ABSTRACT
Lentogenic Newcastle disease virus (lNDV) such as Lasota strain and low pathogenicity avian influenza such as H9N2 virus are two of the most economically important viruses affecting poultry worldwide, and little attention in recent years has been paid to simultaneous infections in chickens with these two viruses for the reason that co-infection do occur but are not easily detected. In the present study, chickens were inoculated with lNDV (Lasota) and LPAIV (A/chicken/Tehran/ZMT-173/99(H9N2)) simultaneously or sequentially three days apart. Oropharyngeal and cloacal swabs were collected from chickens from 1 to 14 days after inoculation. RRT-PCR for AIV and NDV detection was performed. The rate of viral shedding was measured within 14 days. No clinical symptoms were observed during the experiment however the pattern of virus shed was different with co-infection, thus comparing the results obtained from viral shedding showed that AIV is a much stronger agent than NDV in the occurrence of viral interference. This is due to the fact that in simultaneous inoculation, AIV replication delayed and reduced NDV replication, while replication of Lasota in simultaneous or pre-inoculated inoculation could not significantly disrupt H9N2 virus replication. These findings indicate that the infection with one virus can interfere with the replication of another, modifying the pathogenesis of the viruses. So, infection of the host with both viral agents simultaneously causes higher shedding of LPAIV than lNDV in OP and CL areas. In conclusion, co-infection with LPAVI in chickens did not impact clinical signs but affected the replication dynamics of these viruses.
Keywords:
Newcastle disease; Influenza virus (H9N2); viral interference
INTRODUCTION
Avian Influenza (AI) and Newcastle Disease (ND) are two dangerous diseases and the biggest threat to poultry and other avian species recognized worldwide, Ge et al., 2012Ge S, Zheng D, Zhao Y, Liu H, Liu W, Sun Q, et al. Evaluating viral interference between Influenza virus and Newcastle disease virus using real-time reverse transcription-polymerase chain reaction in chicken eggs. Virology Journal 2012;9:128., Costa-Hurtado et al., 2014Costa-Hurtado M, Afonso CL, Miller PJ, Spackman E, Kapczynski DR, Swayne DE, et al. Virus interference between H7N2 low pathogenic avian influenza virus and lentogenic Newcastle disease virus in experimental co-infections in chickens and turkeys. Veterinary Research 2014;45:1.. Viruses of AI and ND cause heavy economic losses to the poultry industry, Monne et al., 2008Monne I, Ormelli S, Salviato A, De Battisti C, Bettini F, Salomoni A, et al. Development and validation of a one-step real-time PCR assay for simultaneous detection of subtype H5, H7, and H9 avian influenza viruses. Journal of Clinical Microbiology 2008;46(5):1769-1773.; Pantin-Jackwood et al., 2015Pantin-Jackwood MJ, Costa-Hurtado M, Miller PJ, Afonso CL, Spackman E, et al. Experimental co-infections of domestic ducks with a virulent Newcastle disease virus and low or highly pathogenic avian influenza viruses. Veterinary Microbiology 2015;177(1-2):7-17.. The genome of both avian influenza virus (AIV) and Newcastle disease virus (NDV) is of single-stranded RNA with a negative antisense, Alexander & Senen., 2008Alexander DJ, Senen DA. Newcastle disease. Newcastle disease, other avian paramyxoviruses and pneumovirus infections. In: Saif YM, Fadly AM, Glisson JR, McDougald LR, Nolan LK, Swayne DE, editors. Diseases of poultry. 12th ed. Ames: Blackwell Publishing; 2008. p.75-100.; Costa-Hurtado et al., 2014. Low pathogenic avian influenza virus (LPAIV) and lentogenic Newcastle disease virus (lNDV) are transmitted from wild birds to domestic birds, and both cause a range of conditions from subclinical infections to diseases with high clinical signs, Wise et al., 2004Wise MG, Suarez DL, Seal BS, Pedersen JC, Senne DA, King DJ, et al. Development of a real-time reverse-transcription PCR for detection of Newcastle disease virus RNA in clinical samples. Journal of Clinical Microbiology 2004;42(1):329-338.; Jinnan et al., 2012; Pantin-Jackwood et al., 2015.
AIV belongs to type A of Orthomyxoviridae family. Based on their differences in surface glycoproteins of hemagglutinin (HA) and neuraminidase (NA) in the cleavage site, these viruses are divided into two groups of low pathogenic (LP) and high pathogenic (HP), Suarez et al., 2007Suarez DL, Das A, Ellis E. Review of rapid molecular diagnostic tools for avian influenza virus. Avian Diseases 2007; (1 Suppl):201-208.; Alexander & Senen, 2008Alexander DJ, Senen DA. Newcastle disease. Newcastle disease, other avian paramyxoviruses and pneumovirus infections. In: Saif YM, Fadly AM, Glisson JR, McDougald LR, Nolan LK, Swayne DE, editors. Diseases of poultry. 12th ed. Ames: Blackwell Publishing; 2008. p.75-100.; Capua & Alexander., 2009Capua I, Alexander DJ. Avian influenza and Newcastle disease: a field and laboratory manual. Rome: Springer Verlag; 2009.; Ben Shabat et al., 2010Ben Shabat M, Meir R, Haddas R, Lapin E, Shkoda I, Raibstein I, et al. Development of a real-time TaqMan RT-PCR assay for the detection of H9N2 avian influenza viruses. Journal of Virological Methods 2010;168(1-2):72-77.; Ge et al., 2012Ge S, Zheng D, Zhao Y, Liu H, Liu W, Sun Q, et al. Evaluating viral interference between Influenza virus and Newcastle disease virus using real-time reverse transcription-polymerase chain reaction in chicken eggs. Virology Journal 2012;9:128.; Pantin-Jackwood et al., 2015Pantin-Jackwood MJ, Costa-Hurtado M, Miller PJ, Afonso CL, Spackman E, et al. Experimental co-infections of domestic ducks with a virulent Newcastle disease virus and low or highly pathogenic avian influenza viruses. Veterinary Microbiology 2015;177(1-2):7-17.. So far, 17 subtypes of HA (H1-H17) and 9 subtypes of NA (N1-N9) have been identified, Sorrell & Perez, 2007Sorrell EM, Perez DR. Adaptation of influenza A/Mallard/Potsdam/178-4/83 H2N2 virus in Japanese quail leads to infection and transmission in chickens. Avian Diseases 2007;264-268.; Alexander, 2007; Lee & Saif, 2009Lee CW, Saif YM. Avian influenza virus. Comparative Immunology, Microbiology & Infectious Diseases 2009;32(4):301-310.; Xishan et al., 2012.
Birds are usually infected with both low pathogenicity AI (LPAIV) and high pathogenicity AI (HPAI); these groups have the potential to cause disease in susceptible birds, De Jong & Hien, 2006De Jong MD, Hien TT. Avian influenza A (H5N1). Journal of Clinical Virology 2006;35:2-13.; Peiris et al., 2007Peiris JS, de Jong MD, Guan Y. Avian influenza virus (H5N1): a threat to human health. Clinical Microbiology Reviews 2007;20(2):243-267.; Monne et al., 2008Monne I, Ormelli S, Salviato A, De Battisti C, Bettini F, Salomoni A, et al. Development and validation of a one-step real-time PCR assay for simultaneous detection of subtype H5, H7, and H9 avian influenza viruses. Journal of Clinical Microbiology 2008;46(5):1769-1773.; EL Bayoumi et al., 2013. Viruses belonging to H9 subtypes are of LPAIV.
Infection with LPAIV has been a major threat to commercial poultry, especially in the Middle East, since 1998, and caused many cases of outbreaks in Iran, Pakistan, Saudi Arabia, and the United Arabic Emirates, Naeem et al., 1999Naeem K, Ullah A, Manvell RJ, Alexander DJ. Avian influenza A subtype H9N2 in poultry in Pakistan. Veterinary Record 1999;145(19):560.; Nilli & Asasi, 2002; Alexander, 2003Alexander DJ. Report on avian influenza in the eastern hemisphere during 1997-2002. Avian Disease 2003;47(s3):792- 797.; Bano et al., 2003Bano S, Naeem K, Malik SA. Evaluation of pathogenic potential of avian influenza virus serotype H9N2 in chickens. Avian Disease 2003;47(s3): 817-822.; Capua & Alexander, 2004Capua I, Alexander DJ. Avian influenza: recent developments. Avian Pathology 2004;33(4):393-404.; Alexander, 2007; Monne et al., 2008Monne I, Ormelli S, Salviato A, De Battisti C, Bettini F, Salomoni A, et al. Development and validation of a one-step real-time PCR assay for simultaneous detection of subtype H5, H7, and H9 avian influenza viruses. Journal of Clinical Microbiology 2008;46(5):1769-1773.; Halvorson, 2008Halvorson DA. Control of low pathogenicity avian influenza. In: Swayne DE, editor. Avian influenza. Ames: Blackwell Publishing; 2008. p.513-536.; Pazani et al., 2008Pazani J, Marandi MV, Ashrafihelan J, Marjanmehr SH, Ghods F. Pathological studies of A/Chicken/Tehran/ZMT-173/99 (H9N2) influenza virus in commercial broiler chickens of Iran. International Journal of Poultry Science 2008;7:502-510.. It is noteworthy that the H9N2 strain of LPAIV is circulating in Iran; it caused a severe epidemic among birds in Tehran province in 1998 and imposed heavy economic losses to the poultry industry, Pazani et al., 2008.
NDVs, known as Paramyxovirus 1 (AMPV-1), belongs to the genus Avulavirus of the Paramyxoviridae family, Wise et al., 2004Wise MG, Suarez DL, Seal BS, Pedersen JC, Senne DA, King DJ, et al. Development of a real-time reverse-transcription PCR for detection of Newcastle disease virus RNA in clinical samples. Journal of Clinical Microbiology 2004;42(1):329-338.; Jang et al., 2011Jang J, Hong SH, Kim IH. Validation of a real-time RT-PCR method to quantify Newcastle disease virus (NDV) titer and comparison with other quantifiable methods. Journal of Microbiology and Biotechnology 2011;21(1):100-108.; Ebrahimi et al., 2012. Based on their ability to increase the severity of the disease, NDV strains are divided into three groups of lentogenic (low virulence), mesogenic (moderate), and velogenic (high virulence), Beard & Hanson, 1984Beard CW, Hanson RP. Newcastle disease. In: Hofstad MS, Barnes HJ, Calnek BW, Reid WM, Yoder HW, editors. Diseases of poultry. 8th ed. Ames: Iowa State University; 1984. p.452-470.; Pham et al., 2005Pham HM , Konnai S, Usui T, Chang KS, Murata S, Mase M, et al. Rapid detection and differentiation of Newcastle disease virus by real-time PCR with melting-curve analysis. Archives of Virology 2005;150(12):2429-2438.; De leeuw et al., 2005De Leeuw OS, Koch G, Hartog L, Ravenshorst N, Peeters BPH. Virulence of Newcastle disease virus is determined by the cleavage site of the fusion protein and by both the stem region and globular head of the haemagglutinin-neuraminidase protein. Journal of General Virology 2005;86(Pt 6):1759-1769.; Jang et al., 2011; Maclachlan & Dubovi, 2011Maclachlan NJ, Dubovi EJ. Paramyxoviridae. In: Maclachlan NJ, Dubovi EJ, editors. Fenner's Veterinary virology. 4th ed. London: Academic Press; 2011. p.299-235.; Ebrahimi et al., 2012; Pantin-Jackwood et al., 2015Pantin-Jackwood MJ, Costa-Hurtado M, Miller PJ, Afonso CL, Spackman E, et al. Experimental co-infections of domestic ducks with a virulent Newcastle disease virus and low or highly pathogenic avian influenza viruses. Veterinary Microbiology 2015;177(1-2):7-17.. Lasota and Hitchner B1 are lentogenic strains of NDV that are jointly used as lNDV live vaccines against ND outbreaks in the developed and developing countries, Alexander & Senen, 2008Alexander DJ, Senen DA. Newcastle disease. Newcastle disease, other avian paramyxoviruses and pneumovirus infections. In: Saif YM, Fadly AM, Glisson JR, McDougald LR, Nolan LK, Swayne DE, editors. Diseases of poultry. 12th ed. Ames: Blackwell Publishing; 2008. p.75-100.; Ebrahimi et al., 2012; Pantin-Jackwood et al., 2015. Both influenza and Newcastle are considered one of the major problems of wild birds in many countries, as they are listed in Class A of poultry diseases by the Office International des Epizootie (OIE), Wise et al., 2004; Perk et al., 2006Perk S, Panshin A, Shihmanter E, Gissin I, Pokamunski S, Pirak M, et al. Ecology and molecular epidemiology of H9N2 avian influenza viruses isolated in Israel during 2000-2004 epizootic. Developmental Biology 2006;124:201-209.; Ge et al., 2012Ge S, Zheng D, Zhao Y, Liu H, Liu W, Sun Q, et al. Evaluating viral interference between Influenza virus and Newcastle disease virus using real-time reverse transcription-polymerase chain reaction in chicken eggs. Virology Journal 2012;9:128.; OIE, 2012; Pantin-Jackwood et al., 2015.
Co-infection of birds with both lNDV and LPAIV has been reported in domestic and waterfowl birds, Halvorson, 2008Halvorson DA. Control of low pathogenicity avian influenza. In: Swayne DE, editor. Avian influenza. Ames: Blackwell Publishing; 2008. p.513-536.; Jahangir et al., 2009Jahangir A, Ruenphet S, Ueda S, Ueno Y, Shoham D, Shindo J, et al. Avian influenza and Newcastle disease viruses from northern pintail in Japan: isolation, characterization and inter-annual comparisons during 2006-2008. Virus Research 2009;143(1):44-52.; Molia et al., 2011Molia S, Samaké K, Diarra A, Sidibé MS, Doumbia L, Camara S, et al. Avian influenza and Newcastle disease in three risk areas for H5N1 highly pathogenic avian influenza in Mali, 2007-2008. Avian Disease 2011;55(4):650-658.; Couacy-Hymann et al., 2012Couacy-Hymann E, Kouakou V A., Aplogan GL., Awoume F, Kouakou CK, Kakpo L, et al. Surveillance for influenza viruses in poultry and Swine, West Africa, 2006-2008. Emerging Infectious Diseases 2012;18(9):1446-1452.; França et al., 2014França M, Howerth EW, Carter D, Byas A, Poulson R, Afonso CL, et al. Co-infection of mallards with low-virulence Newcastle disease virus and low-pathogenic avian influenza virus. Avian Pathology 2014;43(1):96-104.; Pantin-Jackwood et al., 2015Pantin-Jackwood MJ, Costa-Hurtado M, Miller PJ, Afonso CL, Spackman E, et al. Experimental co-infections of domestic ducks with a virulent Newcastle disease virus and low or highly pathogenic avian influenza viruses. Veterinary Microbiology 2015;177(1-2):7-17.. Hence, the infection of a host with a pathogen, including viruses or bacteria, and subsequent exposure to another pathogen lead to a co-infection. In relation to viruses, this phenomenon is called viral interference, Dianzani, 1975Dianzani F. Viral interference and interferon. La Ricerca in Clinica e in Laboratorio 1975;5(3):196-213.; Dapalma et al., 2010; Costa-Hurtado et al., 2014Costa-Hurtado M, Afonso CL, Miller PJ, Spackman E, Kapczynski DR, Swayne DE, et al. Virus interference between H7N2 low pathogenic avian influenza virus and lentogenic Newcastle disease virus in experimental co-infections in chickens and turkeys. Veterinary Research 2014;45:1.; Pantin-Jackwood et al., 2015. From 1949 till now, only six studies have been done on the interference between AIV and NDV. These studies were done by Bang F.B. 1949, Carr J.H. 1960, Florman AL 1984, Kennedy F 1983, Wenbo Liu 2003 and Ge 2012Ge S, Zheng D, Zhao Y, Liu H, Liu W, Sun Q, et al. Evaluating viral interference between Influenza virus and Newcastle disease virus using real-time reverse transcription-polymerase chain reaction in chicken eggs. Virology Journal 2012;9:128.. [Ge et al., 2012]. The results of these studies indicated that LPAIV subtype caused severe interference in the replication of NDV, Ge et al., 2012.
Therefore, co-infection of birds with LPAIV and lNDV can create a complex picture of clinical signs which makes it more difficult and time-consuming to detect and identify both of these viruses, El Zowalaty et al., 2011; Pan et al., 2012Pan Q, Liu A, Zhang F, Ling Y, Ou C, Hou N, et al. Co-infection of broilers with Ornithobacteriumrhinotracheale and H9N2 avian influenza virus. BMC Veterinary Research 2012;8:104.; Stipkovits et al., 2012Stipkovits L, Egyed L, Palfi V, Beres A, Pitlik E, Somogyi M, et al. Effect of low-pathogenicity influenza virus H3N8 infection on Mycoplasma gallisepticum infection of chickens. Avian Pathology 2012;41(1):51-57.. Measurable differences may include changes in tissue permissiveness or tropism, viral replication, patterns of virus progeny production and release, latency, pathology including immunological responses and immunopathology, Dapalma et al., 2010; Costa-Hurtado et al., 2014Costa-Hurtado M, Afonso CL, Miller PJ, Spackman E, Kapczynski DR, Swayne DE, et al. Virus interference between H7N2 low pathogenic avian influenza virus and lentogenic Newcastle disease virus in experimental co-infections in chickens and turkeys. Veterinary Research 2014;45:1.. Nevertheless, there is little information on the interaction of these two viruses when they simultaneously infect chicken species, Ge et al., 2012Ge S, Zheng D, Zhao Y, Liu H, Liu W, Sun Q, et al. Evaluating viral interference between Influenza virus and Newcastle disease virus using real-time reverse transcription-polymerase chain reaction in chicken eggs. Virology Journal 2012;9:128.; Pantin-Jackwood et al., 2015Pantin-Jackwood MJ, Costa-Hurtado M, Miller PJ, Afonso CL, Spackman E, et al. Experimental co-infections of domestic ducks with a virulent Newcastle disease virus and low or highly pathogenic avian influenza viruses. Veterinary Microbiology 2015;177(1-2):7-17.. However, the aim of the present study was to obtain information on the effects of viral interference between Lasota (lNDV) strain and LPAIV (A/chicken/Tehran/ZMT-173/99 (H9N2)) on specific pathogen free (SPF) chickens.
MATERIALS AND METHODS
Virus strains
In the present study, A/chicken/Tehran/ZMT-173/99 (H9N2), which is an isolate obtained from infected birds in Iran, provided by Razi Vaccine and Serum Research Institute- Southern Iran Branch (Shiraz), and Lasota (avirulent NDV strains of chicken) were used. These viruses were propagated in SPF embryonated chicken eggs (ECE) by inoculating 100µl of each virus. According to the predesigned pattern of the experiment, these viruses were inoculated individually or in combination on specified days (Table 1). SPF eggs were purchased from SPF Chicken Centre of Venky, India.
EID 50
Before each experiment, 50% egg infectious dose (EID50) of each virus stock was determined. To calculate the mean embryo infectious dose (EID50), AIV and ND viruses were titrated by preparing 10-fold dilutions over the range 10-1 to10-10 in PBS and inoculating each of five 9-day-old embryonated SPF chickens eggs with 100µl dilution each and incubated at a temperature of 37°C with humidity of 60-65%. All ECE after 5 days were tested for the presence of haemagglutinin activity. Titres of virus in 0.1 ml inoculated material, expressed as median egg infectious doses (EID50), were calculated by Reed and Munch method.
Birds
Although the Maternal Derived Antibodies (MDA) are known as passive immunity in young chickens and primary barriers of antigen-specific protection against pathogens, chickens may still be at risk of infection. According to previous studies, 28-day-old chickens which have the lowest MDA levels are stated to be at higher risk of viral infection, Hamal et al., 2006Hamal KR, Burgess SC, Pevzner IY, Erf GF. Maternal antibody transfer from dams to their egg yolks, egg whites, and chicks in meat lines of chickens. Poultry Science 2006;85(8):1364-1372.; Sasipreeyajan et al., 2012Sasipreeyajan J, Pohuang T, Sirikobkul N. Efficacy of different vaccination programs against Newcastle disease virus challenge in broiler chickens. The Thai Journal of Veterinary Medicine 2012;42:431-437.. Ninety 28-day-old birds SPF white leghorn chickens were used in this study. They were assigned to 6 groups (A to F) of 15, and each group was separately kept in isolator under standard conditions. Feed and water were provided ad libitum.
Experimental design
Chickens were assigned to 6 groups including a control (group A) and 5 groups (B to F) for virus inoculation. Each group consisted of 15 SPF chickens with an age of 4 weeks. Chickens in the control group received 50 µl of normal saline and those in the other groups were inoculated with 50 µl with 106EID50/100µl (single bird inoculation dose) of viruses by the intraocular and intranasal choral cleft routes. Viruses were inoculated separately, simultaneously or alternately (secondary inoculation 3 days after the initial inoculation). Chickens of all groups were daily examined for the observation and control of disease symptoms. In addition, their weight was measured on the first day and third day after the inoculation (dpi). On the fourteenth day, blood sampling was done for serological tests. Swabs of oropharyngeal (OP) and cloacal (CL) were collected from all chickens from the first to the tenth day after inoculation for the study of viral shedding (Table 1).
RNA extraction
OP and CL swabs were collected from all chickens from the first to the tenth day after inoculation for the study of viral shedding. These samples were kept in the liquid medium of PBS-BSA containing antibiotics of different concentrations, such as Penicillin G (2000 units/ml) and Gentamicin (200 μg/ml), in a freezer at -70˚C until the end of the experiment. For extraction of virus RNA from collected samples, High Pure Viral RNA Kit (Roche Applied Science, Mannheim, Germany) was used according to the manufacturer’s recommended protocol. This stage of the experiment was conducted in the Institute Zooprofilattico Seprimentale delle Venezie (IZSVe), Italy, which is a reference laboratory for influenza and Newcastle.
Probes and primers design
Nucleotide sequences of Matrix (M) gene for influenza and L gene for Newcastle virus were considered for the identification of H9N2 and Lasota during a molecular process. Thus, specific primers and probes were developed for the detection of Gene M of H9N2 and Gene L of Lasota. Oligonucleotide sequences of specific primers and probes have been shown below. Primers and probes for the detection of NDV were developed using the European reference center/AHVCA (WEIBRIAGE UK), and those for the identification of AIV were extracted from studies of Spackman (2002Spackman E, Dennis AS, Myers TJ, Bulaga LL, Garber LP, Perdue ML, et al. Development of a real-time reverse transcriptase PCR assay for type A influenza virus and the avian H5 and H7 hemagglutinin subtypes. Journal of Clinical Microbiology 2002;40(9):3256-3260.) and Monne (2008Monne I, Ormelli S, Salviato A, De Battisti C, Bettini F, Salomoni A, et al. Development and validation of a one-step real-time PCR assay for simultaneous detection of subtype H5, H7, and H9 avian influenza viruses. Journal of Clinical Microbiology 2008;46(5):1769-1773.) (Table 2).
All probes were labelled at the 5´ end with the 6-carboxyfluorescein (FAM) reporter dye and at the 3´ end with the 6-carboxytetramethylrhodamine (TAMRA) quencher dye, Spackman et al., 2002Spackman E, Dennis AS, Myers TJ, Bulaga LL, Garber LP, Perdue ML, et al. Development of a real-time reverse transcriptase PCR assay for type A influenza virus and the avian H5 and H7 hemagglutinin subtypes. Journal of Clinical Microbiology 2002;40(9):3256-3260..
Real-Time RT-PCR
The reagents contained in a QuantiTec multiplex RT-PCR kit (Qiagen, Hilden, Germany) were used for Real-Time RT-PCRs (RRT-PCR). The primers targeting the M gene of AIV were applied to the PCR at the optimized concentration of 300nM each. The exceptions were the L gene of NDV primers, which were used at a concentration of 500 nM. Specific fluorescently labeled probes were used at a final concentration of 100 nM for AIV and 200 nM for NDV. The RRT-PCR took place in a final volume of 25µl using a Rotor Gene 6000 apparatus. Each PCR tube contained a single primer/probe set for AIV and NDV. The identical thermal profile was adapted in order to detect both the distinct viruses within the same run. The following protocols were used for M gene primers/probes set: 20 min at 50ºC and 15 min at 95ºC, followed by 40 cycles at 94ºC for 45 sec and 60ºC for 45 sec. Also protocol were used for L gene primers/probes set: 20 min at 50ºC and 15 min at 95ºC, followed by 40 cycles at 94ºC for 45 sec and 50ºC for 45 sec.
Statistical analyses
Data were analyzed using IBM Statistical Package for the Social Sciences (SPSS) v.21 software. One-way ANOVA was used to analyse HI titres and body weights. Two-way ANOVA with Bonferroni multiple comparison analysis was used to evaluate virus titres in CL and OP swabs. Statistical significance was set at p<0.05 unless otherwise stated.
RESULTS
Clinical signs
During the experiment, no clinical sign was observed in the groups under study (groups which were treated with Newcastle or influenza viruses individually or in combination). The results of weighting the chickens showed that there was no significant difference between the groups and all chickens grew naturally (Table 3).
Serology
Blood samples taken from all groups were examined in terms of antibody titres against AI and ND viruses. Hemagglutination inhibition (HI) assay was used to quantify antibody against NDV and AIV viruses. The results indicated that the antibody titre was negative in the control group, while it was positive in all groups to which the secondary virus was administered simultaneously or with an interval of 3 days. The results also demonstrated that in the group that the influenza virus was inoculated individually, the antibody titre of influenza and Newcastle was 4 and 5 Log2 after 14 days, respectively (Figure 1).
Mean HI titers (log2) in SPF chickens. Serum samples were taken at 14 days after inoculation with LPAIV and lNDV. Horizontal axes are the names of groups in both charts as (b: lNDV, c: LPAIV, d: LPAIV+lNDV, e: LPAIV 3 days before lNDV, f: lNDV 3 days before LPAIV). A: this chart shows the titers of HI associated with LPAIV and B: shows the titers of HI associated with lNDV. The highest rate was reported when both viruses were received commonly.
Infection and viral shedding
OP and CL viral shedding were measured using RRT-PCR technique. In all samples of OP and CL swabs, the gene M of matrix area and the gene L were examined for the detection of H9N2 and Lasota viruses, respectively. The results obtained from this technique showed that there was a difference between the groups in terms of viral shedding, while the main route of shedding for LPAIV was detected in both OP and CL areas (p<0.05). In addition, the main route of shedding for lNDV was mostly observed in OP area (p<0.05). In Group F, in which chickens were infected with lNDV and then LPAIV, the number of positive samples in OP and CL areas was higher for LPAIV. This suggested that the shedding rate of influenza in OP and CL areas will be more when the inoculation of NDV precedes the inoculation of LPAIV, compared to the case that influenza virus is inoculated alone. It is noteworthy that the main route of shedding was detected for both lNDV and LPAIV (Figure 2).
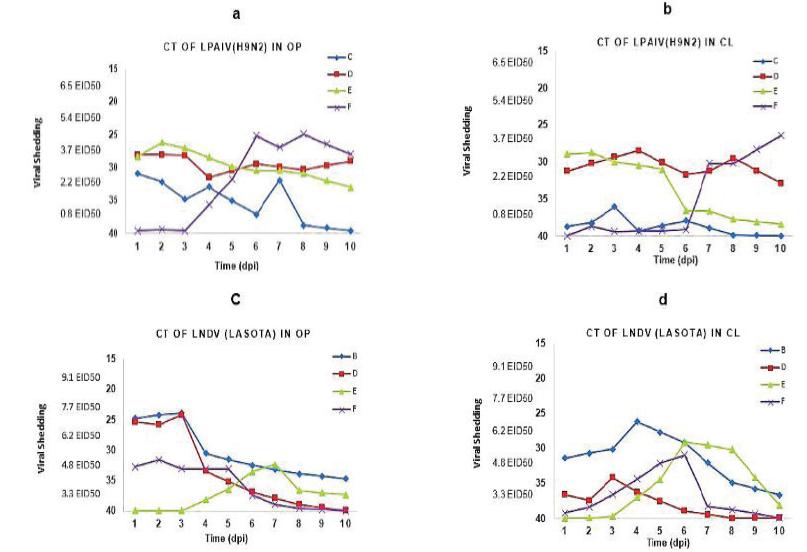
LPAIV and lNDV shedding in SPF chickens. Mean Ct value of real time RT-PCR and equivalent EID50/mL of AIV and NDV are detected in OP and CL swabs at different time points after inoculation. Detection of LPAIV in OP (a) and CL (b) swabs also detect lNDV in OP(c) and CL (d). Different time points show the viral shedding so that the blue square is related to single infected birds (groups : B and C), the red square is related to co-infected birds with both viruses ( group D), the green triangle is related to sequential infected birds ( group E) and the purple multiplication is related to group F.
Influenza viral shedding
By comparing the number of viral shedding particles using RRT-PCR technique at different times after the inoculation, the shedding pattern in OP area showed that the type of infection and exposure of the virus to the host tissue play a major role in virus replication. Based on the results obtained from OP area, the same rate of viral shedding was observed on the first and second day after the inoculation in chickens of groups C (LPAIV alone), D (lNDV and LPAIV simultaneously), and E (LPAIV three days before lNDV) (Figure 2a). However, viral shedding was reported to be distinctly low in the early days in chickens of group F (LPAIV three days after lNDV) and the highest rate of AIV shedding was observed on the fifth day after the secondary inoculation, i.e. on the eighth day after the inoculation (Figure 2a). In group C, where H9N2 was inoculated individually, viral shedding showed a reduction in the fourth to sixth days after the inoculation while such a reduction was not observed in other groups which had been treated with both viruses simultaneously (Figure 2a). The influenza virus shedding in the cloacal area was studied in all groups and the results indicated that the highest shedding rate was related to groups C, D, and E in the second to fourth days after the inoculation, while this peak in group F was found on the tenth day after the inoculation, i.e. 7 days after the secondary inoculation (Figure 2b).
Newcastle viral shedding
According to the results of this study, there was a difference between the groups regarding the shedding rate oflNDV (Figure 2c). Shedding of lNDV was alternatively observed in OP area. As the results indicated, shedding was observed on the second and third day after the inoculation in the group in which lNDV was administered alone (group B) and in the groups where the virus was inoculated simultaneously or with an interval of 3 days (groups D and F). The highest rate of lNDV shedding in OP area was found in groups B and D on the third day and in group F on the second day after the inoculation. On the other hand, the lowest shedding rate was observed in group E where lNDV was injected three days after the administration of LPAIV, as the highest virus titre was measured on the fourth day after the secondary inoculation (the seventh day) (Figure 2c). The lNDV shedding in CL area was recorded from group B on the fourth to the sixth day (three days after the secondary inoculation) while the highest shedding rate in groups E and F was observed only on the sixth day and it had a downward trend gradually until the tenth day in group D (Figure 2d). All control groups were reported to be negative in terms of the presence of the virus in the swab sample of OP and CL.
DISCUSSION
The main objective of the present research was to study the effects of viral interference between lNDV and LPAIV on SPF chickens. These viruses are circulating in poultry in many parts of the world, especially in Iran, and chickens are susceptible tanks of them. Co-infection with NDV and AIV has already been reported in vitro using chicken embryo and the viral interference between lNDV and LPAIV has been demonstrated to prevent the growth of the other virus, Roussan et al., 2008Roussan DA, Haddad R, Khawaldeh G. Molecular survey of avian respiratory pathogens in commercial broiler chicken flocks with respiratory diseases in Jordan. Poultry Science 2008;87(3):444-448.; Pawar et al., 2012Pawar SD, Kale SD, Rawankar AS, Koratkar SS, Raut CG, Pande SA, et al. Avian influenza surveillance reveals presence of low pathogenic avian influenza viruses in poultry during 2009-2011 in the West Bengal State, India. Virology Journal 2012;9:151.; Costa-Hurdato et al., 2014.
It was mentioned that in co-infection with these viruses, one virus prevents the growth of the other, Ge et al., 2012Ge S, Zheng D, Zhao Y, Liu H, Liu W, Sun Q, et al. Evaluating viral interference between Influenza virus and Newcastle disease virus using real-time reverse transcription-polymerase chain reaction in chicken eggs. Virology Journal 2012;9:128.; Pantin-Jackwood et al., 2015Pantin-Jackwood MJ, Costa-Hurtado M, Miller PJ, Afonso CL, Spackman E, et al. Experimental co-infections of domestic ducks with a virulent Newcastle disease virus and low or highly pathogenic avian influenza viruses. Veterinary Microbiology 2015;177(1-2):7-17.. Previous studies have shown that chickens inoculated with both lNDV NDV and LPAIV cause co-infection in other chickens, Ge et al., 2012; Costa-Hurdato et al., 2014; França et al., 2014França M, Howerth EW, Carter D, Byas A, Poulson R, Afonso CL, et al. Co-infection of mallards with low-virulence Newcastle disease virus and low-pathogenic avian influenza virus. Avian Pathology 2014;43(1):96-104.;Pantin-Jackwood et al., 2015. Co-infection and viral interference depend on factors such as bird species, type of virus, timing inoculation, and virus inoculation type, also tissue tropism, latency, dose, virulence and biological properties of pathogens and altered immune response are important, Pantin-Jackwood et al., 2015; Zarkov, 2017Zarkov IV. Avian influenza A virus and Newcastle disease virus mono and co-infections in birds. Trakia Journal of Sciences 2017;2:182-191..
Despite the severity, natural co-infections of lNDV and AIV are anticipated to occur in poultry, and the effects of such co-infections on several host responses such as viral shedding dynamic, seroconversion and clinical signs are not fully known in chickens, Costa-Hurdato et al., 2014. Even with the differences in the type of virus replication, co-infection of chickens with lNDV and LPAIV has not had significant effects on the increase or decrease of clinical symptoms, Pantin-Jackwood et al., 2015Pantin-Jackwood MJ, Costa-Hurtado M, Miller PJ, Afonso CL, Spackman E, et al. Experimental co-infections of domestic ducks with a virulent Newcastle disease virus and low or highly pathogenic avian influenza viruses. Veterinary Microbiology 2015;177(1-2):7-17.. The clinical signs in birds depend on the species, age and virus type, para OIE, 2015. The main clinical signs are nasal discharge, cough, nervous signs, diarrhea, inappetance in laying birds, para OIE, 2015. Therefore, the host species is a factor that can influence the severity of clinical signs and amount of virus replication in such virus co-infections, Costa-Hurtado et al., 2014Costa-Hurtado M, Afonso CL, Miller PJ, Spackman E, Kapczynski DR, Swayne DE, et al. Virus interference between H7N2 low pathogenic avian influenza virus and lentogenic Newcastle disease virus in experimental co-infections in chickens and turkeys. Veterinary Research 2014;45:1.. Costa-Hurtado et al infected chickens and turkeys with lNDV vaccine strain (Lasota) and H7N2 LPAIV simultaneously or sequentially, they reported that none of the chickens infected with LPAIV and lNDV showed clinical signs, while all turkeys exposed with both viruses presented mild clinical symptoms, Costa-Hurtado et al., 2014. According to the previous studies, no effect on clinical signs of co-infection with the two viruses was found in the present study. For the first time in Iran, a report of viral interference between lNDV and LPAIV in SPF chickens using RRT-PCR technique is provided in the present study.
All chickens were infected with lNDV (Lasota) and LPAIV (A/chicken/Tehran/ZMT-173/99 (H9N2)) simultaneously or sequentially. The results clearly showed that viral titre on the OP route was distinctly more than viral titre on the CL route, these results are consistent with a previous study by Costa-Hurtado, 2014Costa-Hurtado M, Afonso CL, Miller PJ, Spackman E, Kapczynski DR, Swayne DE, et al. Virus interference between H7N2 low pathogenic avian influenza virus and lentogenic Newcastle disease virus in experimental co-infections in chickens and turkeys. Veterinary Research 2014;45:1. that stated that LPAIV viral shedding is mainly in the OP route, Costa-Hurtado et al., 2014. In addition, shedding was lower in groups which were treated with viruses individually. Similarly to the previous studies, the findings of the present study prove that LPAIV is more pathogenic for chickens, as the presence of AIV in the host could delay or prohibit the replication of NDV.
Therefore, it is recommended that the severity and degree of interference depend on the amount and pathogenicity of the virus strain, Spackman et al., 2002Spackman E, Dennis AS, Myers TJ, Bulaga LL, Garber LP, Perdue ML, et al. Development of a real-time reverse transcriptase PCR assay for type A influenza virus and the avian H5 and H7 hemagglutinin subtypes. Journal of Clinical Microbiology 2002;40(9):3256-3260.. The present study also showed that lNDV cannot suppress the shedding of LPAIV. Hence, infection with NDV could not always prevent subsequent infections by AIV even if Lasota and H9N2 are simultaneously injected. This suggests that Lasota, if inoculated individually or simultaneously, cannot interfere with the replication and shedding of H9N2 but can only slow down the process of viral replication.
In this study, it was observed that the shedding pattern of both H9N2 and Lasota was strongly influenced by each other, as the viral shedding of LPAIV in OP and CL areas was distinctly delayed in chickens that were infected with lNDV and successively exposed to LPAIV. In other words, due to the inoculation of Lasota before H9N2, the replication of the influenza virus had a slower process and the viral shedding was lower. These values were evaluated in the sixth to the tenth day after the inoculation (Figure 2a, b). By comparing the results of this study with qualitative achievements of previous studies, it can be stated that AIV virus is a more powerful agent than NDV in the interference between these two viruses.
Viral interference is a very important phenomenon in which one cell is infected with a virus, as the virus can prevent the replication of secondary viruses. In other words, it can suppress the shedding of a virus of the homologous or heterologous type which enters the cell, Dianzani, 1975Dianzani F. Viral interference and interferon. La Ricerca in Clinica e in Laboratorio 1975;5(3):196-213..
Several important mechanisms have been proposed to explain the viral interference: Competing by attachment interference. In this mechanism, receptors for the superinfecting virus are reduced or blocked, competing intracellular for replication of the host machinery, virus-induced interferon interference, Kimura et al., 1976Kimura Y, Norrby E, Nagata I, Ito Y, Shimokata K. Homologous interference induced by a temperature-sensitive mutant derived from an HVJ (Sendai virus) carrier culture. Journal of General Virology 1976;33(2):333-343..
lNDV and LPAIV are replicated in the upper area of the respiratory system and epithelial cells of the intestinal tract, where there are trypsin-like enzymes. lNDV binds through the HN glycoprotein to sialic acid receptors on the cell surface, the same as HA glycoprotein does for LPAIV. Therefore, superficial cell receptors for AIV are glycoconjugates containing sialic acid, whereas these receptors for NDV are recommended to be ganglioside and N-glycoprotein, Rott, 1979Rott, R. Molecular basis of infectivity and pathogenicity of myxovirus. Archives of Virology 1979;59(4):285-298.; Ferreira et al., 2004Ferreira L, Villar E, Muñoz-Barroso I. Gangliosides and N-glycoproteins function as Newcastle disease virus receptors. The International Journal of Biochemistry & Cell Biology 2004;36(11):2344-2356.; Ge et al., 2012Ge S, Zheng D, Zhao Y, Liu H, Liu W, Sun Q, et al. Evaluating viral interference between Influenza virus and Newcastle disease virus using real-time reverse transcription-polymerase chain reaction in chicken eggs. Virology Journal 2012;9:128.; Swayne et al., 2013Swayne DE, Suarez DL, Sims LD. Influenza. In: Swayne DE, Glisson JR, McDougald LR, Nair V, Nolan LK, Suarez DL, editors. Diseases of poultry. 13th ed. Ames: Willey-Blackwell; 2013. p.181-218.. These findings suggest that both viruses may compete for the same target cell as Lasota can connect to the HN glycoprotein, LPAIV can connect to HA glycoprotein compounds because both of these compounds contain sialic acid, Costa-Hurtado et al., 2014Costa-Hurtado M, Afonso CL, Miller PJ, Spackman E, Kapczynski DR, Swayne DE, et al. Virus interference between H7N2 low pathogenic avian influenza virus and lentogenic Newcastle disease virus in experimental co-infections in chickens and turkeys. Veterinary Research 2014;45:1..
Previous studies have explained that an inactivated particle of influenza can interfere with the replication of a live virus that later enters the cell. Through the interferon system, these results led to the discovery of interference phenomenon, Ziegler & Horsfall, 1944Ziegler JE, Horsfall FL. Interference between the influenza viruses, I. the effect of active virus upon the multiplication of influenza viruses in the chick embryo. Journal of Experimental Medicine 1944;79(4):361-377.; Henle, 1950Henle W. Interference phenomena between animal viruses: a review. Journal of Immunology 1950;64(3):203-236.. Other studies have also stated that the production of interferon can prevent or suppress the replication of many viruses, particularly AIV and NDV, Issacs & Lindenmann, 1987. Replication of a virus following the activation of anti-viral immune responses, including immunomodulators or the use of immune cells, can influence the replication of other viruses in a similar area, Ge et al., 2012Ge S, Zheng D, Zhao Y, Liu H, Liu W, Sun Q, et al. Evaluating viral interference between Influenza virus and Newcastle disease virus using real-time reverse transcription-polymerase chain reaction in chicken eggs. Virology Journal 2012;9:128.. Stimulation of local interferon generation by lNDV may interfere with the replication of LPAIV because Lasota is known as a poor inducer of interferon, Costa-Hurdato et al., 2014.
As mentioned in the previous studies, Ge et al. (2012Ge S, Zheng D, Zhao Y, Liu H, Liu W, Sun Q, et al. Evaluating viral interference between Influenza virus and Newcastle disease virus using real-time reverse transcription-polymerase chain reaction in chicken eggs. Virology Journal 2012;9:128.) found that if the host is infected with lNDV prior to, H9N2 replication can be suppressed. However, in another study, Costa (2014) stated that high replication of LPAIV in turkeys could prevent the replication of lNDV, Costa-Hurtado et al., 2014Costa-Hurtado M, Afonso CL, Miller PJ, Spackman E, Kapczynski DR, Swayne DE, et al. Virus interference between H7N2 low pathogenic avian influenza virus and lentogenic Newcastle disease virus in experimental co-infections in chickens and turkeys. Veterinary Research 2014;45:1.. França (2014França M, Howerth EW, Carter D, Byas A, Poulson R, Afonso CL, et al. Co-infection of mallards with low-virulence Newcastle disease virus and low-pathogenic avian influenza virus. Avian Pathology 2014;43(1):96-104.) reported that co-infection of poultry and wild birds with NDV and AIV in vivo showed that differences in the pattern of viral shedding depend on the time of co-infection, and that the delay in the peak of viral shedding of LPAIV occurs when lNDV is already inoculated [França et al., 2014]. According to França’s study, although co-infection of chickens with both viruses was observed, no reduction was observed in humoral immune response. This is consistent with the findings of Gelb (2007) in relation to the co-infection of broilers with IBV and lNDV [Gelb et al., 2007; França et al., 2014].
In the present study, the reduction in antibody titre against LPAIV, compared with lNDV, was partially observed (Figure 1). Therefore, in confirmation of the previous studies, it is recommended that humoral response against co-infection may not be much effective in antibody titre courses. In summary, SPF chickens received 50 µl viruses with 106 EID50/ml from Lasota and H9N2 for the study of co-infection and interference between LPAIV and lNDV. The data showed that infection of the host with both viral agents simultaneously caused higher shedding of LPAIV than lNDV in OP and CL areas.
In general, effects of viral interference depend on the adaptation of viruses to a host species, pathogenicity of viruses, time of co-infection, and environmental factors. Assessment and identification of the factors affecting the viral interference or understanding the factors that cause delay in the replication and infection of viruses will help us to better find the path of pathogenicity and transmission of these viruses in chickens. That understanding could also provide new plans of virus control programs, including new tools for identification and more improved protocols of vaccination.
CONCLUSION
Birds are often infected with more than one infectious agents. The co-infection of birds with LPAIV and lNDV is due to the fact that they compete for the same receptors in susceptible cells. AIV have a negative impact on NDV growth if they are inoculated simultaneously or sequentially. According to the results obtained from this study, co-infection of LPAIV and lNDV had no effect on the severity of clinical signs.
ACKNOWLEDGMENT
This study has been supported by Razi Vaccine and Serum Research Institute Shiraz, Iran and Istituto Zooprofilattico Sperimentale delle Venezie (IZSVe) Italy. The authors appreciate the cooperation of Dr Giovanni Cattoli as well as the virology and molecular biology department in IZSVe.
This research was funded by a grant (no-2-18-18-94102) from the Razi Vaccine and Serum Research Institute Iran.
REFERENCES
- Alexander DJ, Senen DA. Newcastle disease. Newcastle disease, other avian paramyxoviruses and pneumovirus infections. In: Saif YM, Fadly AM, Glisson JR, McDougald LR, Nolan LK, Swayne DE, editors. Diseases of poultry. 12th ed. Ames: Blackwell Publishing; 2008. p.75-100.
- Alexander DJ. An overview of the epidemiology of avian influenza. Vaccine 2007;25(30):5637-5644.
- Alexander DJ. Report on avian influenza in the eastern hemisphere during 1997-2002. Avian Disease 2003;47(s3):792- 797.
- Bano S, Naeem K, Malik SA. Evaluation of pathogenic potential of avian influenza virus serotype H9N2 in chickens. Avian Disease 2003;47(s3): 817-822.
- Beard CW, Hanson RP. Newcastle disease. In: Hofstad MS, Barnes HJ, Calnek BW, Reid WM, Yoder HW, editors. Diseases of poultry. 8th ed. Ames: Iowa State University; 1984. p.452-470.
- Ben Shabat M, Meir R, Haddas R, Lapin E, Shkoda I, Raibstein I, et al. Development of a real-time TaqMan RT-PCR assay for the detection of H9N2 avian influenza viruses. Journal of Virological Methods 2010;168(1-2):72-77.
- Capua I, Alexander DJ. Avian influenza and Newcastle disease: a field and laboratory manual. Rome: Springer Verlag; 2009.
- Capua I, Alexander DJ. Avian influenza: recent developments. Avian Pathology 2004;33(4):393-404.
- Costa-Hurtado M, Afonso CL, Miller PJ, Spackman E, Kapczynski DR, Swayne DE, et al. Virus interference between H7N2 low pathogenic avian influenza virus and lentogenic Newcastle disease virus in experimental co-infections in chickens and turkeys. Veterinary Research 2014;45:1.
- Couacy-Hymann E, Kouakou V A., Aplogan GL., Awoume F, Kouakou CK, Kakpo L, et al. Surveillance for influenza viruses in poultry and Swine, West Africa, 2006-2008. Emerging Infectious Diseases 2012;18(9):1446-1452.
- Dapalma T1, Doonan BP, Trager NM, Kasman LM. A systematic approach to virus-virus interactions. Virus Research 2010;149(1):1-9.
- De Jong MD, Hien TT. Avian influenza A (H5N1). Journal of Clinical Virology 2006;35:2-13.
- De Leeuw OS, Koch G, Hartog L, Ravenshorst N, Peeters BPH. Virulence of Newcastle disease virus is determined by the cleavage site of the fusion protein and by both the stem region and globular head of the haemagglutinin-neuraminidase protein. Journal of General Virology 2005;86(Pt 6):1759-1769.
- Dianzani F. Viral interference and interferon. La Ricerca in Clinica e in Laboratorio 1975;5(3):196-213.
- Ebrahimi MM1, Shahsavandi S, Moazenijula G, Shamsara M. Phylogeny and evolution of Newcastle disease virus genotypes isolated in Asia during 2008-2011. Virus Genes 2012;45(1):63-68.
- ELbayoumi KM, Mahgoub K, Mekky HM, Kutkat MA. Molecular detection of H5N1, H9N2 and Newcastle disease viruses isolated from chicken in mixed infection in Egypt. World Applied Sciences Journal 2013;27(1):44-50.
- Ferreira L, Villar E, Muñoz-Barroso I. Gangliosides and N-glycoproteins function as Newcastle disease virus receptors. The International Journal of Biochemistry & Cell Biology 2004;36(11):2344-2356.
- França M, Howerth EW, Carter D, Byas A, Poulson R, Afonso CL, et al. Co-infection of mallards with low-virulence Newcastle disease virus and low-pathogenic avian influenza virus. Avian Pathology 2014;43(1):96-104.
- Ge S, Zheng D, Zhao Y, Liu H, Liu W, Sun Q, et al. Evaluating viral interference between Influenza virus and Newcastle disease virus using real-time reverse transcription-polymerase chain reaction in chicken eggs. Virology Journal 2012;9:128.
- Gelb J Jr, Ladman BS, Licata MJ, Shapiro MH, Campion LR. Evaluating viral interference between infectious bronchitis virus and Newcastle disease virus vaccine strains using quantitative reverse transcription-polymerase chain reaction. Avian Disease 2007;51(4):924-934.
- Halvorson DA. Control of low pathogenicity avian influenza. In: Swayne DE, editor. Avian influenza. Ames: Blackwell Publishing; 2008. p.513-536.
- Hamal KR, Burgess SC, Pevzner IY, Erf GF. Maternal antibody transfer from dams to their egg yolks, egg whites, and chicks in meat lines of chickens. Poultry Science 2006;85(8):1364-1372.
- Henle W. Interference phenomena between animal viruses: a review. Journal of Immunology 1950;64(3):203-236.
- Isaacs A, Lindenmann J. Virus interference. I. The interferon. Journal of Interferon Research 1987;7(5):429-438.
- Jahangir A, Ruenphet S, Ueda S, Ueno Y, Shoham D, Shindo J, et al. Avian influenza and Newcastle disease viruses from northern pintail in Japan: isolation, characterization and inter-annual comparisons during 2006-2008. Virus Research 2009;143(1):44-52.
- Jang J, Hong SH, Kim IH. Validation of a real-time RT-PCR method to quantify Newcastle disease virus (NDV) titer and comparison with other quantifiable methods. Journal of Microbiology and Biotechnology 2011;21(1):100-108.
- Jinnan Li, Haixia H, Qingzhong Y, Diego GD, De-shan L, Patti JM. Generation and characterization of a recombinant Newcastle disease virus expressing the red fluorescent protein for use in co-infection studies. Virology Journal 2012;9:227.
- Kimura Y, Norrby E, Nagata I, Ito Y, Shimokata K. Homologous interference induced by a temperature-sensitive mutant derived from an HVJ (Sendai virus) carrier culture. Journal of General Virology 1976;33(2):333-343.
- Lee CW, Saif YM. Avian influenza virus. Comparative Immunology, Microbiology & Infectious Diseases 2009;32(4):301-310.
- Maclachlan NJ, Dubovi EJ. Paramyxoviridae. In: Maclachlan NJ, Dubovi EJ, editors. Fenner's Veterinary virology. 4th ed. London: Academic Press; 2011. p.299-235.
- Molia S, Samaké K, Diarra A, Sidibé MS, Doumbia L, Camara S, et al. Avian influenza and Newcastle disease in three risk areas for H5N1 highly pathogenic avian influenza in Mali, 2007-2008. Avian Disease 2011;55(4):650-658.
- Monne I, Ormelli S, Salviato A, De Battisti C, Bettini F, Salomoni A, et al. Development and validation of a one-step real-time PCR assay for simultaneous detection of subtype H5, H7, and H9 avian influenza viruses. Journal of Clinical Microbiology 2008;46(5):1769-1773.
- Naeem K, Ullah A, Manvell RJ, Alexander DJ. Avian influenza A subtype H9N2 in poultry in Pakistan. Veterinary Record 1999;145(19):560.
- Nili H, Asasi K. Natural cases and an experimental study of H9N2 avian influenza in commercial broiler chickens of Iran. Avian Pathology 2002;31(3):247-252.
- OIE - World Organisation For Animal Health. Avian influenza: infection with avian influenza viruses [terrestrial manual]. Paris; 2015. chapter 2.3.4, p.1-23.
- OIE - World Organisation For Animal Health. Manual of diagnostic test and vaccines for terrestrial animals: mammals, birds and bees. Paris; 2012.
- Pan Q, Liu A, Zhang F, Ling Y, Ou C, Hou N, et al. Co-infection of broilers with Ornithobacteriumrhinotracheale and H9N2 avian influenza virus. BMC Veterinary Research 2012;8:104.
- Pantin-Jackwood MJ, Costa-Hurtado M, Miller PJ, Afonso CL, Spackman E, et al. Experimental co-infections of domestic ducks with a virulent Newcastle disease virus and low or highly pathogenic avian influenza viruses. Veterinary Microbiology 2015;177(1-2):7-17.
- Pawar SD, Kale SD, Rawankar AS, Koratkar SS, Raut CG, Pande SA, et al. Avian influenza surveillance reveals presence of low pathogenic avian influenza viruses in poultry during 2009-2011 in the West Bengal State, India. Virology Journal 2012;9:151.
- Pazani J, Marandi MV, Ashrafihelan J, Marjanmehr SH, Ghods F. Pathological studies of A/Chicken/Tehran/ZMT-173/99 (H9N2) influenza virus in commercial broiler chickens of Iran. International Journal of Poultry Science 2008;7:502-510.
- Peiris JS, de Jong MD, Guan Y. Avian influenza virus (H5N1): a threat to human health. Clinical Microbiology Reviews 2007;20(2):243-267.
- Perk S, Panshin A, Shihmanter E, Gissin I, Pokamunski S, Pirak M, et al. Ecology and molecular epidemiology of H9N2 avian influenza viruses isolated in Israel during 2000-2004 epizootic. Developmental Biology 2006;124:201-209.
- Pham HM , Konnai S, Usui T, Chang KS, Murata S, Mase M, et al. Rapid detection and differentiation of Newcastle disease virus by real-time PCR with melting-curve analysis. Archives of Virology 2005;150(12):2429-2438.
- Rott, R. Molecular basis of infectivity and pathogenicity of myxovirus. Archives of Virology 1979;59(4):285-298.
- Roussan DA, Haddad R, Khawaldeh G. Molecular survey of avian respiratory pathogens in commercial broiler chicken flocks with respiratory diseases in Jordan. Poultry Science 2008;87(3):444-448.
- Sasipreeyajan J, Pohuang T, Sirikobkul N. Efficacy of different vaccination programs against Newcastle disease virus challenge in broiler chickens. The Thai Journal of Veterinary Medicine 2012;42:431-437.
- Sorrell EM, Perez DR. Adaptation of influenza A/Mallard/Potsdam/178-4/83 H2N2 virus in Japanese quail leads to infection and transmission in chickens. Avian Diseases 2007;264-268.
- Spackman E, Dennis AS, Myers TJ, Bulaga LL, Garber LP, Perdue ML, et al. Development of a real-time reverse transcriptase PCR assay for type A influenza virus and the avian H5 and H7 hemagglutinin subtypes. Journal of Clinical Microbiology 2002;40(9):3256-3260.
- Stipkovits L, Egyed L, Palfi V, Beres A, Pitlik E, Somogyi M, et al. Effect of low-pathogenicity influenza virus H3N8 infection on Mycoplasma gallisepticum infection of chickens. Avian Pathology 2012;41(1):51-57.
- Suarez DL, Das A, Ellis E. Review of rapid molecular diagnostic tools for avian influenza virus. Avian Diseases 2007; (1 Suppl):201-208.
- Swayne DE, Suarez DL, Sims LD. Influenza. In: Swayne DE, Glisson JR, McDougald LR, Nair V, Nolan LK, Suarez DL, editors. Diseases of poultry. 13th ed. Ames: Willey-Blackwell; 2013. p.181-218.
- Wise MG, Suarez DL, Seal BS, Pedersen JC, Senne DA, King DJ, et al. Development of a real-time reverse-transcription PCR for detection of Newcastle disease virus RNA in clinical samples. Journal of Clinical Microbiology 2004;42(1):329-338.
- Xishan Lu, Shi Yi, Gao F, Xiao H, Wang M, Qi J, et al. Insights into avian influenza virus pathogenicity: the hemagglutinin precursor HA0 of subtype H16 has an alpha-helix structure in its cleavage site with inefficient HA1/HA2 cleavage. Journal of Virology 2012;86(23):12861-12870.
- Zarkov IV. Avian influenza A virus and Newcastle disease virus mono and co-infections in birds. Trakia Journal of Sciences 2017;2:182-191.
- Ziegler JE, Horsfall FL. Interference between the influenza viruses, I. the effect of active virus upon the multiplication of influenza viruses in the chick embryo. Journal of Experimental Medicine 1944;79(4):361-377.
Publication Dates
-
Publication in this collection
Jul-Sep 2018
History
-
Received
25 Dec 2017 -
Accepted
08 Apr 2018