS (2001) Status of Proechimys roberti and P. oris (Rodentia, Echimyidae) from Eastern Amazonia and Central Brazil. J Mamm 82:109-122.
- Wilson DE and Reeder DM (2005) Mammal Species of the World. 3rd edition. Johns Hopkins University Press, Baltimore, 2142 pp.
- Yonenaga-Yassuda Y, Souza MJ, Kasahara S, L'Abbate M and Chu HT (1985) Supernumerary system in Proechimys iheringi iheringi (Rodentia, Echimydae) from the state of São Paulo, Brazil. Caryologia 38:179-194.
Send correspondence to:
Publication Dates
-
Publication in this collection
20 Jan 2012 -
Date of issue
2012
History
-
Received
28 July 2011 -
Accepted
04 Nov 2011