ABSTRACT
Despite tremendous advancements in the livestock sector, additional opportunities exist to improve even further livestock production around the globe. Forecasting is not an exact science and it relies heavily on past and current knowledge. Improvements in the nutritional sciences (both human and animal) include a better understanding of agents that cause deterioration of human health, improving the quality of animal products, applying effective fetal programming, developing new feeds and feeding strategies, and revisiting longstanding technologies. Improvements in the understanding of the rumen microbiome will enable scientists to increase the fermentation efficiency and, hopefully, select microbial species of greater interest. Improvements in remote sensing and ground-based instrumentation, telecommunications, and weather forecasting technologies will aid in the continued improvements of early warning systems to assist livestock producers in reducing risk and adapting to the changing environment. Broad utilization of sensor technologies will allow scientists to collect real-time data and, when combined with mathematical modeling, decision support systems will become an indispensable managerial tool for livestock production with the possibility to automate low-level decisions on the farm, such as supplementation schedules, sorting of animals, and early detection of disease and outbreaks. The identification of feed efficient animals may be the single most impactful advancement towards long-term livestock sustainability and the promise of feeding the world animal products. We contend that education across societal levels is the first step to solve current and future challenges of the livestock industry. The dilemma has been who will take the first step forward.
Key Words:
forecasting; livestock; ruminant; solutions; production; vision
Introduction
The only certainty about forecasting the future is that it is a risky practice. Every so often, in many branches of science, we find ourselves predicting the future, but then later explaining what went wrong with our predictions. Planning ahead, however, is necessary to achieve successful stewardship towards the evolution of humankind. Forecasting involves knowledge of the past to predict the future, but this approach has some risks that are too often ignored. These risks exist mainly because even the most accurate and precise explanation of past events does not guarantee flawless future predictions; and yet, current scientific knowledge is the most valuable tool we have for forecasting. The question is: how science will progress in animal nutrition, especially meat production, in the next 40 years? Contemporary issues of a specific science field can, with a certain degree of accuracy, provide a good assessment of what the future holds.
Solutions to ameliorate many contemporary issues include growth-enhancement technologies (implants and hormones), bacteriophages, plant extracts as anaerobic fermentation and metabolic modifiers, improved nutrient requirements, selection for feed efficient animals, intestinal microbiome modification, sensor technologies, integrated modeling and simulation platforms, and the exploration and preservation of global animal biodiversity. However, animal scientists must keep up with cutting edge technology to improve animal production within the sustainable livestock intensification boundaries (i.e., economic, environment, social) (Tedeschi et al., 2015Tedeschi, L. O.; Muir, J. P.; Riley, D. G. and Fox, D. G. 2015. The role of ruminant animals in sustainable livestock intensification programs. International Journal of Sustainable Development & World Ecology 22: 452-465.), without forgetting that the main purpose of livestock production is to efficiently provide affordable, high-quality protein food to an estimated 9.55 billion people by 2050 (United Nations, 2013United Nations, Department of Economic and Social Affairs, Population Division. 2013. World Population Prospects; The 2012 Revision. Economics & Social Affairs. United Nations, New York, NY. Available at: <http://esa.un.org/unpd/wpp/index.htm>. Accessed on: Feb. 1, 2015.
http://esa.un.org/unpd/wpp/index.htm...
), while competing for prime resources (land, water, and energy) under a conceivable climate change scenario (Godfray et al., 2010Godfray, H. C. J.; Beddington, J. R.; Crute, I. R.; Haddad, L.; Lawrence, D.; Muir, J. F.; Pretty, J.; Robinson, S.; Thomas, S. M. and Toulmin, C. 2010. Food security: The challenge of feeding 9 billion people. Science 327: 812-818.).
In a companion paper, Tedeschi et al. (2017)Tedeschi, L. O.; Almeida, A. K.; Atzori, A. S.; Muir, J. P.; Fonseca, M. A. and Cannas, A. 2017. A glimpse of the future in animal nutrition science. 1. Past and future challenges. Revista Brasileira de Zootecnia 46: 438-451. discussed major challenges the animal industry has confronted in the past and challenges that will likely hassle the animal industry despite tremendous advancements and transformations in the livestock business. Briefly, the main public concerns are animal health and welfare, antibiotic resistance as a potential threat to human health, and food safety/quality have further intensified the challenge of producing animal-protein foods in an economic, environmentally, and socially sustainable manner. The Food and Agriculture Organization's (FAO) Climate Change, Agriculture, and Food Security research program (https://ccafs.cgiar.org/) provides additional information on this topic. This paper will focus on identifying possible tools and practices that can be used to solve the past and future challenges in the animal industry, especially in the beef cattle sector.
Current and future solutions
From the nutritional point of view, the principles of energy metabolism, nutrient utilization by animal tissues, and rumen microbiology have not changed since they were discovered 30 to 60 years ago (Ferrell and Oltjen, 2008Ferrell, C. L. and Oltjen, J. W. 2008. ASAS CENTENNIAL PAPER: Net energy systems for beef cattle—Concepts, application, and future models. Journal of Animal Science 86: 2779-2794.; Johnson et al., 2003Johnson, D. E.; Ferrell, C. L. and Jenkins, T. G. 2003. The history of energetic efficiency research: Where have we been and where are we going? Journal of Animal Science 81: E27-E38.; Poppi and McLennan, 2010Poppi, D. P. and McLennan, S. R. 2010. Nutritional research to meet future challenges. Animal Production Science 50: 329-338.), but the ways we collect data to develop and apply them have evolved. However, even with more data in hand, we are failing to use them appropriately. Formation of national committees to summarize scientific data and develop sound recommendations has fallen behind to unforeseen levels. The French feeding standard by the Institut National de la Recherche Agronomique (INRA) took 19 years to be revised (1988 to 2007); the Australian feeding standards by the Standing Committee on Agriculture took 17 years to be updated (1990 to 2007); the revisions of the United States’ Nutrient Requirements of Domestic Animals series by the National Research Council (NRC) and the National Academies of Sciences, Engineering, and Medicine (NASEM) was delayed 26 years for goats (1981 to 2007), 22 years for sheep (1985 to 2007), 20 years for beef (1996 to 2016), and likely 16 years for dairy (2001 to 2017). Many other feeding standards have simply not yet been revised despite tremendous progress in the fields of mathematical modeling and statistical analysis and data availability. In fact, we can collect more data today than ever. Big data has become a reality in all industrial and commercial sectors, including agriculture, and it is transforming how we see the world through sensor technologies, satellite imagery, global positioning system, near infrared, wireless communications, and many more technological advancements (Marr, 2015Marr, B. 2015. Big data: Using smart big data, analytics and metrics to make better decisions and improve performance. John Wiley & Sons Ltd, Chichester, West Sussex, United Kingdom.).
Markets for meat of grain-fed ruminants may face environmental and socio-cultural challenges in the future. Although these markets are currently centered in affluent regions such as North America, Australia, and Europe, they represent a socially and environmentally suspect production mode vis-à-vis grass-fed ruminants or grain-fed monogastrics throughout the world (Van Kernebeek et al., 2016Van Kernebeek, H. R. J.; Oosting, S. J.; De Boer, I. J. M.; Van Ittersum, M. K. and Bikker, P. 2016. Saving land to feed a growing population: consequences for consumption of crop and livestock products. International Journal of Life Cycle Assessment 21: 677-687.) and the possible negative effects on land degradation and biodiversity (Machovina et al., 2015Machovina, B.; Feeley, K. J. and Ripple, W. J. 2015. Biodiversity conservation: The key is reducing meat consumption. Science of The Total Environment 536: 419-431.). Understanding socio-cultural limitations and opportunities to reduce grain in sheep or beef feedlot diets will require multi-disciplinary research and education efforts that focus on socioeconomic questions as much as soil-plant-animal aspects (Steiner et al., 2014Steiner, J. L.; Coleman, S. W.; Starks, P. J.; Engle, D. M.; Xiao, X.; Saleh, A.; Osei, E.; Anandhi, A.; Tomlinson, P.; Rice, C. W.; Devlin, D.; Moffet, C.; Reuter, R.; Basara, J.; Cole, N. A.; Gowda, P.; Todd, R.; Middendorf, G. and Ocshner, T. 2014. Knowledge and tools to enhance resilience of beef grazing systems for sustainable animal protein production. Annals of the New York Academy of Sciences 1328: 10-17.). This could involve stepping back from the current beef and sheep focus on carcass and meat qualities to prioritize instead genetic selection for fiber conversion, pasture-use efficiency, and meat rather than fat content or tenderness.
Food production from grasslands can be far more resilient if the gap among research ideals, land manager realities, and market demands diminishes. The tools to span this gap encompass physical, social, and economic sciences that analyze not just production constraints, but also social and cultural factors (Steiner et al., 2014Steiner, J. L.; Coleman, S. W.; Starks, P. J.; Engle, D. M.; Xiao, X.; Saleh, A.; Osei, E.; Anandhi, A.; Tomlinson, P.; Rice, C. W.; Devlin, D.; Moffet, C.; Reuter, R.; Basara, J.; Cole, N. A.; Gowda, P.; Todd, R.; Middendorf, G. and Ocshner, T. 2014. Knowledge and tools to enhance resilience of beef grazing systems for sustainable animal protein production. Annals of the New York Academy of Sciences 1328: 10-17.). This will require the more direct participation of the land manager in identifying constraints, designing solutions, testing results, and marketing the benefits (Ndove et al., 2004Ndove, T. S.; Whitbread, A. M.; Clark, R. A. and Pengelly, B. C. 2004. Identifying the factors that contribute to the successful adoption of improved farming practices in the smallholder sector of Limpopo Province, South Africa. p. 146-153. In: Tropical legumes for sustainable farming systems in Southern Africa and Australia. ACIAR (Australian Centre for International Agricultural Research) Proceedings Canberra, Australia. ACIAR. Available at: <http://aciar.gov.au/publication/pr115>. Accessed on: May 12, 2016.
http://aciar.gov.au/publication/pr115...
), especially in mixed farming systems that are already important in much of the world and may become more important in other areas that currently focus on monocultures (González-García et al., 2012González-García, E.; Gourdine, J. L.; Alexandre, G.; Archimède, H. and Vaarst, M. 2012. The complex nature of mixed farming systems requires multidimensional actions supported by integrative research and development efforts. Animal 6: 763-777.).
Many of the issues highlighted by Tedeschi et al. (2017)Tedeschi, L. O.; Almeida, A. K.; Atzori, A. S.; Muir, J. P.; Fonseca, M. A. and Cannas, A. 2017. A glimpse of the future in animal nutrition science. 1. Past and future challenges. Revista Brasileira de Zootecnia 46: 438-451. and discussed above behave like a wicked problem; they have “… the essential characteristic that it is not solvable; it can only be managed” (Rittel and Webber, 1973Rittel, H. W. J. and Webber, M. M. 1973. Dilemmas in a general theory of planning. Policy Sciences 4: 155-169.). This type of problem requires the combination of knowledge, innovation, and transdisciplinary scholarship to be dealt with (Peterson, 2013Peterson, H. C. 2013. Sustainability: a wicked problem. p. 1-9. In: Sustainable animal agriculture. Kebreab, E., ed CABI Publishing, Wallingford, UK.). Next, we highlight some ideas that might provide opportunities to deal with these issues.
Nutritional and feeding opportunities
The discussion surrounding nutritional and feed opportunities relies much on what the prospects of future changes within the livestock industry are. For instance, beef productivity has increased substantially in the past 50 years through the adoption of grain-feeding production systems; nutrition, reproductive, and pharmaceutical-based technologies; and crossbreeding and selection programs focused on output traits (Elam and Preston, 2004Elam, T. E. and Preston, R. L. 2004. Fifty years of pharmaceutical technology and its Impact on the beef we provide to consumers. Growth Enhancement Technology Information Team, Johnson, IA. 36p. Available at: <http://www.merck-animal-health-usa.com/binaries/50_Years_of_Technology_and_impact_on_beef_production_tcm96-113484.pdf>. Accessed on: May 21, 2016.
http://www.merck-animal-health-usa.com/b...
). In contrast to poultry and pork industries, advances in beef productivity have been achieved in the absence of direct selection to improve feed efficiency. Poppi and McLennan (2010)Poppi, D. P. and McLennan, S. R. 2010. Nutritional research to meet future challenges. Animal Production Science 50: 329-338. recommended specific areas for nutritional research to meet future challenges such as production systems to meet market weight for age specifications, growth paths and compensatory growth, skeletal growth, parasite control, supplementation of fatty acid isomers, adaptation to low crude protein (CP) diets, rumen microbial ecology, epigenetics, remote data acquisition and animal management, greenhouse gas (GHG) emissions, and C balance of various production systems. In the nutrition research aspects, the NASEM (2016)NASEM - National Academies of Sciences, Engineering, and Medicine. 2016. Nutrient requirements of beef cattle. 8th ed. Nutrient requirements of domestic animals. National Academy Press, Washington, DC. listed many future areas of needed information, including better characterization of dietary carbohydrates and lipids; protein requirements for maintenance and growth and their use efficiency; relationships between energy for maintenance and environmental factors (e.g., global warming, GHG emission); and beef quality and safety. The increased public pressure on governmental agencies to more closely regulate air and water quality, food safety issues, GHG emissions, and animal welfare of the perceived “factory farming” has directed some research related to nutrition and feeding of livestock, such as animal traceability and liability associated with foodborne pathogens, use of pharmacological technologies, and concentrated feeding operations in the United States (Galyean et al., 2011Galyean, M. L.; Ponce, C. and Schutz, J. 2011. The future of beef production in North America. Animal Frontiers 1: 29-36.).
The beef industry in North America may have to increase the grazing period of animals or feed more than the current 10% of forage (Vasconcelos and Galyean, 2007Vasconcelos, J. T. and Galyean, M. L. 2007. Nutritional recommendations of feedlot consulting nutritionists: The 2007 Texas Tech University survey. Journal of Animal Science 85: 2772-2781.) in the feedyards to keep up with “organically produced” beef. This activity, however, would increase methane (CH4) emission and the land required to produce the same amount of beef (Capper and Cady, 2010Capper, J. L. and Cady, R. A. 2010. The environmental impact of corn- fed vs. grass-fed beef finishing systems. p. 686. In: Proceedings of the Joint Annual Meeting of American Society of Animal Science and American Dairy Science Association, Denver, CO. Federation of Animal Science Societies/Federation of Animal Science Societies. Available at: <http://www.jtmtg.org/JAM/2010/toc.asp>. Accessed on: May 21, 2016.
http://www.jtmtg.org/JAM/2010/toc.asp...
). Nonetheless, the notion that the beef industry in this region of the world is based on grain-based diets is not entirely correct; about 81% of the total feed needed to finish one U.S. steer comes from forage when considering the complete production cycle (cow and calf) (NASEM, 2016NASEM - National Academies of Sciences, Engineering, and Medicine. 2016. Nutrient requirements of beef cattle. 8th ed. Nutrient requirements of domestic animals. National Academy Press, Washington, DC.).
Innovative supplementation, especially protein, for grazing animals will become even more important as other industries increase competition for the same resources, raising commodity price and decreasing availability. Novel protein sources, such as on-farm produced algae (Poppi and McLennan, 2010Poppi, D. P. and McLennan, S. R. 2010. Nutritional research to meet future challenges. Animal Production Science 50: 329-338.), might prove worthy of investment by the livestock industry.
Do growth-enhancement additives and other pharmaceutical drugs play a role in the future? Innovative drugs such as 3-nitrooxypropanol (Romero-Perez et al., 2014Romero-Perez, A.; Okine, E. K.; McGinn, S. M.; Guan, L. L.; Oba, M.; Duval, S. M.; Kindermann, M. and Beauchemin, K. A. 2014. The potential of 3-nitrooxypropanol to lower enteric methane emissions from beef cattle. Journal of Animal Science 92: 4682-4693.) are promising opportunities to not only decrease CH4 emission, but also to increase energy availability for growth. Other “natural” products, such as condensed tannins, saponins, and alkaloids, should be further investigated (Tedeschi et al., 2011Tedeschi, L. O.; Callaway, T. R.; Muir, J. P. and Anderson, R. 2011. Potential environmental benefits of feed additives and other strategies for ruminant production. Revista Brasileira de Zootecnia 40: 291-309.). It is worthwhile to revisit “old” technologies to solve current and future problems, such as alkali or probiotic treatment of straw to increase digestibility or feeding hot-climate tolerant plants (e.g., cassava, cactus leaves) to livestock.
Rediscovering the grazing ruminant
There are many distinct morphophysiological attributes that assure ruminants tremendous evolutionary advantage for grazing and browsing when compared to non-ruminants, making them well adapted to thrive on grasslands under many diverse ecosystems (Gordon and Prins, 2008Gordon, I. J. and Prins, H. H. T. 2008. The ecology of browsing and grazing. Ecological studies. vol. 195. Springer, Berlin, Germany.). However, numerous benefits derive from healthy grasslands beyond simple ruminant nutrition. These include stable soils and hydrology, protected biodiversity, climate change mitigation, and even socio-cultural services (Boval and Dixon, 2012Boval, M. and Dixon, R. M. 2012. The importance of grasslands for animal production and other functions: a review on management and methodological progress in the tropics. Animal 6: 748-762.). Our challenge in the future will be to catalogue and quantify these to vouchsafe ecosystems dominated by ruminants, whether native or domesticated. Meat is only one benefit from ecosystems in which ruminants thrive. Researching and divulging fine-tuned grassland management that avoids issues, such as over-grazing or invasive species, can enhance those ecosystem services. The Brazilian generally low input browse-(goats) and grass-fed (beef) production systems (Lobato et al., 2014Lobato, J. F. P.; Freitas, A. K.; Devincenzi, T.; Cardoso, L. L.; Tarouco, J. U.; Vieira, R. M.; Dillenburg, D. R. and Castro, I. 2014. Brazilian beef produced on pastures: Sustainable and healthy. Meat Science 98: 336-345.), if fine-tuned to address other ecosystem concerns such as deforestation or surface-water quality, could provide a useful guide.
Although legumes have long been promoted as a sustainable protein source for accompanying grasses and grazing ruminants in native, rangeland, or cultivated grasslands (Shelton et al., 2005Shelton, H. M.; Franzel, S. and Peters, M. 2005. Adoption of tropical legume technology around the world: analysis of success. Tropical Grasslands 39: 198-209.), their adoption has been sporadic (Muir et al., 2014Muir, J. P.; Pitman, W. D.; Dubeux, J. C. and Foster, J. L. 2014. The future of warm-season, tropical and subtropical forage legumes in sustainable pastures and rangelands. African Journal of Range & Forage Science 31: 187-198.). Challenges that researchers and land managers alike have failed to address include establishment and competition in mixtures with grasses (Muir et al., 2011Muir, J. P.; Pitman, W. D. and Foster, J. L. 2011. Sustainable, low- input, warm-season, grass-legume grassland mixtures: mission (nearly) impossible? Grass and Forage Science 66: 301-315.), as well as persistence under grazing (Muir et al., 2014Muir, J. P.; Pitman, W. D.; Dubeux, J. C. and Foster, J. L. 2014. The future of warm-season, tropical and subtropical forage legumes in sustainable pastures and rangelands. African Journal of Range & Forage Science 31: 187-198.). Future emphasis on supplementing ruminants with forage legumes as protein sources rather than industrial sources (e.g., urea) or pulses (e.g., soybeans) may solve multiple concerns including human food scarcity, fossil fuel consumption, marginal farmland cultivation, water quality, biodiversity, and GHG emissions, to name but a few.
Integrating multiple canopies into native, rangeland, or cultivated pastures can improve ruminant nutrition (Singh and Kundu, 2010Singh, S. and Kundu, S. S. 2010. Effect of tropical browse leaves supplementation on rumen enzymes of sheep and goats fed Dichanthium annulatum grass-based diets. Tropical Animal Health and Production 42: 1181-1187.), especially in the case of legumes. These shrubs and trees not only provide forage that is generally more digestible and greater in CP content than grasses, but they also fill feed gaps in seasons or drought years when shallow-rooted herbaceous plants fail to grow (Dubeux et al., 2015Dubeux, J. C. B., Jr.; Muir, J. P.; Nair, P. K. R.; Sollenberger, L. E.; Silva, H. M. S. and Mello, A. C. L. 2015. The advantages and challenges of integrating tree legumes into pastoral systems. p. 141-164. In: Proceedings of the 1st International Conference on Forages in Warm Climates, Lavras, Brazil. University of Lavras. Available at: <http://www.neforufla.com.br/index.php/en/pg/9/proceedings-confor-2015>. Accessed on: May 13, 2016.
http://www.neforufla.com.br/index.php/en...
). They also provide multiple additional benefits, products, and services to the environment, ruminants, and land managers (Franzel et al., 2014Franzel, S.; Carsan, S.; Lukuyu, B.; Sinja, J. and Wambugu, C. 2014. Fodder trees for improving livestock productivity and smallholder livelihoods in Africa. Current Opinion in Environmental Sustainability 6: 98-103.).
Rouquette et al. (2009)Rouquette, F. M., Jr.; Redmon, L. A.; Aiken, G. E.; Hill, G. M.; Sollenberger, L. E. and Andrae, J. 2009. ASAS Centennial Paper: Future needs of research and extension in forage utilization. Journal of Animal Science 87: 438-446. listed some research needs for the livestock production sector that rely on the animal-plant-soil interactions, including pasture systems and efficiency of production, interface with energy concerns, forage cultivar evaluations and persistence, and environmental impact. We contend that future challenges also include widening the range of germplasm currently available, fine-tuning the interaction among various canopies, and quantifying the nutritive value to ruminant production, including greater digestibility of fibrous, warm-climate C4 grasses to a wider range of species beyond beef cattle. Specific research opportunities include assessing of forage nutritive value and availability to enhance performance of livestock, better understanding of rumen bacteria and fiber components to improve intake and digestion of forages, and developing early detection performance for grazing animals for decision-making purposes (Rouquette et al., 2009Rouquette, F. M., Jr.; Redmon, L. A.; Aiken, G. E.; Hill, G. M.; Sollenberger, L. E. and Andrae, J. 2009. ASAS Centennial Paper: Future needs of research and extension in forage utilization. Journal of Animal Science 87: 438-446.). However, solutions for future livestock production that depend on the animal-plant-soil interactions are far beyond technical issues; they also include increasing the allocation of funding for scientists engaged in animal-forage teaching, research, and extension (Rouquette et al., 2009Rouquette, F. M., Jr.; Redmon, L. A.; Aiken, G. E.; Hill, G. M.; Sollenberger, L. E. and Andrae, J. 2009. ASAS Centennial Paper: Future needs of research and extension in forage utilization. Journal of Animal Science 87: 438-446.).
Early nutritional fetal programming
Many experiments have been conducted to quantify the requirements of energy, protein, and minerals of pregnant animals (Bell et al., 2005Bell, A. W.; Greenwood, P. L. and Ehrhardt, R. A. 2005. Regulation of metabolism and growth during prenatal life. p. 3-34. In: Biology of growing animals. vol. 3. Burrin, D. G. and Mersmann, H. J., eds. Elsevier, Amsterdam.; Ferrell, 1991Ferrell, C. L. 1991. Maternal and fetal influences on uterine and conceptus development in the cow: I. Growth of tissues of the gravid uterus. Journal of Animal Science 69: 1945-53.; Ferrell et al., 1976Ferrell, C. L.; Garrett, W. N. and Hinman, N. 1976. Growth, development and composition of the udder and gravid uterus of beef heifers during pregnancy. Journal of Animal Science 42: 1477-1489.) and the impact of early calfhood nutrition on their reproductive development (Allen et al., 2012Allen, C. C.; Alves, B. R. C.; Li, X.; Tedeschi, L. O.; Zhou, H.; Paschal, J. C.; Riggs, P. K.; Braga-Neto, U. M.; Keisler, D. H.; Williams, G. L. and Amstalden, M. 2012. Gene expression in the arcuate nucleus of heifers is affected by controlled intake of high- and low- concentrate diets. Journal of Animal Science 90: 2222-2232.; Alves et al., 2015Alves, B. R. C.; Cardoso, R. C.; Prezotto, L. D.; Thorson, J. F.; Bedenbaugh, M.; Sharpton, S. M.; Caraty, A.; Keisler, D. H.; Tedeschi, L. O.; Williams, G. L. and Amstalden, M. 2015. Elevated body weight gain during the juvenile alters neuropeptide Y-gonadotropin-releasing hormone circuitry in prepubertal heifers. Biology of Reproduction 92: 1-10.; Amstalden et al., 2014Amstalden, M.; Cardoso, R. C.; Alves, B. R. C. and Williams, G. L. 2014. Reproduction Symposium: Hypothalamic neuropeptides and the nutritional programming of puberty in heifers. Journal of Animal Science 92: 3211-3222.; Cardoso et al., 2014aCardoso, R. C.; Alves, B. R. C.; Prezotto, L. D.; Thorson, J. F.; Tedeschi, L. O.; Keisler, D. H.; Amstalden, M. and Williams, G. L. 2014a. Reciprocal changes in leptin and NPY during nutritional acceleration of puberty in heifers. Journal of Endocrinology 223: 289-298.; Cardoso et al., 2014bCardoso, R. C.; Alves, B. R. C.; Prezotto, L. D.; Thorson, J. F.; Tedeschi, L. O.; Keisler, D. H.; Park, C. S.; Amstalden, M. and Williams, G. L. 2014b. Use of a stair-step compensatory gain nutritional regimen to program the onset of puberty in beef heifers. Journal of Animal Science 92: 2942-2949.). However, not until recently have scientists inquired about controlling the type and quality of the nutrition during fetus development and its long-lasting impacts on the growth and development of the newborn through adulthood: the fetal programming theory. Fetal programming is the process by which a positive (stimulus) or a negative (insult) signal, given during a critical time of the fetus development, will promote permanent changes on the structure, physiology, or metabolism of organs or systems that will ultimately influence the development of offspring (Mossa et al., 2015Mossa, F.; Walsh, S. W.; Ireland, J. J. and Evans, A. C. O. 2015. Early nutritional programming and progeny performance: Is reproductive success already set at birth? Animal Frontiers 5: 18-24.). Gallo et al. (2013)Gallo, L. A.; Tran, M.; Moritz, K. M. and Wlodek, M. E. 2013. Developmental programming: Variations in early growth and adult disease. Clinical & Experimental Pharmacology & Physiology 40: 795-802. believe, however, that in utero adverse exposures not only determines the health and performance of the unborn animal, but it also has far-reaching implications beyond the first, directly exposed generation. In the UK, for example, longer human life expectancy between 1751 and 1930 was attributed to improved childhood living conditions and the analysis of a series of other events led scientists to believe that environmental factors that impaired growth and development in early life resulted in increased risk of heart disease, culminating in the postulation of the “fetal or developmental origins” hypothesis (McMillen and Robinson, 2005McMillen, I. C. and Robinson, J. S. 2005. Developmental origins of the metabolic syndrome: Prediction, plasticity, and programming. Physiological Reviews 85: 571-633.). Since the 1970s, animal scientists have recognized that metabolic alterations during the prenatal nutrition would impact postnatal productivity and possibly the health of livestock (Bell, 2006Bell, A. W. 2006. Prenatal programming of postnatal productivity and health of livestock: a brief review. Australian Journal of Experimental Agriculture 46: 725-732.), but no one has attempted to manipulate the maternal nutrition to “program” the fetus for postnatal performance.
Experimental evidence of fetal programming using livestock has been reported. From an endocrinological perspective, evidence includes predisposition of growth-retarded neonates to develop insulin resistance, high plasma growth hormone concentrations in low-birth-weight lambs, and high plasma leptin concentration in rapidly fattening, low-birth-weight lambs right after postpartum (Bell et al., 2005Bell, A. W.; Greenwood, P. L. and Ehrhardt, R. A. 2005. Regulation of metabolism and growth during prenatal life. p. 3-34. In: Biology of growing animals. vol. 3. Burrin, D. G. and Mersmann, H. J., eds. Elsevier, Amsterdam.). From a practical perspective, as reported by Bell (2006)Bell, A. W. 2006. Prenatal programming of postnatal productivity and health of livestock: a brief review. Australian Journal of Experimental Agriculture 46: 725-732., prolonged ewe undernutrition before 110 days of gestation increases fetal adipose tissue and might increase the propensity of fatter yearling lambs; low-birth-weight lambs also had slower skeletal muscle growth. Epigenetic modifications of the genome through modifications in cytosine methylation in the promoter region of genes that prevent transcription and chromatin (i.e., DNA) remodeling (e.g., methylation, acetylation, and phosphorylation of histone proteins) is responsible for the fetal programming phenomenon, inducing different phenotypes (Gabory et al., 2009Gabory, A.; Attig, L. and Junien, C. 2009. Sexual dimorphism in environmental epigenetic programming. Molecular and Cellular Endocrinology 304: 8-18.; Langley-Evans, 2006Langley-Evans, S. C. 2006. Developmental programming of health and disease. Proceedings of the Nutrition Society 65: 97-105.) and health outcomes in the adult phase (Moritz et al., 2008Moritz, K.; Wintour-Coghlan, E. M.; Black, M. J.; Bertram, J. F. and Caruana, G. 2008. Factors Influencing Mammalian Kidney Development: Implications for Health in Adult Life. Advances in Anatomy, Embryology and Cell Biology. vol. 196. Springer Berlin Heidelberg, New York, NY.).
Implications of fetal programming for the future of livestock production are immense, but feasible and relevant applications have yet to be discovered. Poppi and McLennan (2010)Poppi, D. P. and McLennan, S. R. 2010. Nutritional research to meet future challenges. Animal Production Science 50: 329-338. pointed out that benefits of fetal programming for human biomedical purposes do not necessarily translate into an immediate benefit to livestock production. The latter is more concerned with increased growth performance and carcass composition (Du et al., 2010Du, M.; Tong, J.; Zhao, J.; Underwood, K. R.; Zhu, M.; Ford, S. P. and Nathanielsz, P. W. 2010. Fetal programming of skeletal muscle development in ruminant animals. Journal of Animal Science 88: E51-E60.) and reproductive development and function (Mossa et al., 2015Mossa, F.; Walsh, S. W.; Ireland, J. J. and Evans, A. C. O. 2015. Early nutritional programming and progeny performance: Is reproductive success already set at birth? Animal Frontiers 5: 18-24.) than insulin resistance (Gallo et al., 2013Gallo, L. A.; Tran, M.; Moritz, K. M. and Wlodek, M. E. 2013. Developmental programming: Variations in early growth and adult disease. Clinical & Experimental Pharmacology & Physiology 40: 795-802.) and kidney development (Moritz et al., 2008Moritz, K.; Wintour-Coghlan, E. M.; Black, M. J.; Bertram, J. F. and Caruana, G. 2008. Factors Influencing Mammalian Kidney Development: Implications for Health in Adult Life. Advances in Anatomy, Embryology and Cell Biology. vol. 196. Springer Berlin Heidelberg, New York, NY.). Despite the scientific knowledge gained throughout these years, the discovery and elucidation phase of fetal programming is in its infancy and we will face a long and steep road ahead until we can securely make recommendations.
Selection strategies for efficient meat production
Remarkably, there is little evidence to indicate that genetic merit for feed efficiency or energy requirement maintenance in beef cattle have been favorably altered in the past 50 years (Archer et al., 1999Archer, J. A.; Richardson, E. C.; Herd, R. M. and Arthur, P. F. 1999. Potential for selection to improve efficiency of feed use in beef cattle: a review. Australian Journal of Agricultural Research 50: 147-161.; Johnson et al., 2003Johnson, D. E.; Ferrell, C. L. and Jenkins, T. G. 2003. The history of energetic efficiency research: Where have we been and where are we going? Journal of Animal Science 81: E27-E38.). Since about two-thirds of the cost of producing beef is due to the expense of feed inputs, strategies that improve efficiency of feed utilization will substantially increase the economic viability of beef production systems. In fact, Weaber (apud Perkins et al., 2014Perkins, S. D.; Key, C. N.; Garrett, C. F.; Foradori, C. D.; Bratcher, C. L.; Kriese-Anderson, L. A. and Brandebourg, T. D. 2014. Residual feed intake studies in Angus-sired cattle reveal a potential role for hypothalamic gene expression in regulating feed efficiency. Journal of Animal Science 92: 549-560.) estimated that the U.S. beef industry could save $1 billion annually by reducing residual feed intake (RFI) by 10% (equivalent to reducing daily intake by 0.9 kg per animal). Furthermore, as improvements in feed efficiency will also reduce nutrient excretions and GHG emissions (Waghorn and Hegarty, 2011Waghorn, G. C. and Hegarty, R. S. 2011. Lowering ruminant methane emissions through improved feed conversion efficiency. Animal Feed Science and Technology 166-167: 291-301.), discovery and adoption of technologies to enhance genetic merit for feed efficiency is arguably one of the most cost-effective strategies available to meet future demands for animal protein in a more sustainable manner. While considerable genetic variation in feed efficiency both within and among beef cattle populations is known to exist, the absence of genetic progress is not surprising, given the industry's focus on output traits, cost of measuring feed intake, and the complex interactions that exist between various biotypes and production environments. The lack of an appropriate trait for use in selection programs has also curtailed genetic progress in feed efficiency. In beef cattle, inter-animal variation in RFI has been linked to differences in heat production, CH4, composition of gain, and digestibility, demonstrating that RFI is a complex trait controlled by numerous biological processes and thereby regulated by a large number of divergent genes (Bottje and Carstens, 2009Bottje, W. G. and Carstens, G. E. 2009. Association of mitochondrial function and feed efficiency in poultry and livestock species. Journal of Animal Science 87: E48-63.; Herd and Arthur, 2009Herd, R. M. and Arthur, P. F. 2009. Physiological basis for residual feed intake. Journal of Animal Science 87: E64-71.). Numerous studies have documented that cattle with low RFI phenotypes tend to be leaner. In growing bulls and heifers fed moderate-energy diets, genetic correlations between RFI and carcass backfat depth were weakly positive (Arthur et al., 2001Arthur, P. F.; Archer, J. A.; Johnson, D. J.; Herd, R. M.; Richardson, E. C. and Parnell, P. F. 2001. Genetic and phenotypic variance and covariance components for feed intake, feed efficiency, and other postweaning traits in Angus cattle. Journal of Animal Science 79: 2805-2811.; Lancaster et al., 2009Lancaster, P. A.; Carstens, G. E.; Crews, D. H., Jr.; Welsh Jr., T. H.; Forbes, T. D. A.; Forrest, D. W.; Tedeschi, L. O.; Randel, R. D. and Rouquette, F. M., Jr. 2009. Phenotypic and genetic relationships of residual feed intake with performance and ultrasound carcass traits in Brangus heifers. Journal of Animal Science 87: 3887-3896.; Schenkel et al., 2004Schenkel, F. S.; Devitt, C. J. B.; Wilton, J. W.; Miller, S. P. and Jamrozik, J. 2004. Random regression analyses of feed intake of individually tested beef steers. Livestock Production Science 88: 129-142.), whereas, in steers fed finishing diets, genetic correlations between RFI and backfat depth tended to be more moderate (Robinson and Oddy, 2004Robinson, D. L. and Oddy, V. H. 2004. Genetic parameters for feed efficiency, fatness, muscle area and feeding behaviour of feedlot finished beef cattle. Livestock Production Science 90: 255-270.). Zorzi et al. (2013)Zorzi, K.; Bonilha, S. F. M.; Queiroz, A. C.; Branco, R. H.; Sobrinho, T. L. and Duarte, M. S. 2013. Meat quality of young Nellore bulls with low and high residual feed intake. Meat Science 93: 593-599. found that low-RFI Nellore bulls had higher myofibril fragmentation indexes and were tougher than high-RFI bulls. However, U.S. studies with Bos taurus cattle found that RFI was not associated phenotypically with calpastatin activity, shear force, or sensory panel tenderness scores of loin steaks (Baker et al., 2006Baker, S. D.; Szasz, J. I.; Klein, T. A.; Kuber, P. S.; Hunt, C. W.; Glaze Jr., J. B.; Falk, D.; Richard, R.; Miller, J. C.; Battaglia, R. A. and Hill, R. A. 2006. Residual feed intake of purebred Angus steers: Effects on meat quality and palatability. Journal of Animal Science 84: 938-945.; Behrens et al., 2011Behrens, J. W.; Miller, R. K.; Bailey, J. C. Walter, J. T.; Hafla, A. N.; Mendes, E. D.; Hale, D. S.; Machado, T.; Tedeschi, L. O. and Carstens, G. E. 2011. Effects of residual feed intake classification and breed type on carcass characteristics, tenderness and value in feedlot heifers. p. 761. In: Proceedings of the Joint Annual Meeting of American Society of Animal Science and American Dairy Science Association, 89, New Orleans, LA. Federation of Animal Science Societies. Available at: <http://www.jtmtg.org/JAM/2011/toc.asp>. Accessed on: May 21, 2016.
http://www.jtmtg.org/JAM/2011/toc.asp...
). Hafla et al. (2012b)Hafla, A. N.; Walter, J. T.; Carstens, G. E.; Bailey, J. C.; Tedeschi, L. O.; Moreno, J. G.; Hale, D. S.; Miller, R. K.; Sawyer, J. E. and Anderson, D. P. 2012b. Impact of between-animal variation in performance, carcass-quality and feed efficiency on profitability of Angus-based composite steers. p. 125-126. In: Proceedings of the Plains Nutrition Council Conference, San Antonio, TX. PNC. Available at: <http://amarillo.tamu.edu/facultystaff/tmc/pnc/>. Accessed on: May 9, 2017.
http://amarillo.tamu.edu/facultystaff/tm...
reported that low-RFI steers had lower yield grades (favorable) and quality grades (unfavorable) than high-RFI steers, but grid-formula-based carcass value was not affected by the RFI classification. Use of multi-trait selection indexes to identify terminal sires with superior genetic merit for RFI will improve efficiency and profitability of feedlot progeny with minimal effect on carcass value.
For North American calf-fed and yearling-fed integrated beef production systems, Basarab et al. (2012)Basarab, J.; Baron, V.; López-Campos, Ó.; Aalhus, J.; Haugen- Kozyra, K. and Okine, E. 2012. Greenhouse gas emissions from calf- and yearling-fed beef production systems, with and without the use of growth promotants. Animals 2: 195-220. estimated that the cowherd (cows, bulls, and replacements) requires approximately 82 and 64%, respectively, of total feed inputs. Thus, attempts to improve efficiency of feed utilization and profitability of beef cattle operations will need to focus on reducing cowherd feed inputs relative to system outputs. Archer et al. (2002)Archer, J. A.; Reverter, A.; Herd, R. M.; Johnston, D. J. and Arthur, P. F. 2002. Genetic variation in feed intake and efficiency of mature beef cows and relationships with post-weaning measurements. p. 221-224. In: Proceedings of the 7th World Congress Genetics Applied to Livestock Production, Montepellier, France. measured post-weaning RFI in Angus, Hereford, and Shorthorn heifers and again in the same females as mature cows. Strong genetic correlations were observed between post-weaning RFI of heifers and feed intake and RFI (rg = 0.64 and 0.98) of mature open cows, although the corresponding phenotypic correlations were lower (rp = 0.34 and 0.40, respectively). A low negative genetic correlation between heifer RFI and mature cow weight (rg = −0.22) was observed, indicating that favorable selection based on post-weaning RFI will improve efficiency of feed utilization in cows with minimal effects on mature size. These results and those from more recent studies (Arthur et al., 2005Arthur, P. F.; Herd, R. M.; Wilkins, J. F. and Archer, J. A. 2005. Maternal productivity of Angus cows divergently selected for post- weaning residual feed intake. Australian Journal of Experimental Agriculture 45: 985-993.; Basarab et al., 2007Basarab, J. A.; McCartney, D.; Okine, E. K. and Baron, V. S. 2007. Relationships between progeny residual feed intake and dam productivity traits. Canadian Journal of Animal Science 87: 489-502.; Black et al., 2013Black, T. E.; Bischoff, K. M.; Mercadante, V. R. G.; Marquezini, G. H. L.; Dilorenzo, N.; Chase, C. C., Jr.; Coleman, S. W.; Maddock, T. D. and Lamb, G. C. 2013. Relationships among performance, residual feed intake, and temperament assessed in growing beef heifers and subsequently as 3-year-old, lactating beef cows. Journal of Animal Science 91: 2254-2263.; Hafla et al., 2013Hafla, A. N.; Carstens, G. E.; Forbes, T. D. A.; Tedeschi, L. O.; Bailey, J. C.; Walter, J. T. and Johnson, J. R. 2013. Relationships between postweaning residual feed intake in heifers and forage use, body composition, feeding behavior, physical activity, and heart rate of pregnant beef females. Journal of Animal Science 91: 5353-5365.; MacDonald et al., 2014MacDonald, K. A.; Pryce, J. E.; Spelman, R. J.; Davis, S. R.; Wales, W. J.; Waghorn, G. C.; Williams, Y. J.; Marett, L. C. and Hayes, B. J. 2014. Holstein-Friesian calves selected for divergence in residual feed intake during growth exhibited significant but reduced residual feed intake divergence in their first lactation. Journal of Dairy Science 97: 1427-1435.) demonstrate that post-weaning RFI of heifers is favorably associated phenotypically with efficient feed utilization by gestating and lactating cows, with minimal effects on productivity or reproductive performance. However, favorable selection for RFI may delay the onset of puberty in heifers, thereby increasing the age at first conception without negatively affecting subsequent reproductive performance of cows (Arthur et al., 2005Arthur, P. F.; Herd, R. M.; Wilkins, J. F. and Archer, J. A. 2005. Maternal productivity of Angus cows divergently selected for post- weaning residual feed intake. Australian Journal of Experimental Agriculture 45: 985-993.; Basarab et al., 2007Basarab, J. A.; McCartney, D.; Okine, E. K. and Baron, V. S. 2007. Relationships between progeny residual feed intake and dam productivity traits. Canadian Journal of Animal Science 87: 489-502.; Donoghue et al., 2011Donoghue, K. A.; Arthur, P. F.; Wilkins, J. F. and Herd, R. M. 2011. Onset of puberty and early-life reproduction in Angus females divergently selected for post-weaning residual feed intake. Animal Production Science 51: 183-190.). In support of these findings, Crowley et al. (2011)Crowley, J. J.; Evans, R. D.; McHugh, N.; Kenny, D. A.; McGee, M.; Crews, D. H. and Berry, D. P. 2011. Genetic relationships between feed efficiency in growing males and beef cow performance. Journal of Animal Science 89: 3372-3381. reported that RFI of bulls was not genetically correlated with the interval from calving to first service (rg = −0.03) or calving interval (rg = 0.01), but was negatively correlated (rg = −0.29) with age at first calving. Basarab et al. (2011)Basarab, J. A.; Colazo, M. G.; Ambrose, D. J.; Novak, S.; McCartney, D. and Baron, V. S. 2011. Residual feed intake adjusted for backfat thickness and feeding frequency is independent of fertility in beef heifers. Canadian Journal of Animal Science 91: 573-584. hypothesized that current protocols that measure RFI from 8-to-12-month olds may indirectly favor selection of slightly later maturing animals based on the premise that animals reaching puberty at the beginning of a test will have higher energy expenditures associated with sexual development and activity compared to contemporaries that reach puberty later. Although Basarab et al. (2011)Basarab, J. A.; Colazo, M. G.; Ambrose, D. J.; Novak, S.; McCartney, D. and Baron, V. S. 2011. Residual feed intake adjusted for backfat thickness and feeding frequency is independent of fertility in beef heifers. Canadian Journal of Animal Science 91: 573-584. originally observed lower pregnancy rates in low RFI heifers, these differences were no longer evident when RFI was adjusted for variation in backfat depth and feeding event frequency. These results imply that inter-animal variances in body-fat reserves and activity associated with stage of sexual development may need to be considered in assessing RFI of breeding heifers and bulls (Awda et al., 2013Awda, B. J.; Miller, S. P.; Montanholi, Y. R.; Voort, G. V.; Caldwell, T.; Buhr, M. M. and Swanson, K. C. 2013. The relationship between feed efficiency traits and fertility in young beef bulls. Canadian Journal of Animal Science 93: 185-192.; Hafla et al., 2012aHafla, A. N.; Lancaster, P. A.; Carstens, G. E.; Forrest, D. W.; Fox, J. T.; Forbes, T. D. A.; Davis, M. E.; Randel, R. D. and Holloway, J. W. 2012a. Relationships between feed efficiency, scrotal circumference, and semen quality traits in yearling bulls. Journal of Animal Science 90: 3937-3944.; Wang et al., 2012Wang, Z.; Colazo, M. G.; Basarab, J. A.; Goonewardene, L. A.; Ambrose, D. J.; Marques, E.; Plastow, G.; Miller, S. P. and Moore, S. S. 2012. Impact of selection for residual feed intake on breeding soundness and reproductive performance of bulls on pasture-based multisire mating. Journal of Animal Science 90: 2963-2969.) to ensure that favorable selection for RFI does not negatively affect long-term reproductive performance of beef cattle.
The adoption of appropriate multi-trait selection indexes to identify cattle with superior genetic merit for RFI has considerable potential to improve life-cycle efficiency and profitability of production systems through reductions in costs of feed inputs without compromising the value of carcass outputs and reproductive efficiency. Moreover, substantial reductions in manure N and P excretion and GHG emissions can be achieved through implementation of RFI-based selection indices (Basarab et al., 2012Basarab, J.; Baron, V.; López-Campos, Ó.; Aalhus, J.; Haugen- Kozyra, K. and Okine, E. 2012. Greenhouse gas emissions from calf- and yearling-fed beef production systems, with and without the use of growth promotants. Animals 2: 195-220.). While our understanding of RFI in growing cattle has advanced in recent years, our knowledge of the associations between RFI in growing cattle and efficiency of mature cows is limited. Moreover, the relative ranking of phenotypic RFI in growing cattle can vary when animals are switched from low- to high-energy diets (Durunna et al., 2011Durunna, O. N.; Mujibi, F. D. N.; Goonewardene, L.; Okine, E. K.; Basarab, J. A.; Wang, Z. and Moore, S. S. 2011. Feed efficiency differences and reranking in beef steers fed grower and finisher diets. Journal of Animal Science 89: 158-167.). Given the increasing use of RFI to identify feed-efficient cattle, it will be important to better understand how phenotypic RFI is affected by changes in diet, climatic conditions, and stage of maturity.
Unfortunately, the cost of measuring feed intake remains a key barrier to widespread adoption of selection programs that incorporate RFI, which has prompted numerous candidate-gene-approach (Karisa et al., 2013Karisa, B. K.; Thomson, J.; Wang, Z.; Stothard, P.; Moore, S. S. and Plastow, G. S. 2013. Candidate genes and single nucleotide polymorphisms associated with variation in residual feed intake in beef cattle. Journal of Animal Science 91: 3502-3513.) and genome-wide-association (Rolf et al., 2012Rolf, M. M.; Taylor, J. F.; Schnabel, R. D.; McKay, S. D.; McClure, M. C.; Northcutt, S. L.; Kerley, M. S. and Weaber, R. L. 2012. Genome-wide association analysis for feed efficiency in Angus cattle. Animal Genetics 43: 367-374.) studies to identify RFI quantitative trait locus for marker-assisted selection programs. Although these studies have generated informative single nucleotide polymorphism (SNP), their current utility for selection programs is limited as causative SNP because RFI appear to be breed or population specific (Saatchi et al., 2014Saatchi, M.; Beever, J. E.; Decker, J. E.; Faulkner, D. B.; Freetly, H. C.; Hansen, S. L.; Yampara-Iquise, H.; Johnson, K. A.; Kachman, S. D.; Kerley, M. S.; Kim, J.; Loy, D. D.; Marques, E.; Neibergs, H. L.; Pollak, E. J.; Schnabel, R. D.; Seabury, C. M.; Shike, D. W.; Snelling, W. M.; Spangler, M. L.; Weaber, R. L.; Garrick, D. J. and Taylor, J. F. 2014. QTLs associated with dry matter intake, metabolic mid-test weight, growth and feed efficiency have little overlap across 4 beef cattle studies. BMC Genomics 15: 1-14.). More recent studies (Jami et al., 2014Jami, E.; White, B. A. and Mizrahi, I. 2014. Potential role of the bovine rumen microbiome in modulating milk composition and feed efficiency. PLoS One 9: e85423.; McCann et al., 2014bMcCann, J. C.; Wiley, L. M.; Forbes, T. D.; Rouquette, F. M., Jr. and Tedeschi, L. O. 2014b. Relationship between the rumen microbiome and residual feed intake-efficiency of Brahman bulls stocked on Bermudagrass pastures. PLoS One 9: e91864.; Myer et al., 2015Myer, P. R.; Smith, T. P. L.; Wells, J. E.; Kuehn, L. A. and Freetly, H. C. 2015. Rumen microbiome from steers differing in feed efficiency. PLoS One 10: e0129174.; Roehe et al., 2016Roehe, R.; Dewhurst, R. J.; Duthie, C. -A.; Rooke, J. A.; McKain, N.; Ross, D. W.; Hyslop, J. J.; Waterhouse, A.; Freeman, T. C.; Watson, M. and Wallace, R. J. 2016. Bovine host genetic variation influences rumen microbial methane production with best selection criterion for low methane emitting and efficiently feed converting hosts based on metagenomic gene abundance. PLoS Genetics 12: 1-20.) have highlighted interrelationships that exist between host animals with divergent phenotypes for feed efficiency and their rumen microbiome structure. These recent advances in microbiomics, as well as metabolomics (metabolite profiles; Karisa et al., 2014Karisa, B. K.; Thomson, J.; Wang, Z.; Li, C.; Montanholi, Y. R.; Miller, S. P.; Moore, S. S. and Plastow, G. S. 2014. Plasma metabolites associated with residual feed intake and other productivity performance traits in beef cattle. Livestock Science 165: 200-211.), will drive discovery of more informative genomic markers for more accurate and robust selection for RFI across divergent cattle populations. Finally, advances in sensor technologies to enable individual-animal measurement of physiological (e.g., infrared thermography (IRT); Montanholi et al., 2009Montanholi, Y. R.; Swanson, K. C.; Schenkel, F. S.; McBride, B. W.; Caldwell, T. R. and Miller, S. P. 2009. On the determination of residual feed intake and associations of infrared thermography with efficiency and ultrasound traits in beef bulls. Livestock Science 125: 22-30.) and behavioral metrics (e.g., feeding behavior; Hafla et al., 2012aHafla, A. N.; Lancaster, P. A.; Carstens, G. E.; Forrest, D. W.; Fox, J. T.; Forbes, T. D. A.; Davis, M. E.; Randel, R. D. and Holloway, J. W. 2012a. Relationships between feed efficiency, scrotal circumference, and semen quality traits in yearling bulls. Journal of Animal Science 90: 3937-3944.) associated with RFI will provide additional opportunities to identify phenotypic biomarkers that are predictive of RFI.
Rumen efficiency and rumen microbiome
A rumen that functions efficiently can supply all or most of the absorbed amino acids required by a ruminant animal and allow it to digest high-fiber-based diets. The rumen functions as an independent anaerobic fermentation chamber and the end-products of digestion are volatile fatty acids, used by the ruminant animal as absorbed energy substrates and microbial crude protein (MCP), which is digested and absorbed in the small intestine from the bacteria flushed out of the rumen. The rumen bacteria require a carbohydrate source to ferment and an N source to synthesize amino acids. The carbohydrate source can be plant cell walls (fiber of various fractions), starch, pectins, or sugars, but ruminants have a competitive advantage over monogastrics if they ferment fiber, because fiber cannot be digested by monogastrics. The net outcome to the host is the digestion (or rate of digestion of carbohydrate, often simply measured as dry matter (DM) or organic matter (OM) digestion, but more accurately should be measured as neutral detergent fiber digestion) and MCP production from the total microbial biomass growth either measured as total MCP growth (g/d) or efficiency of MCP production (EMCP) (grams of MCP per Mcal of fermentable metabolizable energy or grams of MCP per kilogram of digestible OM). The host animal is not concerned with what microbial species are present, but rather the net outcome of the fermentation process. Fermentation also produces CH4 and, as discussed above, from an environmental perspective, CH4 production from the rumen is a disadvantage and a waste of energy to the host animal. The key question is whether the microbiome (or rumen ecology) affects the net outcome of the process in terms of yield of energy substrates and MCP or whether the net outcome may be similar from a range of different rumen microbiomes.
The rumen has many microbes made up of bacteria, protozoa, and fungi and the populations per ml of ruminal fluid are very high. Over 50 years of study have shown relationships for the EMCP and Galyean and Tedeschi (2014)Galyean, M. L. and Tedeschi, L. O. 2014. Predicting microbial protein synthesis in beef cattle: Relationship to intakes of total digestible nutrients and crude protein. Journal of Animal Science 92: 5099-5111. recently showed the variability in this value with a mean value of 130 g MCP/kg of total digestible nutrients. This is confirmed in other feeding standards, such as AFRC (1992)AFRC - Agricultural and Food Research Council. 1992. AFRC Technical Committee on Responses to Nutrients, Report 9: Nutritive requirements of ruminant animals: Protein. Nutrition Abstracts and Reviews (Series B: Livestock Feeds and Feeding) 62: 787-835., CSIRO (2007)CSIRO - Commonwealth Scientific and Industrial Research Organization. 2007. Nutrient requirements of domesticated ruminants. Commonwealth Scientific and Industrial Research Organization, Collingwood, VIC., and Detmann et al. (2014)Detmann, E.; Valente, E. E. L.; Batista, E. D. and Huhtanen, P. 2014. An evaluation of the performance and efficiency of nitrogen utilization in cattle fed tropical grass pastures with supplementation. Livestock Science 162: 141-153., which have reported similar values and ranges. Nevertheless, the range of values and accounting for this variability is important to the supply of nutrients to the host and Poppi et al. (1997)Poppi, D. P.; McLennan, S. R.; Bediye, S.; De Vega, A. and Zorrilla- Rios, J. 1997. Forage quality: Strategies for increasing nutritive value of forages. p. 307-322. In: Proceedings of the 18th International Grassland Congress, Winnipeg and Saskatoon, Canada. Canadian Forage Council, Canadian Society of Agronomy, Canadian Society of Animal Science, and Association Management Centre. Available at: <http://www.internationalgrasslands.org/>. Accessed on: May 15, 2016.
http://www.internationalgrasslands.org/...
calculated that the range in EMCP, when N supply to the microbes is adequate, would result in a difference in 300 g/day live weight gain of beef steers. Similarly, differences in digestibility between individual animals and species, although small quantitatively, also result in differences in overall metabolizable energy intake (Ellis et al., 1999Ellis, W. C.; Poppi, D. P.; Matis, J. H.; Lippke, H.; Hill, T. M. and Rouquette, F. M., Jr. 1999. Dietary-digestible-metabolic interactions determining the nutritive potential of ruminant diets. p. 423-481. In: Proceedings of the 5th International Symposium on the Nutrition of Herbivores, San Antonio, TX. American Society of Animal Science.). Thus, the current situation is that we understand responses in MCP and rate of DM digestion when a limiting nutrient, such as N, is supplied. We do not, however, understand the differences that occur between diets supposedly adequate in nutrients for microbial growth and between individuals or species of animals in terms of rate of digestion, EMCP, and CH4 production per unit of fermented substrate. Hypothetically, this variation has generally been related to differences of microbial species within the rumen and this idea has some attraction. To understand whether this is a causative (one variable promotes a change to another variable directly) or an associative effect (variables are correlated, but not necessarily one alters the other), we require some knowledge of the microbiome that is present and if certain microbiome patterns are associated with highly efficient rumens in terms of rate of digestion, EMCP, and CH4 production. In other words, a change in the microbiome does not imply a change in EMCP and vice-versa, EMCP may differ but with the same microbiome. Whether shifts in the microbiome are always associated with or linked to changes in EMCP is not yet certain or perhaps even to be expected.
Only recently have methods become available for studying the rumen microbiome and the methods have developed rapidly from denaturing gradient gel electrophoresis (measuring dominant species only) to 454 pyrosequencing and, now, next generation sequencing. The latter enables thousands of species to be identified from dominant species to those less prevalent, which, nevertheless, might have an important role degrading specific compounds (e.g., mimosine and dihydroxypyridine in Leucaena). The difficulty is employing bioinformatics to make sense of the mass of data (i.e., big data). The important issue here is asking the right question and this must be centered around the concept that we are interested in the rumen output to the host and the species analysis must look for microbiome patterns or causative bacterial species interaction in terms of the cascade of nutrient digestion and flow along a species consortium.
We can make some general statements. Until recently we have only studied ruminant microbial dynamics using in vitro methods and identified in vivo about 15% of the actual species that are present (Mackie et al., 2002Mackie, R. I.; McSweeney, C. S. and Klieve, A. V. 2002. Microbial ecology of the ovine rumen. p. 71-94. In: Sheep nutrition. Freer, M. and Dove, H., eds. CABI Publishing, Cambridge, MA.). We have started to identify major bacterial groupings associated with diet (Larue et al., 2005Larue, R.; Yu, Z.; Parisi, V. A.; Egan, A. R. and Morrison, M. 2005. Novel microbial diversity adherent to plant biomass in the herbivore gastrointestinal tract, as revealed by ribosomal intergenic spacer analysis and rrs gene sequencing. Environmental Microbiology 7: 530-543.) and shown that this can vary at least with dietary extremes of N and carbohydrate type. There are also bacterial patterns associated with high-and low-efficient phenotype animals (Hernandez-Sanabria et al., 2010Hernandez-Sanabria, E.; Guan, L. L.; Goonewardene, L. A.; Li, M.; Mujibi, D. F.; Stothard, P.; Moore, S. S. and Leon-Quintero, M. C. 2010. Correlation of particular bacterial PCR-denaturing gradient gel electrophoresis patterns with bovine ruminal fermentation parameters and feed efficiency traits. Applied and Environmental Microbiology 76: 6338-6350.; McCann et al., 2014bMcCann, J. C.; Wiley, L. M.; Forbes, T. D.; Rouquette, F. M., Jr. and Tedeschi, L. O. 2014b. Relationship between the rumen microbiome and residual feed intake-efficiency of Brahman bulls stocked on Bermudagrass pastures. PLoS One 9: e91864.) and there are archaea patterns associated with high and low CH4-emitting animals (Hegarty et al., 2007Hegarty, R. S.; Goopy, J. P.; Herd, R. M. McCorkell, B. 2007. Cattle selected for lower residual feed intake have reduced daily methane production. Journal of Animal Science 85: 1479-1486.; Ouwerkerk et al., 2008Ouwerkerk, D.; Turner, A. F. and Klieve, A. V. 2008. Diversity of methanogens in ruminants in Queensland. Australian Journal of Experimental Agriculture 48: 722-725.).
If the causative association between patterns of species in the rumen and efficient rumens defined by whatever output of interest (digestion, EMCP, CH4) is established, and it has not been established yet, then this opens up a completely new area for manipulating the rumen. The approach would be to identify the nutrients to which a consortia of bacteria respond and aim to manipulate the consortia of bacteria which are associated with highly efficient rumens. Efficient bacteria (in terms of rumen output) may be limited by the supply of key nutrients, identification of which enables them to be supplied. This raises the question of whether a microbiome pattern responds differently in simple mass action terms (the traditional approach to determining N requirement per Mcal of fermentable ME) or has different mass requirements based on the metabolic interactions of a consortia of bacteria. In addition, does the microbiome change in a predictable fashion to variation in the diet? The association between a microbiome pattern associated with high and low efficient host phenotypes or high and low host CH4 emitters suggests that the host genotype also has an influence over and above that of diet alone. Such observations also beg the question of when after birth these different rumen microbiomes are established and if that is influenced by the host or by a random population initiation which then sets a general lifelong which may be varied to a limited extent but is largely set by the initial pattern. These are all fundamental rumen ecology questions: How is a population established after birth? How is it controlled by the host? How does it respond to diet? And, finally, does it matter to rumen function?
Some examples illustrate these issues. A number of studies illustrate that the rumen microbiome varies with diet. Larue et al. (2005)Larue, R.; Yu, Z.; Parisi, V. A.; Egan, A. R. and Morrison, M. 2005. Novel microbial diversity adherent to plant biomass in the herbivore gastrointestinal tract, as revealed by ribosomal intergenic spacer analysis and rrs gene sequencing. Environmental Microbiology 7: 530-543. conducted one of the first studies and demonstrated a difference between animals fed grass- and those fed starch-containing diets. More recent examples have shown microbiome differences associated with diet, such as carbohydrate source (Shaw et al., 2015Shaw, C. N.; Kim, M.; Eastridge, M. L. and Yu, Z. 2015. Effects of different sources of physically effective fiber on rumen microbial populations. Animal 10: 410-417.) and protein or N source (Bento et al., 2015Bento, C. B. P.; Azevedo, A. C.; Mantovani, H. C.; Gomes, D. I.; Batista, E. D.; Rufino, L. M. A. and Detmann, E. 2015. Effect of protein supplementation on ruminal parameters and microbial community fingerprint of Nellore steers fed tropical forages. Animal 10: 44-54.), novel protein meals such as algae (Harper et al., 2010Harper, K. J.; Klieve, A. V.; Panjaitan, T.; Poppi, D. P. and Ouwerkerk, D. 2010. Changes in rumen bacterial community in steers fed with supplemented tropical pasture. p. 65. In: Proceedings of the Australian Society of Animal Production, 28. Available at: <http://www.asap.asn.au/livestocklibrary/2010/harper065.pdf>. Accessed on: May 9, 2017.
http://www.asap.asn.au/livestocklibrary/...
; McCann et al., 2014aMcCann, J. C.; Drewery, M. L.; Sawyer, J. E.; Pinchak, W. E. and Wickersham, T. A. 2014a. Effect of postextraction algal residue supplementation on the ruminal microbiome of steers consuming low-quality forage. Journal of Animal Science 92: 5063-5075.), and inherent variability between animals (Durso et al., 2010Durso, L. M.; Harhay, G. P.; Smith, T. P. L.; Bono, J. L.; Desantis, T. Z.; Harhay, D. M.; Andersen, G. L.; Keen, J. E.; Laegreid, W. W. and Clawson, M. L. 2010. Animal-to-animal variation in fecal microbial diversity among beef cattle. Applied and Environmental Microbiology 76: 4858-4862.; Martinez et al., 2010Martinez, E. D.; Klieve, A. V.; Ouwerkerk, D.; Swain, T.; Gulino, L. M.; Turnbull, K. and Poppi, D. P. 2010. Between animal variance in ruminal bacteria and protozoal communities from DGGE profiles of steers on a low quality forage diet. p. 63. In: Proceedings of the Australian Society of Animal Production, 28. Available at: <http://www.asap.asn.au/livestocklibrary/2010/martinez063.pdf>. Accessed on: May 9, 2017.
http://www.asap.asn.au/livestocklibrary/...
). It is important to understand that these relationships are still associative and not in all cases has there been a change in efficiency of rumen function, especially in terms of EMCP, but rather an associative change in response to diet. Therefore, different diets may result in diverse microbiomes as the microbiome pattern responds to the supply of fermentative substrates, but not necessarily result in different efficiency of rumen function.
The rumen microbiome also changes in response to host phenotypes, which vary in efficiency, but the extent to which that is causative or associative is not clear (Hernandez-Sanabria et al., 2010Hernandez-Sanabria, E.; Guan, L. L.; Goonewardene, L. A.; Li, M.; Mujibi, D. F.; Stothard, P.; Moore, S. S. and Leon-Quintero, M. C. 2010. Correlation of particular bacterial PCR-denaturing gradient gel electrophoresis patterns with bovine ruminal fermentation parameters and feed efficiency traits. Applied and Environmental Microbiology 76: 6338-6350.). The rumen microbiome is also different between host phenotypes that are high-or low-CH4 emitters (Hegarty et al., 2007Hegarty, R. S.; Goopy, J. P.; Herd, R. M. McCorkell, B. 2007. Cattle selected for lower residual feed intake have reduced daily methane production. Journal of Animal Science 85: 1479-1486.). This is associated with differences in archaea population (Gilbert et al., 2015Gilbert, R. A.; Ouwerkerk, D. and Klieve, A. V. 2015. Phage therapy in livestock methane amelioration. p. 318-335. In: Livestock production and climate change. Malik, P. K.; Bhatta, R.; Takahashi, J., et al., eds. CABI Publishing, Boston, MA.; Ouwerkerk et al., 2008Ouwerkerk, D.; Turner, A. F. and Klieve, A. V. 2008. Diversity of methanogens in ruminants in Queensland. Australian Journal of Experimental Agriculture 48: 722-725.) and, in this case, one could argue for a causal relationship between these species and CH4 production, but it is not understood how these different microbiomes arise, i.e., randomly at birth or determined by the host.
The development of a mature microbiome from birth has been little studied. Recently, Rey et al. (2014)Rey, M.; Enjalbert, F.; Combes, S.; Cauquil, L.; Bouchez, O. and Monteils, V. 2014. Establishment of ruminal bacterial community in dairy calves from birth to weaning is sequential. Journal of Applied Microbiology 116: 245-257. and De Barbieri et al. (2015)De Barbieri, I.; Hegarty, R. S.; Silveira, C.; Gulino, L. M.; Oddy, V. H.; Gilbert, R. A.; Klieve, A. V. and Ouwerkerk, D. 2015. Programming rumen bacterial communities in newborn Merino lambs. Small Ruminant Research 129: 48-59. have shown that there is a pattern of establishment and that this can be influenced by the species that are initiated into the microbiome soon after birth from maternal or other sources. This is a fascinating field of study opening up the possibility of long-term manipulation of the rumen microbiome. Bacteriophages offer another means of manipulating the rumen microbiome (Gilbert and Klieve, 2015Gilbert, R. A. and Klieve, A. V. 2015. Ruminal viruses (Bacteriophages, Archaeaphages). p. 121-141. In: Rumen microbiology: From evolution to revolution. Puniya, K. A.; Singh, R. and Kamra, N. D., eds. Springer India, New Delhi.; Gilbert et al., 2015Gilbert, R. A.; Ouwerkerk, D. and Klieve, A. V. 2015. Phage therapy in livestock methane amelioration. p. 318-335. In: Livestock production and climate change. Malik, P. K.; Bhatta, R.; Takahashi, J., et al., eds. CABI Publishing, Boston, MA.) if we understood what microbiome pattern we need to generate to improve the efficiency of rumen function. This is a completely new field of study in rumen microbiology.
Early warning systems
Given the increasing uncertainty and risk associated with livestock production, early warning systems (EWS) can play a critical role in providing near real-time information needed to assess risk and assist in adaptive management decision making, as well as provide information needed to reduce impacts to ecosystem services. Early warning systems can be defined as a framework for monitoring that is designed to avoid, or at least to minimize, the impact of natural or human-induced threats to humans, property, the environment, or livelihoods (Medina-Cetina and Nadim, 2008Medina-Cetina, Z. and Nadim, F. 2008. Stochastic design of an early warning system. Georisk: Assessment and Management of Risk for Engineered Systems and Geohazards 2: 223-236.). Generally, EWS are comprised of four core components, according to the International Early Warning Programme (http://www.unisdr.org/2006/ppew/iewp/IEWP-brochure.pdf): risk knowledge, monitoring and warning, dissemination and communication, and response capability. The risk knowledge component requires data collection and assessments of hazards and vulnerabilities, in addition to evaluating patterns and trends in the data. Monitoring and warning require that services be put into place to provide warnings in a timely manner and that they have some level of accuracy and scientific basis. The early warning information must also be communicated in a way that provides clear, understandable, and usable information that identifies the risks. Communications and warnings must also be available to all of those who are at risk. Lastly, response capabilities need to be in place at the levels of scale appropriate for the hazards being monitored and stakeholders trained to react to the early warning information provided (Basher, 2006Basher, R. 2006. Global early warning systems for natural hazards: Systematic and people-centred. Philosophical Transactions of the Royal Society of London A: Mathematical, Physical and Engineering Sciences 364: 2167-2182.).
For most cattle, sheep, and goat livestock production systems, drought poses a risk and adaptive management challenges that EWS can provide critical information. Drought impacts that influence livestock production can include reductions in both quantity and quality of forage on grazing lands, as well as reductions in feed crop production and water availability. Using traditional field assessments, near real-time assessment, and characterization of livestock forage and water is problematic, given the time and resources required to conduct accurate assessments of forage productivity and water availability over large landscapes (Angerer, 2012bAngerer, J. P. 2012b. Technologies, tools and methodologies for forage evaluation in grasslands and rangelands. p. 165-200. In: Conducting National Feed Assessments. vol. 15. Coughenour, M. B. and Makkar, H. P. S., eds. Food and Agriculture Organization of the United Nations, Rome, Italy.).
Integrated remote sensing, geographical information system, and simulation modeling approaches generally use climate and remote sensing data as inputs to simulation models and then use geographical information system to synthesize outputs into useful maps and products for monitoring and early warning. In East Africa and Mongolia, regional livestock EWS provide pastoralists, policy makers, and other stakeholders with information on emerging forage conditions to improve risk management and adaptive management decision-making Angerer, 2012aAngerer, J. P. 2012a. Gobi forage livestock early warning system. p. 115-130. In: Conducting National Feed Assessments. vol. 15. Coughenour, M. B. and Makkar, H. P. S., eds. Food and Agriculture Organization of the United Nations, Rome, Italy.; Stuth et al., 2003Stuth, J.; Angerer, J.; Kaitho, R.; Zander, K.; Jama, A.; Heath, C.; Bucher, J.; Hamilton, W.; Conner, R. and Inbody, D. 2003. The livestock early warning system (LEWS): Blending technology and the human dimension to support grazing decisions. Available at: <http://cals.arizona.edu/OALS/ALN/aln53/stuth.html>. Accessed on: May 16, 2016.
http://cals.arizona.edu/OALS/ALN/aln53/s...
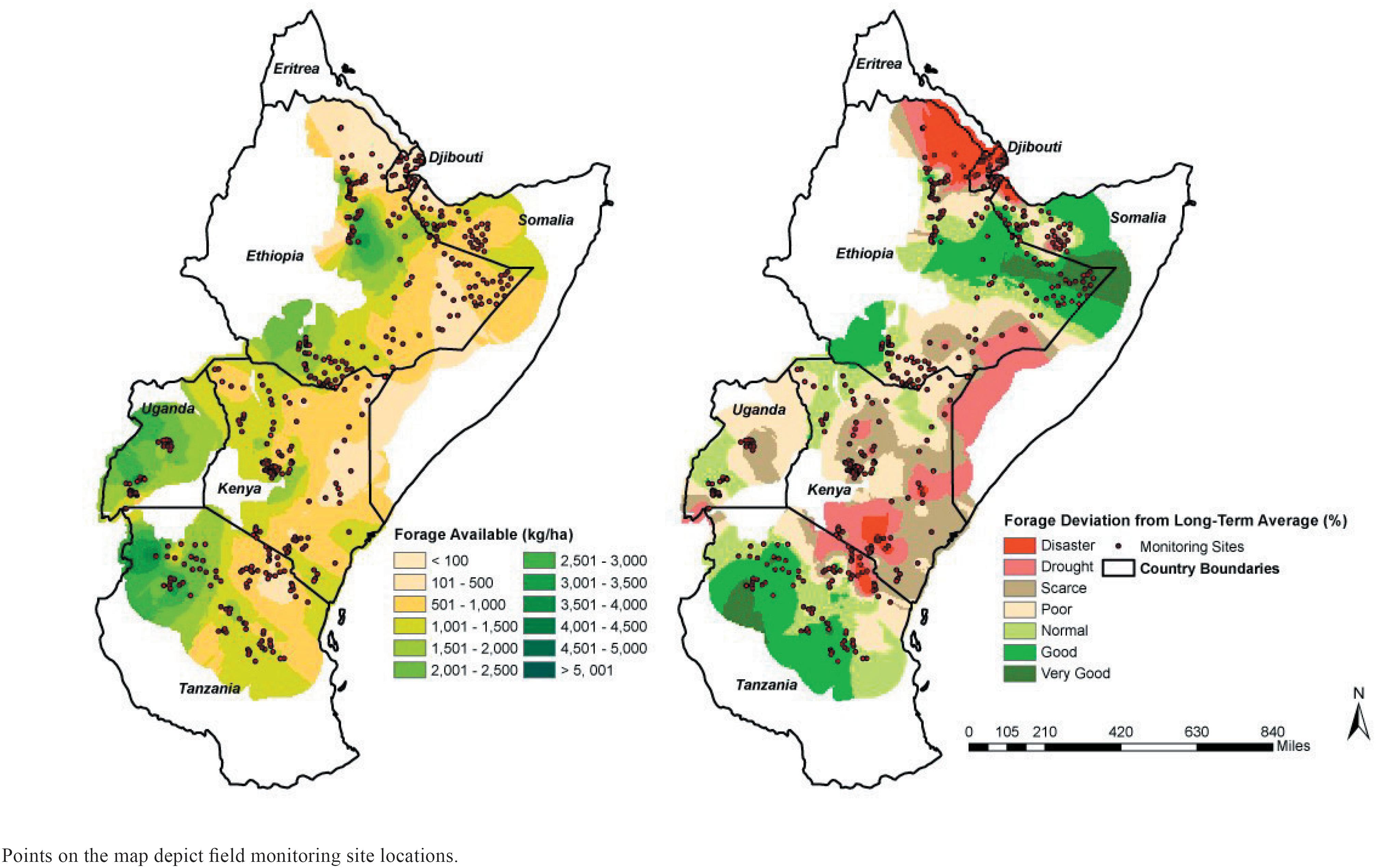
; (Stuth et al., 2005Stuth, J. W.; Angerer, J.; Kaitho, R.; Jama, A. and Marambii, R. 2005. Livestock early warning system for Africa's rangelands. p. 283-294. In: Monitoring and predicting agricultural drought: A Global study. Boken, V. K.; Cracknell, A. P. and Heathcote, R. L., eds. Oxford University Press, Oxford, UK.). These systems use a simulation model driven by remotely sensed climate data and data collected periodically from a series of field monitoring sites across the region. Geographical information system, forecasting, and geostatistical methods are used to produce current and near-term forecast maps (Alhamad et al., 2007Alhamad, M. N.; Stuth, J. and Vannucci, M. 2007. Biophysical modelling and NDVI time series to project near-term forage supply: spectral analysis aided by wavelet denoising and ARIMA modelling. International Journal of Remote Sensing 28: 2513-2548.) of available forage and anomalies (Figure 1) that are coded to early warning categories.
Examples of Livestock Early Warning System maps for East Africa depicting forage availability (left) and early warning status based on deviation from long-term average (right).
Forage quality assessment is as important as the assessment of forage quantity. During the past 20 years, near infrared reflectance spectroscopy (NIRS) scanning of livestock feces has emerged as a reliable tool for assessing the quality of forage grazed by ruminants (Dixon and Coates, 2010Dixon, R. M. and Coates, D. B. 2010. Diet quality estimated with faecal near infrared reflectance spectroscopy and responses to N supplementation by cattle grazing buffel grass pastures. Animal Feed Science and Technology 158: 115-125.; Leite and Stuth, 1995Leite, E. R. and Stuth, J. W. 1995. Fecal NIRS equations to assess diet quality of free-ranging goats. Small Ruminant Research 15: 223-230.; Li et al., 2007Li, H.; Tolleson, D.; Stuth, J.; Bai, K.; Mo, F. and Kronberg, S. 2007. Faecal near infrared reflectance spectroscopy to predict diet quality for sheep. Small Ruminant Research 68: 263-268.; Lyons and Stuth, 1992Lyons, R. K. and Stuth, J. W. 1992. Fecal NIRS equations for predicting diet quality of free-ranging cattle. Journal of Range Management 45: 238-244.; Stuth et al., 1991Stuth, J. W.; Lyons, R. K.; Angerer, J. P. and McKown, C. D. 1991. Role of NIRS-based nutritional monitoring systems for grazing and nutritional management of range livestock. p. 83-93. In: Proceedings of the 2nd Grazing Livestock Nutrition Conference, Steamboat Springs, CO. Oklahoma State University. Available at: <http://coronasc.nmsu.edu/grazing-livestock-nutrit.html>. Accessed on: May 17, 2016.
http://coronasc.nmsu.edu/grazing-livesto...
). Remote sensing-based approaches for forage quality estimation will require additional study to be effective in an EWS framework. Early warning systems for monitoring livestock water are important in arid and semi-arid regions where livestock herding is predominant and herders are dependent on surface tanks and ponds to water livestock. In Sub-Saharan Africa, surface ponds and tanks are delineated using remote sensing image analysis and field data collection (Senay et al., 2013Senay, G. B.; Velpuri, N. M.; Alemu, H.; Md Pervez, S.; Asante, K. O.; Kariuki, G.; Taa, A. and Angerer, J. 2013. Establishing an operational waterhole monitoring system using satellite data and hydrologic modelling: Application in the pastoral regions of East Africa. Pastoralism: Research, Policy and Practice 3: 1-16.; Velpuri et al., 2014Velpuri, N. M.; Senay, G. G.; Alemu, H.; Rowland, J. and Verdin, J. P. 2014. Africa-wide monitoring of small surface water bodies using multisource satellite data: A monitoring system for FEWS NET. p. 69-95. In: Nile River Basin: Ecohydrological challenges, Climate Change and Hydropolitics. Melesse, A. M.; Abtew, W. and Setegn, S. G., eds. Springer, New York, NY.). In Kenya, local monitors also provide information from the field for periodic model verification, as well as to provide information on numbers and kinds of animals using the water source, livestock body condition, and community concerns.
Livestock disease outbreak and spread is another area in which EWS can play an effective role in reducing risk and managing outbreaks. Food and Agriculture Organization, World Organization for Animal Health, and World Health Organization have implemented the Global Early Warning System for Major Animal Diseases Including Zoonosis (GLEWS) program to mitigate potential health threats at the human-animal ecosystems interface. The system uses monitoring data gathered from existing event-based surveillance systems including FAO's Global Animal Disease Information System (EMPRES-I; http://empres-i.fao.org/empres-i/) (FAO et al., 2013FAO - Food and Agriculture Organization; World Organization for Animal Health (OIE) and World Health Organization (WHO). 2013. GLEWS+: The Joint FAO-OIE-WHO Global Early Warning System for Health Threats and Emerging Risks at the Human-Animal-Ecosystems Interface; A Concept Note. Available at: <http://www.glews.net/wp-content/uploads/2013/12/04_GLEWSConcept-20-11.pdf>. Accessed on: May 17, 2016.
http://www.glews.net/wp-content/uploads/...
).
With changing land use and efforts by groups to reduce livestock numbers globally due to potential impacts of livestock on GHG emissions, ecosystem services, and land degradation, an increased vegetation biomass resulting from livestock removal can increase wildfire risk. Because livestock grazes the plant material considered “fine fuels” (i.e., plant material with a high surface area to volume ratio that dries readily and is rapidly consumed by fire when dry), the amount of fine fuels can be reduced. A recent study using targeted grazing found that fire-spread rates could potentially be reduced by 50 to 60% with light grazing (Bruegger et al., 2016Bruegger, R. A.; Varelas, L. A.; Howery, L. D.; Torell, L. A.; Stephenson, M. B. and Bailey, D. W. 2016. Targeted grazing in Southern Arizona: Using cattle to reduce fine fuel loads. Rangeland Ecology & Management 69: 43-51.). Livestock grazing can also create vegetation patchiness through variable levels of grazing intensity across the landscape, thus creating discontinuities in fuel that retard fire spread (Taylor Jr, 2006Taylor Jr, C. A. 2006. Targeted grazing to manage fire risk. p. 107-114. In: Targeted grazing: A natural approach to vegetation management and landscape enhancement. Launchbaugh, K. and Walker, J., eds. American Sheep Industry Association, Englewood, CO.). Advances in technology such as LIDAR (Light Detection And Ranging) can assist in providing high-resolution quantification for biomass and land cover assessments (Bork and Su, 2007Bork, E. W. and Su, J. G. 2007. Integrating LIDAR data and multispectral imagery for enhanced classification of rangeland vegetation: A meta analysis. Remote Sensing of Environment 111: 11-24.; Ku et al., 2012Ku, N. -W.; Popescu, S. C.; Ansley, R. J.; Perotto-Baldivieso, H. L. and Filippi, A. M. 2012. Assessment of Available Rangeland Woody Plant Biomass with a Terrestrial Lidar System. Photogrammetric Engineering & Remote Sensing 78: 349-361.) and shows promise for mapping fuels in a three-dimensional fashion for wildfire risk detection (Mutlu et al., 2008Mutlu, M.; Popescu, S. C.; Stripling, C. and Spencer, T. 2008. Mapping surface fuel models using lidar and multispectral data fusion for fire behavior. Remote Sensing of Environment 112: 274-285.; Peterson et al., 2015Peterson, B.; Nelson, K. J.; Seielstad, C.; Stoker, J.; Jolly, W. M. and Parsons, R. 2015. Automated integration of lidar into the LANDFIRE product suite. Remote Sensing Letters 6: 247-256.).
Bestelmeyer and Briske (2012)Bestelmeyer, B. T. and Briske, D. D. 2012. Grand challenges for resilience-based management of rangelands. Rangeland Ecology & Management 65: 654-663. expressed the need to develop knowledge systems to support adaptation and transformation for resilience-based management on grazing lands. These knowledge systems should bring together, integrate, and mobilize technology and innovations from diverse sources to improve and support decision making. Given the diversity of issues facing livestock producers today and in the future, development of livestock information and knowledge systems that would incorporate data from a variety of monitoring systems, provide early warning and trend assessment, and incorporate scientific and local knowledge, would be beneficial in reducing risks and improving management.
Precision livestock farming
Precision livestock farming (PLF) is often referred to as “smart farming technology” and involves the use of sensor technologies to capture physiological, behavioral, and productivity measurements of individual animals to aid in the implementation of management strategies that can improve the overall performance of livestock production systems (Bewley, 2013Bewley, J. 2013. Exciting dairy breakthroughs: Science fiction of precision dairy farming? p. 1-8. In: Proceedings of the Precision Dairy Conference and Exp, Rochester, MN. University of Minnesota. Available at: <http://www.precisiondairyfarming.com/2015/>. Accessed on: May 23, 2016.
http://www.precisiondairyfarming.com/201...
). The premise of PLF systems is that fully automated continuous monitoring of individual animals will enhance the ability of the producers to detect and manage animal health, productivity/ reproduction, and environmental-impact aspects of their livestock operations (Berckmans, 2015Berckmans, D. 2015. Smart farming for Europe: Value creation through precision livestock farming. p. 25-35. In: Precision livestock farming applications. Halachmi, I., ed Wageningen Academic Publishers, Wageningen, NL.). A non-inclusive list of sensor technologies (Bewley et al., 2015Bewley, J. M.; Russell, R. A.; Dolecheck, K. A. and Borchers, M. R. 2015. Precision dairy monitoring: What have we learned? p. 15-23. In: Precision Livestock Farming Applications. Halachmi, I., ed Wageningen Academic Publishers, Wageningen, NL.; Rutten et al., 2013Rutten, C. J.; Velthuis, A. G. J.; Steeneveld, W. and Hogeveen, H. 2013. Invited review: Sensors to support health management on dairy farms. Journal of Dairy Science 96: 1928-1952.; Theurer et al., 2013aTheurer, M. E.; Amrine, D. E. and White, B. J. 2013a. Remote noninvasive assessment of pain and health status in cattle. Veterinary Clinics of North America: Food Animal Practice 29: 59-74.; Wolfger et al., 2015dWolfger, B.; Timsit, E.; White, B. J. and Orsel, K. 2015d. A systematic review of bovine respiratory disease diagnosis focused on diagnostic confirmation, early detection, and prediction of unfavorable outcomes in feedlot cattle. Veterinary Clinics of North America: Food Animal Practice 31: 351-365.) developed for beef and dairy cattle applications include automated-scale technologies to measure milk yield and body weight; in-line sensors to measure milk components (fat); in-cow sensors to measure rumen temperature and pH (reticulorumen boluses); on-cow sensors to measure physical activity (pedometers, accelerometers), feeding behavior (ultra-wideband radio frequency identification), and rumination (accelerometers, acoustics); and off-cow sensors to measure skin-surface temperature (infrared thermography) and body-fat reserves (automated image analysis). The availability of novel and more cost-effective sensor technologies for PLF applications is expected to grow exponentially in the future due to the ongoing development of new sensor technologies primarily for non-agricultural applications and advances in computing systems (e.g., wireless communications, cloud storage).
Early preclinical detection is critical for effective antimicrobial treatment of bovine respiratory disease (BRD), but current detection methods rely on visual observations of clinical signs by feedlot personnel that are often unreliable as cattle are prey animals with inherent instincts to mask clinical signs of illness. In fact, White and Renter (2009)White, B. J. and Renter, D. G. 2009. Bayesian estimation of the performance of using clinical observations and harvest lung lesions for diagnosing bovine respiratory disease in post-weaned beef calves. Journal of Veterinary Diagnostic Investigation 21: 446-453. found sensitivity of visual observation of BRD detection was only 62%, demonstrating that BRD cases often go undetected or remain undetected until later in the disease process, when successful intervention is less likely and animal suffering has advanced. Moreover, the specificity of BRD detection is also low (63%), resulting in overuse of antimicrobial drugs. Recent advances in metabolomics (e.g., metabolite profiling) offer considerable promise for discovering more accurate preclinical biomarkers of infectious disease (De Buck et al., 2014De Buck, J.; Shaykhutdinov, R.; Barkema, H. W. and Vogel, H. J. 2014. Metabolomic profiling in cattle experimentally infected with Mycobacterium avium subsp. paratuberculosis. PLoS One 9: 1-11.) and metabolic disorders (Hailemariam et al., 2014Hailemariam, D.; Mandal, R.; Saleem, F.; Dunn, S. M.; Wishart, D. S. and Ametaj, B. N. 2014. Identification of predictive biomarkers of disease state in transition dairy cows. Journal of Dairy Science 97: 2680-2693.). However, the utility of these biomarkers will remain dependent upon initial detection of prospective clinically ill animals by livestock personnel, collection of biological samples, and rapid-test chute-side assays to be diagnostically relevant for livestock industries. Development of PLF systems that capture physiological and/or behavioral metrics for accurate preclinical detection of BRD would reduce the economic effect of this disease. Numerous temperature sensors have been deployed to monitor animal health status with reticulorumen boluses, tympanic thermistors, or infrared thermography (IRT). Timsit et al. (2011)Timsit, E.; Assié, S.; Quiniou, R.; Seegers, H. and Bareille, N. 2011. Early detection of bovine respiratory disease in young bulls using reticulo-rumen temperature boluses. The Veterinary Journal 190: 136-142. and Adams et al. (2013)Adams, A. E.; Olea-Popelka, F. J. and Roman-Muniz, I. N. 2013. Using temperature-sensing reticular boluses to aid in the detection of production diseases in dairy cows. Journal of Dairy Science 96: 1549-1555. were able to detect BRD cases 0.5 to four days prior to visual clinical diagnosis with moderate sensitivities. Using a hand-held IRT camera to monitor maximum-orbital temperature of calves, Schaefer et al. (2007)Schaefer, A. L.; Cook, N. J.; Church, J. S.; Basarab, J.; Perry, B.; Miller, C. and Tong, A. K. W. 2007. The use of infrared thermography as an early indicator of bovine respiratory disease complex in calves. Research in Veterinary Science 83: 376-384. found that BRD could be detected four to six days prior to clinical diagnosis, with greater test efficiency (71 vs 55%), then by visual observation. Schaefer et al. (2012)Schaefer, A. L.; Cook, N. J.; Bench, C.; Chabot, J. B.; Colyn, J.; Liu, T.; Okine, E. K.; Stewart, M. and Webster, J. R. 2012. The noninvasive and automated detection of bovine respiratory disease onset in receiver calves using infrared thermography. Research in Veterinary Science 93: 928-935. deployed an automated IRT technology system (camera mounted above water trough) for continuous non-invasive monitoring of orbital temperature and found that test efficiency (93%) was high for preclinical detection of BRD.
Decrease in feed intake is one of the earliest indicators of the onset of clinical illness in cattle. A number of studies have demonstrated reductions in feed intake prior to onset of BRD in beef cattle (Jackson et al., 2016Jackson, K. S.; Carstens, G. E.; Tedeschi, L. O. and Pinchak, W. E. 2016. Changes in feeding behavior patterns and dry matter intake before clinical symptoms associated with bovine respiratory disease in growing bulls. Journal of Animal Science 94: 1644-1652.; Wolfger et al., 2015bWolfger, B.; Schwartzkopf-Genswein, K. S.; Barkema, H. W.; Pajor, E. A.; Levy, M. and Orsel, K. 2015b. Feeding behavior as an early predictor of bovine respiratory disease in North American feedlot systems. Journal of Animal Science 93: 377-385.). While automated systems for measuring individual-animal feed intake are becoming more widely available, the prospects for commercial applications of these technologies for early disease detection are limited due to high investment and maintenance costs. However, changes in behavioral patterns associated with intake of feed and water are also early indicators of disease onset. Daniels et al. (2000)Daniels, T. K.; Bowman, J. G. P.; Sowell, B. F.; Branine, M. E. and Hubbert, M. E. 2000. Effects of metaphylactic antibiotics on behavior of feedlot calves. Professional Animal Scientist 16: 247-253. and Sowell et al. (1999)Sowell, B. F.; Branine, M. E.; Bowman, J. G. P.; Hubbert, M. E.; Sherwood, H. E. and Quimby, W. 1999. Feeding and watering behavior of healty and morbid steers in a commercial feedlot. Journal of Animal Science 77: 1105-1112. found that calves treated for BRD spent 23 to 42% less time at the feed bunk and had 10 to 36% fewer feeding bouts and 12 to 33% fewer drinking bouts compared with healthy calves. Utilizing a real-time location system based on ultra-wideband RFID technology, Theurer et al. (2013b)Theurer, M. E.; Anderson, D. E.; White, B. J.; Miesner, M. D.; Mosier, D. A.; Coetzee, J. F.; Lakritz, J. and Amrine, D. E. 2013b. Effect of Mannheimia haemolytica pneumonia on behavior and physiologic responses of calves during high ambient environmental temperatures. Journal of Animal Science 91: 3917-3929. found that calves challenged with Mannheimia haemolytica spent less time in close proximity to the feed bunk and more time lying down than control calves. Additionally, other technologies for monitoring feeding behavior in cattle that utilize accelerometers attached to ear tags (Wolfger et al., 2015cWolfger, B.; Timsit, E.; Pajor, E. A.; Cook, N.; Barkema, H. W. and Orsel, K. 2015c. Technical note: Accuracy of an ear tag-attached accelerometer to monitor rumination and feeding behavior in feedlot cattle. Journal of Animal Science 93: 3164-3168.) and feed bunk antennas that detect presence of passive RFID transponders that are attached to the front leg of cattle (Wolfger et al., 2015aWolfger, B.; Mang, A. V.; Cook, N.; Orsel, K. and Timsit, E. 2015a. Technical note: Evaluation of a system for monitoring individual feeding behavior and activity in beef cattle. Journal of Animal Science 93: 4110-4114.) are being developed. Real-time behavioral-monitoring systems that accurately quantify feeding and drinking behavior activities have considerable potential for early detection of morbid cattle due to infectious disease and metabolic disorders (González et al., 2008González, L. A.; Tolkamp, B. J.; Coffey, M. P.; Ferret, A. and Kyriazakis, I. 2008. Changes in feeding behavior as possible indicators for the automatic monitoring of health disorders in dairy cows. Journal of Dairy Science 91: 1017-1028.).
Rutten et al. (2013)Rutten, C. J.; Velthuis, A. G. J.; Steeneveld, W. and Hogeveen, H. 2013. Invited review: Sensors to support health management on dairy farms. Journal of Dairy Science 96: 1928-1952. concluded that the performance of the sensor systems is highly variable and dependent on the choice of the gold standards used to confirm specific animal responses, the time resolution and accuracy of the sensors, and the selection and validity of the detection algorithms. In other words, sensor systems will be of limited value and become useless generators of data unless effective detection algorithms are developed to convert sensor data into meaningful animal-status information. De Vries and Reneau (2010De Vries, A. and Reneau, J. K. 2010. Application of statistical process control charts to monitor changes in animal production systems. Journal of Animal Science 88: E11-E24.) and Mertens et al. (2011)Mertens, K.; Decuypere, E.; De Baerdemaeker, J. and De Ketelaere, B. 2011. Statistical control charts as a support tool for the management of livestock production. The Journal of Agricultural Science 149: 369-384. reviewed statistical-process control (SPC) procedures to capture value from automated real-time sensor systems for application in livestock production systems. While SPC procedures have been widely used in non-agricultural industries, few SPC applications to date have been developed for animal agriculture. Quimby et al. (2001)Quimby, W. F.; Sowell, B. F. and Bowman, J. G. P. 2001. Application of feeding behaviour to predict morbidity of newly received calves in a commercial feedlot. Canadian Journal of Animal Science 81: 315-320. used SPC procedures to evaluate deviations in electronic feed bunk attendance data collected from calves at high risk for BRD. Based on control-chart detection of feeding duration, they reported that morbidity events could be predicted three to four days prior to detection of BRD by feedlot personnel, with an accuracy and positive-predictive values of 87 and 91%, respectively. In growing bulls that exhibited a spontaneous outbreak of BRD, Jackson (2015)Jackson, K. S. 2015. Associations of feeding behavior patterns with inter-animal variation in feed efficiency and pre-clinical responses to infectious disease in beef cattle. Thesis. Texas A&M University, College Station, TX. used cumulative sum charts to evaluate individual-animal deviations in feed intake and feeding-behavior patterns relative to the day clinical illness was detected. Detection of SPC-model of BRD based on feed bunk attendance and head-down duration occurred 2.7 and 3.0 days prior to observed BRD diagnosis, with model accuracies of 87 and 89%, respectively. More recently, Moya et al. (2015)Moya, D.; Silasi, R.; McAllister, T. A.; Genswein, B.; Crowe, T.; Marti, S. and Schwartzkopf-Genswein, K. S. 2015. Use of pattern recognition techniques for early detection of morbidity in receiving feedlot cattle. Journal of Animal Science 93: 3623-3638. used nonlinear data-mining analysis of feeding behavior data to develop pattern-recognition-based algorithms to predict morbidity events in beef cattle. When validated against a naive group of calves, they were able to correctly predict the health status of high-risk calves with a model accuracy of 79%.
The integration of decision support systems (DSS) with physiological-and behavioral-based sensor technologies to monitor animal-health status in real time has considerable potential to mitigate the detrimental effects of infectious diseases, promote more judicious use of antimicrobials and improve animal welfare. Additionally, widespread adoption of these technologies would support early-event biosurveillance systems that, when integrated with GLEWS, would substantially reduce the risk of epidemics caused by foreign-animal and emerging disease threats. Beyond the application of PLF-based technologies for preclinical detection and mitigation of disease is their deployment to capture informative phenotypes associated with economically relevant traits such as disease resistance or efficiency of feed utilization by grazing livestock (Greenwood et al., 2016Greenwood, P. L.; Bishop-Hurley, G. J.; González, L. A. and Ingham, A. B. 2016. Development and application of a livestock phenomics platform to enhance productivity and efficiency at pasture. Animal Production Science 56: 1299-1311.). Advances in sensor and computational technologies will facilitate the large-scale real-time collection of phenotypic traits that, when combined with “omics” technologies (Riggs et al., 2017Riggs, P. K.; Tedeschi, L. O.; Turner, N. D.; Braga-Neto, U. and Jayaraman, A. 2017. The role of “omics” technologies for livestock sustainability. Archivos Latinoamericanos de Producción Animal (in press).), will provide the considerable potential to advance genetic merit of livestock and further improve the level of precision of nutritional and management strategies.
Integrating genomics into nutrition modeling
Mathematical modeling is likely the best pragmatic way to integrate accumulated scientific knowledge into decision-smart tools or DSS that can be used to optimize livestock production. Decision support systems are frequently used outside the scope for which they were developed and, in many situations, the necessary inputs are not available. In these situations, DSS might have satisfactory accuracy, but precision is somewhat substandard. Poppi and McLennan (2010)Poppi, D. P. and McLennan, S. R. 2010. Nutritional research to meet future challenges. Animal Production Science 50: 329-338. indicated that the use of (mathematical) models would allow less experienced nutritionists to understand and make recommendations under practical conditions. The data of six feeding experiments were used to evaluate the predictability of the Cornell Net Carbohydrate and Protein System (Fox et al. (2004)Fox, D. G.; Tedeschi, L. O.; Tylutki, T. P.; Russell, J. B.; Van Amburgh, M. E.; Chase, L. E.; Pell, A. N. and Overton, T. R. 2004. The Cornell Net Carbohydrate and Protein System model for evaluating herd nutrition and nutrient excretion. Animal Feed Science and Technology 112: 29-78., currently referred to as the Large Ruminant Nutrition System (http://nutritionmodels.com/lrns.html) of growing young beef cattle fed different types of supplements (Poppi and McLennan, 2010Poppi, D. P. and McLennan, S. R. 2010. Nutritional research to meet future challenges. Animal Production Science 50: 329-338.) in Australia. They reported a general agreement between predicted and observed average daily gain and DM intake, but precision was low (i.e., wide variation surrounding the unity line). Tedeschi et al. (2014)Tedeschi, L. O.; Cavalcanti, L. F. L.; Fonseca, M. A.; Herrero, M. and Thornton, P. K. 2014. The evolution and evaluation of dairy cattle models for predicting milk production: an agricultural model intercomparison and improvement project (AgMIP) for livestock. Animal Production Science 54: 2052-2067. developed a database containing available model inputs of lactating dairy cows published in studies conducted in six regions of the world (n = 19 for Africa, n = 45 for Asia, n = 16 for Europe, n = 12 for Latin America, n = 44 for North America, and n = 37 for Oceania) to compare the predictive adequacy of milk production of four nutrition models. Very few studies had the necessary information needed to properly utilize mechanistic systems and the authors concluded that not all of the models evaluated were suitable for predicting milk production across the diverse conditions (cattle, feed, management, and environment) of these studies. The authors concluded that simpler systems might be more resilient to variations in the quality of inputs and the variable production conditions around the world. Unfortunately, the low awareness and limited knowledge of nutrition models and their usage to design more efficient and profitable animal feeding and management systems are the main factors that further a negative perception of modeling and simulation (Tedeschi et al., 2015Tedeschi, L. O.; Muir, J. P.; Riley, D. G. and Fox, D. G. 2015. The role of ruminant animals in sustainable livestock intensification programs. International Journal of Sustainable Development & World Ecology 22: 452-465.). In hindsight, we have neglected these facts for a long time and very few nutritionists have been trained to use modeling and simulation to solve applied problems.
Within the sustainable livestock intensification theory, Tedeschi et al. (2015)Tedeschi, L. O.; Muir, J. P.; Riley, D. G. and Fox, D. G. 2015. The role of ruminant animals in sustainable livestock intensification programs. International Journal of Sustainable Development & World Ecology 22: 452-465. ascertained that mathematical nutrition models are an important component of forecasting unforeseen variable relationships to quantify expected outcomes that might occur from alternative decisions. The more complex the relationships of the system, the greater the need to use a mathematical model to evaluate alternative solutions more quickly and cheaply.
When inputs are adequate, nutrition models have been used in large scale production scenarios for individual animal management to improve profitability (Guiroy et al., 2001Guiroy, P. J.; Fox, D. G.; Tedeschi, L. O.; Baker, M. J. and Cravey, M. D. 2001. Predicting individual feed requirements of cattle fed in groups. Journal of Animal Science 79: 1983-1995.). Future possibilities to advance individual animal management include the use of genomic markers and remote sensor technologies to capture individual-animal genetic and phenotypic information, respectively, to identify divergent animals in animal performance and growth efficiency. Oltjen et al. (2000)Oltjen, J. W.; Pleasants, A. B.; Soboleva, T. K. and Oddy, V. H. 2000. Second-generation dynamic cattle growth and composition models. p. 197-209. In: Modelling nutrient utilization in farm animals. McNamara, J. P.; France, J. and Beever, D. E., eds. CABI Publishing, New York, NY. integrated genetic parameters (breeding value) into a growth model (Di Marco and Baldwin, 1989Di Marco, O. N. and Baldwin, R. L. 1989. Implementation and evaluation of a steer growth model. Agricultural Systems 29: 247-265.; Di Marco et al., 1989Di Marco, O. N.; Baldwin, R. L. and Calvert, C. C. 1989. Simulation of DNA, protein and fat accretion in growing steers. Agricultural Systems 29: 21-34.; Oltjen et al., 1986Oltjen, J. W.; Bywater, A. C.; Baldwin, R. L. and Garrett, W. N. 1986. Development of a dynamic model of beef cattle growth and composition. Journal of Animal Science 62: 86-97.; Soboleva et al., 1999Soboleva, T. K.; Oddy, V. H.; Pleasants, A. B.; Oltjen, J. W.; Ball, A. J. and McCall, D. G. 1999. A dynamical model of body composition in sheep. p. 275-278. In: Proceedings of the New Zealand Society of Animal Production, 59. Available at: <http://www.nzsap.org/view/biblio/volume/59>. Accessed on: May 25, 2016.
http://www.nzsap.org/view/biblio/volume/...
). Recently, Tedeschi (2015)Tedeschi, L. O. 2015. Integrating genomics with nutrition models to improve the prediction of cattle performance and carcass composition under feedlot conditions. PLoS One 10: e0143483. used SNP panels to improve the predictability of days on feed to reach desired carcass composition of growing beef cattle by the Cattle Value Discovery System (Tedeschi et al., 2004Tedeschi, L. O.; Fox, D. G. and Guiroy, P. J. 2004. A decision support system to improve individual cattle management. 1. A mechanistic, dynamic model for animal growth. Agricultural Systems 79: 171-204.). In their analysis, the inclusion of the molecular breeding value of ribeye area increased by 13% the precision to predict body weight at 28% empty body fat. Despite the development of new concepts for nutrition modeling, revisions of relationships established a long time ago are still needed for three reasons: collection of new data and incorporation with old data (Ellis et al., 2014Ellis, J. L.; Dijkstra, J.; Bannink, A.; Kebreab, E.; Archibeque, S.; Benchaar, C.; Beauchemin, K. A.; Nkrumah, J. D. and France, J. 2014. Improving the prediction of methane production and representation of rumen fermentation for finishing beef cattle within a mechanistic model. Canadian Journal of Animal Science 94: 509-524.; Galyean and Tedeschi, 2014Galyean, M. L. and Tedeschi, L. O. 2014. Predicting microbial protein synthesis in beef cattle: Relationship to intakes of total digestible nutrients and crude protein. Journal of Animal Science 92: 5099-5111.); data used to develop past relationships were deemed deficient and/or were based on assumptions that do not hold anymore (Galyean et al., 2016Galyean, M. L.; Cole, N. A.; Tedeschi, L. O. and Branine, M. E. 2016. BOARD-INVITED REVIEW: Efficiency of converting digestible energy to metabolizable energy and reevaluation of the California Net Energy System maintenance requirements and equations for predicting dietary net energy values for beef cattle. Journal of Animal Science 94: 1329-1341.); or more advanced mathematical and/or statistical analyses have been proposed (Moraes et al., 2014aMoraes, L. E.; Kebreab, E.; Strathe, A. B.; France, J.; Dijkstra, J.; Casper, D. P. and Fadel, J. G. 2014a. Bayesian analysis of energy balance data from growing cattle using parametric and nonparametric modelling. Animal Production Science 54: 2068-2081.; Moraes et al., 2014bMoraes, L. E.; Strathe, A. B.; Fadel, J. G.; Casper, D. P. and Kebreab, E. 2014b. Prediction of enteric methane emissions from cattle. Global Change Biology 20: 2140-2148.).
Conclusions
Achieving future global demands for animal-sourced food in the face of finite natural resources and environmental-impact concerns will demand continued development and adoption of innovative technologies to further improve livestock productivity, refine resource-use efficiency, reduce animal-waste outputs, and foster sustainable use of our ruminant ecosystems. We described current and emerging technologies that can be used to ameliorate or solve the most challenging issues for livestock production that were addressed in a companion paper, including animal health and welfare, antibiotic resistance, and food safety/quality. Adoption of innovative technologies to advance management strategies and more precise diet formulation are needed to mitigate greenhouse gas emissions by ruminants, identify sustainable sources of protein feed (e.g., algae), and more effectively utilize marginal grassland. The continued use of omics technologies will enhance our understanding of fetal programming on animal performance and efficiency (e.g., residual feed intake), and the effect of the rumen microbiome on productivity, and efficiency of livestock to improve the quality of the animal product in sustainable systems. Precision livestock farming technologies have considerable potential to enable producers to make better-informed management decisions, mitigate disease threats, and improve welfare and genetic merit of livestock. We believe that those livestock producers, who successfully implement business plans that adopt proven and novel technological advancements that produce animal protein more efficiently and responsibly, will provide for a growing and hungry human population.
Acknowledgments
The theme was presented at the First International Meeting of Advances in Animal Science held at Jaboticabal, São Paulo, Brazil on June 8 to 10, 2016.
References
- Adams, A. E.; Olea-Popelka, F. J. and Roman-Muniz, I. N. 2013. Using temperature-sensing reticular boluses to aid in the detection of production diseases in dairy cows. Journal of Dairy Science 96: 1549-1555.
- AFRC - Agricultural and Food Research Council. 1992. AFRC Technical Committee on Responses to Nutrients, Report 9: Nutritive requirements of ruminant animals: Protein. Nutrition Abstracts and Reviews (Series B: Livestock Feeds and Feeding) 62: 787-835.
- Alhamad, M. N.; Stuth, J. and Vannucci, M. 2007. Biophysical modelling and NDVI time series to project near-term forage supply: spectral analysis aided by wavelet denoising and ARIMA modelling. International Journal of Remote Sensing 28: 2513-2548.
- Allen, C. C.; Alves, B. R. C.; Li, X.; Tedeschi, L. O.; Zhou, H.; Paschal, J. C.; Riggs, P. K.; Braga-Neto, U. M.; Keisler, D. H.; Williams, G. L. and Amstalden, M. 2012. Gene expression in the arcuate nucleus of heifers is affected by controlled intake of high- and low- concentrate diets. Journal of Animal Science 90: 2222-2232.
- Alves, B. R. C.; Cardoso, R. C.; Prezotto, L. D.; Thorson, J. F.; Bedenbaugh, M.; Sharpton, S. M.; Caraty, A.; Keisler, D. H.; Tedeschi, L. O.; Williams, G. L. and Amstalden, M. 2015. Elevated body weight gain during the juvenile alters neuropeptide Y-gonadotropin-releasing hormone circuitry in prepubertal heifers. Biology of Reproduction 92: 1-10.
- Amstalden, M.; Cardoso, R. C.; Alves, B. R. C. and Williams, G. L. 2014. Reproduction Symposium: Hypothalamic neuropeptides and the nutritional programming of puberty in heifers. Journal of Animal Science 92: 3211-3222.
- Angerer, J. P. 2012a. Gobi forage livestock early warning system. p. 115-130. In: Conducting National Feed Assessments. vol. 15. Coughenour, M. B. and Makkar, H. P. S., eds. Food and Agriculture Organization of the United Nations, Rome, Italy.
- Angerer, J. P. 2012b. Technologies, tools and methodologies for forage evaluation in grasslands and rangelands. p. 165-200. In: Conducting National Feed Assessments. vol. 15. Coughenour, M. B. and Makkar, H. P. S., eds. Food and Agriculture Organization of the United Nations, Rome, Italy.
- Archer, J. A.; Reverter, A.; Herd, R. M.; Johnston, D. J. and Arthur, P. F. 2002. Genetic variation in feed intake and efficiency of mature beef cows and relationships with post-weaning measurements. p. 221-224. In: Proceedings of the 7th World Congress Genetics Applied to Livestock Production, Montepellier, France.
- Archer, J. A.; Richardson, E. C.; Herd, R. M. and Arthur, P. F. 1999. Potential for selection to improve efficiency of feed use in beef cattle: a review. Australian Journal of Agricultural Research 50: 147-161.
- Arthur, P. F.; Archer, J. A.; Johnson, D. J.; Herd, R. M.; Richardson, E. C. and Parnell, P. F. 2001. Genetic and phenotypic variance and covariance components for feed intake, feed efficiency, and other postweaning traits in Angus cattle. Journal of Animal Science 79: 2805-2811.
- Arthur, P. F.; Herd, R. M.; Wilkins, J. F. and Archer, J. A. 2005. Maternal productivity of Angus cows divergently selected for post- weaning residual feed intake. Australian Journal of Experimental Agriculture 45: 985-993.
- Awda, B. J.; Miller, S. P.; Montanholi, Y. R.; Voort, G. V.; Caldwell, T.; Buhr, M. M. and Swanson, K. C. 2013. The relationship between feed efficiency traits and fertility in young beef bulls. Canadian Journal of Animal Science 93: 185-192.
- Baker, S. D.; Szasz, J. I.; Klein, T. A.; Kuber, P. S.; Hunt, C. W.; Glaze Jr., J. B.; Falk, D.; Richard, R.; Miller, J. C.; Battaglia, R. A. and Hill, R. A. 2006. Residual feed intake of purebred Angus steers: Effects on meat quality and palatability. Journal of Animal Science 84: 938-945.
- Basarab, J.; Baron, V.; López-Campos, Ó.; Aalhus, J.; Haugen- Kozyra, K. and Okine, E. 2012. Greenhouse gas emissions from calf- and yearling-fed beef production systems, with and without the use of growth promotants. Animals 2: 195-220.
- Basarab, J. A.; Colazo, M. G.; Ambrose, D. J.; Novak, S.; McCartney, D. and Baron, V. S. 2011. Residual feed intake adjusted for backfat thickness and feeding frequency is independent of fertility in beef heifers. Canadian Journal of Animal Science 91: 573-584.
- Basarab, J. A.; McCartney, D.; Okine, E. K. and Baron, V. S. 2007. Relationships between progeny residual feed intake and dam productivity traits. Canadian Journal of Animal Science 87: 489-502.
- Basher, R. 2006. Global early warning systems for natural hazards: Systematic and people-centred. Philosophical Transactions of the Royal Society of London A: Mathematical, Physical and Engineering Sciences 364: 2167-2182.
- Behrens, J. W.; Miller, R. K.; Bailey, J. C. Walter, J. T.; Hafla, A. N.; Mendes, E. D.; Hale, D. S.; Machado, T.; Tedeschi, L. O. and Carstens, G. E. 2011. Effects of residual feed intake classification and breed type on carcass characteristics, tenderness and value in feedlot heifers. p. 761. In: Proceedings of the Joint Annual Meeting of American Society of Animal Science and American Dairy Science Association, 89, New Orleans, LA. Federation of Animal Science Societies. Available at: <http://www.jtmtg.org/JAM/2011/toc.asp>. Accessed on: May 21, 2016.
» http://www.jtmtg.org/JAM/2011/toc.asp - Bell, A. W. 2006. Prenatal programming of postnatal productivity and health of livestock: a brief review. Australian Journal of Experimental Agriculture 46: 725-732.
- Bell, A. W.; Greenwood, P. L. and Ehrhardt, R. A. 2005. Regulation of metabolism and growth during prenatal life. p. 3-34. In: Biology of growing animals. vol. 3. Burrin, D. G. and Mersmann, H. J., eds. Elsevier, Amsterdam.
- Bento, C. B. P.; Azevedo, A. C.; Mantovani, H. C.; Gomes, D. I.; Batista, E. D.; Rufino, L. M. A. and Detmann, E. 2015. Effect of protein supplementation on ruminal parameters and microbial community fingerprint of Nellore steers fed tropical forages. Animal 10: 44-54.
- Berckmans, D. 2015. Smart farming for Europe: Value creation through precision livestock farming. p. 25-35. In: Precision livestock farming applications. Halachmi, I., ed Wageningen Academic Publishers, Wageningen, NL.
- Bestelmeyer, B. T. and Briske, D. D. 2012. Grand challenges for resilience-based management of rangelands. Rangeland Ecology & Management 65: 654-663.
- Bewley, J. 2013. Exciting dairy breakthroughs: Science fiction of precision dairy farming? p. 1-8. In: Proceedings of the Precision Dairy Conference and Exp, Rochester, MN. University of Minnesota. Available at: <http://www.precisiondairyfarming.com/2015/>. Accessed on: May 23, 2016.
» http://www.precisiondairyfarming.com/2015/ - Bewley, J. M.; Russell, R. A.; Dolecheck, K. A. and Borchers, M. R. 2015. Precision dairy monitoring: What have we learned? p. 15-23. In: Precision Livestock Farming Applications. Halachmi, I., ed Wageningen Academic Publishers, Wageningen, NL.
- Black, T. E.; Bischoff, K. M.; Mercadante, V. R. G.; Marquezini, G. H. L.; Dilorenzo, N.; Chase, C. C., Jr.; Coleman, S. W.; Maddock, T. D. and Lamb, G. C. 2013. Relationships among performance, residual feed intake, and temperament assessed in growing beef heifers and subsequently as 3-year-old, lactating beef cows. Journal of Animal Science 91: 2254-2263.
- Bork, E. W. and Su, J. G. 2007. Integrating LIDAR data and multispectral imagery for enhanced classification of rangeland vegetation: A meta analysis. Remote Sensing of Environment 111: 11-24.
- Bottje, W. G. and Carstens, G. E. 2009. Association of mitochondrial function and feed efficiency in poultry and livestock species. Journal of Animal Science 87: E48-63.
- Boval, M. and Dixon, R. M. 2012. The importance of grasslands for animal production and other functions: a review on management and methodological progress in the tropics. Animal 6: 748-762.
- Bruegger, R. A.; Varelas, L. A.; Howery, L. D.; Torell, L. A.; Stephenson, M. B. and Bailey, D. W. 2016. Targeted grazing in Southern Arizona: Using cattle to reduce fine fuel loads. Rangeland Ecology & Management 69: 43-51.
- Capper, J. L. and Cady, R. A. 2010. The environmental impact of corn- fed vs. grass-fed beef finishing systems. p. 686. In: Proceedings of the Joint Annual Meeting of American Society of Animal Science and American Dairy Science Association, Denver, CO. Federation of Animal Science Societies/Federation of Animal Science Societies. Available at: <http://www.jtmtg.org/JAM/2010/toc.asp>. Accessed on: May 21, 2016.
» http://www.jtmtg.org/JAM/2010/toc.asp - Cardoso, R. C.; Alves, B. R. C.; Prezotto, L. D.; Thorson, J. F.; Tedeschi, L. O.; Keisler, D. H.; Amstalden, M. and Williams, G. L. 2014a. Reciprocal changes in leptin and NPY during nutritional acceleration of puberty in heifers. Journal of Endocrinology 223: 289-298.
- Cardoso, R. C.; Alves, B. R. C.; Prezotto, L. D.; Thorson, J. F.; Tedeschi, L. O.; Keisler, D. H.; Park, C. S.; Amstalden, M. and Williams, G. L. 2014b. Use of a stair-step compensatory gain nutritional regimen to program the onset of puberty in beef heifers. Journal of Animal Science 92: 2942-2949.
- CSIRO - Commonwealth Scientific and Industrial Research Organization. 2007. Nutrient requirements of domesticated ruminants. Commonwealth Scientific and Industrial Research Organization, Collingwood, VIC.
- Crowley, J. J.; Evans, R. D.; McHugh, N.; Kenny, D. A.; McGee, M.; Crews, D. H. and Berry, D. P. 2011. Genetic relationships between feed efficiency in growing males and beef cow performance. Journal of Animal Science 89: 3372-3381.
- Daniels, T. K.; Bowman, J. G. P.; Sowell, B. F.; Branine, M. E. and Hubbert, M. E. 2000. Effects of metaphylactic antibiotics on behavior of feedlot calves. Professional Animal Scientist 16: 247-253.
- De Barbieri, I.; Hegarty, R. S.; Silveira, C.; Gulino, L. M.; Oddy, V. H.; Gilbert, R. A.; Klieve, A. V. and Ouwerkerk, D. 2015. Programming rumen bacterial communities in newborn Merino lambs. Small Ruminant Research 129: 48-59.
- De Buck, J.; Shaykhutdinov, R.; Barkema, H. W. and Vogel, H. J. 2014. Metabolomic profiling in cattle experimentally infected with Mycobacterium avium subsp. paratuberculosis. PLoS One 9: 1-11.
- De Vries, A. and Reneau, J. K. 2010. Application of statistical process control charts to monitor changes in animal production systems. Journal of Animal Science 88: E11-E24.
- Detmann, E.; Valente, E. E. L.; Batista, E. D. and Huhtanen, P. 2014. An evaluation of the performance and efficiency of nitrogen utilization in cattle fed tropical grass pastures with supplementation. Livestock Science 162: 141-153.
- Di Marco, O. N. and Baldwin, R. L. 1989. Implementation and evaluation of a steer growth model. Agricultural Systems 29: 247-265.
- Di Marco, O. N.; Baldwin, R. L. and Calvert, C. C. 1989. Simulation of DNA, protein and fat accretion in growing steers. Agricultural Systems 29: 21-34.
- Dixon, R. M. and Coates, D. B. 2010. Diet quality estimated with faecal near infrared reflectance spectroscopy and responses to N supplementation by cattle grazing buffel grass pastures. Animal Feed Science and Technology 158: 115-125.
- Donoghue, K. A.; Arthur, P. F.; Wilkins, J. F. and Herd, R. M. 2011. Onset of puberty and early-life reproduction in Angus females divergently selected for post-weaning residual feed intake. Animal Production Science 51: 183-190.
- Du, M.; Tong, J.; Zhao, J.; Underwood, K. R.; Zhu, M.; Ford, S. P. and Nathanielsz, P. W. 2010. Fetal programming of skeletal muscle development in ruminant animals. Journal of Animal Science 88: E51-E60.
- Dubeux, J. C. B., Jr.; Muir, J. P.; Nair, P. K. R.; Sollenberger, L. E.; Silva, H. M. S. and Mello, A. C. L. 2015. The advantages and challenges of integrating tree legumes into pastoral systems. p. 141-164. In: Proceedings of the 1st International Conference on Forages in Warm Climates, Lavras, Brazil. University of Lavras. Available at: <http://www.neforufla.com.br/index.php/en/pg/9/proceedings-confor-2015>. Accessed on: May 13, 2016.
» http://www.neforufla.com.br/index.php/en/pg/9/proceedings-confor-2015 - Durso, L. M.; Harhay, G. P.; Smith, T. P. L.; Bono, J. L.; Desantis, T. Z.; Harhay, D. M.; Andersen, G. L.; Keen, J. E.; Laegreid, W. W. and Clawson, M. L. 2010. Animal-to-animal variation in fecal microbial diversity among beef cattle. Applied and Environmental Microbiology 76: 4858-4862.
- Durunna, O. N.; Mujibi, F. D. N.; Goonewardene, L.; Okine, E. K.; Basarab, J. A.; Wang, Z. and Moore, S. S. 2011. Feed efficiency differences and reranking in beef steers fed grower and finisher diets. Journal of Animal Science 89: 158-167.
- Elam, T. E. and Preston, R. L. 2004. Fifty years of pharmaceutical technology and its Impact on the beef we provide to consumers. Growth Enhancement Technology Information Team, Johnson, IA. 36p. Available at: <http://www.merck-animal-health-usa.com/binaries/50_Years_of_Technology_and_impact_on_beef_production_tcm96-113484.pdf>. Accessed on: May 21, 2016.
» http://www.merck-animal-health-usa.com/binaries/50_Years_of_Technology_and_impact_on_beef_production_tcm96-113484.pdf - Ellis, J. L.; Dijkstra, J.; Bannink, A.; Kebreab, E.; Archibeque, S.; Benchaar, C.; Beauchemin, K. A.; Nkrumah, J. D. and France, J. 2014. Improving the prediction of methane production and representation of rumen fermentation for finishing beef cattle within a mechanistic model. Canadian Journal of Animal Science 94: 509-524.
- Ellis, W. C.; Poppi, D. P.; Matis, J. H.; Lippke, H.; Hill, T. M. and Rouquette, F. M., Jr. 1999. Dietary-digestible-metabolic interactions determining the nutritive potential of ruminant diets. p. 423-481. In: Proceedings of the 5th International Symposium on the Nutrition of Herbivores, San Antonio, TX. American Society of Animal Science.
- Ferrell, C. L. 1991. Maternal and fetal influences on uterine and conceptus development in the cow: I. Growth of tissues of the gravid uterus. Journal of Animal Science 69: 1945-53.
- Ferrell, C. L.; Garrett, W. N. and Hinman, N. 1976. Growth, development and composition of the udder and gravid uterus of beef heifers during pregnancy. Journal of Animal Science 42: 1477-1489.
- Ferrell, C. L. and Oltjen, J. W. 2008. ASAS CENTENNIAL PAPER: Net energy systems for beef cattle—Concepts, application, and future models. Journal of Animal Science 86: 2779-2794.
- FAO - Food and Agriculture Organization; World Organization for Animal Health (OIE) and World Health Organization (WHO). 2013. GLEWS+: The Joint FAO-OIE-WHO Global Early Warning System for Health Threats and Emerging Risks at the Human-Animal-Ecosystems Interface; A Concept Note. Available at: <http://www.glews.net/wp-content/uploads/2013/12/04_GLEWSConcept-20-11.pdf>. Accessed on: May 17, 2016.
» http://www.glews.net/wp-content/uploads/2013/12/04_GLEWSConcept-20-11.pdf - Fox, D. G.; Tedeschi, L. O.; Tylutki, T. P.; Russell, J. B.; Van Amburgh, M. E.; Chase, L. E.; Pell, A. N. and Overton, T. R. 2004. The Cornell Net Carbohydrate and Protein System model for evaluating herd nutrition and nutrient excretion. Animal Feed Science and Technology 112: 29-78.
- Franzel, S.; Carsan, S.; Lukuyu, B.; Sinja, J. and Wambugu, C. 2014. Fodder trees for improving livestock productivity and smallholder livelihoods in Africa. Current Opinion in Environmental Sustainability 6: 98-103.
- Gabory, A.; Attig, L. and Junien, C. 2009. Sexual dimorphism in environmental epigenetic programming. Molecular and Cellular Endocrinology 304: 8-18.
- Gallo, L. A.; Tran, M.; Moritz, K. M. and Wlodek, M. E. 2013. Developmental programming: Variations in early growth and adult disease. Clinical & Experimental Pharmacology & Physiology 40: 795-802.
- Galyean, M. L.; Cole, N. A.; Tedeschi, L. O. and Branine, M. E. 2016. BOARD-INVITED REVIEW: Efficiency of converting digestible energy to metabolizable energy and reevaluation of the California Net Energy System maintenance requirements and equations for predicting dietary net energy values for beef cattle. Journal of Animal Science 94: 1329-1341.
- Galyean, M. L.; Ponce, C. and Schutz, J. 2011. The future of beef production in North America. Animal Frontiers 1: 29-36.
- Galyean, M. L. and Tedeschi, L. O. 2014. Predicting microbial protein synthesis in beef cattle: Relationship to intakes of total digestible nutrients and crude protein. Journal of Animal Science 92: 5099-5111.
- Gilbert, R. A. and Klieve, A. V. 2015. Ruminal viruses (Bacteriophages, Archaeaphages). p. 121-141. In: Rumen microbiology: From evolution to revolution. Puniya, K. A.; Singh, R. and Kamra, N. D., eds. Springer India, New Delhi.
- Gilbert, R. A.; Ouwerkerk, D. and Klieve, A. V. 2015. Phage therapy in livestock methane amelioration. p. 318-335. In: Livestock production and climate change. Malik, P. K.; Bhatta, R.; Takahashi, J., et al., eds. CABI Publishing, Boston, MA.
- Godfray, H. C. J.; Beddington, J. R.; Crute, I. R.; Haddad, L.; Lawrence, D.; Muir, J. F.; Pretty, J.; Robinson, S.; Thomas, S. M. and Toulmin, C. 2010. Food security: The challenge of feeding 9 billion people. Science 327: 812-818.
- González-García, E.; Gourdine, J. L.; Alexandre, G.; Archimède, H. and Vaarst, M. 2012. The complex nature of mixed farming systems requires multidimensional actions supported by integrative research and development efforts. Animal 6: 763-777.
- González, L. A.; Tolkamp, B. J.; Coffey, M. P.; Ferret, A. and Kyriazakis, I. 2008. Changes in feeding behavior as possible indicators for the automatic monitoring of health disorders in dairy cows. Journal of Dairy Science 91: 1017-1028.
- Gordon, I. J. and Prins, H. H. T. 2008. The ecology of browsing and grazing. Ecological studies. vol. 195. Springer, Berlin, Germany.
- Greenwood, P. L.; Bishop-Hurley, G. J.; González, L. A. and Ingham, A. B. 2016. Development and application of a livestock phenomics platform to enhance productivity and efficiency at pasture. Animal Production Science 56: 1299-1311.
- Guiroy, P. J.; Fox, D. G.; Tedeschi, L. O.; Baker, M. J. and Cravey, M. D. 2001. Predicting individual feed requirements of cattle fed in groups. Journal of Animal Science 79: 1983-1995.
- Hafla, A. N.; Carstens, G. E.; Forbes, T. D. A.; Tedeschi, L. O.; Bailey, J. C.; Walter, J. T. and Johnson, J. R. 2013. Relationships between postweaning residual feed intake in heifers and forage use, body composition, feeding behavior, physical activity, and heart rate of pregnant beef females. Journal of Animal Science 91: 5353-5365.
- Hafla, A. N.; Lancaster, P. A.; Carstens, G. E.; Forrest, D. W.; Fox, J. T.; Forbes, T. D. A.; Davis, M. E.; Randel, R. D. and Holloway, J. W. 2012a. Relationships between feed efficiency, scrotal circumference, and semen quality traits in yearling bulls. Journal of Animal Science 90: 3937-3944.
- Hafla, A. N.; Walter, J. T.; Carstens, G. E.; Bailey, J. C.; Tedeschi, L. O.; Moreno, J. G.; Hale, D. S.; Miller, R. K.; Sawyer, J. E. and Anderson, D. P. 2012b. Impact of between-animal variation in performance, carcass-quality and feed efficiency on profitability of Angus-based composite steers. p. 125-126. In: Proceedings of the Plains Nutrition Council Conference, San Antonio, TX. PNC. Available at: <http://amarillo.tamu.edu/facultystaff/tmc/pnc/>. Accessed on: May 9, 2017.
» http://amarillo.tamu.edu/facultystaff/tmc/pnc/ - Hailemariam, D.; Mandal, R.; Saleem, F.; Dunn, S. M.; Wishart, D. S. and Ametaj, B. N. 2014. Identification of predictive biomarkers of disease state in transition dairy cows. Journal of Dairy Science 97: 2680-2693.
- Harper, K. J.; Klieve, A. V.; Panjaitan, T.; Poppi, D. P. and Ouwerkerk, D. 2010. Changes in rumen bacterial community in steers fed with supplemented tropical pasture. p. 65. In: Proceedings of the Australian Society of Animal Production, 28. Available at: <http://www.asap.asn.au/livestocklibrary/2010/harper065.pdf>. Accessed on: May 9, 2017.
» http://www.asap.asn.au/livestocklibrary/2010/harper065.pdf - Hegarty, R. S.; Goopy, J. P.; Herd, R. M. McCorkell, B. 2007. Cattle selected for lower residual feed intake have reduced daily methane production. Journal of Animal Science 85: 1479-1486.
- Herd, R. M. and Arthur, P. F. 2009. Physiological basis for residual feed intake. Journal of Animal Science 87: E64-71.
- Hernandez-Sanabria, E.; Guan, L. L.; Goonewardene, L. A.; Li, M.; Mujibi, D. F.; Stothard, P.; Moore, S. S. and Leon-Quintero, M. C. 2010. Correlation of particular bacterial PCR-denaturing gradient gel electrophoresis patterns with bovine ruminal fermentation parameters and feed efficiency traits. Applied and Environmental Microbiology 76: 6338-6350.
- Jackson, K. S. 2015. Associations of feeding behavior patterns with inter-animal variation in feed efficiency and pre-clinical responses to infectious disease in beef cattle. Thesis. Texas A&M University, College Station, TX.
- Jackson, K. S.; Carstens, G. E.; Tedeschi, L. O. and Pinchak, W. E. 2016. Changes in feeding behavior patterns and dry matter intake before clinical symptoms associated with bovine respiratory disease in growing bulls. Journal of Animal Science 94: 1644-1652.
- Jami, E.; White, B. A. and Mizrahi, I. 2014. Potential role of the bovine rumen microbiome in modulating milk composition and feed efficiency. PLoS One 9: e85423.
- Johnson, D. E.; Ferrell, C. L. and Jenkins, T. G. 2003. The history of energetic efficiency research: Where have we been and where are we going? Journal of Animal Science 81: E27-E38.
- Karisa, B. K.; Thomson, J.; Wang, Z.; Li, C.; Montanholi, Y. R.; Miller, S. P.; Moore, S. S. and Plastow, G. S. 2014. Plasma metabolites associated with residual feed intake and other productivity performance traits in beef cattle. Livestock Science 165: 200-211.
- Karisa, B. K.; Thomson, J.; Wang, Z.; Stothard, P.; Moore, S. S. and Plastow, G. S. 2013. Candidate genes and single nucleotide polymorphisms associated with variation in residual feed intake in beef cattle. Journal of Animal Science 91: 3502-3513.
- Ku, N. -W.; Popescu, S. C.; Ansley, R. J.; Perotto-Baldivieso, H. L. and Filippi, A. M. 2012. Assessment of Available Rangeland Woody Plant Biomass with a Terrestrial Lidar System. Photogrammetric Engineering & Remote Sensing 78: 349-361.
- Lancaster, P. A.; Carstens, G. E.; Crews, D. H., Jr.; Welsh Jr., T. H.; Forbes, T. D. A.; Forrest, D. W.; Tedeschi, L. O.; Randel, R. D. and Rouquette, F. M., Jr. 2009. Phenotypic and genetic relationships of residual feed intake with performance and ultrasound carcass traits in Brangus heifers. Journal of Animal Science 87: 3887-3896.
- Langley-Evans, S. C. 2006. Developmental programming of health and disease. Proceedings of the Nutrition Society 65: 97-105.
- Larue, R.; Yu, Z.; Parisi, V. A.; Egan, A. R. and Morrison, M. 2005. Novel microbial diversity adherent to plant biomass in the herbivore gastrointestinal tract, as revealed by ribosomal intergenic spacer analysis and rrs gene sequencing. Environmental Microbiology 7: 530-543.
- Leite, E. R. and Stuth, J. W. 1995. Fecal NIRS equations to assess diet quality of free-ranging goats. Small Ruminant Research 15: 223-230.
- Li, H.; Tolleson, D.; Stuth, J.; Bai, K.; Mo, F. and Kronberg, S. 2007. Faecal near infrared reflectance spectroscopy to predict diet quality for sheep. Small Ruminant Research 68: 263-268.
- Lobato, J. F. P.; Freitas, A. K.; Devincenzi, T.; Cardoso, L. L.; Tarouco, J. U.; Vieira, R. M.; Dillenburg, D. R. and Castro, I. 2014. Brazilian beef produced on pastures: Sustainable and healthy. Meat Science 98: 336-345.
- Lyons, R. K. and Stuth, J. W. 1992. Fecal NIRS equations for predicting diet quality of free-ranging cattle. Journal of Range Management 45: 238-244.
- MacDonald, K. A.; Pryce, J. E.; Spelman, R. J.; Davis, S. R.; Wales, W. J.; Waghorn, G. C.; Williams, Y. J.; Marett, L. C. and Hayes, B. J. 2014. Holstein-Friesian calves selected for divergence in residual feed intake during growth exhibited significant but reduced residual feed intake divergence in their first lactation. Journal of Dairy Science 97: 1427-1435.
- Machovina, B.; Feeley, K. J. and Ripple, W. J. 2015. Biodiversity conservation: The key is reducing meat consumption. Science of The Total Environment 536: 419-431.
- Mackie, R. I.; McSweeney, C. S. and Klieve, A. V. 2002. Microbial ecology of the ovine rumen. p. 71-94. In: Sheep nutrition. Freer, M. and Dove, H., eds. CABI Publishing, Cambridge, MA.
- Marr, B. 2015. Big data: Using smart big data, analytics and metrics to make better decisions and improve performance. John Wiley & Sons Ltd, Chichester, West Sussex, United Kingdom.
- Martinez, E. D.; Klieve, A. V.; Ouwerkerk, D.; Swain, T.; Gulino, L. M.; Turnbull, K. and Poppi, D. P. 2010. Between animal variance in ruminal bacteria and protozoal communities from DGGE profiles of steers on a low quality forage diet. p. 63. In: Proceedings of the Australian Society of Animal Production, 28. Available at: <http://www.asap.asn.au/livestocklibrary/2010/martinez063.pdf>. Accessed on: May 9, 2017.
» http://www.asap.asn.au/livestocklibrary/2010/martinez063.pdf - McCann, J. C.; Drewery, M. L.; Sawyer, J. E.; Pinchak, W. E. and Wickersham, T. A. 2014a. Effect of postextraction algal residue supplementation on the ruminal microbiome of steers consuming low-quality forage. Journal of Animal Science 92: 5063-5075.
- McCann, J. C.; Wiley, L. M.; Forbes, T. D.; Rouquette, F. M., Jr. and Tedeschi, L. O. 2014b. Relationship between the rumen microbiome and residual feed intake-efficiency of Brahman bulls stocked on Bermudagrass pastures. PLoS One 9: e91864.
- McMillen, I. C. and Robinson, J. S. 2005. Developmental origins of the metabolic syndrome: Prediction, plasticity, and programming. Physiological Reviews 85: 571-633.
- Medina-Cetina, Z. and Nadim, F. 2008. Stochastic design of an early warning system. Georisk: Assessment and Management of Risk for Engineered Systems and Geohazards 2: 223-236.
- Mertens, K.; Decuypere, E.; De Baerdemaeker, J. and De Ketelaere, B. 2011. Statistical control charts as a support tool for the management of livestock production. The Journal of Agricultural Science 149: 369-384.
- Montanholi, Y. R.; Swanson, K. C.; Schenkel, F. S.; McBride, B. W.; Caldwell, T. R. and Miller, S. P. 2009. On the determination of residual feed intake and associations of infrared thermography with efficiency and ultrasound traits in beef bulls. Livestock Science 125: 22-30.
- Moraes, L. E.; Kebreab, E.; Strathe, A. B.; France, J.; Dijkstra, J.; Casper, D. P. and Fadel, J. G. 2014a. Bayesian analysis of energy balance data from growing cattle using parametric and nonparametric modelling. Animal Production Science 54: 2068-2081.
- Moraes, L. E.; Strathe, A. B.; Fadel, J. G.; Casper, D. P. and Kebreab, E. 2014b. Prediction of enteric methane emissions from cattle. Global Change Biology 20: 2140-2148.
- Moritz, K.; Wintour-Coghlan, E. M.; Black, M. J.; Bertram, J. F. and Caruana, G. 2008. Factors Influencing Mammalian Kidney Development: Implications for Health in Adult Life. Advances in Anatomy, Embryology and Cell Biology. vol. 196. Springer Berlin Heidelberg, New York, NY.
- Mossa, F.; Walsh, S. W.; Ireland, J. J. and Evans, A. C. O. 2015. Early nutritional programming and progeny performance: Is reproductive success already set at birth? Animal Frontiers 5: 18-24.
- Moya, D.; Silasi, R.; McAllister, T. A.; Genswein, B.; Crowe, T.; Marti, S. and Schwartzkopf-Genswein, K. S. 2015. Use of pattern recognition techniques for early detection of morbidity in receiving feedlot cattle. Journal of Animal Science 93: 3623-3638.
- Muir, J. P.; Pitman, W. D.; Dubeux, J. C. and Foster, J. L. 2014. The future of warm-season, tropical and subtropical forage legumes in sustainable pastures and rangelands. African Journal of Range & Forage Science 31: 187-198.
- Muir, J. P.; Pitman, W. D. and Foster, J. L. 2011. Sustainable, low- input, warm-season, grass-legume grassland mixtures: mission (nearly) impossible? Grass and Forage Science 66: 301-315.
- Mutlu, M.; Popescu, S. C.; Stripling, C. and Spencer, T. 2008. Mapping surface fuel models using lidar and multispectral data fusion for fire behavior. Remote Sensing of Environment 112: 274-285.
- Myer, P. R.; Smith, T. P. L.; Wells, J. E.; Kuehn, L. A. and Freetly, H. C. 2015. Rumen microbiome from steers differing in feed efficiency. PLoS One 10: e0129174.
- NASEM - National Academies of Sciences, Engineering, and Medicine. 2016. Nutrient requirements of beef cattle. 8th ed. Nutrient requirements of domestic animals. National Academy Press, Washington, DC.
- Ndove, T. S.; Whitbread, A. M.; Clark, R. A. and Pengelly, B. C. 2004. Identifying the factors that contribute to the successful adoption of improved farming practices in the smallholder sector of Limpopo Province, South Africa. p. 146-153. In: Tropical legumes for sustainable farming systems in Southern Africa and Australia. ACIAR (Australian Centre for International Agricultural Research) Proceedings Canberra, Australia. ACIAR. Available at: <http://aciar.gov.au/publication/pr115>. Accessed on: May 12, 2016.
» http://aciar.gov.au/publication/pr115 - Oltjen, J. W.; Bywater, A. C.; Baldwin, R. L. and Garrett, W. N. 1986. Development of a dynamic model of beef cattle growth and composition. Journal of Animal Science 62: 86-97.
- Oltjen, J. W.; Pleasants, A. B.; Soboleva, T. K. and Oddy, V. H. 2000. Second-generation dynamic cattle growth and composition models. p. 197-209. In: Modelling nutrient utilization in farm animals. McNamara, J. P.; France, J. and Beever, D. E., eds. CABI Publishing, New York, NY.
- Ouwerkerk, D.; Turner, A. F. and Klieve, A. V. 2008. Diversity of methanogens in ruminants in Queensland. Australian Journal of Experimental Agriculture 48: 722-725.
- Perkins, S. D.; Key, C. N.; Garrett, C. F.; Foradori, C. D.; Bratcher, C. L.; Kriese-Anderson, L. A. and Brandebourg, T. D. 2014. Residual feed intake studies in Angus-sired cattle reveal a potential role for hypothalamic gene expression in regulating feed efficiency. Journal of Animal Science 92: 549-560.
- Peterson, B.; Nelson, K. J.; Seielstad, C.; Stoker, J.; Jolly, W. M. and Parsons, R. 2015. Automated integration of lidar into the LANDFIRE product suite. Remote Sensing Letters 6: 247-256.
- Peterson, H. C. 2013. Sustainability: a wicked problem. p. 1-9. In: Sustainable animal agriculture. Kebreab, E., ed CABI Publishing, Wallingford, UK.
- Poppi, D. P. and McLennan, S. R. 2010. Nutritional research to meet future challenges. Animal Production Science 50: 329-338.
- Poppi, D. P.; McLennan, S. R.; Bediye, S.; De Vega, A. and Zorrilla- Rios, J. 1997. Forage quality: Strategies for increasing nutritive value of forages. p. 307-322. In: Proceedings of the 18th International Grassland Congress, Winnipeg and Saskatoon, Canada. Canadian Forage Council, Canadian Society of Agronomy, Canadian Society of Animal Science, and Association Management Centre. Available at: <http://www.internationalgrasslands.org/>. Accessed on: May 15, 2016.
» http://www.internationalgrasslands.org/ - Quimby, W. F.; Sowell, B. F. and Bowman, J. G. P. 2001. Application of feeding behaviour to predict morbidity of newly received calves in a commercial feedlot. Canadian Journal of Animal Science 81: 315-320.
- Rey, M.; Enjalbert, F.; Combes, S.; Cauquil, L.; Bouchez, O. and Monteils, V. 2014. Establishment of ruminal bacterial community in dairy calves from birth to weaning is sequential. Journal of Applied Microbiology 116: 245-257.
- Riggs, P. K.; Tedeschi, L. O.; Turner, N. D.; Braga-Neto, U. and Jayaraman, A. 2017. The role of “omics” technologies for livestock sustainability. Archivos Latinoamericanos de Producción Animal (in press).
- Rittel, H. W. J. and Webber, M. M. 1973. Dilemmas in a general theory of planning. Policy Sciences 4: 155-169.
- Robinson, D. L. and Oddy, V. H. 2004. Genetic parameters for feed efficiency, fatness, muscle area and feeding behaviour of feedlot finished beef cattle. Livestock Production Science 90: 255-270.
- Roehe, R.; Dewhurst, R. J.; Duthie, C. -A.; Rooke, J. A.; McKain, N.; Ross, D. W.; Hyslop, J. J.; Waterhouse, A.; Freeman, T. C.; Watson, M. and Wallace, R. J. 2016. Bovine host genetic variation influences rumen microbial methane production with best selection criterion for low methane emitting and efficiently feed converting hosts based on metagenomic gene abundance. PLoS Genetics 12: 1-20.
- Rolf, M. M.; Taylor, J. F.; Schnabel, R. D.; McKay, S. D.; McClure, M. C.; Northcutt, S. L.; Kerley, M. S. and Weaber, R. L. 2012. Genome-wide association analysis for feed efficiency in Angus cattle. Animal Genetics 43: 367-374.
- Romero-Perez, A.; Okine, E. K.; McGinn, S. M.; Guan, L. L.; Oba, M.; Duval, S. M.; Kindermann, M. and Beauchemin, K. A. 2014. The potential of 3-nitrooxypropanol to lower enteric methane emissions from beef cattle. Journal of Animal Science 92: 4682-4693.
- Rouquette, F. M., Jr.; Redmon, L. A.; Aiken, G. E.; Hill, G. M.; Sollenberger, L. E. and Andrae, J. 2009. ASAS Centennial Paper: Future needs of research and extension in forage utilization. Journal of Animal Science 87: 438-446.
- Rutten, C. J.; Velthuis, A. G. J.; Steeneveld, W. and Hogeveen, H. 2013. Invited review: Sensors to support health management on dairy farms. Journal of Dairy Science 96: 1928-1952.
- Saatchi, M.; Beever, J. E.; Decker, J. E.; Faulkner, D. B.; Freetly, H. C.; Hansen, S. L.; Yampara-Iquise, H.; Johnson, K. A.; Kachman, S. D.; Kerley, M. S.; Kim, J.; Loy, D. D.; Marques, E.; Neibergs, H. L.; Pollak, E. J.; Schnabel, R. D.; Seabury, C. M.; Shike, D. W.; Snelling, W. M.; Spangler, M. L.; Weaber, R. L.; Garrick, D. J. and Taylor, J. F. 2014. QTLs associated with dry matter intake, metabolic mid-test weight, growth and feed efficiency have little overlap across 4 beef cattle studies. BMC Genomics 15: 1-14.
- Schaefer, A. L.; Cook, N. J.; Bench, C.; Chabot, J. B.; Colyn, J.; Liu, T.; Okine, E. K.; Stewart, M. and Webster, J. R. 2012. The noninvasive and automated detection of bovine respiratory disease onset in receiver calves using infrared thermography. Research in Veterinary Science 93: 928-935.
- Schaefer, A. L.; Cook, N. J.; Church, J. S.; Basarab, J.; Perry, B.; Miller, C. and Tong, A. K. W. 2007. The use of infrared thermography as an early indicator of bovine respiratory disease complex in calves. Research in Veterinary Science 83: 376-384.
- Schenkel, F. S.; Devitt, C. J. B.; Wilton, J. W.; Miller, S. P. and Jamrozik, J. 2004. Random regression analyses of feed intake of individually tested beef steers. Livestock Production Science 88: 129-142.
- Senay, G. B.; Velpuri, N. M.; Alemu, H.; Md Pervez, S.; Asante, K. O.; Kariuki, G.; Taa, A. and Angerer, J. 2013. Establishing an operational waterhole monitoring system using satellite data and hydrologic modelling: Application in the pastoral regions of East Africa. Pastoralism: Research, Policy and Practice 3: 1-16.
- Shaw, C. N.; Kim, M.; Eastridge, M. L. and Yu, Z. 2015. Effects of different sources of physically effective fiber on rumen microbial populations. Animal 10: 410-417.
- Shelton, H. M.; Franzel, S. and Peters, M. 2005. Adoption of tropical legume technology around the world: analysis of success. Tropical Grasslands 39: 198-209.
- Singh, S. and Kundu, S. S. 2010. Effect of tropical browse leaves supplementation on rumen enzymes of sheep and goats fed Dichanthium annulatum grass-based diets. Tropical Animal Health and Production 42: 1181-1187.
- Soboleva, T. K.; Oddy, V. H.; Pleasants, A. B.; Oltjen, J. W.; Ball, A. J. and McCall, D. G. 1999. A dynamical model of body composition in sheep. p. 275-278. In: Proceedings of the New Zealand Society of Animal Production, 59. Available at: <http://www.nzsap.org/view/biblio/volume/59>. Accessed on: May 25, 2016.
» http://www.nzsap.org/view/biblio/volume/59 - Sowell, B. F.; Branine, M. E.; Bowman, J. G. P.; Hubbert, M. E.; Sherwood, H. E. and Quimby, W. 1999. Feeding and watering behavior of healty and morbid steers in a commercial feedlot. Journal of Animal Science 77: 1105-1112.
- Steiner, J. L.; Coleman, S. W.; Starks, P. J.; Engle, D. M.; Xiao, X.; Saleh, A.; Osei, E.; Anandhi, A.; Tomlinson, P.; Rice, C. W.; Devlin, D.; Moffet, C.; Reuter, R.; Basara, J.; Cole, N. A.; Gowda, P.; Todd, R.; Middendorf, G. and Ocshner, T. 2014. Knowledge and tools to enhance resilience of beef grazing systems for sustainable animal protein production. Annals of the New York Academy of Sciences 1328: 10-17.
- Stuth, J.; Angerer, J.; Kaitho, R.; Zander, K.; Jama, A.; Heath, C.; Bucher, J.; Hamilton, W.; Conner, R. and Inbody, D. 2003. The livestock early warning system (LEWS): Blending technology and the human dimension to support grazing decisions. Available at: <http://cals.arizona.edu/OALS/ALN/aln53/stuth.html>. Accessed on: May 16, 2016.
» http://cals.arizona.edu/OALS/ALN/aln53/stuth.html - Stuth, J. W.; Angerer, J.; Kaitho, R.; Jama, A. and Marambii, R. 2005. Livestock early warning system for Africa's rangelands. p. 283-294. In: Monitoring and predicting agricultural drought: A Global study. Boken, V. K.; Cracknell, A. P. and Heathcote, R. L., eds. Oxford University Press, Oxford, UK.
- Stuth, J. W.; Lyons, R. K.; Angerer, J. P. and McKown, C. D. 1991. Role of NIRS-based nutritional monitoring systems for grazing and nutritional management of range livestock. p. 83-93. In: Proceedings of the 2nd Grazing Livestock Nutrition Conference, Steamboat Springs, CO. Oklahoma State University. Available at: <http://coronasc.nmsu.edu/grazing-livestock-nutrit.html>. Accessed on: May 17, 2016.
» http://coronasc.nmsu.edu/grazing-livestock-nutrit.html - Taylor Jr, C. A. 2006. Targeted grazing to manage fire risk. p. 107-114. In: Targeted grazing: A natural approach to vegetation management and landscape enhancement. Launchbaugh, K. and Walker, J., eds. American Sheep Industry Association, Englewood, CO.
- Tedeschi, L. O. 2015. Integrating genomics with nutrition models to improve the prediction of cattle performance and carcass composition under feedlot conditions. PLoS One 10: e0143483.
- Tedeschi, L. O.; Almeida, A. K.; Atzori, A. S.; Muir, J. P.; Fonseca, M. A. and Cannas, A. 2017. A glimpse of the future in animal nutrition science. 1. Past and future challenges. Revista Brasileira de Zootecnia 46: 438-451.
- Tedeschi, L. O.; Callaway, T. R.; Muir, J. P. and Anderson, R. 2011. Potential environmental benefits of feed additives and other strategies for ruminant production. Revista Brasileira de Zootecnia 40: 291-309.
- Tedeschi, L. O.; Cavalcanti, L. F. L.; Fonseca, M. A.; Herrero, M. and Thornton, P. K. 2014. The evolution and evaluation of dairy cattle models for predicting milk production: an agricultural model intercomparison and improvement project (AgMIP) for livestock. Animal Production Science 54: 2052-2067.
- Tedeschi, L. O.; Fox, D. G. and Guiroy, P. J. 2004. A decision support system to improve individual cattle management. 1. A mechanistic, dynamic model for animal growth. Agricultural Systems 79: 171-204.
- Tedeschi, L. O.; Muir, J. P.; Riley, D. G. and Fox, D. G. 2015. The role of ruminant animals in sustainable livestock intensification programs. International Journal of Sustainable Development & World Ecology 22: 452-465.
- Theurer, M. E.; Amrine, D. E. and White, B. J. 2013a. Remote noninvasive assessment of pain and health status in cattle. Veterinary Clinics of North America: Food Animal Practice 29: 59-74.
- Theurer, M. E.; Anderson, D. E.; White, B. J.; Miesner, M. D.; Mosier, D. A.; Coetzee, J. F.; Lakritz, J. and Amrine, D. E. 2013b. Effect of Mannheimia haemolytica pneumonia on behavior and physiologic responses of calves during high ambient environmental temperatures. Journal of Animal Science 91: 3917-3929.
- Timsit, E.; Assié, S.; Quiniou, R.; Seegers, H. and Bareille, N. 2011. Early detection of bovine respiratory disease in young bulls using reticulo-rumen temperature boluses. The Veterinary Journal 190: 136-142.
- United Nations, Department of Economic and Social Affairs, Population Division. 2013. World Population Prospects; The 2012 Revision. Economics & Social Affairs. United Nations, New York, NY. Available at: <http://esa.un.org/unpd/wpp/index.htm>. Accessed on: Feb. 1, 2015.
» http://esa.un.org/unpd/wpp/index.htm - Van Kernebeek, H. R. J.; Oosting, S. J.; De Boer, I. J. M.; Van Ittersum, M. K. and Bikker, P. 2016. Saving land to feed a growing population: consequences for consumption of crop and livestock products. International Journal of Life Cycle Assessment 21: 677-687.
- Vasconcelos, J. T. and Galyean, M. L. 2007. Nutritional recommendations of feedlot consulting nutritionists: The 2007 Texas Tech University survey. Journal of Animal Science 85: 2772-2781.
- Velpuri, N. M.; Senay, G. G.; Alemu, H.; Rowland, J. and Verdin, J. P. 2014. Africa-wide monitoring of small surface water bodies using multisource satellite data: A monitoring system for FEWS NET. p. 69-95. In: Nile River Basin: Ecohydrological challenges, Climate Change and Hydropolitics. Melesse, A. M.; Abtew, W. and Setegn, S. G., eds. Springer, New York, NY.
- Waghorn, G. C. and Hegarty, R. S. 2011. Lowering ruminant methane emissions through improved feed conversion efficiency. Animal Feed Science and Technology 166-167: 291-301.
- Wang, Z.; Colazo, M. G.; Basarab, J. A.; Goonewardene, L. A.; Ambrose, D. J.; Marques, E.; Plastow, G.; Miller, S. P. and Moore, S. S. 2012. Impact of selection for residual feed intake on breeding soundness and reproductive performance of bulls on pasture-based multisire mating. Journal of Animal Science 90: 2963-2969.
- White, B. J. and Renter, D. G. 2009. Bayesian estimation of the performance of using clinical observations and harvest lung lesions for diagnosing bovine respiratory disease in post-weaned beef calves. Journal of Veterinary Diagnostic Investigation 21: 446-453.
- Wolfger, B.; Mang, A. V.; Cook, N.; Orsel, K. and Timsit, E. 2015a. Technical note: Evaluation of a system for monitoring individual feeding behavior and activity in beef cattle. Journal of Animal Science 93: 4110-4114.
- Wolfger, B.; Schwartzkopf-Genswein, K. S.; Barkema, H. W.; Pajor, E. A.; Levy, M. and Orsel, K. 2015b. Feeding behavior as an early predictor of bovine respiratory disease in North American feedlot systems. Journal of Animal Science 93: 377-385.
- Wolfger, B.; Timsit, E.; Pajor, E. A.; Cook, N.; Barkema, H. W. and Orsel, K. 2015c. Technical note: Accuracy of an ear tag-attached accelerometer to monitor rumination and feeding behavior in feedlot cattle. Journal of Animal Science 93: 3164-3168.
- Wolfger, B.; Timsit, E.; White, B. J. and Orsel, K. 2015d. A systematic review of bovine respiratory disease diagnosis focused on diagnostic confirmation, early detection, and prediction of unfavorable outcomes in feedlot cattle. Veterinary Clinics of North America: Food Animal Practice 31: 351-365.
- Zorzi, K.; Bonilha, S. F. M.; Queiroz, A. C.; Branco, R. H.; Sobrinho, T. L. and Duarte, M. S. 2013. Meat quality of young Nellore bulls with low and high residual feed intake. Meat Science 93: 593-599.
Publication Dates
-
Publication in this collection
May 2017
History
-
Received
20 Dec 2016 -
Accepted
25 Mar 2017