ABSTRACT
The application of beneficial bacteria has recently been used for sustainable agriculture. In current research, 71 bacterial isolates were obtained from rice plant and the rhizosphere soil of different paddy fields in Guilan province, Iran. After primitive investigation, 40 bacteria with typical predominant characteristics were selected. By PCR-RFLP of their 16S r-DNA gene, 8 Operational Taxonomic Units (OTUs) totally consisted of 33 isolates were obtained. From all of them, 8 isolates were selected for rice seed germination experiment, then, effective isolates were used for pot experiment to evaluate their ability for promoting rice growth. All of them were able to increase rice growth and yield, but in different potential. These tested isolates were identified as Alcaligenes faecalis (DEp8, O1R4), Pantoea ananatis (AEn1), Bacillus vietnamensis (MR5), Bacillus idriensis (MR2) and Stenotrophomonas maltophilia by partial sequencing of their 16S r-DNA gene. Among them, AEn1 and MR5 produced indole-3- acetic acid (IAA) in larger amounts than the other isolates and the isolates AEn1 and O1R4 were able to solubilize phosphate in higher amounts. According to the results obtained, it can be concluded that AEn1, O1R4 and MR5 can be considered as bacterial inoculants to use as alternatives for chemical fertilizers.
Key words:
Rice associated bacteria; Plant growth promoting bacteria; PCR-RFLP of 16S r-DNA; IAA production; Phosphate solubilizing bacteria
INTRODUCTION
Rice, which is the most important cereal crop and the main food in many countries, provides more than 20% of the consumers’ daily calories. About 90% of rice is cultivated in 200 million paddy fields in Asia 1919 IRRI, AfricaRice, CIAT. Global Rice Science Partnership (GRISP); 2010. 267 p.. In Iran, most rice farms are in the Northern provinces including Guilan and Mazandaran.
Due to the growing demand for rice, one of the increasing concern about rice production in the world especially in Iran is the extra use of chemical fertilizers, which have negative effects on human health and environment 9. So, there is an urgent need to use the substitutes without or with less harmful side effects. Thus, recently, microorganisms as an inoculants to reduce the use of chemicals are being a matter of interest 1313 Gerhardson B. Biological substitutes for pesticides. Trnds Biotechnol. 2002; 20: 338-343.. For stimulating rice seed germination and rice growth promotion biologically, there are many attempts to find successful bacteria 33 Ahmed B, Midrarullah Sajjad Mirza M. Effects of inoculation with plant growth promoting rhizobacteria (PGPRs) on different growth parameters of cold area rice variety, Fakre Malakand. Afr J Microbiol Res. 2013; 7: 1651-1656.,55 Baset Mia MA, Shamsuddin ZH, Mahmood M. Effects of rhizobia and plant growth promoting bacteria inoculation on germination and seedling vigor of lowland rice. Afr J Biotechnol . 2012; 11: 3758-3765.,2121 Jha Y, Subramanian RB. Isolation of root associated bacteria from the local variety of rice GJ-17. World Res J Geoinformatics. 2012; 1: 21-26.,4242 Sharma A, Shankhdhar D, Sharma A, Shankhdhar SC. Growth promotion of the rice genotypes by PGPRs isolated from rice rhizosphere. J Soil Sci Plant Nutr. 2014; 14: 505-517..
Rice seed priming with plant growth promoting bacteria (PGPB), as an alternative for chemicals, can be considered to enhance seed germination, vigor index and germination speed 1111 Doni F, Anizan I, Che Radziah CMZ, Salman AH, Rodzihan MH, Wan yusoff WM. Enhancement of rice seed germination and vigor by trichoderma spp. Res J App Sci Eng Technol. 2014; 7:4547-4552.. Because germination stage is the most susceptible period in rice life cycle 3939 Rajjou L, Duval M, Gallardo K, Catusse J, Bally J, Job C. Seed germination and vigor. Annu Rev Plant Biol. 2012; 63: 507-533.. Rice growth promotion can be achieved by beneficial bacteria but the exact mechanism by which PGPB induce their effects is not clearly understood, however, several hypotheses such as producing phytohormones, activation of phosphate solubilization and promotion of the mineral nutrient uptake are usually considered to be involved 88 Bowen GD, Rovira AD. The rhizosphere and its management to improve plant growth. Adv Agron. 1999; 66: 1-102.,1414 Glick BR. The enhancement of plant growth by free-living bacteria. Can J Microbiol Res. 1995; 159: 371-394.,2626 Lalande R, Bisssonnette N, Coutlee D, Antoun H. Identification of rhizobacteria from maize and determination of their plant-growth promoting potential. Plant Soil. 1989; 115: 7-11.,2929 Liu ST, Lee LY, Tai CY, Hung CH, Chang YS, Wolfram H J, Rogers R, Goldstein AH. Cloning of an Erwinia herbicola gene necessary for gluconic acid production and enhanced mineral phosphate solubilization in Escherichia coli HB101. J Bacteriol. 1992; 174: 5814-5819.. Production of phytohormones is one of the important mechanism and indole-3 acetic acid (IAA), which is a member of native auxin, is a common metabolism produced by several beneficial bacteria 3131 Lynch JM. Origin, nature and biological activity of alphabetic substances and growth hormones found in soil. In: Vaughan D, Malcom RE, editors. 9th ed. Soil organic matter and biological activity. Netherlands: Springer; 1985. P. 151-174.. Indole-3 acetic acid stimulates cell elongation through enhancement of cell osmotic contents and cell wall synthesis, permeability of water into cell, decrease in wall pressure and inducing protein synthesis, flowering and fruiting 22 Ahao Y. Auxin biosynthesis and its role in plant development. Annua Rev Plant Biol. 2010; 6: 49-64.. Solubilization of phosphate is one of the other important mechanism the PGPB uses to enhance growth of host plant. A great part of soil phosphorus is in the form of insoluble phosphate and cannot be used by plants 2222 Kannaiyan S, Kumar K, Govindarajan K. Biofertilizer technology for rice based cropping system. India: scientific publishers Jodhpur; 2004. 450 p. and phosphorus deficiency results in small leaves, weak stem and slow development 4040 Ranjan A, Rajan Mahalakshmi M, Sridevi M. Isolation and characterization of phosphate-solubilizing bacterial species from different crop fields of Salem, Tamil Nuda, India. I J N P N D. 2013; 3: 29-33.. By considering these facts, the aim of current study is to isolate rice associated bacteria from Guilan’s paddy fields, the evaluation of their ability to enhance rice seed germination and plant growth, and investigation of the mechanisms such as IAA production and phosphate solubilization in order to introduce any possible native beneficial bacteria for rice growth promotion as a biological inoculant to use in paddy fields instead of hazardous chemical fertilizers.
MATERIAL AND METHODS
Sample Collection and Isolation of Bacteria
Rice plants (variety Hashemi) including stem, leaf, root and rhizosphere soil were collected from different paddy fields of 16 sites in Guilan province, (37.268̊ N, 49.589̊ E, 2 m a.s.l), Iran, including Astara, Talesh, Rezvanshahr, Masal, Fooman, Shaft, Anzali, Rasht, Astaneh Ashrafieh, Lahijan, Langerood, Roodsar, Amlash, Rostam Abaad, Some Sara and Siahkal from June to July 2015. The condition of rice cultivation in paddy fields were continuous irrigation or continuous flow of water (ponding) and the soil type was clayey with the average pH of 6.7. Samples were kept in plastic bags and immediately brought to the laboratory of Rice Research Institute of Iran, Guilan. Samples were kept at 4 ̊C in refrigerator.
The epiphytic, endophytic and rhizospheric culturable bacteria were isolated according to Lindow et al. (1978), Sturz et al. (1997) and Mohite (2013), respectively. For isolation of epiphytic bacteria, rice samples of leaves and stems were cut in to small sizes and separately put in 500 ml flasks with 200 ml of 0.1 M phosphate buffer (pH:7.0) and 1 g/l peptone. Flasks were shaken for about 2h at 150 rpm. For endophytic isolation, stems, leaves and roots of rice were washed thoroughly under tap water and rinsed with deionized water. Surface sterilization of samples was done with 5% sodium hypochlorite solution for 5 minutes followed by 3 times washing with sterile distilled water. Then samples were cut into small pieces with sterile scalpel and were ground well with a sterile pestle and mortar. Rhizospheric bacteria were isolated by keeping the clay firmly adhering the roots and removing the rest. Ten grams of this clay from each sample were suspended in 90 ml of sterile distilled water in a 250 ml flasks and shaken for about 20 min at 120 rpm. Serial dilutions of all suspensions were made up to 1/105 and an aliquot of 100 µl from each dilutions were spread on Nutrient Agar (NA) medium. Single colonies were picked up after 3 days of incubation at 28 ̊C and streaked on fresh plates to get pure colonies.
Colony Characteristics
Colony characteristics such as shape, color, margins and gram staining were carefully studied. The final isolated bacteria were selected from the bacteria with typical predominant characteristics on NA medium. Thus, the redundant isolates with different characteristics but in few population were discarded
DNA Extraction and Amplification of 16S rDNA
Total genomic DNA of bacteria in each group was extracted from freshly grown cultures on NA medium according to the method described by Murry & Thompson (1980). Quantification of extracted DNA was done on 1% agarose gel electrophoresis. Universal primers 27f: 5′-AGAGTTTGATCCTGGCTCAG-3′ and 1492r: 5′- GGTTACCTTGTTACGACTT- 3′ were used for amplification of 16S rDNA region. The final reaction mixture was 10 µl, contained of 0.5 µl of magnesium chloride [MgCl2], 1 µl of PCR buffer, 1.2 µl of dNTPs, 0.6 µl of each forward and reverse primers, 2.96 µl sterile deionized water and 0.14 µl of Taq DNA Polymerase (Cinnagen, Iran). Polymerase Chain Reaction (PCR) amplification were carried out on 3 µl of DNA in an automated thermal cycler (T Gradient-Biometra, Germany) with following program: an initial denaturation in 94̊C for 4 min, 35 cycles of denaturation (94̊C for 45s), annealing (60̊C for 1 min), extension (72̊C for 2 min) and final extension in 72̊C for 7 min. PCR product size was confirmed on 1.5% agarose gel electrophoresis and photographed under UV light.
PCR-Restriction Fragment Length Polymorphism (PCR-RFLP) of 16S rDNA
This method was used to cluster bacterial isolates in groups with most similarity according to their restricted bands. Digestion of the amplified 16S rDNA was carried out by two recognizing enzymes: Mse I and Hinf I. The final reaction mixture was 15 µl, contained of 8 µl of sterile deionized water, 1.5 µl of 10X buffer with 5 µl of the PCR products subjected to digest with 5 units (5 µl) of each enzyme at 37 ̊ C for 30 to 60 min. The restricted DNA fragments were separated on 10% polyacrylamide gel (140 V for 2 h) stained in ethidium bromide and photographed under UV light. The results were converted into a binary matrix indicating the presence and absence of restricted bands. The analysis of this matrix was done using SM (Simple Matching) correlation index and clustering analysis was performed using Complete method. The correlation index and clustering analysis method was performed according to the highest cophenetic correlation coefficient (0.8486). The software NTSYSpc Ver2 was used for these analyses
Sequencing of 16S r-DNA
The 16S r-DNA gene of tested bacterial isolates, were partially sequenced using forward or reverse universal primers 27f and 1492r (used in PCR above) to identify the isolates. The sequencing reaction were done at Macrogen Sequencing Service, Republic of Korea and sequences were submitted to the National Center for Biotechnology Information database.
Rice Seed Germination Test
Rice seed (Variety Hashemi), obtained from Rice Research Institute of Iran, Guilan, were surface sterilized by 5% sodium hypochlorite for 5 min and rinsed three times with sterile distilled water. Then, they were soaked in different bacterial suspensions including 4 bacteria isolated from rhizosphere (O1R4, MR5, MR2 and KR1), 3 from rice endophyte (AEn1, AEn4 and FEn1) and 1 from rice epiphyte (DEp8), with sufficient population density of 106 to 107 cfu/ml and shaken overnight in 150 rpm. Seeds were soaked in sterilized distill water considered as control. For each isolate, 45 seeds in 3 replications were allocated and in each replication, 15 seeds were put on sterile filter paper in 100 mm petri dishes. They were irrigated by sterilized distill water during 6 days of incubation at 28±2 ̊C
In the stage of imperfect leaf emergence (code 09) 6, shoot and root length of all seedlings in each replication were measured in mm scale. For weight measurements, shoot and root were separated and weighted as fresh weight and dry weight after keeping at 48 ̊C for 72 h in an electric oven. Weight measurements was done by digital weight scales and noted in gram
Germination rate or seedling emergence was calculated using the following formula proposed by International Rice Research Institute (2011) on last day of experiment
Vigor index of seedling were measured on last day of experiment (day 6) according to the formula proposed by Abul-Baki & Anderson (1973):
The speed of germination was calculated by counting the number of emerged seedlings every day of experiment according to Gupta (1993):
The experiment was performed in a completely randomized design. Data were statistically analyzed with SAS version 9.0 and the comparison of means was done by LSD.
Growth Promotion in Pot Experiment
More effective isolates on enhancement of rice seed germination were selected for evaluation of their effects on growth and yield of rice. Seedling trays were sterilized with 5% sodium hypochlorite and washed thoroughly by sterile distilled water. They were filled with autoclaved soil and surface sterilized rice seed (Variety Hashemi) were sown for nursery raising. Seedling trays were irrigated by sterile distilled water until the stage of unfolded 3 leaves (code 13) 66 Biologische Bundesanstallt für Land-und Forstwirtschaft, Meier, U. Growth Stages of Mono-and Dicotyledonous Plants: BBCH Monograph. Berlin: Blackwell Wissenschafts-Verlag; 1997.. Rice pots sterilized with 5% sodium hypochlorite were filled with autoclaved paddy field soil and irrigated by sterilized water. For each isolate 6 pots (replications) and for each pot, 3 seedlings were allocated. Before planting, rice seedling roots were dipped and shaken in selected rhizobacterial suspension (the bacteria isolated from rice rhizosphere) with a population density of 106 to 107 cfu/ml for one hour in 150 rpm to let bacteria colonize roots (Sharma et al. 2014) and the phyllosphere spraying of bacterial suspension (the epiphytic and endophyte bacteria isolated from rice leaf and stem) was done after planting for epiphyte and endophyte bacterial colonization. Sprayed inoculated seedlings were wrapped up with plastic bags for 24 hours to make a humid condition for better colonization of bacteria. Seedlings without bacterial inoculation considered as control. Data collection of rice plant growth including stem and root length and weight were done by tillering stage (code 29) 66 Biologische Bundesanstallt für Land-und Forstwirtschaft, Meier, U. Growth Stages of Mono-and Dicotyledonous Plants: BBCH Monograph. Berlin: Blackwell Wissenschafts-Verlag; 1997., for 3 pots (replications) and the other 3 pots were remained to evaluate the bacterial effects on rice yield. Yield data collection including plant height and weight, flag leaf area, number of tillers, number of panicle, panicle length, weight of 1000 grain and number of full grain were done according to standard evaluation system for rice 2020 International Network for Genetic Evaluation of Rice, International Rice Research Institute. Standard evaluation system for rice, 4th ed. Manila Philippines, The International Rice Research Institute; 1996. 52 p. at ripening stage (code 89) 66 Biologische Bundesanstallt für Land-und Forstwirtschaft, Meier, U. Growth Stages of Mono-and Dicotyledonous Plants: BBCH Monograph. Berlin: Blackwell Wissenschafts-Verlag; 1997..
The pot experiment was done in a randomized complete block design and the data obtained were subjected to analysis with SAS version 9.0. The significance of differences between mean values were evaluated by LSD
Phosphate Solubilizing Ability
The phosphate solubilizing ability was studied by plate assay using the National Botanical Research Institute’s Phosphate (NBRIP) growth medium 38. The medium was consisted of 10g glucose, 5g calcium phosphate [Ca3(PO4)2], 5g magnesium chloride [MgCl2], 0.25g magnesium sulfate [MgSO4], 0.2g potassium chloride [KCl], 0.1g ammonium sulfate [(NH4)2SO4] and 15 g/l agar in 1 liter. The pH was adjusted to 7.0 before autoclave. Plates were kept at 28 ̊C and formation of visible halo zones in NBRIP media around bacterial colonies after 7 days showed the ability of solubilizing phosphate.
Production of IAA
The presence of IAA in bacterial metabolites and the amount of IAA production was colorimetrically determined by Salkowski reagent 1515 Glickmann E, Dessaux Y. A critical examination of the specificity of the salkowski reagent for indolic compounds produced by phytopatogenic bacteria. Appl Environ Microbiol . 1994; 61: 793-796.. Bacterial isolates were inoculated in broth containing 20g/l bacteriological peptone, 1.15g/l dipotassium phosphate [K2HPO4], 1.5g/l magnesium sulfate [MgSO4.7H2O] and the same broth supplemented by tryptophan (0.5g/l). Bacterial grown cultures were centrifuged after 48 hours incubation at 28 ̊C and 1ml of supernatant was mixed with 1ml of Salkowski reagent (12 g/l ferric chloride [FeCl3] in sulfuric acid [H2SO4, 7.9 M]). After 30 min of dark incubation at room temperature, the optical densities were measured at 530 nm by spectrophotometer (CECIL, England). The concentration of IAA production was estimated by a standard IAA curve.
RESULT
Isolation and Colony Characteristics
A total of 71 pure bacterial isolates were obtained and their morphological study showed typical bacterial characters. Their color varied from yellow to pink, milky and orange. Their shape was mostly rounded with some smooth or undulate margins and gram test separated them in to two major groups. Due to these typical predominant characteristics the total number of bacteria reduced to 40 and the rest with many diverse characteristics but in fewer population were removed
Amplification of 16S r-DNA and PCR-RFLP
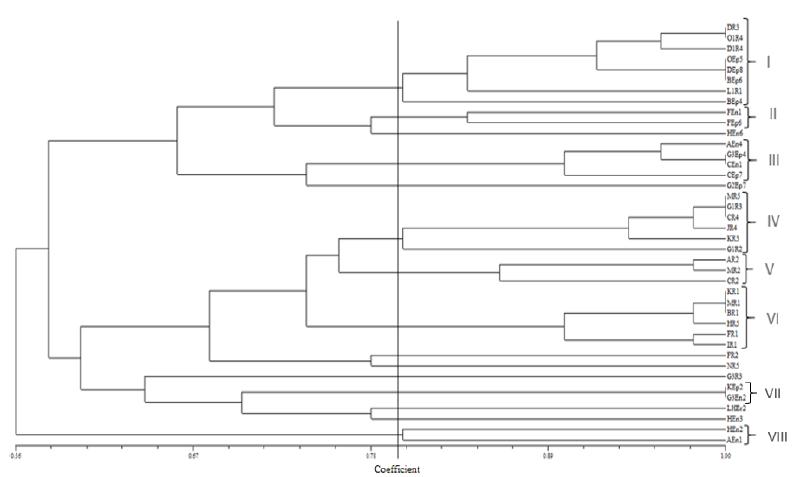
Amplification of all 40 bacterial 16S r-DNA genes produced single band around 1500 bp. After digestion of this amplified genes with restriction enzymes, the bacterial isolates were grouped according to their restricted patterns. Eight Operational Taxonomic Units (OTUs) including 33 bacterial isolates were found, whereas 7 isolates showed unique patterns (Fig 1). According to PCR-RFLP results, from OTU (I), with higher number of isolates, two representatives (DEp8 and OR4) and from each one of II, III, IV, V, VI and VIII OTUs, one representative isolate including (FEn1, AEn4, MR5, MR2, KR1 and AEn1) were selected respectively for evaluation of their ability to enhance rice seed germination.
Dendrogram based on UPGMA cluster analysis of PCR-RFLP pattern of 16S r-DNA profiles showing the relationships among the isolated bacteria
Rice Seed Germination
Seed inoculation with 8 bacterial isolates from 8 groups resulted in increased seedlings’ length and vigor index compared to un-inoculated control. In all treatments, the increase in shoot length were higher than control but in root elongation, just the isolate KR1 showed similar effect with control. The highest effect on shoot and root length obtained from the seedlings inoculated with isolates DEp8 and AEn1 respectively followed by O1R4 in shoot length and FEn1 in root length. Isolates KR1, MR2 and AEn4 could not increase the shoot fresh and dry weight more than control and O1R4 was the isolate which had the most effect on enhancement of shoot fresh and dry weight. The root fresh and dry weight of seedlings inoculated by MR2 were the highest but AEn4 could not increase fresh weight of roots. Comparing the treated seeds and non-treated seeds, there was not a considerable difference in germination rate and the result of data analysis showed that MR5, MR2, O1R4, DEp8 and FEn1 treated seeds had the same germination rate but some of the seeds in KR1, AEn1 and AEn4 treatments did not germinate. Speed of germination which computed by counting the number of germinated seeds at an interval of 24 h to the day number of counting from the beginning day of experiment, were faster in seeds inoculated by isolates MR2, OR4 and FEn1 respectively than un-inoculated seeds and vigor index of all seeds were enhanced by all bacterial treatments except the isolate KR1 (table 1).
Growth Promotion in Pot Experiment
From 8 isolates studied in seed germination test, the isolates KR1 and AEn4 which had less effects on evaluated traits, were discarded and the rest were used for pot experiment. At tillering stage (code 29) 66 Biologische Bundesanstallt für Land-und Forstwirtschaft, Meier, U. Growth Stages of Mono-and Dicotyledonous Plants: BBCH Monograph. Berlin: Blackwell Wissenschafts-Verlag; 1997., 3 pots were collected and the other 3 pots were left for yield increasing evaluation. The results of seedling inoculation with selected bacteria showed significant increase in all investigated characters including root and stem length and weight compared to un-inoculated seedlings. The isolate O1R4 showed the highest consistency in ability to enhance growth in almost all traits except the root length which the highest was for isolate MR2 followed by FEn1. The isolates MR2, DEp8 and AEn1 were in second place after O1R4 in increasing each evaluation of root and stem fresh and dry weight (table 2).
The evaluation of rice yield in 3 left pots were studied at the end of ripening stage. Many traits were measured and the inoculated bacteria were different in their potential to enhance rice yield. Plant height, which were measured from the soil surface up to the end of the longest panicle, were the highest in plants inoculated by the isolates AEn1 and DEp8 followed by MR5. The isolate AEn1 also produced the most number of full grains in inoculated rice plants compared to other isolates and un-inoculated control. Plants inoculated by MR5 produced the highest weight of 1000 grain and also the highest plant and panicle fresh and dry weight but in the evaluation of increasing flag leaf area, MR5 could not increase the area of flag leaf more than control. The number of panicles and tillers were significantly the highest for isolate DEp8. The isolate O1R4 showed the highest consistency in ability to increase panicle length and flag leaf area, but O1R4 with 3 other isolates, AEn1, FEn1 and DEp8 could not increase plant and panicles fresh and dry weight more than control (table 3).
Identification of Bacterial Isolates
Identification of tested bacteria in pot experiment were done through partial sequencing of their 16S r-DNA genes. They were identified as Alcaligenes facealis (O1R4 and DEp8 from the same OTU), Stenotrophomonas maltophilia (FEn1), Pantoea ananatis (AEn1), Bacillus vietnamensis (MR5) and Bacillus idriensis (MR2). The information of these isolates with related accession numbers are presented in table 4.
Phosphate Solubilizing Ability
All of six bacterial isolates were able to solubilize phosphate and produced clear halo zones around bacterial colonies on NBRIP medium. According to the halo zone size the isolates AEn1 (P. ananatis), O1R4 and DEp8 (A. facealis) were high solubilizer, but FEn1 (S. maltophilia), MR2 (B. idriensis) and MR5 (B. vietnamensis) solubilize phosphate in lower amount (table 5).
Production of IAA
The bacterial ability to produce IAA was determined by changing the color of bacterial broth after addition of Salkowski reagent and this color change was measured by spectrophotometer. All tested bacteria produced IAA in both media, but, tryptophan supplemented media had higher readings than the media without tryptophan. The amount of IAA was the highest in B. vietnamensis (MR5) which followed by P.ananatis (AEN1). The results are shown in table 5.
DISCUSSION
Different bacteria can be found in association with rice plant. They have the ability to enhance rice growth and because of this potential they can be used as biofertilizers 3333 Meunchang S, Janzen HH, Rovira AD. Phylogenetic and physiological characterization of indigenous Azospirillum isolates in Thailand. Soil Sci Plant Nutr. 2004; 50: 413-421.. In current research, many bacterial isolates were obtained from rice plants including endophytes, epiphyte and rhizosphere bacteria. At first, primitive investigations of bacteria, which is a common way of partial bacterial characterization 44 Ashrafuzzaman M, Hossen FA, Razi Ismail M, Anamul Hoque MD, Zahurul Islam M, Shahidullah SM Efficiency of plant growth-promoting rhizobacteria (PGPR) for the enhancement of rice growth. Afr J Biotechnol. 2009; 8: 1247-1252.,3737 Mwajita MR, Muage H, Tani A, Kahangi EM. Evaluation of rhizosphere, rhizoplane and phyllosphere bacteria and fungi isolated from rice in Kenya for plant growth promoters. Springer plus. 2013; doi: 10.1186/2193-1801-2-606
https://doi.org/10.1186/2193-1801-2-606...
,4242 Sharma A, Shankhdhar D, Sharma A, Shankhdhar SC. Growth promotion of the rice genotypes by PGPRs isolated from rice rhizosphere. J Soil Sci Plant Nutr. 2014; 14: 505-517.was done. In this way, the bacterial population density with very different characteristics can be discarded and the total population density is consisted of some nearly homogeneous bacterial groups that share same color, shape or gram reaction
The studied bacteria were from different geographical locations and from different parts of rice plants and this was reflected in 16S r-DNA digestion with restriction enzymes. The final phylogenetic tree showed that the isolates were rather divers and from 40 isolates, 7 OTUs representing 31 isolates at the similarity of about 80% were obtained. Fingerprinting of 16S r-DNA has been used by many researchers to discriminate bacterial isolates genetically 77 Biswas JC, Ladha JK, Dazzo FB. Rhizobia inoculation improves nutrient uptake and growth of lowland rice. Soil Sci Soc Am J. 2000; 164: 1644-1650.,1212 Fischer SE, Sandra I, Fischer SM, Gladys BM. Isolation and characterization of bacteria from the rhizosphere of wheat. World J Microbiol Biot. 2007; 23: 895-903.,4545 Tripathi AK, Verma SC, Ron EZ. Molecular characterization of a salt tolerant bacterial community in the rice rhizosphere. Res Microbiol. 2000; 153: 579-584.. In such a similar practice, Loaces et al. (2011), found 13 OTUs consisting of 55 isolates from 109 total isolates, whereas the rest showed unique restricted patterns indicating that the population density was quite variable.
To find bacteria with growth promoting effects on rice plants, one or two isolates from each OTU were selected for rice seed germination test. Seed priming or seed inoculation by beneficial bacteria has become a newly acceptable substitute for sustainable agriculture 11. The result of seed germination in current study showed that seedling length and weight can be increased by the inoculation with most of tested bacteria. Two isolates including KR1 and AEn4 could not affect rice seed germination beneficially significant compared to control and the other isolates. In many literatures, the increase in seedling length and weight is attributed to bacterial phytohormones 55 Baset Mia MA, Shamsuddin ZH, Mahmood M. Effects of rhizobia and plant growth promoting bacteria inoculation on germination and seedling vigor of lowland rice. Afr J Biotechnol . 2012; 11: 3758-3765.,1111 Doni F, Anizan I, Che Radziah CMZ, Salman AH, Rodzihan MH, Wan yusoff WM. Enhancement of rice seed germination and vigor by trichoderma spp. Res J App Sci Eng Technol. 2014; 7:4547-4552.,4141 Rosenblueth M, Martinez Romero E. Bacterial endophytes and their interactions with hosts. Mol Plant Microb Interct. 2006; 19: 827-837.. Among different phytohormones, indole-3-acetic acid (IAA) is usually considered as the most important native auxin. Production of IAA can vary in different species and strains and it is influenced by culture condition, growth stage and substrate availability 3434 Mirza MS, Ahmad W, Latif F, Haurat J, Bally R, Normand P. Isolation, partial characterization and the effect of plant-growth promoting bacteria (PGPB) on micro-propagated sugarcane in vitru. Plant Soil . 2001; 237: 47-54.. Tryptophan is considered as a precursor for IAA biosynthesis, because IAA producing bacterial culture supplemented by tryptophan produced more concentration of IAA 1010 Costacurta A, Vanderleyden J. Synthesis of phytohormones by plant associated bacteria. Crit Rev Microbiol. 1995; 21: 1-18.. Tryptophan dependent IAA biosynthesis had been also reported in many other bacteria. Tien et al. (1979) reported that Azospirillum was able to produce IAA when exposed to tryptophan or Karnwal (2009) tested fluorescent pseudomonas production of IAA in the absence and presence of tryptophan and showed that indole production enhanced by increasing tryptophan concentration. The assessment of IAA production in current research showed different concentration of IAA produced from all tested bacteria but the addition of tryptophan to the bacterial broth, increased IAA production considerably. It showed all of tested bacteria preferred tryptophan dependent way to produce IAA. There was not a significant differences between rhizospheric and epiphytic or bacterial endophyte tested, which is in agreement with issued publication of Lindow & Brandl (2003) who said that bacterial epiphytes are also known to produce phytohormnes like endophytes and rhizospheric bacteria.
Seed vigor is mainly determined based on the seedling length 18. Vigor index reflects the seedlings health, establishment and the state of final productivity of the plant 3232 Mcdonald MB, Copeland LO. Seed production: principles and practices. Chapman & hall, London; 1997. 719 p.. According to results obtained here, since the seedling length were significantly increased by bacterial inoculations, the seed vigor enhancement was anticipated, as it was. The speed of seed germination was not affected considerably by seed priming compared to control. In some bacterial treatments, the rate of seed germination was slowly during the experiment time. The isolates MR2 (B. idriensis) and O1R4 (A. faecalis) were able to colonize rice seeds in a shorter time and therefore can be more successful biofertilizers. This has been approved by other authors who said that to be an effective plant growth promoting bacteria, they must be able to establish themselves and colonize plant to reach at an appropriate density sufficient for producing beneficial effects 44 Ashrafuzzaman M, Hossen FA, Razi Ismail M, Anamul Hoque MD, Zahurul Islam M, Shahidullah SM Efficiency of plant growth-promoting rhizobacteria (PGPR) for the enhancement of rice growth. Afr J Biotechnol. 2009; 8: 1247-1252.
Vegetative growth is an important growth phase in many crops as it determines the amount of biomass production and in rice it is important for development of tillers. A strong vegetative growth of rice plants reflects a higher plant height and greater plant biomass, larger number of tillers and panicles 4242 Sharma A, Shankhdhar D, Sharma A, Shankhdhar SC. Growth promotion of the rice genotypes by PGPRs isolated from rice rhizosphere. J Soil Sci Plant Nutr. 2014; 14: 505-517.. In current investigations, after seed germination experiment, more successful bacteria were selected for pot experiment to evaluate bacterial growth promoting effects on vegetative growth phase. At tillering stage (code 29) 66 Biologische Bundesanstallt für Land-und Forstwirtschaft, Meier, U. Growth Stages of Mono-and Dicotyledonous Plants: BBCH Monograph. Berlin: Blackwell Wissenschafts-Verlag; 1997., rice growth in all measured traits including root and stem length and weight was higher than control and it showed the beneficial effects of bacteria treated. One reason of this enhancement can result from phytohormones which led to higher root and stem length. Production of IAA from all of treated bacteria in different concentrations can explain the different effectiveness of beneficial bacteria on rice growth. Other than phytohormones, solubilizing of phosphate can be other reason for growth enhancement of rice. Phosphorus is one of the major nutrient requirements for plant growth. Phosphate-solubilizing microorganisms, in addition to provide phosphorus for plants, provide growth promoting substances like hormones, vitamins and amino acids. It is reported that mechanism of solubilizing phosphate is associated with release of low molecular weight organic acids which chelate the cation bound to phosphate and convert it into soluble form 1616 Goldstein AH. Recent progress in understanding the molecular genetics and biochemistry of calcium phosphate solubilization by gram negative bacteria. Bio Agrc Hort. 1995; 12: 185-93.,2424 Kim k Y, Jordan D, Mcdonald GA. Enterobacter agglomerans, phosphate solubilizing bacteria, and microbial activity in soil: effect of carbon sources. Soil Bio Biochem. 1998; 30: 995-1003.,2525 Kpomblekou K, Tabatabai MA. Effect of organic acids production and phosphorus from phosphate rocks. Soil Sci. 1994; 158: 442-53.. According to our result from phosphate solubilizing ability of studied bacteria on NBRIP medium, almost all of them could solubilize phosphate but in different amount. The isolates ability of phosphate solubilizing was detected by measuring the halo zone size around bacteria. This halo zone was quite small in isolates FEn1 (S. maltophilia), MR2 (B. idriensis) and MR5 (B. vietnamensis) which was showed by a single (+) as low solubilizers, but the isolates AEn1 (P. ananatis), O1R4 and DEp8 (A. facealis), were considered as high phosphate solubilizers (+++). According to our results, there was not a significant differences between tested rhizospheric and epiphytic or bacterial endophyte for solubilizing phosphate which is not completely in agreement by Vazquez et al. (2000) who stated that high proportion of phosphate solubilizing bacteria are concentrated in the rhizosphere and they are metabolically more active than other sources. In current research, the isolate AEn1 (P. ananatis) which isolated from rice as a bacterial endophyte or DEp8 (A. faecalis) as a bacterial epiphyte were considered as high solubilizers since the diameter of the halo zone made by these isolates were more than the others, but this privilege was not reflected in the result of rice growth enhancement, because the rhizospheric isolates, MR5 (B. vietnamensis) and MR2 (B. idriensis) with low ability of phosphate solubilizing showed considerable enhancement of rice growth and yield. The same results obtained from the experiment conducted by Mwajita et al. (2013) in which over half of the bacterial phyllosphere isolates were able to solubilize phosphate, but generally the bacterial isolates from the rhizosphere and rhizoplane were more efficient in plant growth enhancement.
The increase in shoot and root dry matter directly effects on the plant higher productivity 4242 Sharma A, Shankhdhar D, Sharma A, Shankhdhar SC. Growth promotion of the rice genotypes by PGPRs isolated from rice rhizosphere. J Soil Sci Plant Nutr. 2014; 14: 505-517., which is clear in our results and the investigation of bacterial effects on rice yield at the end of growth season showed higher productivity than un-inoculated control including tiller number, panicle number, the weight of 1000 grain and the number of full grain. It is evident that many different species colonize the tissues of the plants, but their persistence, activity and their interaction in this microbial community is not completely cleared. The understanding of the interaction between bacteria-bacteria and bacteria-plant would help us to modify the beneficial effects that bacteria may have on host plants. In other words, introducing a PGPB and its inoculation to plant cannot guarantee the enhancement of plant yield. The success of a PGPB inoculation depends on different factors including soil and agricultural conditions, plant properties and the characteristics of the bacteria involved.
CONCLUSION
The current study showed that rice plants collected from different paddy fields in Guilan province, harbour diverse bacterial epiphytic, endophytic and rhizospheric population which some of them had the ability of plant growth promoting, but in different potential. All of tested bacteria were able to produce IAA and solubilize phosphate. Among them, the isolates Alcaligenes faecalis, Pantoea ananatis and Bacillus vietnamensis showed high potential of IAA production and phosphate solubilizing. From this study, it can be concluded that the above bacterial isolates can be promising as PGPB, but the final conclusion will be obtained when their potential of being a successful PGPB is investigated in paddy field conditions and when they are identified as safe microorganisms for human life in field trials.
REFERENCES
-
1Abul-Baki AA, Anderson JD. Vigor determination in soybean by multiple criteria. Crop sci .1973; 3: 630-637.
-
2Ahao Y. Auxin biosynthesis and its role in plant development. Annua Rev Plant Biol. 2010; 6: 49-64.
-
3Ahmed B, Midrarullah Sajjad Mirza M. Effects of inoculation with plant growth promoting rhizobacteria (PGPRs) on different growth parameters of cold area rice variety, Fakre Malakand. Afr J Microbiol Res. 2013; 7: 1651-1656.
-
4Ashrafuzzaman M, Hossen FA, Razi Ismail M, Anamul Hoque MD, Zahurul Islam M, Shahidullah SM Efficiency of plant growth-promoting rhizobacteria (PGPR) for the enhancement of rice growth. Afr J Biotechnol. 2009; 8: 1247-1252.
-
5Baset Mia MA, Shamsuddin ZH, Mahmood M. Effects of rhizobia and plant growth promoting bacteria inoculation on germination and seedling vigor of lowland rice. Afr J Biotechnol . 2012; 11: 3758-3765.
-
6Biologische Bundesanstallt für Land-und Forstwirtschaft, Meier, U. Growth Stages of Mono-and Dicotyledonous Plants: BBCH Monograph. Berlin: Blackwell Wissenschafts-Verlag; 1997.
-
7Biswas JC, Ladha JK, Dazzo FB. Rhizobia inoculation improves nutrient uptake and growth of lowland rice. Soil Sci Soc Am J. 2000; 164: 1644-1650.
-
8Bowen GD, Rovira AD. The rhizosphere and its management to improve plant growth. Adv Agron. 1999; 66: 1-102.
-
9Compant S, Duffy B, Nowak J, Clement CH, Barka EA. Use of plant growth-promoting bacteria for biocontrol of plant diseases: principles, mechanisms of action, and future prospects. Appl Environ Microbiol. 2005; 71: 4951-4959.
-
10Costacurta A, Vanderleyden J. Synthesis of phytohormones by plant associated bacteria. Crit Rev Microbiol. 1995; 21: 1-18.
-
11Doni F, Anizan I, Che Radziah CMZ, Salman AH, Rodzihan MH, Wan yusoff WM. Enhancement of rice seed germination and vigor by trichoderma spp. Res J App Sci Eng Technol. 2014; 7:4547-4552.
-
12Fischer SE, Sandra I, Fischer SM, Gladys BM. Isolation and characterization of bacteria from the rhizosphere of wheat. World J Microbiol Biot. 2007; 23: 895-903.
-
13Gerhardson B. Biological substitutes for pesticides. Trnds Biotechnol. 2002; 20: 338-343.
-
14Glick BR. The enhancement of plant growth by free-living bacteria. Can J Microbiol Res. 1995; 159: 371-394.
-
15Glickmann E, Dessaux Y. A critical examination of the specificity of the salkowski reagent for indolic compounds produced by phytopatogenic bacteria. Appl Environ Microbiol . 1994; 61: 793-796.
-
16Goldstein AH. Recent progress in understanding the molecular genetics and biochemistry of calcium phosphate solubilization by gram negative bacteria. Bio Agrc Hort. 1995; 12: 185-93.
-
17Gupta PC. Seed vigor testing. In: Agarwal kP, editor. Handbook of seed testing. New Delhi; National Seed Corporation; 1993. P. 245-246.
-
18International Rice Research Institute. Rice Knowledge Bank. Measuring seed germination [Internet]. 2011 [cited 2016 March 1]; Available from: Available from: http://www.knowledgebank.irri.org/index.php?option=com_zoo&view=item&layout=item&Itemid=468
» http://www.knowledgebank.irri.org/index.php?option=com_zoo&view=item&layout=item&Itemid=468 -
19IRRI, AfricaRice, CIAT. Global Rice Science Partnership (GRISP); 2010. 267 p.
-
20International Network for Genetic Evaluation of Rice, International Rice Research Institute. Standard evaluation system for rice, 4th ed. Manila Philippines, The International Rice Research Institute; 1996. 52 p.
-
21Jha Y, Subramanian RB. Isolation of root associated bacteria from the local variety of rice GJ-17. World Res J Geoinformatics. 2012; 1: 21-26.
-
22Kannaiyan S, Kumar K, Govindarajan K. Biofertilizer technology for rice based cropping system. India: scientific publishers Jodhpur; 2004. 450 p.
-
23Karnwal A. Production of indol acetic acid by fluorescent pseudomonas in the presence of l-tryptophan and rice root exudates. J plant pathol. 2009; 91: 61-63.
-
24Kim k Y, Jordan D, Mcdonald GA. Enterobacter agglomerans, phosphate solubilizing bacteria, and microbial activity in soil: effect of carbon sources. Soil Bio Biochem. 1998; 30: 995-1003.
-
25Kpomblekou K, Tabatabai MA. Effect of organic acids production and phosphorus from phosphate rocks. Soil Sci. 1994; 158: 442-53.
-
26Lalande R, Bisssonnette N, Coutlee D, Antoun H. Identification of rhizobacteria from maize and determination of their plant-growth promoting potential. Plant Soil. 1989; 115: 7-11.
-
27Lindow SE, Arny DC, Upper CD. Distribution of ice-nucleation-active bacteria on plants in nature. Appl Environ Microbial. 1978; 36: 831-838.
-
28Lindow SE, Brandl MT. Microbiology of the phyllosphere. Appl Environ Microbial . 2003; 69: 1875-1883.
-
29Liu ST, Lee LY, Tai CY, Hung CH, Chang YS, Wolfram H J, Rogers R, Goldstein AH. Cloning of an Erwinia herbicola gene necessary for gluconic acid production and enhanced mineral phosphate solubilization in Escherichia coli HB101. J Bacteriol. 1992; 174: 5814-5819.
-
30Loaces I, Ferrando L, Scavino AF. Dynamics, diversity and function of endophytic siderophore-producing bacteria in rice. Microbial Ecol. 2011; 61: 606-618.
-
31Lynch JM. Origin, nature and biological activity of alphabetic substances and growth hormones found in soil. In: Vaughan D, Malcom RE, editors. 9th ed. Soil organic matter and biological activity. Netherlands: Springer; 1985. P. 151-174.
-
32Mcdonald MB, Copeland LO. Seed production: principles and practices. Chapman & hall, London; 1997. 719 p.
-
33Meunchang S, Janzen HH, Rovira AD. Phylogenetic and physiological characterization of indigenous Azospirillum isolates in Thailand. Soil Sci Plant Nutr. 2004; 50: 413-421.
-
34Mirza MS, Ahmad W, Latif F, Haurat J, Bally R, Normand P. Isolation, partial characterization and the effect of plant-growth promoting bacteria (PGPB) on micro-propagated sugarcane in vitru. Plant Soil . 2001; 237: 47-54.
-
35Mohite B. Isolation and characterization of indole acetic acid (IAA) producing bacteria from rhizosphere soil and its effect on plant growth. J Soil Sci Plant Nutr. 2013; 13: 638-649.
-
36Murry MG, Thompson WF. Rapid isolation of high molecular weight plant DNA. Nucleic acids Res. 1980; 8: 4321-4325.
-
37Mwajita MR, Muage H, Tani A, Kahangi EM. Evaluation of rhizosphere, rhizoplane and phyllosphere bacteria and fungi isolated from rice in Kenya for plant growth promoters. Springer plus. 2013; doi: 10.1186/2193-1801-2-606
» https://doi.org/10.1186/2193-1801-2-606 -
38Nautiyal SC. An efficient microbiological growth medium for screening phosphate solubilizing microorganisms. FEMS Microbiol Lett. 1999; 170: 265-270.
-
39Rajjou L, Duval M, Gallardo K, Catusse J, Bally J, Job C. Seed germination and vigor. Annu Rev Plant Biol. 2012; 63: 507-533.
-
40Ranjan A, Rajan Mahalakshmi M, Sridevi M. Isolation and characterization of phosphate-solubilizing bacterial species from different crop fields of Salem, Tamil Nuda, India. I J N P N D. 2013; 3: 29-33.
-
41Rosenblueth M, Martinez Romero E. Bacterial endophytes and their interactions with hosts. Mol Plant Microb Interct. 2006; 19: 827-837.
-
42Sharma A, Shankhdhar D, Sharma A, Shankhdhar SC. Growth promotion of the rice genotypes by PGPRs isolated from rice rhizosphere. J Soil Sci Plant Nutr. 2014; 14: 505-517.
-
43Sturz AV, Christie BR, Matheson BG, Nowak J. Biodiversity of endophytic bacteria which colonize red clover nodules, roots, stems and foliage and their influence on host growth. Biol Fert Soils. 1997; 25: 13-19.
-
44Tien TM, Gaskinsa MH, Hubbell NDDH. Plant growth substances produced by Azospirillum brasilense and their effect on the growth of pearl millet (pennisetum americanum l.). Appl Environ Microbiol . 1979; 37: 1016-1024.
-
45Tripathi AK, Verma SC, Ron EZ. Molecular characterization of a salt tolerant bacterial community in the rice rhizosphere. Res Microbiol. 2000; 153: 579-584.
-
46Vazquez P, Holguin G, Puente M, Cortes AE, Basham Y. Phosphate solubilizing microorganisms associated with the rhizosphere of mangroves in a semi-arid coastal lagoon. Biol Fert Soils . 2000; 30: 460-8.
Publication Dates
-
Publication in this collection
2017
History
-
Received
03 Feb 2016 -
Accepted
14 July 2016