Abstract
The aim of this work was to describe the phenotypic and genotypic variability related to iris color for the population of Buenos Aires province (Argentina), and to assess the usefulness of current methods of analysis for this country. We studied five Single Nucleotide Polymorphisms (SNPs) included in the IrisPlex kit, in 118 individuals, and we quantified eye color with Digital Iris Analysis Tool. The markers fit Hardy-Weinberg equilibrium for the whole sample, but not for rs12913832 within the group of brown eyes (LR=8.429; p=0.004). We found a remarkable association of HERC2 rs12913832 GG with blue color (p < 0.01) but the other markers did not show any association with iris color. The results for the Buenos Aires population differ from those of other populations of the world for these polymorphisms (p < 0,01). The differences we found might respond to the admixed ethnic composition of Argentina; therefore, methods of analysis used in European populations should be carefully applied when studying the population of Argentina. These findings reaffirm the importance of this investigation in the Argentinian population for people identification based on iris color.
Keywords:
Population genetics; Iris color; forensic genetics; phenotype determination; Argentinian population
Introduction
Iris color is a conspicuous phenotypic trait in humans, and its variation is one of the most relevant manifestations of the physical aspect among individuals. This phenotypic trait is polygenic, and its genetic background is variable across different populations. Eye color is subjected to certain changes along the lifetime (Sun et al., 2014Sun HP, Lin Y and Pan CW (2014) Iris color and associated pathological ocular complications: A review of epidemiologic studies. Int J Ophthalmol 7:872-878.). However, this phenotype is mainly influenced by genetic factors acting together as a quantitative trait, giving rise to a range of colors from blue to dark brown (Brues, 1975Brues AM (1975) Rethinking human pigmentation. Am J Phys Anthropol 43:387-391.) generated by complex genetic interactions (Kanetsky et al., 2002Kanetsky PA, Swoyer J, Panossian S, Holmes R, Guerry D and Rebbeck TR (2002) A polymorphism in the Agouti Signaling Protein gene is associated with human pigmentation. Am J Hum Genet 70:770-775.). The importance of its study relies in its potential application in Forensic Genetics, as a tool for predicting Externally Visible Characteristics (Tully, 2007Tully G (2007) Genotype versus phenotype: Human pigmentation. Forensic Sci Int Genet 1:105-110.; Kayser and Schneider, 2009Kayser M and Schneider P (2009) DNA-based prediction of human externally visible characteristics in forensics: Motivations, scientific challenges, and ethical considerations. Forensic Sci Int Genet 3:154-161.).
In recent years the principal genes involved in phenotype determination of iris color have been identified, including HERC2, IRF4, SLC24A4, SLC45A2 and TYR among the most important ones. Within these genes, particular SNPs have been reported as responsible for the pigmentation variation, allowing the determination of iris color in European populations with small levels of error. These SNPs are included in the IrisPlex (Walsh et al., 2011aWalsh S, Liu F, Ballantyne K, van Oven M, Lao O and Kayser M (2011a) IrisPlex: A sensitive DNA tool for accurate prediction of blue and brown eye colour in the absence of ancestry information. Forensic Sci Int Genet 5:170-180.) genotyping system for eye color prediction, which is currently being used in different countries for identification purposes. However, the available data has been collected mainly from European populations (Sulem et al., 2007Sulem P, Gudbjartsson DF, Stacey SN, Helgason A, Rafnar T, Magnusson KP, Manolescu A, Karason A, Palsson A, Thorleifsson G, et al. (2007) Genetic determinants of hair, eye and skin pigmentation in Europeans. Nat Genet 39:1443-1452.; Liu et al., 2009Liu F, van Duijn K, Vingerling J, Hofman A, Uitterlinden AG, Janssens AC and Kayser M (2009) Eye color and the prediction of complex phenotypes from genotypes. Curr Biol 19:R192-R193.; Mengel-From et al., 2009Mengel-From J, Wong TH, Morling N, Rees JL and Jackson IJ (2009) Genetic determinants of hair and eye colours in the Scottish and Danish populations. BMC Genetics 10:88.; Kastelic and Drobnic, 2012Kastelic V and Drobnic K (2012) A single-nucleotide polymorphism (SNP) multiplex system: The association of five SNPs with human eye and hair color in the Slovenian population and comparison using a Bayesian network and logistic regression model. Croat Med J 53:401-408.), while the information about the variability of these SNPs in other populations of the world, and especially of Latin America, is yet incomplete (Beleza et al,. 2013Beleza S, Johnson NA, Candille SI, Absher DM, Coram MA, Lopes J, Campos J, Inês Araújo I, Anderson TM, Vilhjálmsson BJ, et al. (2013) Genetic architecture of skin and eye color in an African-European admixed population. PLoS Genet 9:e1003372.; Edwards et al., 2015Edwards M, Cha D, Krithika S, Johnson M, Cook G and Parra EJ (2015) Iris pigmentation as a quantitative trait: Variation in populations of European, East Asian and South Asian ancestry and association with candidate gene polymorphisms. Pigment Cell Melanoma Res 29:141-162.).
Concerning phenotypic variation, until recently, the assignment of iris color was determined only by qualitative methods (Grigore and Avram, 2015Grigore M and Avram A (2015) Iris colour classification scales - Then and now. Rom J Ophthalmol 59:29-33.). Nevertheless, the qualitative description of this phenotypic trait by assigning each eye subjectively to a group (blue, green or brown), could result in some bias in the classification. This bias usually hinders to reproduce the results, especially when the study attempts to relate these phenotypes with certain genetic markers that could be associated to minor variations in eye color. By using a quantitative method, the bias in determinations given by human subjectivity can be diminished, facilitating the comparison of studies on eye color genetics (Andersen et al., 2013Andersen JD, Johansen P, Harder S, Christoffersen SR, Delgado MC, Henriksen ST, Nielsen MM, Sørensen E, Ullum H, Hansen T, et al. (2013) Genetic analyses of the human eye colours using a novel objective method for eye colour classification. Forensic Sci Int Genet 7:508-515.). At present, digital classification of iris color through software made possible a more objective quantification. Different programs were specifically designed for this purpose, such as the Digital Iris Analysis Tool (DIAT), developed by Jeppe Andersen (Andersen et al., 2013Andersen JD, Johansen P, Harder S, Christoffersen SR, Delgado MC, Henriksen ST, Nielsen MM, Sørensen E, Ullum H, Hansen T, et al. (2013) Genetic analyses of the human eye colours using a novel objective method for eye colour classification. Forensic Sci Int Genet 7:508-515.), and a web application developed by David Cha (Edwards et al., 2015Edwards M, Cha D, Krithika S, Johnson M, Cook G and Parra EJ (2015) Iris pigmentation as a quantitative trait: Variation in populations of European, East Asian and South Asian ancestry and association with candidate gene polymorphisms. Pigment Cell Melanoma Res 29:141-162.). In particular, the DIAT software counts the number of blue and brown pixels in the iris image and calculates a Pixel Index of the Eye (PIE-score) that describes the eye color quantitatively. The PIE-score ranges from -1 to 1 (brown to blue, respectively). As the understanding of genes and markers responsible for intermediate iris color is still limited at present (Edwards et al., 2015Edwards M, Cha D, Krithika S, Johnson M, Cook G and Parra EJ (2015) Iris pigmentation as a quantitative trait: Variation in populations of European, East Asian and South Asian ancestry and association with candidate gene polymorphisms. Pigment Cell Melanoma Res 29:141-162.), the use of software for quantitative methods may improve the analysis.
In Argentina, immigrations and admixture played a fundamental role in the constitution of the current genetic composition of the population. Before the 16th century, Native Americans were the original and only people living in the region. Between the 16th and 18th centuries a relatively small number of Spanish settlers arrived, and then African individuals were forced to migrate between the 17th and 18th centuries. Later, a great European immigration wave took place between 1850 and 1955, mainly composed of Spanish and Italians, and a smaller amount of Germans, Polish and Russians. Also, a small number of Jews and inhabitants from Asia, Middle and Near East, arrived (Devoto, 2007Devoto F (2007) La inmigración de ultramar. In: Torrado S (ed) Población y Bienestar en la Argentina del Primero al Segundo Centenario, Tomo I. Edhasa, Buenos Aires, pp 531-548., Salas et al., 2008Salas A, Jaime JC, Álvarez-Iglesias V and Carracedo Á (2008) Gender bias in the multiethnic genetic composition of central Argentina. J Hum Genet 53:662-674.). Consistently, Avena et al. (2012)Avena SA, Goycoechea AS, Dugoujon JM, Slepoy MG, Slepoy AS and Carnese FR (2001) Análisis antropogenético de los aportes indígena y africano en muestras hospitalarias de la Ciudad de Buenos Aires. Rev Arg Antrop Biol 3:79-99. reported an average autosomal ancestry of 65% European, 31% Native American and 4% African for Argentinians, although a gradient of different distribution of frequencies was found along the country. As a consequence, the genotypic features of Native Americans and Africans have diminished (Avena et al., 2001Avena SA, Goycoechea AS, Dugoujon JM, Slepoy MG, Slepoy AS and Carnese FR (2001) Análisis antropogenético de los aportes indígena y africano en muestras hospitalarias de la Ciudad de Buenos Aires. Rev Arg Antrop Biol 3:79-99.), and the European traits of facial appearance, skin, eyes and hair color may predominate in some regions of Argentina (Merenson, 2015Merenson S (2015) Between brotherhood and exceptionalism: Processes of identification, social marking, and justification in Uruguayan immigration in Buenos Aires. Migraciones internacionales 8:9-37.). According to official data (INDEC, 2010INDEC (2010) Instituto Nacional de Estadísticas y Censos, http://www.indec.gob.ar/censos_total_pais.asp?id_tema_1=2&id_tema_2=41&id_tema_3=135&t=3&s=8&c=2010 (accessed July 2016).
http://www.indec.gob.ar/censos_total_pai...
), from a total of approximately 40 million individuals in the country, 150,000 are Afro-descendant and approximately one million are Native or predominantly of Native ancestry. Besides, since the middle of 20th century, immigration from neighboring South American countries, mainly Paraguay and Bolivia, increased, modifying the genetic composition of Argentinians, contributing more to the Amerindian genetic proportion (Avena et al., 2006Avena SA, Goicochea AS, Rey J, Dugoujon JM, Dejean CB and Carnese FR (2006) Mezcla génica en una muestra poblacional de la ciudad de Buenos Aires. Medicina (Buenos Aires) 66:113-118.).
Populations of different ethnic and geographic origin could carry distinct allele variants of the above mentioned genes, with different effects on phenotype (Frudakis et al., 2003Frudakis T, Thomas M, Gaskin Z, Venkateswarlu K, Suresh Chandra K, Ginjupalli S, Gunturi S, Natrajan S, Ponnuswamy VK and Ponnuswamy KN (2003) Sequences associated with human iris pigmentation. Genetics 165:2071-2083.; Sturm and Frudakis, 2004Sturm RA and Frudakis TN (2004) Eye colour: Portals into pigmentation genes and ancestry. Trends Genet 20:327-332.; Duffy et al,. 2007Duffy DL, Montgomery GW, Chen W, Zhao Z, Le L, James MR, Hayward NK, Martin NG and Sturm RA (2007) A three-single-nucleotide polymorphism haplotype in intron 1 of OCA2 explains most human eye-color variation. Am J Hum Genet 80:241-252.; Sulem et al., 2007Sulem P, Gudbjartsson DF, Stacey SN, Helgason A, Rafnar T, Magnusson KP, Manolescu A, Karason A, Palsson A, Thorleifsson G, et al. (2007) Genetic determinants of hair, eye and skin pigmentation in Europeans. Nat Genet 39:1443-1452.; Mengel-From et al., 2009Mengel-From J, Wong TH, Morling N, Rees JL and Jackson IJ (2009) Genetic determinants of hair and eye colours in the Scottish and Danish populations. BMC Genetics 10:88.; Nan et al,. 2009Nan H, Kraft P, Hunter D and Han J (2009) Genetic variants in pigmentation genes, pigmentary phenotypes, and risk of skin cancer in Caucasians. Int J Cancer 125:909-917.; Walsh et al., 2011aWalsh S, Liu F, Ballantyne K, van Oven M, Lao O and Kayser M (2011a) IrisPlex: A sensitive DNA tool for accurate prediction of blue and brown eye colour in the absence of ancestry information. Forensic Sci Int Genet 5:170-180.; Cerqueira et al., 2012Cerqueira CCS, Paixao-Cortes VR, Zambra FMB, Salzano FM, Hünemeier T and Bortolini MC (2012) Predicting Homo pigmentation phenotype through genomic data: From Neanderthal to James Watson. Am J Hum Biol 24:705-709., 2014Cerqueira CCS, Hünemeier T, Gomez-Valdés J, Ramallo V, Volasko-Krause CD, Leal BAA, Vargas-Pinilla P, Ciconet Dornelles R, Longo D, Rothhammer F, et al. (2014) Implications of the admixture process in skin color molecular assessment. PLoS One 9:e96886.; Kastelic and Drobnic, 2012Kastelic V and Drobnic K (2012) A single-nucleotide polymorphism (SNP) multiplex system: The association of five SNPs with human eye and hair color in the Slovenian population and comparison using a Bayesian network and logistic regression model. Croat Med J 53:401-408.; Beleza et al., 2013Beleza S, Johnson NA, Candille SI, Absher DM, Coram MA, Lopes J, Campos J, Inês Araújo I, Anderson TM, Vilhjálmsson BJ, et al. (2013) Genetic architecture of skin and eye color in an African-European admixed population. PLoS Genet 9:e1003372.; Martinez-Cadenas et al,. 2013Martinez-Cadenas C, Peña-Chilet M, Ibarrola-Villava M and Ribas G (2013) Gender is a major factor explaining discrepancies in eye colour prediction based on HERC2/OCA2 genotype and the IrisPlex model. Forensic Sci Int Genet 7:453-460.; Kastelic et al., 2013Kastelic V, Pospiech E, Draus-Barini J, Branicki W and Drobnic K (2013) Prediction of eye color in the Slovenian population using the IrisPlex SNPs. Croat Med J 54:381-386.; Lim and Oh, 2013Lim J and Oh B (2013) Allelic frequencies of 20 visible phenotype variants in the Korean population. Genomics Inform 11:93-96.; Ruiz et al., 2013Ruiz Y, Phillips C, Gomez-Tato A, Alvarez-Dios J, Casares de Cal M, Cruz R, O. Maroñas O, Söchtig J, Fondevila M, Rodriguez-Cid MJ, et al. (2013). Further development of forensic eye color predictive tests. Forensic Sci Int Genet 7:28-40.; Wilde et al., 2014Wilde S, Timpson A, Kirsanow K, Kaiser E, Kayser M, Unterländer M, Hollfelder N, Potekhina ID, Schier W, Thomas MG, et al. (2014) Direct evidence for positive selection of skin, hair, and eye pigmentation in Europeans during the last 5,000 years. Proc Natl Acad Sci U S A 111:4832-4837.). In this way, the availability of the genetic information responsible for iris color determination in Argentina will provide new tools for its future application in individual identification. In our investigation, we studied a sample from Buenos Aires province in Argentina. The Buenos Aires province (Buenos Aires from now on) population is composed by individuals from all around the country, and they can be considered representative of the Argentinian population. The aim of this work was to describe the genetic component related to iris color variation in Argentina, and to assess the usefulness of current methods of analysis for a Latin American population, not only by checking the relationship between IrisPlex markers and eye color, but also by measuring the phenotypic variation quantitatively. Our results, the first for the country, showed genotype differentiation in the Buenos Aires population, thus contributing to phenotypic individual identification in forensic cases.
Materials and Methods
Sampling, DNA extraction, and phenotypic classification
We collected samples of 118 volunteer adult donors (69 females and 49 males) from Buenos Aires. This project was approved by the Ethics Committee for Biomedical Research at IMBICE, and donors provided individual written informed consent. The age of participants ranged between 18 and 50 years to avoid changes in iris color due to late age of individuals (Sun et al., 2014Sun HP, Lin Y and Pan CW (2014) Iris color and associated pathological ocular complications: A review of epidemiologic studies. Int J Ophthalmol 7:872-878.). We obtained DNA from mouthwash through a standard technique using Proteinase K (Promega, USA) and LiCl (Gemmel and Akiyama, 1996Gemmel N and Akiyama S (1996) An efficient method for the extraction of DNA from vertebrate tissues. Trends Genet 12:338-339.), and we registered iris color variation with a digital camera (Nikon Coolpix s3400, 20 megapixels) using fill-flash. We avoided the variation of environmental light incidence and added a flashlight with a standardized constant light intensity. We were able to categorize 86 iris phenotypes out of the 118 individuals following Andersen et al. (2013)Andersen JD, Johansen P, Harder S, Christoffersen SR, Delgado MC, Henriksen ST, Nielsen MM, Sørensen E, Ullum H, Hansen T, et al. (2013) Genetic analyses of the human eye colours using a novel objective method for eye colour classification. Forensic Sci Int Genet 7:508-515., by using a custom designed software Digital Iris Analysis Tool (DIAT), gently provided by Dr. Jeppe D. Andersen (Dept. of Forensic Medicine, Faculty of Health and Medical Sciences, University of Copenhagen).
Genotyping
We designed oligonucleotides for genotyping the SNPs rs12896399, rs1393350, rs12913832 and rs1689198, while those for rs12203592 were obtained from a previous publication (Walsh et al., 2013Walsh S, Liu F, Wollstein A, Kovatsi L, Ralf A, Kosiniak-Kamysz A, Branicki W and Kayser M (2013) The HIrisPlex system for simultaneous prediction of hair and eye colour from DNA. Forensic Sci Int Genet 7:98-115.). PCR reactions were performed using recombinant Taq polymerase (T-Plus from Inbio-Highway, Tandil, Argentina) in a thermocycler (MPI, La Plata, Argentina). For rs12203592, rs12896399 and rs1393350, we performed allele-specific PCR followed by electrophoretic separation in 1.8% agarose (Promega, USA) electrophoresis and staining with GelRedTM (Biotium, USA). The SNPs rs12913832 and rs16891982 were genotyped by PCR-RFLP, using DdeI (New England Biolabs, USA) and HinfI enzymes (Promega, USA) respectively for restriction digestion, followed by 8% polyacrylamide (Promega, USA) electrophoresis. Bands were recorded in a Gel DocTM XR+ system (Bio-Rad Laboratories, California, USA).
Data analysis
We used data from the five SNPs included in this work for calculating allele and genotype frequencies, and for testing Hardy-Weinberg equilibrium (HWE) by comparison of observed and expected heterozygosity values through exact test (Markov chain) with Arlequin software 3.5 (Excoffier and Lischer, 2010Excoffier L and Lischer HEL (2010) Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564-567.). We also computed nucleotide diversity and Fis for the Buenos Aires population using the same software and determined the distribution of iris color variability as well. The mean, median and standard deviation for color and markers were calculated. We used a median regression analysis adding each SNPs as dummy variable to assess the association among genetic polymorphism. Additionally, a possible link between sex and iris color was verified, as proposed by Martinez-Cadenas et al. (2013)Martinez-Cadenas C, Peña-Chilet M, Ibarrola-Villava M and Ribas G (2013) Gender is a major factor explaining discrepancies in eye colour prediction based on HERC2/OCA2 genotype and the IrisPlex model. Forensic Sci Int Genet 7:453-460. by Chi-square test with Yates correction.
Fst and AMOVA were analyzed with Arlequin 3.5 for comparison with previous literature data for Native Americans (3 countries: Columbia, Brazil, and Mexico, all native populations, total 61 individuals), Africans (6 native populations: Mbuti Pygmies, Biaka Pygmies, Mandenka, Yoruba, San, and Bantu, total 102 individuals), East Asians (3 countries: Japan, China and Russia (Siberia), total 230 individuals) and Europeans (4 countries: France, Italy, United Kingdom (Orkney Islands), and Russia, total 151 individuals) (Walsh et al., 2011aWalsh S, Liu F, Ballantyne K, van Oven M, Lao O and Kayser M (2011a) IrisPlex: A sensitive DNA tool for accurate prediction of blue and brown eye colour in the absence of ancestry information. Forensic Sci Int Genet 5:170-180.). For Fst, a Bonferroni correction was used (Armstrong 2014Armstrong RA (2014) When to use the Bonferroni correction. Ophthalmic Physiol Opt 34:502-508.). For AMOVA analysis, populations were grouped by continent, either including data from Africa, or comparing data more closely related by origin.
Population structure and ancestry estimation were done with Structure 2.3 (Pritchard et al., 2000Pritchard JK, Stephens M and Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945-959.), assuming an admixture model, for K values between 2 and 5 (burn-in period: 10,000), and we compared Buenos Aires population to those above mentioned (Walsh et al., 2011aWalsh S, Liu F, Ballantyne K, van Oven M, Lao O and Kayser M (2011a) IrisPlex: A sensitive DNA tool for accurate prediction of blue and brown eye colour in the absence of ancestry information. Forensic Sci Int Genet 5:170-180.) by non-metric multidimensional scaling (MDS) using the software package PAST (Hammer et al., 2001Hammer Ø, Harper DAT and Ryan PD (2001) PAST: Paleontological Statistics Software Package for Education and Data Analysis. Palaeontol Electronica, v. 4, p 9.).
Results
We observed a non normal distribution of iris color with major prevalence of brown (Figure 1). The vast majority of DIAT results were consistent with the expected PIE-score (80.2% agreement for the whole sample). Nevertheless, not all results from DIAT could be included in our statistical analysis because of non-concordance of some PIE-scores with visual categorization of irises. Errors were mainly present in the group of irises visually considered as brown (20.9%), with lower influence in the group visually considered as blue or green (14.7%).
Histogram of distribution of iris color variation. X axis: PIE-score (Pixel Index of the Eye) ranged from -1 to 1 (i.e. from brown to blue, respectively). Y axis: Number of individuals with each PIE-score value.
Table 1 shows the genotype frequencies that were obtained. The markers fitted HWE for the whole sample. When analyzed separately by iris color, HERC2 rs12913832 did not fit HWE within the group of irises visually determined as brown eyes (LR=8.429; p=0.004), while TYR rs1373350 also showed a tendency to HW disequilibrium within the blue group (LR=5.091; p=0.078).
Nucleotide diversity for the five SNPs was 0.3306 ± 0.2287; the mean observed heterozygosity was 0.3178 ± 0.0758, and the mean expected heterozygosity was 0.3335 ± 0.1277. Fis values were under 15% and non-significant, with rs12203592 in the limit of significance (Fis=19%; p=0.06264 ± 0.00025), which could be explained by the low frequency of the T allele for this marker.
When calculating genotype-phenotype association by a median regression, after discarding the underrepresented genotypes (Table 2), we only found a significant association of HERC2 rs12913832 GG with blue iris color (p < 0.01). No association with iris color was found for the other markers.
Results from median regression. REF: reference variable, sd: standard deviation, IC95: 95% confidence interval. *: removed from the analysis for being unrepresentative.
Considering the sex distribution, the Chi-square test with Yates correction was non-significant (0.811 with p=0.367), leading to conclude that for this sample size it was not possible to confirm an association between sex and iris color as has been previously proposed (Martinez-Cadenas et al., 2013Martinez-Cadenas C, Peña-Chilet M, Ibarrola-Villava M and Ribas G (2013) Gender is a major factor explaining discrepancies in eye colour prediction based on HERC2/OCA2 genotype and the IrisPlex model. Forensic Sci Int Genet 7:453-460.).
According to the Fst results (Table 3) and after applying Bonferroni correction, the Buenos Aires population should be considered as different from the other populations of the world for the analyzed polymorphisms, being most distant from the African populations and closest to the Native American ones here included for comparison. The AMOVA values were not significant (data not shown).
Fst values obtained from interpopulation comparison. *p=0.00000. All values are significative, except for **p=0.01802 ± 0.0121.
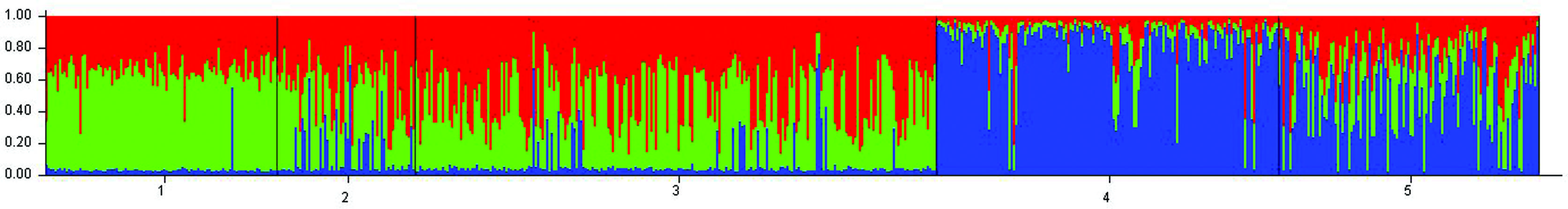
A population structure analysis was made assuming mixed ancestry. Structure Harvester (Earl and vonHoldt, 2012Earl DA and vonHoldt BM (2012) STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359-361.) was used to estimate the most probable K (Table 4). The bar plot grouped by population for K=3 shown in Figure 2 presented an evident differentiation of the Buenos Aires population from African, Native American, East Asian, and European populations.
Most probable K (in bold letter) of the Structure analysis shown above. Performed with Structure Harvester
Structure analysis of different populations: 1-African, 2-Native American, 3-East Asian, 4-European, 5-Buenos Aires. Each color represents an assumed ancestral population. Each individual is represented by a single vertical line divided into K segments of different colors.
The MDS matrix put the Buenos Aires population at similar distances from Europeans and East Asians, closer to Native Americans, and less related to African populations (Figure 3). Stress value analysis performed with PAST resulted in 0. For confirmation of this value the analysis was rerun using SSPS software, and this resulted in 0.00788.
MDS matrix performed with Past. The distance among populations is calculated from Fst. Stress value=0.
Discussion
Eye color is a polygenic trait and requires the analysis of multiple genes for phenotype assignment in forensic practice or for other purposes. In this study we present data on the phenotypic variation of iris and the genetic variation of five polymorphisms related to eye color, localized in different genes, and we assessed the reliability of applying the methods for eye color prediction in individuals from Buenos Aires with the same criteria as those used in other populations of the world. At present, these polymorphisms and genotype-phenotype relationship have not been studied yet in our population.
Good quality images and direct observation of iris color agreed with the PIE-scores resulting from DIAT. According to Andersen et al. (2013)Andersen JD, Johansen P, Harder S, Christoffersen SR, Delgado MC, Henriksen ST, Nielsen MM, Sørensen E, Ullum H, Hansen T, et al. (2013) Genetic analyses of the human eye colours using a novel objective method for eye colour classification. Forensic Sci Int Genet 7:508-515., the effect of light reflection in dark-brown eyes is more likely to confound the analysis (showing values near 0) than in blue eyes. Hence, those images were not included in the statistical analysis. In our sample, as in the Latin American population in general, the majority of the individuals are brown-eyed. Therefore, the use of this software in our population needs careful attention to avoid this problem, by reducing to minimum the iris light incidence when the photograph is taken.
As reported by Andersen et al. (2013)Andersen JD, Johansen P, Harder S, Christoffersen SR, Delgado MC, Henriksen ST, Nielsen MM, Sørensen E, Ullum H, Hansen T, et al. (2013) Genetic analyses of the human eye colours using a novel objective method for eye colour classification. Forensic Sci Int Genet 7:508-515., the software does not accurately distinguish among different tonalities, and not only images of dark and light-brown eyes, but also images of dark and light-blue irises might result in similar PIE-scores.
The HW disequilibrium seen for rs12913832 can be explained by association of homozygous genotype GG with blue iris color, resulting in a lower observed proportion of this genotype among brown-eyed individuals. This is in accordance with reports for this SNP as one of the most important markers for determination of iris color in the great majority of populations (Liu et al., 2009Liu F, van Duijn K, Vingerling J, Hofman A, Uitterlinden AG, Janssens AC and Kayser M (2009) Eye color and the prediction of complex phenotypes from genotypes. Curr Biol 19:R192-R193.; Mengel-From et al., 2009Mengel-From J, Wong TH, Morling N, Rees JL and Jackson IJ (2009) Genetic determinants of hair and eye colours in the Scottish and Danish populations. BMC Genetics 10:88.; Pospiech et al., 2011Pospiech E, Draus-Barini J, Kupiec T, Wojas-Pelc A and Branicki W (2011) Gene-gene interactions contribute to eye color variation in humans. J Hum Genet 56:447-455.; Walsh et al., 2012Walsh S, Wollstein A, Liu F, Chakravarthy U, Rahu M, Seland JH, Soubrane G, Tomazzoli L, Topouzis F, Vingerling JR, et al. (2012) DNA-based eye colour prediction across Europe with the IrisPlex system. Forensic Sci Int Genet 6:330-340.; Kastelic et al., 2013Kastelic V, Pospiech E, Draus-Barini J, Branicki W and Drobnic K (2013) Prediction of eye color in the Slovenian population using the IrisPlex SNPs. Croat Med J 54:381-386.; Ruiz et al., 2013Ruiz Y, Phillips C, Gomez-Tato A, Alvarez-Dios J, Casares de Cal M, Cruz R, O. Maroñas O, Söchtig J, Fondevila M, Rodriguez-Cid MJ, et al. (2013). Further development of forensic eye color predictive tests. Forensic Sci Int Genet 7:28-40.; Wilde et al., 2014Wilde S, Timpson A, Kirsanow K, Kaiser E, Kayser M, Unterländer M, Hollfelder N, Potekhina ID, Schier W, Thomas MG, et al. (2014) Direct evidence for positive selection of skin, hair, and eye pigmentation in Europeans during the last 5,000 years. Proc Natl Acad Sci U S A 111:4832-4837.).
As expected, the relationship genotype-phenotype for rs12913832 was the same as that observed in other populations (Sulem et al., 2007Sulem P, Gudbjartsson DF, Stacey SN, Helgason A, Rafnar T, Magnusson KP, Manolescu A, Karason A, Palsson A, Thorleifsson G, et al. (2007) Genetic determinants of hair, eye and skin pigmentation in Europeans. Nat Genet 39:1443-1452.; Eiberg et al., 2008Eiberg H, Troelsen J, Nielsen M, Mikkelsen A, Mengel-From J, Kjaeret K and Hansen L (2008) Blue eye color in humans may be caused by a perfectly associated founder mutation in a regulatory element located within the HERC2 gene inhibiting OCA2 expression. Hum Genet 123:177-187.; Sturm et al., 2008Sturm RA, Duffy DL, Zhao ZZ, Leite FPN, Stark MS, Hayward NK, Martin NG and Montgomery GW (2008) A single SNP in an evolutionary conserved region within intron 86 of the HERC2 gene determines human blue-brown eye color. Am J Hum Genet 82:424-431.; Nan et al., 2009Nan H, Kraft P, Hunter D and Han J (2009) Genetic variants in pigmentation genes, pigmentary phenotypes, and risk of skin cancer in Caucasians. Int J Cancer 125:909-917.; Liu et al., 2010Liu F, Wollstein A, Hysi PG, Ankra-Badu GA, Spector TD, Park D, Zhu G, Larsson M, Duffy DL, Montgomery GW, et al. (2010) Digital quantification of human eye color highlights genetic association of three new loci. PLoS Genet 6:e1000934.; Valenzuela et al., 2010Valenzuela RK, Henderson MS, Walsh MH, Garrison NA, Kelch JT, Cohen-Barak O, Erickson DT, Meaney FJ, Walsh JB, Cheng KC, et al. (2010) Predicting phenotype from genotype: Normal pigmentation. J Forensic Sci 55:315-322.; Spichenok et al., 2011Spichenok O, Budimlija ZM, Mitchell AA, Jenny A, Kovacevic L, Marjanovic D, Caragine T, Prinz M and Wurmbach E (2011) Prediction of eye and skin color in diverse populations using seven SNPs. Forensic Sci Int Genet 5:472-478.; Sturm and Duffy, 2012Sturm RA and Duffy DL (2012) Human pigmentation genes under environmental selection. Genome Biol 13:248.; Walsh et al., 2011aWalsh S, Liu F, Ballantyne K, van Oven M, Lao O and Kayser M (2011a) IrisPlex: A sensitive DNA tool for accurate prediction of blue and brown eye colour in the absence of ancestry information. Forensic Sci Int Genet 5:170-180.,bWalsh S, Lindenbergh A, Zuniga SB, Sijen T, de Knijff P, Kayser M and Ballantyne KN (2011b) Developmental validation of the IrisPlex system: Determination of blue and brown iris colour for forensic intelligence. Forensic Sci Int Genet 5:464-471., 2012Walsh S, Wollstein A, Liu F, Chakravarthy U, Rahu M, Seland JH, Soubrane G, Tomazzoli L, Topouzis F, Vingerling JR, et al. (2012) DNA-based eye colour prediction across Europe with the IrisPlex system. Forensic Sci Int Genet 6:330-340.; Kastelic and Drobnic, 2012Kastelic V and Drobnic K (2012) A single-nucleotide polymorphism (SNP) multiplex system: The association of five SNPs with human eye and hair color in the Slovenian population and comparison using a Bayesian network and logistic regression model. Croat Med J 53:401-408.; Beleza et al., 2013Beleza S, Johnson NA, Candille SI, Absher DM, Coram MA, Lopes J, Campos J, Inês Araújo I, Anderson TM, Vilhjálmsson BJ, et al. (2013) Genetic architecture of skin and eye color in an African-European admixed population. PLoS Genet 9:e1003372.; Draus-Barini et al., 2013Draus-Barini J, Walsh S, Pospiech E, Kupiec T, Glab H, Branicki W and Kayser M (2013) Bona fide colour: DNA prediction of human eye and hair colour from ancient and contemporary skeletal remains. Investig Genet 4:3.; Hart et al., 2013Hart KL, Kimura SL, Mushailov V, Budimlija ZM, Prinz M and Wurmbach E (2013) Improved eye- and skin-color prediction based on 8 SNPs. Croat Med J 54:248-256.; Kastelic et al., 2013Kastelic V, Pospiech E, Draus-Barini J, Branicki W and Drobnic K (2013) Prediction of eye color in the Slovenian population using the IrisPlex SNPs. Croat Med J 54:381-386.; Lim and Oh, 2013Lim J and Oh B (2013) Allelic frequencies of 20 visible phenotype variants in the Korean population. Genomics Inform 11:93-96.; Ruiz et al., 2013Ruiz Y, Phillips C, Gomez-Tato A, Alvarez-Dios J, Casares de Cal M, Cruz R, O. Maroñas O, Söchtig J, Fondevila M, Rodriguez-Cid MJ, et al. (2013). Further development of forensic eye color predictive tests. Forensic Sci Int Genet 7:28-40.; Wilde et al., 2014Wilde S, Timpson A, Kirsanow K, Kaiser E, Kayser M, Unterländer M, Hollfelder N, Potekhina ID, Schier W, Thomas MG, et al. (2014) Direct evidence for positive selection of skin, hair, and eye pigmentation in Europeans during the last 5,000 years. Proc Natl Acad Sci U S A 111:4832-4837.). The G allele is associated to blue iris color, and the A allele to brown color. The G allele has been reported as a consequence of a founder mutation in HERC2 (SNP rs12913832) which emerged at the East or Northwest of the Black Sea, about 6,000 to 10,000 years ago (Cavalli-Sforza and Feldman, 2003Cavalli-Sforza LL and Feldman MW (2003) The application of molecular genetic approaches to the study of human evolution. Nat Genet 33:266-275.; Eiberg et al,. 2008Eiberg H, Troelsen J, Nielsen M, Mikkelsen A, Mengel-From J, Kjaeret K and Hansen L (2008) Blue eye color in humans may be caused by a perfectly associated founder mutation in a regulatory element located within the HERC2 gene inhibiting OCA2 expression. Hum Genet 123:177-187.) and extended along European populations due to sexual selection (Cavalli-Sforza and Feldman, 2003Cavalli-Sforza LL and Feldman MW (2003) The application of molecular genetic approaches to the study of human evolution. Nat Genet 33:266-275.; Eiberg et al., 2008Eiberg H, Troelsen J, Nielsen M, Mikkelsen A, Mengel-From J, Kjaeret K and Hansen L (2008) Blue eye color in humans may be caused by a perfectly associated founder mutation in a regulatory element located within the HERC2 gene inhibiting OCA2 expression. Hum Genet 123:177-187.; Rodrigues Nogueira Forti and Young, 2016Forti IRN and Young RJ (2016) Human commercial models’ eye colour shows negative frequency-dependent selection. PLoS One 11:e0168458.). The blue-eye associated homozygote rs12913832-GG genotype, as well as the heterozygote genotype, are much more frequent in Europe (G frequency=0.79 according to Beleza et al. (2013)Beleza S, Johnson NA, Candille SI, Absher DM, Coram MA, Lopes J, Campos J, Inês Araújo I, Anderson TM, Vilhjálmsson BJ, et al. (2013) Genetic architecture of skin and eye color in an African-European admixed population. PLoS Genet 9:e1003372., and 0.71 according to Wilde et al. (2014)Wilde S, Timpson A, Kirsanow K, Kaiser E, Kayser M, Unterländer M, Hollfelder N, Potekhina ID, Schier W, Thomas MG, et al. (2014) Direct evidence for positive selection of skin, hair, and eye pigmentation in Europeans during the last 5,000 years. Proc Natl Acad Sci U S A 111:4832-4837. in the surrounding areas, such as the Middle East and West Asia, where blue and intermediate eye colors are expected. In contrast, populations of African origin totally lack this allele. On the other hand, the brown eye associated the rs12913832-AA genotype is present everywhere across the continents, and it is nearly the only one found in certain areas of the world where no blue eye color is expected (Walsh et al., 2011aWalsh S, Liu F, Ballantyne K, van Oven M, Lao O and Kayser M (2011a) IrisPlex: A sensitive DNA tool for accurate prediction of blue and brown eye colour in the absence of ancestry information. Forensic Sci Int Genet 5:170-180.). In the Buenos Aires population, the AA genotype is more frequent than in Europe, in accordance with a differential transoceanic migratory contribution to the native genetic background of our population (Avena et al., 2006Avena SA, Goicochea AS, Rey J, Dugoujon JM, Dejean CB and Carnese FR (2006) Mezcla génica en una muestra poblacional de la ciudad de Buenos Aires. Medicina (Buenos Aires) 66:113-118.), and our results agreed with the varied ethnic composition of our sample. With regard to migratory processes, the presence of rs12913832-G allele can be attributed to migrations and, as a consequence, blue and green eyes in the Argentinian population, since these iris colors have the major occurrence among Europeans and individuals descending from them (Sturm and Frudakis, 2004Sturm RA and Frudakis TN (2004) Eye colour: Portals into pigmentation genes and ancestry. Trends Genet 20:327-332.). However, the migratory component coming from Europe was mainly from regions with a lower proportion of blue eyes (Spain and Italy), compared to whole Europeans (Devoto, 2007Devoto F (2007) La inmigración de ultramar. In: Torrado S (ed) Población y Bienestar en la Argentina del Primero al Segundo Centenario, Tomo I. Edhasa, Buenos Aires, pp 531-548.). Additionally, populations from neighboring countries also contributed with brown-eyed individuals to the Buenos Aires population.
The other markers fit HWE, and Fis values were non-significant, suggesting that important genetic drift, selection or inbreeding processes are not occurring at present in the population under study.
As the Buenos Aires population has a certain Native American contribution (Avena et al., 2012Avena SA, Via M, Ziv E, Pérez-Stable EJ, Gignoux CR, Dejean CB, Huntsman S, Torres-Mejía G, Dutil J, Matta JL, et al. (2012) Heterogeneity in genetic admixture across different regions of Argentina. PLoS One 7:e34695.), we searched in our sample for the presence of male Native American markers, such as M3 (Underhill et al., 1996Underhill P, Jin L, Zemans R, Oefner PJ and Cavalli-Sforza LL (1996) A pre-Columbian Y chromosome-specific transition and its implications for human evolutionary history. Proc Natl Acad Sci U S A 93:196-200.), M242 and M346 in the Y-chromosome, following Jurado-Medina et al. (2014)Jurado-Medina LS, Muzzio M, Schwab M, Costantino ML, Barreto G and Bailliet G (2014) Human Y-chromosome SNP characterization by multiplex amplified product-length polymorphism analysis. Electrophoresis 35:1-4.. We only found 2% of M3 (1 brown-eyed man among 48 men), a result less common than the frequency observed in previous data (4%) (Martínez Marignac et al., 2004Martínez-Marignac VL, Bertoni B, Parra E and Bianchi NO (2004) Characterization of admixture in an urban sample from Buenos Aires, Argentina, using uniparentally and biparentally inherited genetic markers. Hum Biol 76:543-557.) although still present. In this work, we did not perform an analysis of individual mitochondrial ancestry, which could result in a higher percentage of a Native component, as previously reported (Corach et al., 2010Corach D, Lao O, Bobillo C, van Der Gaag K, Zuniga S and Vermeulen M (2010) Inferring continental ancestry of Argentineans from autosomal, Y-chromosomal and mitochondrial DNA. Ann Hum Genet 74:65-76.). Further research focused on Native American contribution to ancestry and genetic association with iris color could reveal whether other mutations are influencing these results.
In this work, an association with iris color was not detected for four markers, contrary to previous reports from other authors (Sulem et al,. 2007Sulem P, Gudbjartsson DF, Stacey SN, Helgason A, Rafnar T, Magnusson KP, Manolescu A, Karason A, Palsson A, Thorleifsson G, et al. (2007) Genetic determinants of hair, eye and skin pigmentation in Europeans. Nat Genet 39:1443-1452.; Liu et al., 2009Liu F, van Duijn K, Vingerling J, Hofman A, Uitterlinden AG, Janssens AC and Kayser M (2009) Eye color and the prediction of complex phenotypes from genotypes. Curr Biol 19:R192-R193.; Valenzuela et al., 2010Valenzuela RK, Henderson MS, Walsh MH, Garrison NA, Kelch JT, Cohen-Barak O, Erickson DT, Meaney FJ, Walsh JB, Cheng KC, et al. (2010) Predicting phenotype from genotype: Normal pigmentation. J Forensic Sci 55:315-322.; Spichenok et al., 2011Spichenok O, Budimlija ZM, Mitchell AA, Jenny A, Kovacevic L, Marjanovic D, Caragine T, Prinz M and Wurmbach E (2011) Prediction of eye and skin color in diverse populations using seven SNPs. Forensic Sci Int Genet 5:472-478.; Walsh et al., 2011aWalsh S, Liu F, Ballantyne K, van Oven M, Lao O and Kayser M (2011a) IrisPlex: A sensitive DNA tool for accurate prediction of blue and brown eye colour in the absence of ancestry information. Forensic Sci Int Genet 5:170-180.; Kastelic and Drobnic, 2012Kastelic V and Drobnic K (2012) A single-nucleotide polymorphism (SNP) multiplex system: The association of five SNPs with human eye and hair color in the Slovenian population and comparison using a Bayesian network and logistic regression model. Croat Med J 53:401-408.; Hart et al., 2013Hart KL, Kimura SL, Mushailov V, Budimlija ZM, Prinz M and Wurmbach E (2013) Improved eye- and skin-color prediction based on 8 SNPs. Croat Med J 54:248-256.; Lim and Oh, 2013Lim J and Oh B (2013) Allelic frequencies of 20 visible phenotype variants in the Korean population. Genomics Inform 11:93-96.; Ruiz et al,. 2013Ruiz Y, Phillips C, Gomez-Tato A, Alvarez-Dios J, Casares de Cal M, Cruz R, O. Maroñas O, Söchtig J, Fondevila M, Rodriguez-Cid MJ, et al. (2013). Further development of forensic eye color predictive tests. Forensic Sci Int Genet 7:28-40.; Fernandes Durso et al., 2014Fernandes Durso D, Bydlowski SP, Hutz MH, Magalhães TR, Suarez-Kurtz G and Junho Pena SD (2014) Association of genetic variants with self-assessed color categories in Brazilians. PLoS One 9:e83926.; Wilde et al., 2014Wilde S, Timpson A, Kirsanow K, Kaiser E, Kayser M, Unterländer M, Hollfelder N, Potekhina ID, Schier W, Thomas MG, et al. (2014) Direct evidence for positive selection of skin, hair, and eye pigmentation in Europeans during the last 5,000 years. Proc Natl Acad Sci U S A 111:4832-4837.). In the case of the SNP rs12203592 from IRF4 gene, the reduced frequency observed for the T allele might interfere in the possibility of finding an association with iris color. Another factor might be the sample size, and we also have to consider that, in the population under study, the number of individuals with blue eyes is much lower than individuals with brown eyes. It is important to note that iris color depends, in part, on light scattering and absorptive properties of extracellular components, especially in lightly pigmented irises (Eagle, 1994Eagle RC (1994) Congenital, developmental and degenerative disorders of the iris and ciliary body. In: Albert DM and Jakobiec FA (eds) Principles and Practice of Ophthalmology. Saunders, Philadephia, pp 367-388.), so this fact can be a complication for defining a clear relationship between each polymorphism and iris color.
In contrast to other studies (Martinez-Cadenas, 2013Martinez-Cadenas C, Peña-Chilet M, Ibarrola-Villava M and Ribas G (2013) Gender is a major factor explaining discrepancies in eye colour prediction based on HERC2/OCA2 genotype and the IrisPlex model. Forensic Sci Int Genet 7:453-460., 2014Martinez-Cadenas C, Peña-Chilet M, Llorca-Cardeñosa MJ, Cervera R, Ibarrola-Villava M and Ribas G (2014) Gender and eye colour prediction discrepancies: A reply to criticisms. Letter to the Editor. Forensic Sci Int Genet 9:e7-e9.), it was not possible to demonstrate an association between sex and iris color in our population. Our results were consistent with the lack of association, as it was proposed by Liu et al. (2014)Liu F, Walsh S and Kayser M (2014) Of sex and IrisPlex eye colour prediction: A reply to Martinez-Cadenas et al. Letter to the Editor. Forensic Sci Int Genet 9:e5-e6..
Results obtained for Fst consistently showed that the Buenos Aires population is genotypically different from other populations of the world, reaffirming the importance of studying these genes in our population for people identification based on iris color. The structure analysis also pointed out this differentiation. The results for the Buenos Aires population were more similar to native populations from other American countries, and in second place, to European populations, suggesting a considerable contribution of Native Americans to the current genetic composition of the Buenos Aires population. However, the percentage of Native contribution in this province is much lower than in other regions of the country (Marino et al., 2007Marino M, Bobillo C, Sala A and Corach D (2007) Evaluación de la subestructura genética en la población argentina evaluada por STRs autosómicos e Y-STRs. Rev Arg Antrop Biol 9:47., Avena et al., 2012Avena SA, Via M, Ziv E, Pérez-Stable EJ, Gignoux CR, Dejean CB, Huntsman S, Torres-Mejía G, Dutil J, Matta JL, et al. (2012) Heterogeneity in genetic admixture across different regions of Argentina. PLoS One 7:e34695., Motti et al., 2013Motti JMB, Muzzio M, Ramallo V, Kladniew BR, Alfaro EL, Dipierri JE, Bailliet G and Bravi CM (2013) Origen y distribución espacial de linajes maternos nativos en el noroeste y centro oeste argentinos. Rev Arg Antrop Biol 15:3-14.). Hence, these results, which are in contrast with values of the demographic composition (2.5% of Native American descendants for the whole country, according to INDEC, 2010INDEC (2010) Instituto Nacional de Estadísticas y Censos, http://www.indec.gob.ar/censos_total_pais.asp?id_tema_1=2&id_tema_2=41&id_tema_3=135&t=3&s=8&c=2010 (accessed July 2016).
http://www.indec.gob.ar/censos_total_pai...
), could be explained by recent migration from neighboring countries (Avena et al., 2006Avena SA, Goicochea AS, Rey J, Dugoujon JM, Dejean CB and Carnese FR (2006) Mezcla génica en una muestra poblacional de la ciudad de Buenos Aires. Medicina (Buenos Aires) 66:113-118.) searching for work opportunities (Oyhenart et al., 2013Oyhenart EE, Garraza M, Bergel ML, Torres MF, Castro LE, Luis MA, Forte LM, Gamboa MI, Zonta ML, Cesani MF, et al. (2013) Caracterización del estado nutricional, enteroparasitosis y condiciones socio-ambientales de la población infanto-juvenil del partido de La Plata. Rev Arg Antrop Biol 15:47-60.). Moreover, it has been reported that a considerable proportion of the Argentinian population has at least one Indigenous American ancestor, in contrast to the perception of most Argentine people, who identify themselves as of European-descent, with only 1% of the total population self-identifying as descendants of an indigenous group (Avena et al., 2012Avena SA, Via M, Ziv E, Pérez-Stable EJ, Gignoux CR, Dejean CB, Huntsman S, Torres-Mejía G, Dutil J, Matta JL, et al. (2012) Heterogeneity in genetic admixture across different regions of Argentina. PLoS One 7:e34695.).
According to our results, Africans were the most distant from Native American, European, East Asian and Buenos Aires populations, in agreement with the age of divergence of African populations from the rest of the world (Cavalli-Sforza and Feldman, 2003Cavalli-Sforza LL and Feldman MW (2003) The application of molecular genetic approaches to the study of human evolution. Nat Genet 33:266-275.; Liu et al., 2006Liu H, Prugnolle F, Manica A and Balloux F (2006) A geographically explicit genetic model of worldwide human-settlement history. Am J Hum Genet 79:230-237.; Reyes-Centeno et al., 2015Reyes-Centeno H, Hubbe M, Hanihara T, Stringer C and Harvati K (2015) Testing modern human out-of-Africa dispersal models and implications for modern human origins. J Hum Evol 87:95-106.). Similarities between Asians and Native Americans could be explained by an ancient common origin of both populations (Cavalli-Sforza and Feldman, 2003Cavalli-Sforza LL and Feldman MW (2003) The application of molecular genetic approaches to the study of human evolution. Nat Genet 33:266-275.) that share an absence of blue eyes (Beals and Hoijer, 1965Beals RL and Hoijer H (1965) An Introduction to Anthropology. Macmillan, New York, 788 p.).
Our results suggest that iris color SNPs should be analyzed for the population in which the system IrisPlex is going to be implemented. Another option is to select the SNPs to use in identification after an evaluation of the population under consideration. The differentiation found among populations is consistent with the fact that these markers are related to a phenotype with an important regional adaptive value.
We conclude that the genetic background for iris color in the Buenos Aires population is genetically different to that of other countries. The differences we found might be due to the admixed ethnic composition of Argentina. Buenos Aires presents a population identity that differentiates it from other populations of the world, and for that reason, the methods of analysis used in European populations should be carefully applied in this case. Our results emphasize the importance of analyzing the variation of these genes and their relationship with iris color, not only in this province, but along the Argentinian country for people identification based on iris color.
Acknowledgments
This work was supported by grants from CONICET (Argentina, PIP 2012-2015-0051, and PIP 2015-2017-00930) and UNLP (Argentina, Proyecto de Incentivos 2017). Diana Hohl was a fellow from Comisión de Investigaciones Científicas de la Provincia de Buenos Aires (April 2016 – March 2017) and is a fellow of CONICET. Brenda Bezus is a fellow from the Comisión de Investigaciones Científicas de la Provincia de Buenos Aires (Argentina). Cecilia Catanesi is a researcher from CONICET (Argentina). We thank Gabriela Paula Di Santo Meztler and Dr. Laura Glesmann (Lab. Diversidad Genética, IMBICE (CONICET-UNLP-CIC), La Plata, Argentina) for helping in collecting samples and technical support. Many thanks to Dr. Jeppe D. Andersen (Dept. of Forensic Medicine, Faculty of Health and Medical Sciences, University of Copenhagen, Denmark) for kindly supplying the files for running DIAT software, to Dr. Michael Hout (Dept. of Psychology, New Mexico State University) for kind help with statistical analysis, and to Dr. Marisa I. Bauzá (St. Joseph’s University, USA) and Dr. Alejandro J. Román (University of Pennsylvania, Dept. of Ophtalmology, Philadelphia, PA, USA) for revision of the manuscript.
References
- Andersen JD, Johansen P, Harder S, Christoffersen SR, Delgado MC, Henriksen ST, Nielsen MM, Sørensen E, Ullum H, Hansen T, et al. (2013) Genetic analyses of the human eye colours using a novel objective method for eye colour classification. Forensic Sci Int Genet 7:508-515.
- Armstrong RA (2014) When to use the Bonferroni correction. Ophthalmic Physiol Opt 34:502-508.
- Avena SA, Goycoechea AS, Dugoujon JM, Slepoy MG, Slepoy AS and Carnese FR (2001) Análisis antropogenético de los aportes indígena y africano en muestras hospitalarias de la Ciudad de Buenos Aires. Rev Arg Antrop Biol 3:79-99.
- Avena SA, Goicochea AS, Rey J, Dugoujon JM, Dejean CB and Carnese FR (2006) Mezcla génica en una muestra poblacional de la ciudad de Buenos Aires. Medicina (Buenos Aires) 66:113-118.
- Avena SA, Via M, Ziv E, Pérez-Stable EJ, Gignoux CR, Dejean CB, Huntsman S, Torres-Mejía G, Dutil J, Matta JL, et al. (2012) Heterogeneity in genetic admixture across different regions of Argentina. PLoS One 7:e34695.
- Beals RL and Hoijer H (1965) An Introduction to Anthropology. Macmillan, New York, 788 p.
- Beleza S, Johnson NA, Candille SI, Absher DM, Coram MA, Lopes J, Campos J, Inês Araújo I, Anderson TM, Vilhjálmsson BJ, et al. (2013) Genetic architecture of skin and eye color in an African-European admixed population. PLoS Genet 9:e1003372.
- Brues AM (1975) Rethinking human pigmentation. Am J Phys Anthropol 43:387-391.
- Cavalli-Sforza LL and Feldman MW (2003) The application of molecular genetic approaches to the study of human evolution. Nat Genet 33:266-275.
- Cerqueira CCS, Paixao-Cortes VR, Zambra FMB, Salzano FM, Hünemeier T and Bortolini MC (2012) Predicting Homo pigmentation phenotype through genomic data: From Neanderthal to James Watson. Am J Hum Biol 24:705-709.
- Cerqueira CCS, Hünemeier T, Gomez-Valdés J, Ramallo V, Volasko-Krause CD, Leal BAA, Vargas-Pinilla P, Ciconet Dornelles R, Longo D, Rothhammer F, et al. (2014) Implications of the admixture process in skin color molecular assessment. PLoS One 9:e96886.
- Corach D, Lao O, Bobillo C, van Der Gaag K, Zuniga S and Vermeulen M (2010) Inferring continental ancestry of Argentineans from autosomal, Y-chromosomal and mitochondrial DNA. Ann Hum Genet 74:65-76.
- Devoto F (2007) La inmigración de ultramar. In: Torrado S (ed) Población y Bienestar en la Argentina del Primero al Segundo Centenario, Tomo I. Edhasa, Buenos Aires, pp 531-548.
- Draus-Barini J, Walsh S, Pospiech E, Kupiec T, Glab H, Branicki W and Kayser M (2013) Bona fide colour: DNA prediction of human eye and hair colour from ancient and contemporary skeletal remains. Investig Genet 4:3.
- Duffy DL, Montgomery GW, Chen W, Zhao Z, Le L, James MR, Hayward NK, Martin NG and Sturm RA (2007) A three-single-nucleotide polymorphism haplotype in intron 1 of OCA2 explains most human eye-color variation. Am J Hum Genet 80:241-252.
- Eagle RC (1994) Congenital, developmental and degenerative disorders of the iris and ciliary body. In: Albert DM and Jakobiec FA (eds) Principles and Practice of Ophthalmology. Saunders, Philadephia, pp 367-388.
- Earl DA and vonHoldt BM (2012) STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359-361.
- Edwards M, Cha D, Krithika S, Johnson M, Cook G and Parra EJ (2015) Iris pigmentation as a quantitative trait: Variation in populations of European, East Asian and South Asian ancestry and association with candidate gene polymorphisms. Pigment Cell Melanoma Res 29:141-162.
- Eiberg H, Troelsen J, Nielsen M, Mikkelsen A, Mengel-From J, Kjaeret K and Hansen L (2008) Blue eye color in humans may be caused by a perfectly associated founder mutation in a regulatory element located within the HERC2 gene inhibiting OCA2 expression. Hum Genet 123:177-187.
- Excoffier L and Lischer HEL (2010) Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564-567.
- Fernandes Durso D, Bydlowski SP, Hutz MH, Magalhães TR, Suarez-Kurtz G and Junho Pena SD (2014) Association of genetic variants with self-assessed color categories in Brazilians. PLoS One 9:e83926.
- Frudakis T, Thomas M, Gaskin Z, Venkateswarlu K, Suresh Chandra K, Ginjupalli S, Gunturi S, Natrajan S, Ponnuswamy VK and Ponnuswamy KN (2003) Sequences associated with human iris pigmentation. Genetics 165:2071-2083.
- Gemmel N and Akiyama S (1996) An efficient method for the extraction of DNA from vertebrate tissues. Trends Genet 12:338-339.
- Grigore M and Avram A (2015) Iris colour classification scales - Then and now. Rom J Ophthalmol 59:29-33.
- Hart KL, Kimura SL, Mushailov V, Budimlija ZM, Prinz M and Wurmbach E (2013) Improved eye- and skin-color prediction based on 8 SNPs. Croat Med J 54:248-256.
- Hammer Ø, Harper DAT and Ryan PD (2001) PAST: Paleontological Statistics Software Package for Education and Data Analysis. Palaeontol Electronica, v. 4, p 9.
- Jurado-Medina LS, Muzzio M, Schwab M, Costantino ML, Barreto G and Bailliet G (2014) Human Y-chromosome SNP characterization by multiplex amplified product-length polymorphism analysis. Electrophoresis 35:1-4.
- Kanetsky PA, Swoyer J, Panossian S, Holmes R, Guerry D and Rebbeck TR (2002) A polymorphism in the Agouti Signaling Protein gene is associated with human pigmentation. Am J Hum Genet 70:770-775.
- Kastelic V and Drobnic K (2012) A single-nucleotide polymorphism (SNP) multiplex system: The association of five SNPs with human eye and hair color in the Slovenian population and comparison using a Bayesian network and logistic regression model. Croat Med J 53:401-408.
- Kastelic V, Pospiech E, Draus-Barini J, Branicki W and Drobnic K (2013) Prediction of eye color in the Slovenian population using the IrisPlex SNPs. Croat Med J 54:381-386.
- Kayser M and Schneider P (2009) DNA-based prediction of human externally visible characteristics in forensics: Motivations, scientific challenges, and ethical considerations. Forensic Sci Int Genet 3:154-161.
- Lim J and Oh B (2013) Allelic frequencies of 20 visible phenotype variants in the Korean population. Genomics Inform 11:93-96.
- Liu F, van Duijn K, Vingerling J, Hofman A, Uitterlinden AG, Janssens AC and Kayser M (2009) Eye color and the prediction of complex phenotypes from genotypes. Curr Biol 19:R192-R193.
- Liu F, Wollstein A, Hysi PG, Ankra-Badu GA, Spector TD, Park D, Zhu G, Larsson M, Duffy DL, Montgomery GW, et al. (2010) Digital quantification of human eye color highlights genetic association of three new loci. PLoS Genet 6:e1000934.
- Liu F, Walsh S and Kayser M (2014) Of sex and IrisPlex eye colour prediction: A reply to Martinez-Cadenas et al. Letter to the Editor. Forensic Sci Int Genet 9:e5-e6.
- Liu H, Prugnolle F, Manica A and Balloux F (2006) A geographically explicit genetic model of worldwide human-settlement history. Am J Hum Genet 79:230-237.
- Marino M, Bobillo C, Sala A and Corach D (2007) Evaluación de la subestructura genética en la población argentina evaluada por STRs autosómicos e Y-STRs. Rev Arg Antrop Biol 9:47.
- Martinez-Cadenas C, Peña-Chilet M, Ibarrola-Villava M and Ribas G (2013) Gender is a major factor explaining discrepancies in eye colour prediction based on HERC2/OCA2 genotype and the IrisPlex model. Forensic Sci Int Genet 7:453-460.
- Martinez-Cadenas C, Peña-Chilet M, Llorca-Cardeñosa MJ, Cervera R, Ibarrola-Villava M and Ribas G (2014) Gender and eye colour prediction discrepancies: A reply to criticisms. Letter to the Editor. Forensic Sci Int Genet 9:e7-e9.
- Martínez-Marignac VL, Bertoni B, Parra E and Bianchi NO (2004) Characterization of admixture in an urban sample from Buenos Aires, Argentina, using uniparentally and biparentally inherited genetic markers. Hum Biol 76:543-557.
- Mengel-From J, Wong TH, Morling N, Rees JL and Jackson IJ (2009) Genetic determinants of hair and eye colours in the Scottish and Danish populations. BMC Genetics 10:88.
- Merenson S (2015) Between brotherhood and exceptionalism: Processes of identification, social marking, and justification in Uruguayan immigration in Buenos Aires. Migraciones internacionales 8:9-37.
- Motti JMB, Muzzio M, Ramallo V, Kladniew BR, Alfaro EL, Dipierri JE, Bailliet G and Bravi CM (2013) Origen y distribución espacial de linajes maternos nativos en el noroeste y centro oeste argentinos. Rev Arg Antrop Biol 15:3-14.
- Nan H, Kraft P, Hunter D and Han J (2009) Genetic variants in pigmentation genes, pigmentary phenotypes, and risk of skin cancer in Caucasians. Int J Cancer 125:909-917.
- Oyhenart EE, Garraza M, Bergel ML, Torres MF, Castro LE, Luis MA, Forte LM, Gamboa MI, Zonta ML, Cesani MF, et al. (2013) Caracterización del estado nutricional, enteroparasitosis y condiciones socio-ambientales de la población infanto-juvenil del partido de La Plata. Rev Arg Antrop Biol 15:47-60.
- Pospiech E, Draus-Barini J, Kupiec T, Wojas-Pelc A and Branicki W (2011) Gene-gene interactions contribute to eye color variation in humans. J Hum Genet 56:447-455.
- Pritchard JK, Stephens M and Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945-959.
- Reyes-Centeno H, Hubbe M, Hanihara T, Stringer C and Harvati K (2015) Testing modern human out-of-Africa dispersal models and implications for modern human origins. J Hum Evol 87:95-106.
- Forti IRN and Young RJ (2016) Human commercial models’ eye colour shows negative frequency-dependent selection. PLoS One 11:e0168458.
- Ruiz Y, Phillips C, Gomez-Tato A, Alvarez-Dios J, Casares de Cal M, Cruz R, O. Maroñas O, Söchtig J, Fondevila M, Rodriguez-Cid MJ, et al. (2013). Further development of forensic eye color predictive tests. Forensic Sci Int Genet 7:28-40.
- Salas A, Jaime JC, Álvarez-Iglesias V and Carracedo Á (2008) Gender bias in the multiethnic genetic composition of central Argentina. J Hum Genet 53:662-674.
- Spichenok O, Budimlija ZM, Mitchell AA, Jenny A, Kovacevic L, Marjanovic D, Caragine T, Prinz M and Wurmbach E (2011) Prediction of eye and skin color in diverse populations using seven SNPs. Forensic Sci Int Genet 5:472-478.
- Sturm RA and Duffy DL (2012) Human pigmentation genes under environmental selection. Genome Biol 13:248.
- Sturm RA and Frudakis TN (2004) Eye colour: Portals into pigmentation genes and ancestry. Trends Genet 20:327-332.
- Sturm RA, Duffy DL, Zhao ZZ, Leite FPN, Stark MS, Hayward NK, Martin NG and Montgomery GW (2008) A single SNP in an evolutionary conserved region within intron 86 of the HERC2 gene determines human blue-brown eye color. Am J Hum Genet 82:424-431.
- Sulem P, Gudbjartsson DF, Stacey SN, Helgason A, Rafnar T, Magnusson KP, Manolescu A, Karason A, Palsson A, Thorleifsson G, et al. (2007) Genetic determinants of hair, eye and skin pigmentation in Europeans. Nat Genet 39:1443-1452.
- Sun HP, Lin Y and Pan CW (2014) Iris color and associated pathological ocular complications: A review of epidemiologic studies. Int J Ophthalmol 7:872-878.
- Tully G (2007) Genotype versus phenotype: Human pigmentation. Forensic Sci Int Genet 1:105-110.
- Underhill P, Jin L, Zemans R, Oefner PJ and Cavalli-Sforza LL (1996) A pre-Columbian Y chromosome-specific transition and its implications for human evolutionary history. Proc Natl Acad Sci U S A 93:196-200.
- Valenzuela RK, Henderson MS, Walsh MH, Garrison NA, Kelch JT, Cohen-Barak O, Erickson DT, Meaney FJ, Walsh JB, Cheng KC, et al. (2010) Predicting phenotype from genotype: Normal pigmentation. J Forensic Sci 55:315-322.
- Walsh S, Liu F, Ballantyne K, van Oven M, Lao O and Kayser M (2011a) IrisPlex: A sensitive DNA tool for accurate prediction of blue and brown eye colour in the absence of ancestry information. Forensic Sci Int Genet 5:170-180.
- Walsh S, Lindenbergh A, Zuniga SB, Sijen T, de Knijff P, Kayser M and Ballantyne KN (2011b) Developmental validation of the IrisPlex system: Determination of blue and brown iris colour for forensic intelligence. Forensic Sci Int Genet 5:464-471.
- Walsh S, Wollstein A, Liu F, Chakravarthy U, Rahu M, Seland JH, Soubrane G, Tomazzoli L, Topouzis F, Vingerling JR, et al. (2012) DNA-based eye colour prediction across Europe with the IrisPlex system. Forensic Sci Int Genet 6:330-340.
- Walsh S, Liu F, Wollstein A, Kovatsi L, Ralf A, Kosiniak-Kamysz A, Branicki W and Kayser M (2013) The HIrisPlex system for simultaneous prediction of hair and eye colour from DNA. Forensic Sci Int Genet 7:98-115.
- Wilde S, Timpson A, Kirsanow K, Kaiser E, Kayser M, Unterländer M, Hollfelder N, Potekhina ID, Schier W, Thomas MG, et al. (2014) Direct evidence for positive selection of skin, hair, and eye pigmentation in Europeans during the last 5,000 years. Proc Natl Acad Sci U S A 111:4832-4837.
Internet Resources
- INDEC (2010) Instituto Nacional de Estadísticas y Censos, http://www.indec.gob.ar/censos_total_pais.asp?id_tema_1=2&id_tema_2=41&id_tema_3=135&t=3&s=8&c=2010 (accessed July 2016).
» http://www.indec.gob.ar/censos_total_pais.asp?id_tema_1=2&id_tema_2=41&id_tema_3=135&t=3&s=8&c=2010
-
Associate Editor: Francisco Mauro Salzano
Publication Dates
-
Publication in this collection
Jan-Mar 2018
History
-
Received
07 June 2017 -
Accepted
31 Aug 2017