ABSTRACT
Newcastle disease is a highly contagious disease responsible for major outbreaks and considerable economic losses in the poultry industry in China. There is still little information available regarding gene characterization of the NDV, especially in ducks and pigeons. Therefore, the aim of this study was to investigate NDV isolated from ducks and pigeons in Hubei, China. In this study, three NDVs from ducks and pigeons were isolated between 2013 and 2015.The fusion protein (F) gene of the NDV isolates was sequenced and phylogenetically analyzed. The clinical signs and gross histopathological lesions were examined. Phylogenetic analysis of these strains indicated that all the sequences are classified as genotype II. The isolates shared a 112 G-R-Q-G-R-L 117motif at the F protein cleavage site, indicating that these three isolates strains are lentogenic. Necropsy and histopathology showed the typical pathological changes. It was concluded that commercial ducks and pigeons in Hubei province carry lentogenic NDV strains with regular genetic divergence, indicating that these species may act as the main reservoirs of NDV in poultry. Therefore, strategies and surveillance should be undertaken to reduce the risk of ND outbreaks.
Keywords:
Newcastle disease virus; phylogenetic analysis; pathology; ducks; pigeons.
INTRODUCTION
Newcastle disease is recognized as one of the highly contagious and lethal infectious disease of poultry industry worldwide. NDV is the member of genus Avulavirus, family Paramyxoviridae, with six transcriptional proteins, including haemagglutinin-neuraminidase protein, fusion protein, phosphoprotein, nucleocapsid protein, matrix protein and polymerase protein (Mayo 2002aMayo MA. A summary of taxonomic changes recently approved by ICTV. Archives of Virology 2002a;147:1655-1663.; Mayo 2002b; Chukwudi et al., 2012Chukwudi OE, Chukwuemeka ED, Mary U. Newcastle disease virus shedding among healthy commercial chickens and its epidemiological importance. Pakistan Veterinary Journal 2012;32(3):354-356.). It is mainly characterized by diarrhea, expiratory dyspnea, neurological symptoms, cloacal hemorrhages, focal glandular gastric bleeding or ulcers, intestinal mucosal bleeding, and necrosis of the pancreas or the spleen. This virus was recognized in chickens for the first time in 1926, and then in geese, ducks, and pigeons (Seal et al., 2005Seal BS, Wise MC, Pedersen JC, Senne DA, Alvarez R, et al. Genomic sequences of low-virulence avian paramyxovirus-1 (Newcastle disease virus) isolates obtained from live-bird markets in North America not related to commonly utilized commercial vaccine strains. Veterinary Microbiology 2005;106:7-16.; Erickson et al., 1980Erickson GA, Brugh M, Beard CW. Viscerotropic velogenic Newcastle disease in pigeons: clinical disease and immunization. Avian Diseases 1980;24:256-267.).
Fusion (F) protein is the key protein that makes NDV pathogenic. The phylogenetic analysis of the F genes was carried out in different studies in recent decades (Samal et al., 2012Samal S, Khattar SK, Kumar S, Collins PL, Samal SK. Coordinate deletion of N-glycans from the heptad repeats of the fusion F protein of Newcastle disease virus yields a hyperfusogenic virus with increased replication, virulence, and immunogenicity. Journal of Virology 2012;86:2501-2511.). According to F genes, NDVs can be categorized into classes I and II (Diel et al., 2012Diel DG, Silva LHAD, Liu H, Wang Z, Miller PJ, et al. Genetic diversity of avian paramyxovirus type 1: proposal for a unified nomenclature and classification system of Newcastle disease virus genotypes. Infection Genetics and Evolution 2012;12(8):1770-1779.). Class I has nine genotypes, while class II has at least 18 genotypes identified so far (Kim et al., 2007Kim LM, King DJ, Curry PE, Suarez DL, Swayne DE, et al. Phylogenetic diversity among low-virulence Newcastle disease viruses from waterfowl and shorebirds and comparison of genotype distributions to those of poultry-origin isolates. Journal of Virology 2007;81:12641-12653.; Snoeck et al., 2013Snoeck CJ, Owoade AA, Couacyhymann E, Alkali BR, Okwen MP, et al. High genetic diversity of newcastle disease virus in poultry in west and Central Africa: Cocirculation of genotype XIV and newly defined genotypes XVII and XVIII. Journal of Clinical Microbiology 2013;51:2250-2260.). Virulence intensity of NDVs can be classified according to basic amino acid sequences from 112 to 117 of the F protein as virulent (112R/K-R-Q-K/R-R 117) or with low virulence (112 G/E-K/R-Q-G/E-R-L 117), according to Kim et al.(2008).
In recent years, NDV has caused large-scale outbreaks in many countries, including China (Qin et al., 2008Qin ZM, Tan LT, Xu HY, Ma BC, Wang YL, et al. Pathotypical characterization and molecular epidemiology of Newcastle disease virus isolates from different hosts in China from 1996 to 2005. Journal of Clinical Microbiology 2008;46:601-611.; Zhang et al., 2010Zhang R, Pu J, Su JL, Zhao J, Wang X, et al. Phylogenetic characterization of Newcastle disease virus isolated in the mainland of China during 2001-2009. Veterinary Microbiology 2010;141:246-257.; Kang et al., 2016Kang Y, Xiang B, Yuan R, Zhao X, Feng M, et al. Phylogenetic and Pathotypic Characterization of Newcastle disease viruses circulating in South China and transmission in different birds. Frontiers in Microbiology 2016;7:119.), Kazakhstan (Bogoyavlenskiy et al., 2009Bogoyavlenskiy A, Berezin V, Prilipov A, Usachev E, Lyapina O, et al. Newcastle disease outbreaks in Kazakhstan and Kyrgyzstan during 1998, 2000, 2001, 2003, 2004, and 2005 were caused by viruses of the genotypes VIIb and VIId. Virus Genes 2009;39:94-101.), Indonesia (Xiao et al., 2012Xiao S, Paldurai A, Nayak B, Samuel A, Bharoto EE, et al. Complete genome sequences of Newcastle disease virus strains circulating in chicken populations of Indonesia. Journal of Virology 2012;86:5969-5970.), and in North America (Kang et al., 2014). China is the world’s largest waterfowl producing and consuming country, and accounts for than 70% and 90% of global duck and goose production, respectively; therefore, it is also known as “king of waterfowl world”. Moreover, duck- or pigeon-origin NDV isolates can infect chickens and maybe transmitted to naïve chickens (Kang et al., 2014). Newcastle disease virus infections have been reported all over the world, but the infection types and the virulence strength vary among regions. However, there are no information on NDV pathogenesis and transmission among ducks and pigeons reared in Hubei. The present study reports for the first time the characterization of NDV isolated from ducks and pigeons in Hubei, China.
MATERIALS AND METHODS
Sample collection
The present study was carried out on commercial poultry farms of different regions of Wuhan, Hubei province, China (Figure 1) between 2013 and 2015. Two suspected ND outbreaks in pigeon farms and one outbreak in a duck farm were studied. Dead pigeons and ducks were packed in ice bags and transported to Huazhong Agricultural University.
Clinical examination of signs, gross lesions, and histopathology
After the clinical examination of pigeons and ducks by researcher, intestine, heart, liver, and spleen specimens were fixed in 10% formalin. After 72h, the samples were dehydrated, embedded in paraffin, cut in 4-µm thick sections, and stained with hematoxylin and eosin (Bio-Swamp, Wuhan, China).
NDV isolation and identification
The selected organs (brain, heart, liver, spleen, lungs) were collected from dead pigeons and ducks, while virus isolation was carried out according to the guidelines and standards provided by the World Organization for Animal Health (OIE, 2013). Briefly, organ suspensions were homogenized and placed in viral transportation media (VTM) overnight at 4°C. The organ suspensions were then inoculated into the allantoic cavity of 9- to 11-day-old specific pathogen free (SPF) chicken embryos, which were incubated at 37 °C (Alexander, 2010Alexander DJ. Newcastle disease and other avian paramyxoviruses. Revue Scientifique Et Technique 2010;19:443-462.) for three days. After 72 h, the allantoic fluid was harvested, and NDV presence was tested using hemagglutination inhibition (HI) and hemagglutination (HA) tests. The Lasota strain was used as a positive control in this study. The positive allantoic fluids were stored at -70 °C until subsequent use and further analysis.
Pathogenicity analysis
The infective titer was expressed as EID50 (50% egg infective dose). Virus virulence was confirmed from same embryos inoculated for virus isolation by mean death time (MDT) and intracerebral pathogenicity index (ICPI) using standard procedures (Alexander, 1989Alexander DJ. Newcastle disease. In: Purchase HG, Arp LH, Domermuth CH, Pearson JE, editors. A laboratory manual for the isolation and identification of avian pathogens. 3rd ed. Kenneth Square: American Association of Avian Pathologists; 1989. p.114-120.; Kim et al., 2012Kim SH, Xiao S, Shive H, Collins PL, Samal SK. Replication, neurotropism, and pathogenicity of avian paramyxovirus serotypes 1-9 in chickens and ducks. PLoS One 2012;7:e34927.).
Viral RNA purification and RT-PCR of F gene
Total RNA of the three isolated strains (named: HP20131125, JX20131014 and XZ20151225) were extracted using the TIANamp Virus DNA/RNA kit (TianGen, China). RT-PCR amplification approach was used to amplify a fragment (1662 bp) of the F gene. For this purpose, the forward primer ATGGGCTCCAGACCTTCTACCA and reverse primer CATTTTTGTAGTGGCTCTCATCTGAT were used, and reverse transcription was done using a commercial RT-PCR Kit (TianGen, China). The RT-PCR was performed as previously described (Khan et al., 2016Khan AZ, Kumbhar S, Hamid M, Afzal S, Parveen F, et al. Effects of selenium-enriched robiotics on heart lesions by influencing the mRNA expressions of selenoproteins and heat shock proteins in heat stressed broiler chickens. Pakistan Veterinary Journal 2016;36(4):460-464.; Zhuang et al., 2016Zhuang Y, Liu P, Zhang CY, Ye S, Hu G, Cao HB. Effect of cadmium on the concentration of ceruloplasmin and its mRNA expression in goats under molybdenum stress. Pakistan Veterinary Journal 2016;36(2):209-213.; Zhu et al., 2017). Briefly, the reaction consisted of 35 cycles, denaturation at 95 °C for 40 s and 72 °C for 1.5 min, annealing at 55 °C for 50 s, and final extension at 72 °C for 10 min. PCR products were separated on agarose gel (1%). The products were purified using a TaKaRa DNA Extraction Kit Ver.4.0 (Takara Biotechnology Co., Ltd, Dalian, China) according to manufacturer’s instructions. The obtained positive products were sequenced by a commercial company (Sangon Biosciences, Wuhan, China).
Phylogenetic tree and Sequence analysis of the F protein cleavage site
The nucleotide sequences editing of the F gene (1662bp) of the three isolates in this study were compared with previously reported NDV sequences available at NCBI database. Phylogenetic analysis, amino acid sequence prediction, and nucleotide sequence similarity were performed using Molecular Evolutionary Genetic Analysis (MEGA 6.0) and the DNASTAR (version 3.3.8; DNASTAR) software. The phylogenetic tree and evolutionary distances were constructed using the neighbor-joining method and Kimura two-parameter method, respectively. The evolutionary distances among genotype or sub-genotypes were analyzed using the Maximum Likelihood method with 1000 bootstraps.
RESULTS
Virus isolation
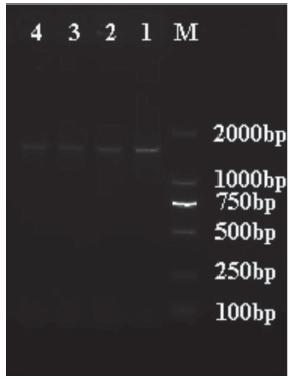
Three Newcastle disease viruses (NDVs) were isolated from ducks (HP20131125) and pigeons (JX20131014 and XZ20151225) on SPF embryonated chicken eggs in Hubei province from 2013 to 2015 (Table 1). Their identification and characterization was based on clinical symptoms, pathological observation and molecular investigations as shown in Figure 2.
The RT-PCR amplification results of the F gene of NDV. (M: Marker; 1: HP20131125; 2: JX20131014; 3: XZ20151225; 4,5: Positive control)
Pathogenicity tests of the NDV isolates
The pathogenicity tests revealed that all the three isolates had a MDT >120 h and 0.2 ICPI value (Table 1), and these three NDV isolates shared the motif of 112-GRQGRL-117 at the F protein cleavage site of the amino acid sequences (Table 2); therefore, they were classified as lentogenic NDV.
Clinical examination of signs, gross lesion and histopathology
The visual examination revealed depression and body condition of the chickens with dehydration, emaciation, enlarged abdomen, flush on the eyelids and eye secretions. The gross lesions revealed severe necrosis in the lung, multifocal hemorrhages in the mucosa of the proventriculus and small intestine, necrosis of tracheal mucosa, and neck subcutaneous hemorrhage, as shown in Figure 3.
Gross organ lesions in ducks and pigeons infected with NDV. (A,D) Severe necrosis in the lung; (B) Multifocal hemorrhages in the mucosa of the proventriculus; (C, F) Multifocal hemorrhages in the mucosa of the small intestine; (E) Hemorrhage and necrosis of the mucosa of the trachea and subcutaneous neck hemorrhage.
Tissue samples were subjected to histopathological examination by H&E staining. Liver showed hepatocyte necrosis, caryolysis, and fatty degeneration (Figure 4A, D). Lung revealed severe pulmonary hemorrhages, necrosis, alveolar cavity edema and serous effusion around the blood vessels (Figure 4B, E). Glandular gas-trorrhagia with large number of red blood cells was observed in ducks (Figure 4C), while the spleen revealed hemorrhage, congestion, and structure destruction in pigeons (Figure 4F).
Histopathological detection of NDV in duck and pigeon tissues. (A) Dropoutnecrosis, and caryolysis of hepatocytes in ducks; (B) Severe pulmonary hemorrhage, vascular necrosis in the lung; (C) glandular gastrorrhagia with large number of red blood cells; (D) Hepatocyte fatty degeneration, necrosis, and some caryolysis in pigeons; (E) Severe pulmonary hemorrhage, alveolar cavity edema, and serous effusion around blood vessel in the lung; (F) Spleen hemorrhage, congestion, and destroyed structure.
Phylogenetic analysis
In order to determine the evolutionary relationships between the isolated strains and representative NDVs, the phylogenetic analysis was performed based on the F gene (Figure 5). As determined from the phylogenetic tree, based on the nucleotide sequences 47nt to 420nt of the F gene, all the NDV isolates were clustered within the genotype II of the viruses collected from ducks and pigeons in Hubei from 2013 to 2015. All the NDV isolates shared 98.5% to 99.9% nucleotide identity with LaSota, Clone 30 and B1 (Accession number: AY845400, Y18898, and AF309418, respectively), among which, close homology of nucleotide sequences was determined between the isolated strains and some reference strains (LaSota, Clone 30 and B1).
Phylogenetic analysis of the F gene of the three isolates obtained in this study were compared with previously reported NDV sequences available at the NCBI database. The phylogenetic tree was constructed by the neighbor-joining method with 1,000 bootstrap replicates. Evolutionary analysis was conducted by MEGA 6.0 software.
DISCUSSION
Newcastle disease can cause high mortality and morbidity rates, which may pose a significant potential threat to the poultry industry in terms of serious economic losses in China (Kim et al., 2016Kim SH, Xiao S, Collins PL, Samal SK. LaSota fusion (F) cleavage motif-mediated fusion activity is affected by other regions of the F protein from different genotype Newcastle disease virus in a chimeric virus: implication for virulence attenuation. Journal of General Virology 2016;97:1-8.). The ND virus has been one of the most devastating pathogen of poultry industry for many years, and mainly causes Newcastle disease in chickens, avian paramyxovirus type 1 (APMV-1) infections in geese, etc. In recent years, APMV-1 infections in pigeons and ducks have been reported in a growing number of provinces in China, as well as ND infection. However, farmers and researchers are more concerned with the care and treatment of this disease than with its molecular characterization, transmission characteristics, and phylo-genetic analysis of NDV. Hubei province is located in the middle of China, and it is considered a large poultry-producing and consuming province, with large-scale poultry farms, a multitude of small backyard farms and live poultry markets. This study provides the important information to elucidate three NDVs isolated from commercial ducks and pigeons in Hubei to aid the prevention and control of this disease.
In China, most research has focused on NDV isolated from chickens because of its importance as a food-producing bird. However, there are few studies on NDV in ducks and pigeons. Ducks and pigeons show variable susceptibility to different NDV strains (Kim et al., 2012Kim SH, Xiao S, Shive H, Collins PL, Samal SK. Replication, neurotropism, and pathogenicity of avian paramyxovirus serotypes 1-9 in chickens and ducks. PLoS One 2012;7:e34927.; Carrasco et al., 2008Carrasco ADOT, Seki MC, Raso TDF, Paulillo AC, Pinto AA. Experimental infection of Newcastle disease virus in pigeons (Columba livia): Humoral antibody response, contact transmission and viral genome shedding. Veterinary Microbiology 2008;129(1/2):89-96.; Guo et al., 2014Guo H, Liu X, Xu Y, Han Z, Shao Y, et al. A comparative study of pigeons and chickens experimentally infected with PPMV-1 to determine antigenic relationships between PPMV-1 and NDV strains. Veterinary Microbiology 2014;168:88-97.). Molecular epidemiology research indicates that most NDVs isolated from ducks and pigeons in general belong to genotypes II, III, VI, and IX (Diel et al., 2012Diel DG, Silva LHAD, Liu H, Wang Z, Miller PJ, et al. Genetic diversity of avian paramyxovirus type 1: proposal for a unified nomenclature and classification system of Newcastle disease virus genotypes. Infection Genetics and Evolution 2012;12(8):1770-1779.; Gaikwad et al., 2016Gaikwad S, Kim JY, Lee HJ, Jung SC, Choi KS. Genetic characterization and evolutionary analysis of Newcastle disease virus isolated from domestic duck in South Korea. Gene 2016;579:34-40.).
The phylogenetic analysis results suggest that all the three NDV strains isolated in the present study belong to genotype II in Class II. Class II NDVs have been reported in many countries, including China, and in many species (Gaikwad et al., 2016Gaikwad S, Kim JY, Lee HJ, Jung SC, Choi KS. Genetic characterization and evolutionary analysis of Newcastle disease virus isolated from domestic duck in South Korea. Gene 2016;579:34-40.). We also found that three new isolated viruses bear close genetic relationship with the strains LaSota, Clones 30 and B1 (Accession numbers: AY845400, Y18898 and AF309418, respectively), which belong to genotype II of class II in terms of F genome analysis. The NDV strains LaSota, Clone-30 and B1 are still used to produce attenuated vaccines on a large scale in China (Kim et al., 2016Kim SH, Xiao S, Collins PL, Samal SK. LaSota fusion (F) cleavage motif-mediated fusion activity is affected by other regions of the F protein from different genotype Newcastle disease virus in a chimeric virus: implication for virulence attenuation. Journal of General Virology 2016;97:1-8.; Hu et al., 2009Hu S, Ma H, Wu Y, Liu W, Wang X, et al. A vaccine candidate of attenuated genotype VII Newcastle disease virus generated by reverse genetics. Vaccine 2009;27:904-910.). Our results revealed that the NDVs isolated in Hubei and LaSota, Clone 30 and B1 strains present close phylogenetic relationship and evolutionary distance. The vaccine-like viruses indicate that transmission from vaccinated to non-vaccinated poultry may be recurrent in Hubei. In addition, some reports also found the same phenomenon in a surveillance study with ducks in Guangdong province, China (Kang et al., 2016Kang Y, Xiang B, Yuan R, Zhao X, Feng M, et al. Phylogenetic and Pathotypic Characterization of Newcastle disease viruses circulating in South China and transmission in different birds. Frontiers in Microbiology 2016;7:119.). These findings suggest that the NDV viruses used in attenuated live vaccines in domestic ducks may exhibit reversion to virulence.
According to the OIE, lentogenic NDV strains have an ICPI of 0.2 and an MDT >90h, or carry a lentogenic amino acid sequence motif from 112 to 117 of F protein. The F gene cleavage site motif sequence was 112 G/E-K/R-Q-G/E-R-L 117 in all of the three isolates, which indicates that they are lentogenic, and the pathogenicity test (ICPI and MDT) also confirmed these findings. This indicates that the cleavage site motifs can be used to analyze the virulence of NDV strains, while pathogenicity tests (MDT, ICPI) can be used for further evidence of the cleavage site to precisely predict the pathogenicity and virulence of NDV strains.
In conclusion, our results showed that the duck and pigeon populations in Hubei province carry lentogenic NDVs. Furthermore, the genetic characterization provided clear evidence that non-standard use of vaccines may be the important reason for epidemic outbreaks of NDV in some parts of Hubei province, China. Such knowledge may be valuable for future studies to improve the control and diagnostic strategies of this disease.
ACKNOWLEDGMENT
Public sector (agriculture) scientific research (201303044)
REFERENCES
- Alexander DJ. Newcastle disease and other avian paramyxoviruses. Revue Scientifique Et Technique 2010;19:443-462.
- Alexander DJ. Newcastle disease. In: Purchase HG, Arp LH, Domermuth CH, Pearson JE, editors. A laboratory manual for the isolation and identification of avian pathogens. 3rd ed. Kenneth Square: American Association of Avian Pathologists; 1989. p.114-120.
- Bogoyavlenskiy A, Berezin V, Prilipov A, Usachev E, Lyapina O, et al. Newcastle disease outbreaks in Kazakhstan and Kyrgyzstan during 1998, 2000, 2001, 2003, 2004, and 2005 were caused by viruses of the genotypes VIIb and VIId. Virus Genes 2009;39:94-101.
- Carrasco ADOT, Seki MC, Raso TDF, Paulillo AC, Pinto AA. Experimental infection of Newcastle disease virus in pigeons (Columba livia): Humoral antibody response, contact transmission and viral genome shedding. Veterinary Microbiology 2008;129(1/2):89-96.
- Chukwudi OE, Chukwuemeka ED, Mary U. Newcastle disease virus shedding among healthy commercial chickens and its epidemiological importance. Pakistan Veterinary Journal 2012;32(3):354-356.
- Diel DG, Silva LHAD, Liu H, Wang Z, Miller PJ, et al. Genetic diversity of avian paramyxovirus type 1: proposal for a unified nomenclature and classification system of Newcastle disease virus genotypes. Infection Genetics and Evolution 2012;12(8):1770-1779.
- Erickson GA, Brugh M, Beard CW. Viscerotropic velogenic Newcastle disease in pigeons: clinical disease and immunization. Avian Diseases 1980;24:256-267.
- Gaikwad S, Kim JY, Lee HJ, Jung SC, Choi KS. Genetic characterization and evolutionary analysis of Newcastle disease virus isolated from domestic duck in South Korea. Gene 2016;579:34-40.
- Guo H, Liu X, Xu Y, Han Z, Shao Y, et al. A comparative study of pigeons and chickens experimentally infected with PPMV-1 to determine antigenic relationships between PPMV-1 and NDV strains. Veterinary Microbiology 2014;168:88-97.
- Hu S, Ma H, Wu Y, Liu W, Wang X, et al. A vaccine candidate of attenuated genotype VII Newcastle disease virus generated by reverse genetics. Vaccine 2009;27:904-910.
- Kang Y, Li Y, Yuan R, Li X, Sun M, et al. Phylogenetic relationships and pathogenicity variation of two Newcastle disease viruses isolated from domestic ducks in Southern China. Virology Journal 2014;11:147.
- Kang Y, Xiang B, Yuan R, Zhao X, Feng M, et al. Phylogenetic and Pathotypic Characterization of Newcastle disease viruses circulating in South China and transmission in different birds. Frontiers in Microbiology 2016;7:119.
- Khan AZ, Kumbhar S, Hamid M, Afzal S, Parveen F, et al. Effects of selenium-enriched robiotics on heart lesions by influencing the mRNA expressions of selenoproteins and heat shock proteins in heat stressed broiler chickens. Pakistan Veterinary Journal 2016;36(4):460-464.
- Kim LM, King DJ, Curry PE, Suarez DL, Swayne DE, et al. Phylogenetic diversity among low-virulence Newcastle disease viruses from waterfowl and shorebirds and comparison of genotype distributions to those of poultry-origin isolates. Journal of Virology 2007;81:12641-12653.
- Kim LM, King DJ, Guzman H, Tesh RB, Travassos DA, et al. Biological and phylogenetic characterization of pigeon paramyxovirus serotype 1 circulating in wild North American pigeons and doves. Journal of Clinical Microbiology 2008;46:3303-3310.
- Kim SH, Xiao S, Collins PL, Samal SK. LaSota fusion (F) cleavage motif-mediated fusion activity is affected by other regions of the F protein from different genotype Newcastle disease virus in a chimeric virus: implication for virulence attenuation. Journal of General Virology 2016;97:1-8.
- Kim SH, Xiao S, Shive H, Collins PL, Samal SK. Replication, neurotropism, and pathogenicity of avian paramyxovirus serotypes 1-9 in chickens and ducks. PLoS One 2012;7:e34927.
- Mayo MA. A summary of taxonomic changes recently approved by ICTV. Archives of Virology 2002a;147:1655-1663.
- Mayo MA. Virus taxonomy-Houston 2002. Archives of Virology 2002b;147:1071-1076.
- Qin ZM, Tan LT, Xu HY, Ma BC, Wang YL, et al. Pathotypical characterization and molecular epidemiology of Newcastle disease virus isolates from different hosts in China from 1996 to 2005. Journal of Clinical Microbiology 2008;46:601-611.
- Samal S, Khattar SK, Kumar S, Collins PL, Samal SK. Coordinate deletion of N-glycans from the heptad repeats of the fusion F protein of Newcastle disease virus yields a hyperfusogenic virus with increased replication, virulence, and immunogenicity. Journal of Virology 2012;86:2501-2511.
- Seal BS, Wise MC, Pedersen JC, Senne DA, Alvarez R, et al. Genomic sequences of low-virulence avian paramyxovirus-1 (Newcastle disease virus) isolates obtained from live-bird markets in North America not related to commonly utilized commercial vaccine strains. Veterinary Microbiology 2005;106:7-16.
- Snoeck CJ, Owoade AA, Couacyhymann E, Alkali BR, Okwen MP, et al. High genetic diversity of newcastle disease virus in poultry in west and Central Africa: Cocirculation of genotype XIV and newly defined genotypes XVII and XVIII. Journal of Clinical Microbiology 2013;51:2250-2260.
- Xiao S, Paldurai A, Nayak B, Samuel A, Bharoto EE, et al. Complete genome sequences of Newcastle disease virus strains circulating in chicken populations of Indonesia. Journal of Virology 2012;86:5969-5970.
- Zhang R, Pu J, Su JL, Zhao J, Wang X, et al. Phylogenetic characterization of Newcastle disease virus isolated in the mainland of China during 2001-2009. Veterinary Microbiology 2010;141:246-257.
- Zhu H, Zhang H, Wang Y, Ciren D, Dong H, et al. Genetic characterization and evolutionary analysis of emerging Newcastle disease virus isolated from Tibetan chickens. Kafkas Universitesi Veteriner Fakultesi Dergersi Journal 2017;23(3):385-389.
- Zhuang Y, Liu P, Zhang CY, Ye S, Hu G, Cao HB. Effect of cadmium on the concentration of ceruloplasmin and its mRNA expression in goats under molybdenum stress. Pakistan Veterinary Journal 2016;36(2):209-213.
Publication Dates
-
Publication in this collection
Jan-Mar 2018
History
-
Received
25 Dec 2016 -
Accepted
02 Apr 2017