Abstract
BACKGROUND: Red blood group genes are highly polymorphic and the distribution of alleles varies among different populations and ethnic groups. AIM: To evaluate allele polymorphisms of the Rh, Kell, Duffy and Kidd blood group systems in a population of the State of Paraná METHODS: Rh, Kell, Duffy and Kidd blood group polymorphisms were evaluated in 400 unrelated blood or bone marrow donors from the northwestern region of Paraná State between September 2008 and October 2009. The following techniques were used: multiplex-polymerase chain reaction genotyping for the identification of the RHD gene and RHCE*C/c genotype; allele-specific polymerase chain reaction for the RHDΨ and restriction fragment length polymorphism polymerase chain reaction for the RHCE*E/e, KEL, FY-GATA and JK alleles. RESULTS: These techniques enabled the evaluation of the frequencies of Rh, Kell, Duffy and Kidd polymorphisms in the population studied, which were compared to frequencies in two populations from the eastern region of São Paulo State. CONCLUSION: The RHCE*c/c, FY*A/FY*B, GATA-33 T/T, JK*B/JK*B genotypes were more prevalent in the population from Paraná, while RHCE*C/c, FY*B/FY*B, GATA-33 C/C, JK*A/JK*B genotypes were more common in the populations from São Paulo.
Genotype; Polymorphism, genetic; Blood group; Brazil
ORIGINAL ARTICLES
Genetic polymorphisms of Rh, Kell, Duffy and Kidd systems in a population from the State of Paraná, southern Brazil
Gláucia Andréia Soares GuelsinI; Ana Maria SellI; Lilian CastilhoII; Viviane Lika MasakiI; Fabiano Cavalcante de MeloI; Margareth Naomi HashimotoI; Loide Souza HirleI; Jeane Eliete Laguila VisentainerI
IUniversidade Estadual de Maringá UEM, Maringá (PR), Brazil
IIUniversidade Estadual de Campinas UNICAMP, Campinas (SP), Brazil
Correspondence Correspondence: Jeane Eliete Laguila Visentainer Universidade Estadual de Maringá - Departamento de Ciências Básicas da Saúde Av. Colombo, 5790 87020-900 Maringá ( PR) Brazil Phone: 55 44 3011-4864/Fax: 55 44 3011-4931 jelvisentainer@uem.br jelvisentainer@gmail.com
ABSTRACT
BACKGROUND: Red blood group genes are highly polymorphic and the distribution of alleles varies among different populations and ethnic groups.
AIM: To evaluate allele polymorphisms of the Rh, Kell, Duffy and Kidd blood group systems in a population of the State of Paraná
METHODS: Rh, Kell, Duffy and Kidd blood group polymorphisms were evaluated in 400 unrelated blood or bone marrow donors from the northwestern region of Paraná State between September 2008 and October 2009. The following techniques were used: multiplex-polymerase chain reaction genotyping for the identification of the RHD gene and RHCE*C/c genotype; allele-specific polymerase chain reaction for the RHDΨ and restriction fragment length polymorphism polymerase chain reaction for the RHCE*E/e, KEL, FY-GATA and JK alleles.
RESULTS: These techniques enabled the evaluation of the frequencies of Rh, Kell, Duffy and Kidd polymorphisms in the population studied, which were compared to frequencies in two populations from the eastern region of São Paulo State.
CONCLUSION: The RHCE*c/c, FY*A/FY*B, GATA-33 T/T, JK*B/JK*B genotypes were more prevalent in the population from Paraná, while RHCE*C/c, FY*B/FY*B, GATA-33 C/C, JK*A/JK*B genotypes were more common in the populations from São Paulo.
Keywords: Genotype; Polymorphism, genetic; Blood group; Brazil
Introduction
According to the International Society for Blood Transfusion (ISBT), there are currently 308 red blood cell antigens classified into 30 blood group systems.(1) The genes of all these systems have been cloned except for one: the gene responsible for the P1/P2 polymorphism.(2,3)
Knowledge of the molecular basis of these genes enabled the development of molecular biology typing methods, the identification of new mutations, an understanding of polymorphisms and the discovery of new alleles. Genes of red blood cell groups are highly polymorphic and the distribution of allelic frequencies in these systems varies between different regions of the world.(4,5)
The red blood cell groups are composed of numerous antigens, of which the main ones are: ABO, Rh, Kell, Duffy, Kidd and MNS. ABO and Rh are the most important systems in transfusion medicine.
The Rh system is the most complex of all blood group systems with 50 well-established antigens currently known. The antigens of this system are encoded by two genes located on chromosome 1: RHD and RHCE.(5)
The Kell system consists of 31 antigens, including six pairs of antithetical antigens. The Kell protein is associated with other proteins on the red blood cell membrane.(6) The classic Kell antigen (KEL1) is important in transfusion medicine: anti-K antibodies can cause severe or even fatal transfusion reactions and perinatal hemolytic disease (PHD). The gene encoding the antigens of this system (KEL) is located on chromosome 7q33 and its expression occurs in erythroid cells and in some tissues, such as the brain, lymphoid organs, the heart and muscles.(7)
The Duffy gene (FY or DARC), the first human gene recognized as being autosomal, consists of two exons, and its locus is on chromosome 1q22-q23. Fya and Fyb are encoded by the FY*A and FY*B alleles, and are responsible for the Fy (a+b-), Fy (a-b+) and Fy (a+b+) phenotypes.(8,9)
The common phenotypes of the Kidd blood group system [Jk (a+b-), Jk (a-b+) and Jk (a+b+)] are encoded by two codominant alleles: JK*A and JK*B, located in the 18q12,3 region.(10-12)
The aim of the present study was to evaluate allele polymorphisms of the Rh, Kell, Duffy and Kidd blood group systems in a population from the northwestern region of the State of Paraná, southern Brazil.
Methods
Donors
After receiving approval from the Human Research Ethics Committee and after signing informed consent terms, peripheral blood was randomly collected from 400 voluntary blood and bone marrow donors from the city of Maringá in the northwest of Paraná State. Of these 400 donors, 217 were male and 183 female, with a mean age of 31.16 ± 11.34 years (20-42 years). The donors were of mixed ethnic origin, but predominantly of Caucasian descent.
Genomic DNA extraction
To obtain the DNA, 5 mL of peripheral blood was collected in EDTA and centrifuged (2500 rpm for 10 min) to obtain the Buffy-coat. The DNA was extracted using the EZ-DNA (Biological Industries®, Kibbutz Beit Haemek, Israel) extraction kit following the manufacturer's protocol.
RHD, RHDY and RHCE*C/c genotyping
Genotyping of the alleles of the Rh system was performed according to the protocol previously described in the literature,(13) with some modifications: For the amplification of the RHCE*C/c alleles and to test for the presence of the RHD gene, the PCR (polymerase chain reaction)-multiplex technique was used. This technique amplifies RHCE*C/c alleles and tests for the presence of RHD through the analysis of the exon 7 and intron 4 of this gene. Detection of a 37-bp duplication in the RHD gene known as RHDΨ or RHD pseudogene, was achieved using the AS-PCR (allele-specific polymerase chain reaction) technique with a pair of specific primers for this genetic region and a pair of primers (HGH - human growth hormone) as an internal control. For both reactions (PCR-multiplex and AS-PCR), 200 ng of DNA, 50 pmol of each primer, 2 nmol of each dNTP, 1.0 U of Taq DNA polymerase (Invitrogen Life Technologies®, Grand Island, NY, USA) and buffer to make up a final volume of 50 µL were used. The PCR-multiplex amplification cycles were performed in a System 9700 PCR thermocycler (Applied Biosystems®, Foster City, CA, USA) and consisted of: denaturation at 95ºC for 15 minutes and 30 cycles of one minute at 94ºC, one minute at 65ºC and 3.5 minutes at 72ºC, followed by extension for ten minutes at 72ºC. For AS-PCR, the amplification cycles consisted of: denaturation at 95ºC for five minutes and 28 cycles of one minute at 94ºC, one minute at 60ºC and one minute at 72ºC, followed by extension for seven minutes at 72ºC. The analysis of the PCR products obtained was performed after electrophoresis in 2% agarose gel, stained with SYBR Green (Invitrogen Life Technologies®, Grand Island, NY, USA) in a micro cube SSP gel system (One Lambda®, San Diego, CA, USA).
RHCE*E/e, KEL, FY-GATA and JK genotyping
RHCE*E/e, KEL*1/KEL*2, FY*A/FY*B-GATA and JK*A/JK*B genotyping was performed by the RFLP-PCR (restriction fragment length polymorphism-polymerase chain reaction) technique according to protocols previously described in the literature,(14,15) with some modifications. The PCR reaction used 200 ng of DNA, 50 pmol of each primer, 2 nmol of each dNTP, 1.0 U of Taq DNA polymerase (Invitrogen Life Technologies®, Grand Island, NY, USA) and buffer to make up a final volume of 50 µL. The amplification cycles were performed in a System 9700 PCR thermocycler (Applied Biosystems®, Foster City, CA, USA) and consisted of: denaturation at 95ºC for 15 minutes and 35 cycles of twenty seconds at 94ºC, twenty seconds at 62ºC and twenty seconds at 72ºC, followed by extension for ten minutes at 72ºC. The products obtained from the RHCE*E/e, KEL*1/KEL*2, FY*A/FY*B-GATA and JK*A/JK*B PCR were digested overnight with their respective restriction enzymes (MBI Fermentas®, Canada): Mnl I, Bsml, Ban I, Sty I and Mnl I, in final volumes of 20 µL, using 10 µL of PCR product and 10 µL of the enzyme/buffer mixture, following the protocol recommended by the manufacturer. Analysis of mutations in the GATA-box region was performed after electrophoresis in 6% polyacrylamide gel stained with silver. Three percent agarose gel stained with SYBR Green (Invitrogen Life Technologies®, Grand Island, NY, USA) was used to evaluate the other alleles (RHCE*E/e, KEL*1/KEL*2, FY*A/FY*B and JK*A/JK*B).
Statistical analysis
The genotypic and allelic frequencies were obtained by direct counting on Excel spreadsheets (Microsoft® Office Excel 2003). A comparison of the genetic and allelic frequencies in blood groups from the population of blood and bone marrow donors in this study and the two previously studied populations of blood donors from Campinas, São Paulo, was carried out.(16,17) The comparison was performed using the Chi-square test or Fisher's exact test, where appropriate, using a 2x2 contingency table.(18) The Arlequin computer program version 3.1 available at: http://cmpg.unibe.ch/software/arlequin3/ was used to see if the distributions of genes and alleles were in Hardy-Weinberg equilibrium.
Results
The present study is the first to report the frequency of HR, KEL, FY and JK gene polymorphisms in a population from the northwestern region of the State of Paraná (Table 1). The population was found to be in Hardy-Weinberg equilibrium for all the genes analyzed.
The Rh system
Of the 400 samples analyzed, 345 (86.25%) were RHD positive (presence of the RHD gene) and 55 (13.75%) RHD negative (absence of the RHD gene). The RHD pseudogene (RHDΨ) was present in only four heterozygous samples (1.0%) and so none of the samples presented the RhD negative phenotype. The frequencies for the RHCE gene were: 17.50% for the RHCE*C/C genotype, 42.75% for RHCE*C/c and 39.75% for RHCE*c/c. For the RHCE gene the frequencies were: 2.25% for RHCE*E/E, 25.75% for RHCE*E/e and 72.0% for RHCE*e/e.
The Kell system
The KEL*2/KEL*2 genotype was observed in 379 samples (94.75%), KEL*1/KEL*2 was observed in 20 samples (5.0%) and only one had the rare KEL*1/KEL*1 genotype (0.25%).
The Kidd system
The genotype distribution for this system was as follows: JK*A/JK*A was observed in 109 (27.25%) samples, JK*A/JK*B in 192 (48.0%) and JK*B/JK*B in 99 (24.75%).
The Duffy system
The FY*A/FY*A genotype was observed in 50 (12.60%) samples, FY*A/FY*B in 192 samples (48.0%) and FY*B/FY*B in 158 (39.50%) samples. The mutation (-33T>C), which occurs in the GATA-box erythroid promoter region and prevents the expression of the FY*B allele on the surface of red blood cells, was evaluated. For this polymorphism, 10 (2.5%) samples were genotyped as being homozygous (GATA-33C/C), 78 (19.5%) as heterozygous (GATA-33T/C) and the vast majority, 312 samples (78.0%), without this mutation (GATA-33T/T). All of the homozygous samples (GATA-33C/C) had the FY*B/FY*B genotype, which corresponds to the Fy(a-b-) phenotype. Among the heterozygous samples (GATA-33T/C), 35 (44.87%) had the FY*A/FY*B genotype, i.e. they expressed the Fy(a+b-) phenotype and 43 (55.13%) had FY*B/FY*B, which corresponds to the Fy(a-b+) phenotype which is characterized by a reduced expression of the Fyb antigen.
Allele frequencies and comparison of red blood cell gene frequencies in populations from Paraná and São Paulo
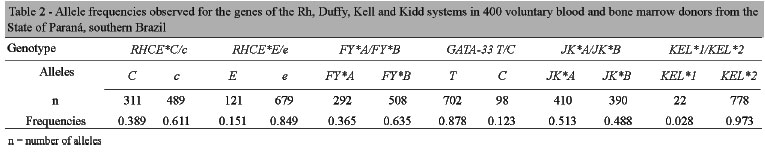
Allele frequencies were calculated and the results are shown in Table 2.
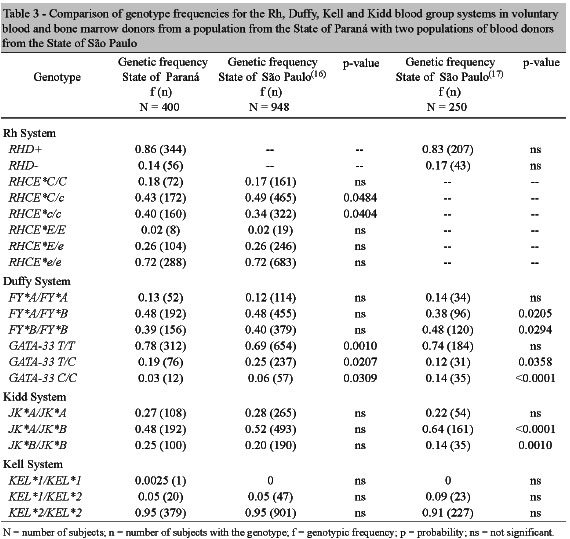
The gene frequencies observed in the voluntary blood and bone marrow donors evaluated in this study were compared with those found in two populations of voluntary blood donors from the city of Campinas in the State of São Paulo (Table 3).(16,17)
Several statistical differences were observed in red blood cell gene frequencies. The RHCE*C/c, FY*B/FY*B, GATA-33 C/C and JK*A/JK*B genotypes were more prevalent in the populations from São Paulo, while the RHCE*c/c, FY*A/FY*B, GATA-33 T/T and JK*B/JK*B genotypes were more common in the population from Paraná. There was a difference in the frequency of the GATA-33 T/C genotype between the two populations from São Paulo: in one of the populations the frequency was 0.25,(16) while in the other it was 0.12.(17) In the population from Paraná, the frequency for this genotype was 0.19.
Discussion
The population of Brazil is one of the most heterogeneous in the world, resulting from interbreeding of races from three continents over a period of five centuries: the European colonizers (mainly from Portugal and Italy), African slaves and local Amerindian populations.(19,20)
The samples evaluated in the present study from the State of Paraná, in the south of Brazil, represent an interracial mixture of European, African and Amerindian genes, as the region is predominantly populated by European immigrants (80.5%), with small but significant numbers of Africans (12.5%) and Amerindians (7.0%).(21) For this reason, in Brazil, skin color, determined by physical evaluation, is a poor predictor of genomic African ancestry.(19)
In a study carried out on a population from Amazonia in the north of Brazil, the frequency of the FY*A allele was 0.365, of FY*B (adding together the frequencies of the FY*B and FY*B-33 alleles) was 0.62 and of the FY*X allele the frequency was 0.014.(22) These allele frequencies are similar to those found in the population from Paraná in the current study.
According to the Brazilian Census Institute (Instituto Brasileiro de Geografia Estatística http://www.ibge.gov.br), the population of the State of São Paulo can also be considered as being of mixed ethnic origin, with a predominance of Europeans (70.0%) and Africans (28.0%). When the gene frequencies of the samples from Paraná were compared with those of the two populations from São Paulo, statistically significant differences were observed for the RHCE*c/c, FY*A/FY*B, GATA-33 T/T, JK*B/JK*B genotypes, which were more common in the population from Paraná, and for the RHCE*C/c, FY*B/FY*B, GATA-33 C/C, JK*A/JK*B genotypes, which were more prevalent in the populations from São Paulo.
A previous study carried out with populations from the same geographical regions (the northwest of Paraná State and the east of São Paulo State) compared the polymorphisms of cytokine genes and found no statistical differences between the two populations for the genes studied.(23) The genetic similarity observed between these two populations could be due to a similar pattern of miscegenation. However, the differences observed for some red blood cell genes demonstrates that, although similar, they may have been subject to a greater influence of one or another ethnic group.
Knowledge of the molecular basis of blood groups has made the study of red blood cell gene frequencies possible in different parts of the world. However, until now, few studies have been carried out on Brazilian populations. Further research must be carried out in all regions of the country in order to gain a greater understanding of the genetic variability and distribution of these systems to discover new alleles and to design new studies and strategies to study rare genotypes and phenotypes. This would enhance the search for compatible blood when carrying out transfusions in different geographical regions. These studies should include other blood group systems that are relevant to transfusion medicine, such as the MNS, Diego, Dombrock, Colton and Lutheran systems, among others. In the near future, DNA array technology could be widely used for this type of research, as it allows the identification of various polymorphisms within a single platform, as well as the simultaneous analysis of large numbers of samples.
Genotyping techniques used in this study enabled the evaluation of the genetic frequencies of Rh, Kell, Duffy and Kidd polymorphisms in a population from the northwestern region of the State of Paraná, and the results obtained can be used for anthropological comparisons, as well as for association studies for different diseases. Furthermore, this study enabled the evaluation of the genotyping method for the identification of red blood cell genes; this was considered to be a reliable alternative to resolve transfusion complications, where conventional immunohematological techniques have limitations.
Acknowledgements
The authors would like to thank the voluntary donors who participated in this study and everyone that contributed to its completion. This research was financially supported by LIG-UEM and by the Fundação Araucária do Estado do Paraná. The manuscript was linguistically revised by Peter Grimshaw.
Submitted: 4/7/2010
Accepted: 11/21/2010
Conflict-of-interest disclosure: The authors declare no competing financial interest
- 1. Daniels G, Castilho L, Flegel WA, Fletcher A, Garratty G, Levene C, et al. International Society of Blood Transfusion Committee on Terminology for red blood cell surface antigens: Macao report. Vox Sang. 2009;96(2):153-6.
- 2. Hellberg A, Chester MA, Olsson ML. Two previously proposed P1/P2-differentiating and nine novel polymorphisms at the A4GALT (Pk) locus do not correlate with the presence of the P1 blood group antigen. BMC Genet. 2005;6:49.
- 3. Tilley L, Green C, Daniels G. Sequence variation in the 5' untranslated region of the human A4GALT gene is associated with, but does not define, the P1 blood group polymorphism. Vox Sang. 2006;90 (3):198-203.
- 4. Yip SP. Sequence variation at the human ABO locus. Ann Hum Genet. 2002;66(Pt1):1-27.
- 5. Daniels G. The molecular genetics of blood group polymorphism. Hum Genet. 2009;126(6):729-42.
- 6. Lee S. Molecular basis of Kell blood group phenotypes. Vox Sang. 1997;73(1):1-11. Erratum in: Vox Sang 1998;74(1):58.
- 7. Lee S, Zambas ED, Marsh WL, Redman CM. The human Kell blood group gene maps to chromosome 7q33 and its expression is restricted to erythroid cells. Blood. 1993;81(10):2804-9.
- 8. Pruenster M, Rot A. Throwing light on DARC. Biochem Soc Trans. 2006;34(Pt6):1005-8.
- 9. Miller LH, Mason SJ, Dvorak JA, McGinniss MH, Rothman IK. Erythrocyte receptors for (Plasmodium knowlesi) malaria: Duffy blood group determinants. Science. 1975;189(4202): 561-3.
- 10. You G, Smith CP, Kanai Y, Lee W-S, Stelzner M, Hediger MA. Cloning and characterization of the vasopressin-regulated urea transporter. Nature. 1993;365(6449):844-7.
- 11. Olivès B, Merriman M, Bailly P, Bain S, Barnett A, Todd J, et al. The molecular basis of the Kidd blood group polymorphism and its lack of association with type 1 diabetes susceptibility. Hum Mol Genet. 1997;6(7):1017-20.
- 12. Lin Y, Pavenski K, Saidenberg E, Branch DR. Blood group antigens and normal red blood cell physiology: a Canadian blood services research and development symposium. Transfus Med Rev. 2009; 23(4):292-309.
- 13. Singleton BK, Green CA, Avent ND, Martin PG, Smart E, Daka A, et al. The presence of an RHD pseudogene containing a 37 base pair duplication phenotype and a nonsense mutation in africans with the Rh D-negative blood group. Blood. 2000;95(1):12-8.
- 14. Rios M, Cash K, Strupp A, Uehlinger J, Reid ME. DNA from urine sediment or buccal cells can be used for blood group molecular genotyping. Immunohematology. 1999;15(2):61-5.
- 15. Reid ME, Rios M, Yazdanbakhsh K. Applications of molecular biology techniques to transfusion medicine. Semin Hematol. 2000;37(2):166-76.
- 16. Ribeiro KR, Guarnieri MH, da Costa DC, Costa FF, Pellegrino J Jr, Castilho L. DNA array analysis for red blood cell antigens facilitates the transfusion support with antigen-matched blood in patients with sickle cell disease. Vox Sang. 2009;97(2):147-52.
- 17. Pellegrino J Jr, Castilho L, Rios M, De Souza CA. Blood group genotyping in a population of highly diverse ancestry. J Clin Lab Anal. 2001;15(1):8-13.
- 18. Svejgaard A, Jersild C, Nielsen S, Bodmer WF. HLA and disease. Statistical genetic consideration. Tissue Antigens. 1974;4(2):95-105.
- 19. Parra FC, Amado RC, Lambertucci JR, Rocha J, Antunes CM, Pena SDJ. Color and genomic ancestry in Brazilians. Proc Natl Acad Sci USA. 2003;100(1):177-82.
- 20. Carvalho-Silva DR, Santos FR, Rocha J, Pena SD. The phylogeography of Brazilian Y-chromosome lineages. Am J Hum Genet. 2001;68(1):281-6.
- 21. Probst CM, Bompeixe EP, Pereira NF, de O Dalalio MM, Visentainer JE, Tsuneto LT, et al. HLA polymorphism and evaluation of European, African, and Amerindian contribution to the white and mulatto populations from Paraná, Brazil. Hum Biol. 2000;72(4): 597-617.
- 22. Cavasini CE, de Mattos LC, Couto AA, Couto VS, Gollino Y, Moretti LJ, et al. Duffy blood group gene polymorphisms among malaria vivax patients in four areas of the Brazilian Amazon region. Malar J. 2007;6:167-74.
- 23. Visentainer JE, Sell AM, da Silva GC, Cavichioli AD, Franceschi DS, Lieber SR, et al. TNF, IFNG, IL6, IL10 and TGFB1 gene polymorphisms in South and Southeast Brazil. Int J Immunogenet. 2008;35(4-5):287-93.
Correspondence:
Publication Dates
-
Publication in this collection
05 May 2011 -
Date of issue
Feb 2011
History
-
Accepted
21 Nov 2010 -
Received
04 July 2010