Abstracts
Random Amplified Polymorphic DNA (RAPD) procedure was used to examine the genetic variability among fourteen isolates of Trichoderma and their ability to antagonize Rhizoctonia solani using a dual-culture assay for correlation among RAPD products and their hardness to R. solani. Seven oligodeoxynucleotide primers were selected for the RAPD assays which resulted in 197 bands for 14 isolates of Trichoderma. The data were entered into a binary matrix and a similarity matrix was constructed using DICE similarity (SD) index. A UPGMA cluster based on SD values was generated using NTSYS (Numerical Taxonomy System, Applied Biostatistics) computer program. A mean coefficient of similarity obtained for pairwise comparisons among the most antagonics isolates was around 40%. The results presented here showed that the variability among the isolates of Trichoderma was very high. No relationship was found between the polymorphism showed by the isolates and their hardness, origin and substrata.
RAPD; Trichoderma; Rhizoctonia solani
A técnica de RAPD (Random Amplified Polymorphic DNA) foi utilizada para examinar a variabilidade genética em quatorze isolados de Trichoderma além de sua capacidade de antagonizar o fungo fitopatogênico Rhizoctonia solani usando pareamento in vitro, e a possível relação entre perfís de RAPD e agressividade dos isolados de Trichoderma a R. solani. Foram selecionados sete primers para os ensaios de RAPD, os quais produziram 197 bandas. Os dados foram introduzidos no programa de computador NTSYS (Numerical Taxonomy System, Applied Biostatistics)na forma de uma matrix binária, sendo construída uma matriz de similaridade utilizando-se o coeficiente de similaridade de DICE (SD) e baseado nos valores SD, pelo método de agrupamento UPGMA um dendrograma. Observou-se que o grau de similaridade das amostras que apresentaram melhor desempenho antagônico foi bastante baixo, em torno de 40%. Os resultados demonstraram que a variabilidade entre os isolados de Trichoderma é muito alta. Aparentemente não foi demonstrada nenhuma relação entre o perfil de RAPD obtido e o nível de antagonismo dos isolados de Trichoderma nem agrupamento em função da origem dos isolados ou substrato.
Randomly Amplified Polymorphic DNA of Trichoderma Isolates and Antagonism Against Rhizoctonia solani
Larissa Brandão Góes; Ana Bolena Lima da Costa; Laurineide Lopes de Carvalho Freire and Neiva Tinti de Oliveira* * Author for correspondence
Universidade Federal de Pernambuco-UFPE; CCB; Depto. Micologia; CEP 50.670-420; Recife - PE, Brasil
ABSTRACT
Random Amplified Polymorphic DNA (RAPD) procedure was used to examine the genetic variability among fourteen isolates of Trichoderma and their ability to antagonize Rhizoctonia solani using a dual-culture assay for correlation among RAPD products and their hardness to R. solani. Seven oligodeoxynucleotide primers were selected for the RAPD assays which resulted in 197 bands for 14 isolates of Trichoderma. The data were entered into a binary matrix and a similarity matrix was constructed using DICE similarity (SD) index. A UPGMA cluster based on SD values was generated using NTSYS (Numerical Taxonomy System, Applied Biostatistics) computer program. A mean coefficient of similarity obtained for pairwise comparisons among the most antagonics isolates was around 40%. The results presented here showed that the variability among the isolates of Trichoderma was very high. No relationship was found between the polymorphism showed by the isolates and their hardness, origin and substrata.
Key words: RAPD; Trichoderma; Rhizoctonia solani
INTRODUCTION
R. solani Kühn [anamorph of Thanatephorus cucumeris (Frank) Donk], is a common pathogen for several commercial plant crops in Brazil (Bolkan & Ribeiro, 1985; Brancão & Martins, 1983). The measures preconized for its control, such as rotation of cultures, use of resistant varieties and treatment of seeds and/or soil with fungicides, a lot of times become unsuitable or not effective, mainly due to the genetic variability presented by the pathogen to the hosts range, capacity to survive in the soil and in the seeds, and physiologic flexibility to infect different hosts (Leach & Garber, 1970).
The use of antagonistic microrganisms to R. solani has been investigated as an alternative control method, species of Trichoderma, applied as treatment of seeds or soil, have been demonstrated to control the pathogen in a variety of cultures in the greenhouse and field studies (Melo, 1991; Nelson, 1991; Papavizas, 1985; Papavizas, 1992).
The ability of Trichoderma to control R. solani varies considerably, and it is possible to improve its biological control efficiency by the selection of isolates with high antagonistic potential and adapted to certain ecological or geographical areas (Noronha et al. 1996).
The Random Amplified Polymorphic DNA (RAPD) procedure developed by Williams et al. (1990) and Welsh & Mc Clelland (1990) that involves simultaneous amplification of several anonymous loci in the genome using primers of arbitrary sequence has been used for genetic, taxonomic and ecological studies of several fungi (Crowhurst et al., 1995; Fungaro et al. ,1996; Manulis et al., 1993; Zinno et al., 1998; Abbasi, et al., 1999; Paavanem-Huhtala et al., 2000).
Fourteen strains of Trichoderma were studied by Santos (1992), using molecular markers of RAPD among other techniques, and it was possible to verify the natural genetic variability and to divide the strains in similarity groups, as well as differentiating the original strains of different regions. Muthumeenakshi et al. (1998) characterized genetically 15 strains of T. harzianum aggressive for edible mushrooms in the United States and England using RAPD, wich were designated "T. harzianum group 4", that presented a high homogeneity degree. Comparison of molecular data of group 4 with group 2 (the causal agent of the epidemic green mold in industrial mushrooms in England) indicated that the isolates of T. harzianum group 4 were different from that of group 2. Gomez et al. (1997) analyzed the RAPD profiles of strains of T. harzianum and classified them in different groups according to their capacity for control of plant pathogenic fungi. Arisan-Atac et al. (1995) studied 11 strains of T. viride, 2 strains of Hypocrea rufa and 9 other species of Trichoderma with relation to the RAPD profile and their ability for controlling Cryphonectria parasitica through pairing in vitro.
The results allowed the construction of a dendrogram where the strains characterized with potential for antagonize C. parasitica were gathered in very different groups. The isolate of H. rufa was among T. viride strains with negative biocontrol capacity, what was in agreement with their non ability for antagonizing C. parasitica. Zimand et al. (1994) used RAPD markers obtained from 9 arbitrary primers to distinguish strains of Trichoderma. Ten of the strains identified as T. harzianum exhibited similarities, and it was possible to distinguish the isolate T-39, used commercially as biocontrol agent of Botrytis cinerea. For efficient selection of strains of Trichoderma with taxonomic finalities, Fujimori & Okuda (1993) examined 74 strains of Trichoderma by RAPD profiles and the results were consistent with the morphological, physiological and ecological data of these strains, what suggests that the technique can aid to eliminate strains duplicated in a program for microbial selection. Using RAPD Schlick et al. (1994) analyzed strains of T. harzianum and mutants induced by gamma radiation originated from one wild isolate, verifying that with RAPD it was possible to differentiate all the mutants strains for at least one primer and concluding that the method was valuable for identification and fast differentiation of strains.
The genetic variability of Trichoderma isolates using the RAPD technique and their antagonistic potential against R. solani, as well as the relationship between their antagonistic capacity and RAPD profiles were analysed.
MATERIAL AND METHODS
Isolates used
Fourteen isolates of Trichoderma (six of T. viride, six of T. harzianum, one T. polysporum, one of T. pseudokoningii) and one of R. solani (isolate registered under no4014 from Micoteca URM,UFPE, Recife,PE,Brazil) were obtained from the Culture Collection (URM) of the Federal University of Pernambuco (UFPE), Recife, PE, Brazil and from Culture Collection of the Federal Rural University of Pernambuco (UFRPE).
Extraction and amplification of the DNA of the isolates of Trichoderma
The extraction of total genomic DNA of each isolate of Trichoderma was made as described by Raeder & Broda (1985). The primers OPA, OPR, OPX and OPW, as well as Taq DNA polimerase and the dNTPs were supplied by Operon Technologies, CA, USA. Amplification reactions were carried out in a total volume of 25 m l containing approximately 20-25 ng of template DNA, primer 0,4 mmol l-1 and MgCl2 3,0 mmol l-1. The control reaction consisted of all the components, but the genomic DNA, none of the used primers presented detectable amplified products. The amplification process was accomplished in thermal cycler (MJ Research), programmed to accomplish an initial denaturing of 5 minutes at 92 °C, 40 cycles of 1 minute at 92 °C (denaturing), 1 minute and 30 seconds at 39 °C (annealing), 2 minutes at 72 °C (extension) and finally 5 minutes to 72 °C for a final extension, during 40 cycles. After the process, samples of 22 m l of the amplification products were assayed by electrophoresis in 1,4% agarose gels, at 3 V/cm-1 of distance among the electrodes, running with TBE buffer. Gels were stained with ethidium bromide, and photographed under U.V. light. The l DNA cut by HindIII was used as molecular weight size markers.
The NTSYS.PC (Numerical Taxonomy System Applied Biostatistics, Setauket, New York) computer program was used for data analysis. The data (band presence or absence) were introduced in the form of a binary matrix and a pairwise similarity matrix,was constructed using the DICE coefficient (SD) (Sneath & Sokal, 1973). SD values were obtained by the double number of shared bands between two patterns divided by the sum of all the bands in the same pattern (value 1 indicates identical patterns for two individuals and value 0 indicates completely different patterns). The UPGMA (Unweighted pair-group method with arithmetical averages) grouping of the SD values was generated using the NTSYS program.
Antagonistic capacity of Trichoderma isolates
The antagonism experiments were accomplished by the method described by Dennis & Webster (1971). The isolates were cultivated in Petri dishes with PDA media for seven days. Disks of 5mm of diameter were cut and removed from the growing borders of the colonies and transferred to another Petri dish with PDA. Each plate received two disks, one with Trichoderma mycelium and another with Rhizoctonia mycelium, placed at a distance of 7 cm from each other, in time intervals according the growth speed test organisms. The plates were incubated at 26°C. The experiment was entirely at random with three replicates.
Classification of the isolates was based on the Bell et al. (1982) scale of antagonism classes.
RESULTS AND DISCUSSION
Primers selection
Eighty primers from the kits OPA, OPR, OPW and OPX were tested. Forty primers from OPR and OPW did not provide amplification products. Some primers of the kits A and X amplified the tested DNA, but only seven were chosen, because they presented strong and defined bands, and consistent fragments patterns: OPA5 (5'-AGGGGTCTTG), OPA13 (5'-AGCACCCAC), OPX1 (5'-CTGGGCACGA), OPX2 (5'-TTCCGCCACC), OPX3 (5'-GGCGCAGTG), OPX7 (5'-GAGCGAGGCT) and OPX12 (5'-TCGCCAGCCA).
DNA Polimorphism analysis of the Trichoderma isolates
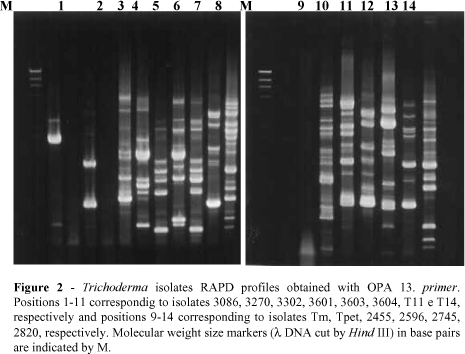
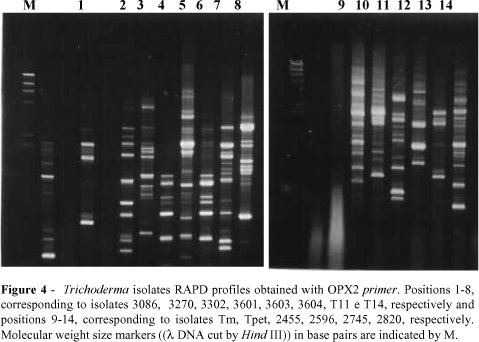
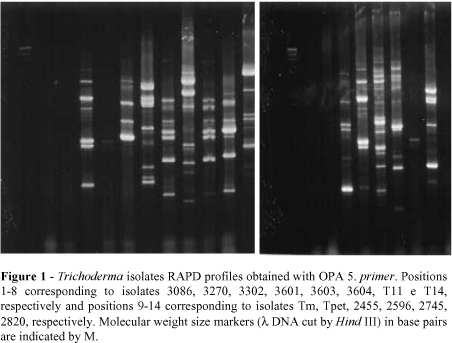
A total of 197 bands were produced with the 7 primers used (Figs. 1-7). The dendrogram constructed through the method UPGMA is presented in Fig 8. For elaboration of the dendrogram, it was necessary to use lost portion for the isolate 2745, amplified by the primer OPA-5, in the attempt of avoiding a false positive result. No monomorphic bands were found, while polimorphic bands were evident.
Classification of Trichoderma isolates antagonistic to R. solani, in vitro
Trichoderma isolates demonstrated high antagonistic action over R. solani; the isolates of T. harzianum were the most aggressive. The isolates of Trichoderma overlaped the colony of R. solani with 120 hours except for the isolates 2596, 3086 and 2745. The isolates 3302, 3601and Tm, started overlaping the colony of R. solani earlier than the other isolates. The isolate T14 (Trichoderma pseudokoningii) was not able of overlaping the colony of R. solani, but inhibited its growth (Table 1). The classification of the isolates with relation to the antagonism, and based on the overlaping of the colony, according to Bell et al. (1982) is shown in Table 2.
RAPD Profiles and the antagonism of isolates of Trichoderma against R. solani
The intraspecific genetic variation among the isolates was clear (Fig. 8). The isolate Tm (isolated from corn seed) was grouped in the dendrogram with the isolate 2820 (isolated from sugar-cane). Although belonging to the same species, the isolates presented low similarity, around 55%. The 3603 isolate presented stonger antagonistic action than the isolate T14, in agreement with the classification of antagonism (Table 2). The isolate T14 and 3603 were in the same group in the dendrogram, although they were of different species. Arisan-Atac et al. (1995) working with RAPD of species of Trichoderma and two strains of Hypocrea rufa antagonistic against C. parasitica, also obtained this type of contradictory result, verifying that H. rufa and T. viride were in the same group in the dendrogram. This suggested that doubts in the classic classification of species of Trichoderma existed and should be investigated with complementary information being achieved by other methods.
The technique of RAPD was efficient in demonstrating the DNA polymorphism of these isolates of Trichoderma, showing hight intraspecific genetic variability.
The similarity degree of among the samples that presented better antagonistic acting was around 40%. The dendrogram did not show grouping of the isolates by the level of antagonism. Relationships among the polymorphism existent, the aggressiveness and the origin of the isolates were not found.
RESUMO
A técnica de RAPD (Random Amplified Polymorphic DNA) foi utilizada para examinar a variabilidade genética em quatorze isolados de Trichoderma além de sua capacidade de antagonizar o fungo fitopatogênico Rhizoctonia solani usando pareamento in vitro, e a possível relação entre perfís de RAPD e agressividade dos isolados de Trichoderma a R. solani. Foram selecionados sete primers para os ensaios de RAPD, os quais produziram 197 bandas. Os dados foram introduzidos no programa de computador NTSYS (Numerical Taxonomy System, Applied Biostatistics)na forma de uma matrix binária, sendo construída uma matriz de similaridade utilizando-se o coeficiente de similaridade de DICE (SD) e baseado nos valores SD, pelo método de agrupamento UPGMA um dendrograma. Observou-se que o grau de similaridade das amostras que apresentaram melhor desempenho antagônico foi bastante baixo, em torno de 40%. Os resultados demonstraram que a variabilidade entre os isolados de Trichoderma é muito alta.
Aparentemente não foi demonstrada nenhuma relação entre o perfil de RAPD obtido e o nível de antagonismo dos isolados de Trichoderma nem agrupamento em função da origem dos isolados ou substrato.
Received: May 22, 2000;
Revised: October 30, 2000;
Accepted: June 27, 2001.
- Abbasi, P. A.; Miller, S. A.; Meulia, T.; Hoitink, H. A. J. and Kim, J. M. (1999) Precise detection and tracing of Trichoderma hamatum 382 in compost-amended potting mixes by using molecular markers. Applied and Environmental Microbiology, 65 : (12), 5421-5426.
- Arisan-Atac, I.; Heidenreich, E. and Kubicek, C. P. (1995), Randomly amplified polymorphic DNA fingerprinting identifies subgroups of Trichoderma viride and other Trichoderma sp. capable of chestnut blight biocontrol. FEMS Microbiology Letters, 126, 249-256.
- Bell, D. K.; Well, H. D. Markhabell, D. K. and Wells, C. R. (1982), In vitro antagonism of Trichoderma species against six fungal plant pathogens. Phythopatology, 72 : (4), 379-382.
- Bolkan, H. A. and Ribeiro, W. R. C. (1985), Anastomosis groups and pathogenicity of Rhizoctonia isolates from Brasil. Plant Disease, 69 : (7), 599-601.
- Brancão, N. and Martins, R. M. (1983), Levantamento de doenças de beterraba açucareira (Beta vulgaris L.). Fitopatologia Brasileira, 8, 592.
- Chet, I. (1990), Biological control of soil-borne plant pathogens with fungal antagonists in combination with soil treatments. In:. Biological control of soil-borne plant pathogens Hornby, D. (Ed.)Wallingford: CAB International, pp. 15-25.
- Crowhurst, R. N.; King, F. Y.; Hawthorne, B. T.; Sanderson, F.R. and Choi-Pheng, Y. (1995), Rapd characterization of Fusarium oxysporum associated with wilt of angsana (Pterocarpus indicus) in Singapore. Mycological Research, 99 : (1), 14-18.
- Dennis, C. J. and Webster, J. (1971), Antagonism properties of species groups of Trichoderma, III. Hyphal interaction. Transactions British Mycological Society, 57, 363-369.
- Fungaro, M. H. P.; Vieira, M. L. C.; Pizzarani-Kleiner, A. A. and Azevedo, J. L. (1996), Diversity among soil and insect isolates of Metarhizium anisopliae var. anisopliae detected by RAPD. Letters in Applied Microbiology, 22, 389-392.
- Fugimori, F. and Okuda, T. (1993), Aplication of the random amplified polymorphic DNA using the polymerase chain reaction for efficient elimination of duplicate strains in microbial screening. The Journal of antibiotics, 47 : (2), 173-182.
- Gomez, I.; Chet, I. and Herreraestrela, A., (1997), Genetic diversity and vegetative compatibility among Trichoderma harzianum isolates. Molecular & General Genetics, 256, 127-135.
- Leach, L. D. and Garber, R. H. (1970), Control of Rhizoctonia solani In - Rhizoctonia solani: Biology and pathology Parmeter, J. R., (ed.), Berkeley: The University of California Press, pp. 189-199.
- Manulis, S.; Kogan, N.; Reuven, M. and Ben-Yephet, Y. (1993), Use of the RAPD technique for identification of Fusarium oxysporum f. sp. dianthi from carnation. Phytopathology, 84 : (1), 98-101.
- Melo, I. S. (1991), Potencialidade da utilização de Trichoderma spp., no controle biológico de doenças de plantas. In - Controle biológico de doenças de plantas Bettiol, W., (Org.) Jaguariúna: EMBRAPA-CNPDA, pp. 135-156. (EMBRAPA-CNPDA. Documentos; 15).
- Muthumeenakshi, S., Brown, A. E. and Mills, P. R. (1998), Genetic comparision of aggressive weed mould strains of Trichoderma harzianum from mushroom compost in North America and the British Isles. Mycological Research, 4, 385-390.
- Nelson, E. B. (1991), Current limits to biological control of fungal phytopathogens. In: Handbook of applied mycology-Soil and plants Arora, D. K.; Rai, B.; Mukerji, K. G.; Knudsen, G., (Eds.) New York: Marcell Dekker, 1, 327-355.
- Noronha, M. A.; Sobrinho, S. A.; Silveira, N. S. S; Michereff, S. J.; Mariano, R. L. R. and Maranhão, E. (1996), Seleção de isolados de Trichoderma spp. para o controle de Rhizoctonia solani em feijoeiro. Summa Phytopathologica, 22, 156-162.
- Paavanem-Huhtala, S.; Avikainem, H. and Yli-Mattila, T. (2000), Development of strain-specific primers for a strain of Gliocladium catenulatum used in biological control, 106 : (2), 187-198.
- Papavizas, G. C. (1985), Trichoderma and Gliocladium: Biology and potential for biocontrol. Annual Review of Phytopathology, 23, 23-54.
- Papavizas, G. C. (1992), Biological control of selected soilborne plant pathogens with Gliocladium and Trichoderma In - Biological control of plant diseases Tjamos, E. C., Papavizas, G. C., Cook, R. J., (Eds.). New York: Plenum Press, pp 223-230,
- Raeder, U. and Broda, P. (1985), .Rapid preparation of DNA from filamentous fungi. Letters in Applied Microbiology, 1, 17-20.
- Santos, I. C. (1992), Avaliação da variabilidade genética de Trichoderma por isoenzimas, RFLP e RAPD. Piracicaba-SP,. thesis- São Paulo University ,Escola Superior de Agricultura "Luiz de Queiroz" (ESALQ), Brazil.
- Schlick, A.; Kuhls, K.;Meyer, W.; Lieckfeldt, E.; Börner, T. and Messner, K. (1994), Fingerprinting reveals gamma-ray induced mutations in fungal DNA: implications for identification of patent strains of Trichoderma harzianum Current Genetic, 26, 74-78.
- Sneath, P. H. A. and Sokal, R. R. (1973), Numerical Taxonomy: The Principles and Pratice of Numerical Classification San Francisco: Freeman, 573 pp.
- Welsh, J. and McClelland, M. (1990), Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Research, 18, 7213-7218.
- Zimand, G.; Valinsky, L.; Elad, Y.; Chet, I. and Manulis, S. (1994), Use of the RAPD procedure for the identification of Trichoderma strains. Mycological Research, 98 : (5), 531-534.
- Zinno, T. D.; Longrée, H. and Maraite, H. (1998), Diversity of Pyrenophora tritici-repentis isolates from warm wheat growin atreas: pathogenicity, toxin production and RAPD analysis. In - Helminthosporium blights of wheat: spot blotch and tan spot Duveiller et al. (eds.), Mexico, D.F.: CIMMYT, pp. 302-311.
Publication Dates
-
Publication in this collection
27 Sept 2002 -
Date of issue
June 2002
History
-
Received
22 May 2000 -
Reviewed
30 Oct 2000 -
Accepted
27 June 2001