ABSTRACT
Monoaromatics, such as benzene, toluene, ethylbenzene and xylene (BTEX), are simple aromatic compound that are highly toxic due to their high solubility nature. Many chemical and physical methods for their degradation and breakup into nontoxic products are available, but still use of microorganism is preferred over these processes. In this present study Bacillus pumilus MVSV3 (Accession number JN089707), a less explored bacteria in the field of BTEX degradation, isolated from petroleum contaminated soil is utilized for BTEX degradation. At optimized conditions the isolate degraded 150 mg/L of BTEX completely within 48 h. GC-MS analysis revealed that the microorganism produces catechol and muconic acid during degradation indicating an ortho pathway of degradation. Enzyme assays were carried out to identify and characterize catechol 1, 2- dioxygenase (C12D). The optimal temperature and pH for the enzyme activity was identified as 35 (C and 7.5, respectively. SDS-PAGE revealed the molecular weight of the enzyme to be approximately 35,000 Da. Zymography analysis indicated the presence of three isoforms of the enzyme. Hence Bacillus pumilus MVSV3 and the isolated C12D, proved to be efficient in degrading the toxic aromatic compounds.
Key words:
Bacillus pumilus MVSV3; catechol 1; 2-dioxygenase; ortho pathway; BTEX.
INTRODUCTION
Simple monoaromatic compounds benzene, toluene, ethylbenzene and isomers of xylenes, together are called as BTEX. These compounds are in the European priority lists and are found commonly as groundwater contaminant due to their extensive use in petroleum industry or as solvent11. Jindrova E, Chocova M, Demnerova K, et al. Bacterial aerobic degradation of benzene, toluene, ethylbenzene and Xylene. Folia Microbiol. 2002; 47: 83-93.. They are toxic and produce significant health issues when exposed to concentrations higher than the EPA (Environment Protection Agency) maximum contaminant levels22. Cao B, Nagarajan K, Loh KC. Biodegradaion of aromatic compounds: current status and opportunities for biomolecule approaches. Appl Microbiol Biotechnol. 2009; 85: 207-228.. Though carcinogenic and harmful in nature, some organisms degrade these compounds into simpler and nontoxic substrates. It is because these compounds act as a source of carbon for the microorganism themselves, making the process more cost effective. Hence major research interest is now towards the biodegradation of BTEX using microorganism, as one among the concern is the contamination of drinking water sources by the water soluble and mobile petroleum components BTEX33. Payel S, Andrew RR, Shilpi G. Degradation of Aromatic Hydrocarbons (BTEX) by a solvent tolerant bacterial consortium. Journal of Urban and Environmental Engineering. 2013; 7: 274-279.. Many researches with microorganism on aromatic compounds have been tested for their degradation, but still search for a better microorganism continues. It is because that affinity of bacteria towards a particular hydrocarbon will ensure a wide choice of degradation. In many research, it was determined that aerobic conditions permit a rapid and complete mineralization of hydrocarbons. The aromatic structure of BTEX makes them inert towards simple oxidation or reduction and so requires an elaborate degradation strategy. Oxic conditions provide O2 as a powerful means to oxidize and cleave the aromatic ring44. Georg F, Matthias B, Johann H. Microbial degradation of aromatic compounds- from one strategy to four. Nature Reviews, Microbiology. 2011; 9: 803-816..
Catabolic pathways of volatile aromatic hydrocarbon degradation by microorganisms can vary in many chemical reactions. The end products produced are later incorporated in the metabolic pathways as sources of carbon and energy55 . Machnicka A, Suschka J. Activity of selected microorganism and mixtures in BTX biodegradation. Pol. J. Environ. Stud. 2001; 10: 341-346.. Metabolic pathway for BTEX degradation is carried out specifically by two enzymes: dioxygenases and monooxygenases. All pathways converge in the formation of catechol intermediates. Subsequently catechol is then mineralized by catechol 1, 2dioxygenase (via ortho cleavage pathway) or by catechol 2, 3 dioxygenase (C23D) (via meta cleavage pathway). Finally, producing low molecular mass compounds such as aldehydes and pyruvate66. Muftah HEN, Janice AA, Ayat EET. Aerobic biodegradation of BTEX: progresses and prospects. Journal of Environmental Chemical Engineering. 2012; 2: 1104-1122.. The ortho pathway of catechol degradation is also called as the β- ketoadipate pathway, and it has been studied in both gram negative and positive bacteria77. Emmanuel VPP, Dayananda S. Purification and characterization of catechol 1,2 dioxygenase from Acinetobacter sp. DS002 and cloning , sequencing of partial cat A gene. Indian J. Microbiol. 2011; 51: 312-318.. Although the importance of gram positive BTEX degrading bacteria is well known, only limited information is available about their detection in contaminated environment88. Tancsics A, Scoboszlay S, Kriszt, B, et al. Applicability of the functional gene catechol 1,2- dioxygenase as a biomarker in the detection of BTEX degrading Rhodococcus species. J. Appl. Microbiol. 2008; 105: 1026-1033..
The present study reports the degradation of BTEX by an isolated bacteria Bacillus pumilus MVSV3 with the characterization of C12D isolated from the bacteria. GC-MS were carried out to discuss the BTEX degradation pathway, which resulted in the production of catechol, muconic acid and other intermediates of ortho pathway.
MATERIALS AND METHODS
MICROORGANISM
Petroleum contaminated soil sample was collected from Ennore, Tamilnadu. BTEX degrading bacterial strains were isolated from the soil sample by enriching the soil in MSM (Mineral Salt mineral) media containing (g/L): 1.0g KH2PO4, 1.25g Na2HPO4. 2 H2O, 0.5g (NH4)2SO4, 0.5g MgSO4. 7 H2O, 0.5g CaCl2. 2 H2O, 0.005g FeSO4. 7 H2O supplemented with BTEX. All the experimental contents were sterilized for 20 min at 121 °C and 15 lbs in an autoclave before inoculation. Pure cultures were obtained by serially diluting 1 mL culture 105 times and spreading each 100 µL of aliquots on solid MSM (supplemented with BTEX)9. After incubation for 48 h, prominent colonies grown on MSM were selected as BTEX degrading bacteria for further studies. The pure culture was acclimatized in MSM with BTEX as sole source of carbon for 4 weeks with several transfers of acclimatized bacteria from low BTEX concentration (10 mg/L) to high concentration (300 mg/L). The bacteria isolated were identified by morphological, biochemical and 16S rRNA gene sequence analysis.
DEGRADATION STUDIES
Inoculum from BTEX acclimatized cells were used in this study. Optimized conditions were determined by keeping all variables constant except one for which optimization value was to be determined. The optimized value for each variable (such as Temperature, pH, Inoculum and substrate concentration) was selected and kept constant for further studies. The degradation studies by Bacillus pumilus MVSV3 were carried out in MSM supplemented with BTEX (individually) as sole carbon source at constant pH 7, temperature 30 ºC, inoculum concentration 3% and substrate concentration 150 mg/L of benzene, toluene, ethylbenzene and o-xylene respectively. The degradation set up was kept at 150 rpm in the orbital shaker for 48 h. Biomass concentration was determined by dry cell weight.
ANALYSIS OF BTEX AND THEIR METABOLITES
Optimized parameters were set and degradation studies were carried out. The samples to be analyzed were collected at regular interval. Centrifugation was carried out at 10000 rpm for 10 min. The supernatant was extracted thrice with the organic solvent hexane. The amount of BTEX present in the sample was then determined by HPLC (Shimaduzu, Japan) equipped with UV-Vis detector set at 254 nm, C18 column (Phenomenex, USA). The mobile phase consisted of acetonitrile and water (75:25), respectively. The flow rate was maintained at 1 mL/min. The degradation percentage was calculated as following: degrading efficiency = (Ci -Cf / Ci )×100; where, Ci - initial concentration of substrate, Cf - final concentration of substrate1010. Brinda LM, Velan, M. Biodegradation of the toxic polycyclic aromatic hydrocarbons, Phenanthrene by an indigenously isolated Alkaligenes feacalis MVMB1 strain. IPCBEE, 2nd international conference in environmental science and technology, IACSIT PRESS, Singapore; 2011.. GC-MS analysis was carried out to determine the intermetabolites produced during biodegradation and the equipment used was JEOL GC MATE II (USA). The flow rate was preset to 1 mL/min. The oven temperature was varied from 50 to 250 ºC at 10 ºC/ min, equipped with mass analyzer (quadruple double focusing mass analyzer), the detector used was photon multiplier tube with front inlet temperature of 220 (C. All metabolites were identified with authentic standard and NIST (National Institute of Standards and Technology) library data.
ENZYME EXTRACTION AND PURIFICATION
Bacterial strain was grown in 150 mL MSM (Minimal Salts Medium supplemented with BTEX) till exponential phase of growth. The cell pellets obtained by centrifugation at 7000 rpm for 15 min were washed with 10 mM phosphate buffer (pH 7.0) and resuspended in the same buffer. Disruption of the cells was done using cold sonication for 30 s at 70 % amplitude, cellular debris were removed again by centrifugation at 4 (C at 10000 rpm for 10 min. The supernatant obtained was used as crude enzyme and was maintained at 4 (C and used immediately. This crude was subjected to partial purification by dialysis and ammonium sulphate precipitation. Partial purification was carried out by dialysis with 0.1 M phosphate buffer (pH 8.0). Protein estimation was carried out using Lowry method using BSA (Bovine serum albumin) as standard1111. Lowry OH, Rosebrough NJ, Farr L, et al. Protein measurement with the folin phenol reagent. J.Biol.Chem. 1951; 193: 265-275..
ENZYME ASSAY
Enzyme activity was assayed spectrophotometrically using a Hitachi U-2800spectrophotometer. The standard assay of enzyme activity was performed by making an assay mixture containing 5 µL of catechol 1, 2 dioxygenase, 20 µL of 10 mM catechol as a substrate and final volume adjusted to 1 mL with 50 mM sodium phosphate buffer (pH 7.0). The enzyme activity was monitored by measuring the formation of cis, cis-muconic acid at 260 nm. One unit (U) of the enzyme activity was defined as the amount of the enzyme required to catalyze the formation of 1 μmol of product per min at 25 °C. Protein estimation was carried out using Lowry method using BSA as standard.
DETERMINATION OF MOLECULAR WEIGHT
The molecular weight of the enzyme C12D was determined by SDS-PAGE12. The analysis of molecular weight of partially purified enzyme was carried out by using different protein molecular weight markers1313. Matsumura E, Ooi S, Murakami S, Takenaka S, et al. Constitutive synthesis, purification and characterization of catechol 1,2-dioxygenase from the aniline- assimilating bacterium Rhodococcus sp.AN-22. J. Biosci. Bioeng. 2004; 98: 71-76..
CATECHOL ZYMOGRAPHY
Catechol was added to standard Laemmli acrylamide polymerization mixture, at a final concentration of 3 %, under non-reducing conditions. Partially purified protein extract was mixed in the ratio 3:1 with substrate gel sample buffer. About 20-30 mL of sample was loaded into Laemmli acrylamide stacking gel on a cast vertical mini gel. Gel was run at 15 mA and 20 mA during stacking and separating phase at 4 °C. Following electrophoresis, the gel was soaked in 2.5 % Triton X-100 with gentle shaking for 30 min at room temperature. The gel was then rinsed and incubated overnight at 37 °C in Tris-HCl (0.05 M, pH 7.5). The sample lane was later subjected to activity staining with slight modifications1414. Piyali M, Pranab R, Copper enhanced monooxygenase activity and FT-IR spectroscopic characterization of biotransformation products in trichloroethylene degrading bacterium : Stenotrophomonas maltophilia PM102, Hindawi Publishing corporation, BioMed Research International. 2013; Article ID 723680, Available from: http://dx.doi.org/10.1155/2013/723680.
http://dx.doi.org/10.1155/2013/723680...
. In brief the stock solution of p-toluidine was prepared in N,N-dimethylformamide and store over a prolonged period at 5°C. The proteins resolved on the Catechol incorporated acrylamide gels were stained with 50 µg/ml of p-toluidine and were incubated at room temperature in a gel rocker till the Catechol in the gels were stained black1515. Parke D.Application of p-toluidine in chromogenic detection of catechol and protocatechuate, diphenolic intermediates in catabolism of aromatic compounds. Applied And Environmental Microbiology, 1992; 58;2694-2697..
pH AND TEMPERATURE OPTIMA OF C12D
The effect of pH on the activity of enzyme was determined by assaying the enzyme activity at various pH (4.0 to 9.0 ) by using 0.5 M citrate phosphate buffer (for pH 4.0 to 5.5) and 0.5 M sodium - phosphate buffer (for pH 6.0 to 8.5) and 0.5 M glycine -NaOH buffer (for pH 9.0). The effect of temperature on enzyme activity was assayed by varying the temperature between 15 (C to 60 (C. Activity was checked at different pH and temperature after 30 min of incubation1616. Tsai SC, Li YK. Purification and characterization of catechol 1,2- dioxygenase from a phenol degrading Candida albicans TL3. Arch Microbiol. 2007; 187: 199-206..
FFECT OF METAL IONS
Enzyme sample (1 mL) was incubated with 50 mM of sodium phosphate buffer having 1.0 mM and 10 mM concentration of different metal ions. The ions tested were Cu2+, Mn2+, Mg2+, NH4 + and Ag+ at pH 7 and at 35 (C. The residual enzyme activity was determined by standard enzyme assay1717. Silva AS, Jacques RJS, Andreazza R, et al. Properties of catechol 1,2-dioxygenase in the cell free extract and immobilized extract of Mycobacterium fortuitum. Braz. J. Microbiol. 2013; 44: 291-297.. Ions were not added to the control. The chemicals used were copper sulphate, manganese chloride, magnesium chloride, ammonium sulphate and silver nitrate.
RESULTS AND DISCUSSION
BTEX degrading Bacillus pumilus MVSV3
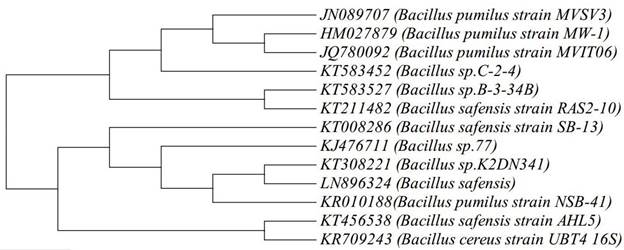
A total of five strains MVSV3, MVSV4, MVSV5, MVSV6 and MVSV7 were isolated from petroleum contaminated soil near Ennore, Tamilnadu. The isolated samples were determined as Bacillus pumilus Sp MVSV3, Bacillus cereus Sp MVSV4, Alcaligenes faecalis MVSV5, Ochrobactrum Sp MVSV6 and Alcaligenes faecalis MVSV7 by 16S rRNA gene sequence. These isolates have also been deposited in NCBI, under the following accession numbers - JN089707, JN089708, JN089709, JN089710 and JN089711, respectively. Initial Analysis revealed that the maximum degradation of all the four compounds were achieved with Bacillus pumilus MVSV3 than the other bacterial strains99. Sheeba VS, Brinda LM, Velan M. Isolation and characterization of non-adapted BTEX degrading bacterial strains from petroleum contaminated environment.. J. Pure, Appl. Microbio. 2015; 9: 3161-3169..The bacterial isolate MVSV3 formed unpigmented, irregular and wrinkled colonies on MSM supplemented with BTEX as sole source (Table 1). The isolated bacterial cells were gram positive, motile and aerobic rods. The nearly complete 988-bp-long16S rRNA gene sequence of the isolate MVSV3 was found to be 99% identical to that of Bacillus pumilus strain MW-1 (HM027879.1). After pairwise alignment of the sequences of 16S rRNA of the isolate and the EMBL database sequences, the sequences which gave the highest scores with Bacillus pumilus MVSV3 were retrieved to construct the phylogenetic tree by neighbor-joining analysis method (Figure 1).
Degradation of BTEX by isolate MVSV3
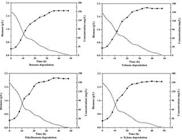
Bacillus pumilus MVSV3 degraded all the four mono aromatic compounds completely at optimized conditions. It could also degrade other substrates like phenanthrene, phenol and catechol. Isolate MVSV3 completely degraded 150 mg/L concentration of benzene, toluene, ethylbenzene and o-xylene. Growth inhibition and lesser degradation occurred at higher concentration and no degradation and no growth was seen at the concentration of 500 mg/L. The optimized conditions for maximum degradation of BTEX were found to be at pH 7.0, temperature 30 ºC and at 3% inoculum. A lag phase of 12 h was observed during the degradation studies and complete degradation of BTEX (Individually) occurred within 48 h (Figure 2A-2D).
Biodegradation and Growth of Bacillus pumilus MVSV3 on A) benzene B) toluene C) ethylbenzene and D) xylene, at pH 7, temperature 30 (C, inocula 3%, substrate concentration 150 mg/L.
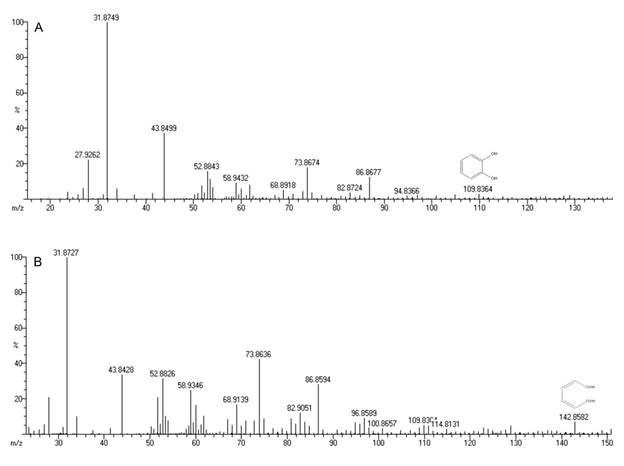
The degradation rates were estimated as the ratio of BTEX removed to the corresponding time after the lag period. Biodegradation rates of benzene, toluene, ethylbenzene and o-xylene by Bacillus pumilus MVSV3 were determined to be 1.77, 2.1, 1.85 and 1.70 mg/Lh, respectively. Benzene and o-xylene showed higher resistance to degradation when compared to toluene and ethylbenzene that degraded rapidly. These findings were similar to the study of Goudar and Strevett1818. Goudar CT, Strevett KA. Comparision of relative rates of BTEX biodegradation using respirometry. J. Ind. Microbiol. Biotechnol. 1998; 21: 11-18. who degraded BTEX using a respirometry. In this study biodegradation rate increased in the order of B<X<E<T, this finding slightly varied from You et al1919. You Y, Shim J, Cho CH, et al. Biodegradation of BTEX mixture by Psuedomonas putida YNS1 isolated from oil contaminated soil. J. Basic Microbiol. 2013; 53: 469-475. who showed degradation by Psuedomonas putida YNS1 increased in the order of X<B<E<T and stated that this difference may be due to strains that can degrade BTEX in different pathway. Maximum growth yield of the isolate MVSV3 was found to be 2.6 g/L. The initial pH (7.0) decreased during the experiment and was found to be in the range of 6.2 to 5.9 pH for the four substrates. The GC-MS analysis of biodegradation samples detected the presence of catechol (109.82 m/z) muconic acid (142.85 m/z) (Figure 3A-3B) and many other inter metabolites as listed in Table 2.
A) Mass spectrum of catechol (m/z identification - 109.82) and B) Mass spectrum of muconic acid (m/z identification - 142.85).
BTEX degradation intermetabolites
Degradation of BTEX will occur only when the microorganism has the ability to uptake these aromatic compounds as sole carbon source for growth. However, it is not easy, as benzene forms the basic structure for the other BTEX compounds. Benzene has six carbon - carbon bond and the electron density is evenly distributed above and below the ring. This is called as delocalization. For any organism to degrade BTEX, it must first convert the inert aromatic structure by simple oxidation or reduction. BTEX is initially attacked by O2 by oxygenases, which produces catechol. This may occur via either ortho or meta pathway, by C12D or by C23D, respectively. In this present study, C12D enzyme is predicted to carry out the ortho ring cleavage pathway. The suggested catabolic pathway of Bacillus pumilus MSV3 in the degradation of BTEX is shown in Figure 4. The major transformation involved in BTEX degradation is formation of catechol by ring activation followed by ring cleavage. Bacillus pumilus MVSV3 being an aerobic bacteria, the oxic environment provides O2 to oxidize and cleave the aromatic ring. This requires C12D to activate oxygen and rearrange it such that it produces catechol after rearrangement. These are called as central intermediates which get activated for ring cleavage. During Benzene degradation, the enzyme initially provides an O2 molecule; oxidation is carried out due to which benzene dihydradiol is formed, which is highly unstable. This then rearranges itself to form catechol. Carboxylation takes place in the next step, adjacent hydrogen gets cleaved and shifting of electrons occurs. Addition of O2 molecule in the 2nd position further give rise to muconic acid. Dissociation of molecules leads to formation of succinic acid and acetic acid.
Toluene degradation also begins with insertion of O2 molecule to form benzyl alcohol. Further attack by dioxygenase leads to the formation of benzoic acid which then decarboxylates to form catechol. Aromatic ring cleavage occurs to form muconic acid and succinate. Ethylbenzene can be degraded by many aerobic bacteria. Degradation starts with oxygenase reaction, either with alkyl group oxidation or with ring oxidation. Here Bacillus pumilus MVSV3 initiated the degradation by alkyl group oxidation which resulted in the formation of phenylethanol. Similar results were observed in Rhodococcus rhodochrous NRRLB-2153 by Hou et al.2020. Hou CT, Jackson MA, Bagby MO, et al. Microbial oxidation of cumene by octane grown cells. Appl Microbiol Biotechnol. 1994; 1: 178-182.. Further oxidation transformed it to acetophenone before forming the central intermetabolites catechol. It was sequentially oxidized into muconic acid along with propionate. Usually it is considered that xylene degrading bacteria are more prominent in soil, isolate MVSV3 being a soil microorganism, it efficiently degrades xylene into nontoxic end products. The metabolic route started with demethylation and addition of hydroxl group of o-xylene forming 4-methyl catechol. Carboxylation of the ring structure results in the formation of muconic acid and 1, propanol. In this study, the proposed pathway is assumed to undergo many other cycles of reactions to produce CO2 and H2O. The production of intermetabolites such as acetic acid, succinate, propionate and 1- propanol from BTEX respectively were in agreement with Martin et al.21 who had also reported that these intermetabolites are further metabolized to smaller intermediates. Overall BTEX degradation and transformation of it to catechol have been based on reactions that are catalyzed by C12D.
Purification of the enzyme
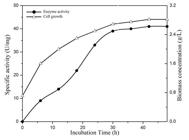
Partial purification of the enzyme was carried out using ammonium precipitation and dialysis method. The specific activity of C12D and growth of the microorganism was observed during cultivation on BTEX supplemented media for 48 h (Figure 5). A maximum biomass of 2.82 g/L and a maximum specific activity of 41 U/mg was achieved when BTEX was used as a sole carbon source.
Production of C12D of Bacillus pumilus MVSV3on MSM (supplemented with BTEX and SDS). The bacterium was incubated at 30 (C with shaking. At appropriate intervals, cell growth (Triangles) and enzyme activity (Circles) were assayed.
Effect of pH on enzyme activity
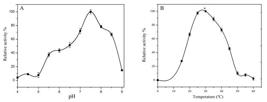
Enzyme activity was tested on different pH. The highest relative activity of C12D was obtained in the phosphate buffer at the pH 7.5 (Figure 6A). This finding was similar to the enzyme isolated by Kou and Li2222. Kou X, Li Q. The preparation and properties of catechol-1,2- dioxygenase from Psuedomonas putida. Acta Microbiologica Sinica. 1990; 30: 397-399. who reported the C12D isolated from Pseudomonas putida 84103 to exhibit highest activity at the optimum pH 7.5-8.0. Also, on the other hand, C12D isolated from Psuedomonas putida strain N6 showed highest activity at pH 7.42323. Guzik U, Gren I, Kocurek KH, et al. Catechol 1,2-dioxygenase from the new aromatic compounds - degrading Psuedomonas putida strain N6. Int. Biodeterior. Biodegrad. 2011; 65: 504-512.. The enzyme isolated from Bacillus pumilus lost about 90% of its enzyme activity at pH below 5.0 and 80% was lost at pH 9.0. This proved that the enzyme was sensitive to different pH of the solution. Silva et al.1717. Silva AS, Jacques RJS, Andreazza R, et al. Properties of catechol 1,2-dioxygenase in the cell free extract and immobilized extract of Mycobacterium fortuitum. Braz. J. Microbiol. 2013; 44: 291-297. have identified C12D from Gordonia polyisoprenivorans which showed maximum activity of the enzyme at pH 7.0 and pH 8.0 at cell free and immobilized state of the enzyme.
Effect of Temperature on enzyme activity
Experiments were carried out at different temperature to determine the optimum temperature for the enzyme activity (Figure 6B). The highest relative activity of the enzyme was obtained at 35 (C. Interestingly the enzyme lost 73% of activity at 15 (C and 28 % at 40 (C. The activity of the enzyme rapidly decreased after 50-60 (C 2424. Miguzez CB, Greer CW, Ingram JM. Purification and properties of chlorocatechol 1,2-dioxygenase from Alcaligenes denitrifcans BRI 6011. Can. J. Microbiol. 1993; 39: 1-5.. Similar findings were discussed by Tsai & Li 1616. Tsai SC, Li YK. Purification and characterization of catechol 1,2- dioxygenase from a phenol degrading Candida albicans TL3. Arch Microbiol. 2007; 187: 199-206. who stated that C12D from candida albicans TL 3 were stable when it was kept at a temperature below 40 (C.
Effect of metal ions on enzyme activity
The enzyme activity was differently affected at different metal ion concentrations. Complete inhibition of the enzyme C12D was seen in the presence of Cu2+ and Ag+ ions (Figure 7). Even the minimum concentration of 1.0 mM was found to be highly toxic to the activity of the enzyme. Highest relative activity was shown in the presence of Mn2+ions. At 1.0 mM concentration of Mn2+, Mg2+ and NH 4 +, relative activity of 95, 82 and 70 % were observed. Similar results were found by Matsumura et al.1313. Matsumura E, Ooi S, Murakami S, Takenaka S, et al. Constitutive synthesis, purification and characterization of catechol 1,2-dioxygenase from the aniline- assimilating bacterium Rhodococcus sp.AN-22. J. Biosci. Bioeng. 2004; 98: 71-76., in the aniline assimilating bacterium Rhodococcus sp. Hg2+and Ag+ showed inhibition of the C12D enzyme of Rhodococcus sp. whereas other metal ions did not show any evident effect on enzyme activity.
Molecular properties of C12D
A basic step protocol was used to partially purify C12D. The molecular weight was estimated to be approximately 35 KDa from the relative mobility on a denaturing SDS-PAGE (Figure8). Emmanuel et al.77. Emmanuel VPP, Dayananda S. Purification and characterization of catechol 1,2 dioxygenase from Acinetobacter sp. DS002 and cloning , sequencing of partial cat A gene. Indian J. Microbiol. 2011; 51: 312-318. had also discussed that mostly C12D occurs as a homodimer with the subunit molecular weight of 30-34 kDa. A subunit molecular weight of 34,500 was estimated from SDS- PAGE by Chen et al.2525. Chen YP, Lovell R. Purification and properties of catechol 1,2- dioxygenase from Rhizobium leguminosarum biovar viceae USDA 2370. Appl. Environ. Microbiol. 1990; 56: 1971-1973. from Rhizobium leguminosarum biovar viceae USDA 2370. Zymography analysis revealed three bands in lane 2 (Figure 9). This indicated the presence of isoforms of C12D1414. Piyali M, Pranab R, Copper enhanced monooxygenase activity and FT-IR spectroscopic characterization of biotransformation products in trichloroethylene degrading bacterium : Stenotrophomonas maltophilia PM102, Hindawi Publishing corporation, BioMed Research International. 2013; Article ID 723680, Available from: http://dx.doi.org/10.1155/2013/723680.
http://dx.doi.org/10.1155/2013/723680...
. Dorn and Knackmuss2626. Dorn E, knackmuss HJ. Chemical structure and biodegradability of halogenated aromatic compounds - Two catechol 1,2-dioxygenases from a 3-Chlorobenzoate- grown Pseudomonas. Biochem. J. 1978; 174: 73-84. also identified two C12D pyrocatechase I and II in Pseudomonas sp B13 when grown on 3-chlrobenzoate. Isoforms of C12D from 16 bacterial isolates obtained from soil and grown on aniline as a sole source of C and N have been reported by Murakami et al.27 in 199127. Murakami A, Nakanishi Y, Shinke R, et al. Catechol 1,2- dioxygenase isoenzymes in soil bacteria metabolizing aromatic compounds. Soil. Biol. Biochem. 1991; 23: 815-819.. A multiform enzyme has the ability to select distinctly and also play different physiological roles or regulation that allows fine-tuning of cell metabolism2828. Kalcheva EO, Faiziev MM, Shanskaya VO, et al. Regulation of two aspartokinase isoenzymes in Streptococcus bovis. Can. J. Microbiol. 1994; 40: 224-227.. Multiple forms of enzymes increase the capabilities of the organism to adapt and cope with a wide range of environmental change. It also offers a physiological advantage to the producing organism2929. Naessens M, Vandamme EJ. Multiple forms of microbial enzymes. Biotechnol. Lett. 2003; 25: 1119-1124.Hence the observation of three bands during Zymography suggested that Bacillus pumilus MVSV3 might synthesis isoforms of C12D when metabolizing BTEX. Further study of these isoforms needs to be investigated.
The presence of C12D in four Bacillus sp., isolated from soil, namely Bacillus pumilus (TN42), Bacillus megaterium (TN41), Bacillus cereus (PS1) and Bacillus simplex (PS11) were detected with the C12D enzyme activity during the degradation of phenol, toluene, biphenyl, naphthalene and other aromatic compounds3030. Dokic L, Narancic T, Runic JN, et al. Four Bacillus sp. Soil isolates capable of degrading phenol, toluene, biphenyl, naphthlene and other aromatic compounds exhibit different aromatic catabolic potentials. Arch. Biol. Sci. 2011; 63: 1057-1067..Similarly guaiacol glyceryl ether degradation by Bacillus subtilis was also carried out by C12D via ortho cleavage pathway3131. Gurujeyalakshmi G, Mahadevan A. Degradation of guaiacol glyceryl ether (GCE) by Bacillus subtilis. Appl Microbiol Biotechnol. 1987; 26: 289-293..
SDS-PAGE of C12D: Marker (L-1) size of markers are at the left, crude sample (Grown in BTEX supplemented MSM) (L-2), crude sample (Grown in nutrient broth) (L-3).
CONCLUSION
BTEX compounds mineralize under both aerobic and anaerobic conditions. Use of microorganism is considered to be the best suitable way for the clean up of BTEX contaminants from gasoline contaminated soil and water11. Jindrova E, Chocova M, Demnerova K, et al. Bacterial aerobic degradation of benzene, toluene, ethylbenzene and Xylene. Folia Microbiol. 2002; 47: 83-93.. Use of bacteria and its enzyme to degrade BTEX also helps in overcoming the difficulties and constrains with other conventional chemical methods3232. El-Naas M, Acio JA, El-Telib AE. Aerobic biodegradation of BTEX: progresses and prospects, Journal of Environmental Chemical Engineering , 2014, 2: 1104-1122.. In this present study the isolated bacterial strain Bacillus pumilus MVSV3 was evaluated for its potential to degrade 150 mg/L of BTEX (individually) in batch reaction setup. Higher concentrations of BTEX inhibited both growth and degradation of the organism and compound respectively. Since BTEX were structurally similar, much difference in growth pattern or degradation rates was not so different. The metabolic pathway involved in B, T, E and X degradation were different3333. Lee EH, Cho KS. Effect of substrate interaction on the degradation of methyl tert- butyl ether, benzene, toluene, ethylbenzene and xylene by Rhodococcus sp. J. Hazard. Mater. 2009; 167: 669-674..The GC-MS analysis identified some of the intermediates which were useful for the prediction of the pathway followed by Bacillus pumilus MVSV3. In accordance with the data obtained, benzene and o-xylene were less preferred than toluene, this was in good agreement with that of Shim et al.3434. Shim H, Yang ST. Biodegrad ation of benzene, toluene, ethylbenzene and o-xylene by a coculture of Pseudomonas putida and Pseudomonas fluorescence immobilized in a fibrous- bed bioreactor. J. Biotechnol. 1999; 67: 99-112. who degraded BTEX with a cocultutre of Pseudomonas putida and Pseudomonas flurescens. They had also achieved 0.88, 3.61, 0.29 and 0.80 mg/Lh of biodegradation rate for free cells on B(135mg/L), T(542 mg/L), E(186 mg/L) and X (212 mg/L), respectively. Whereas Alvarez and Vogel3535. Alvarez PJJ, Vogel TM, Degradation of BTEX and their aerobic metabolites by indigenous microorganisms under nitrate reducing conditions. Wat. Sci. Tech. 1995; 31 : 15-28. reported that benzene, ethylbenzene and catechol were not degraded even till four months by indigenous microorganism under nitrate reducing conditions. Similar reports were reported for BTEX at 70 mg/L, where over all percentage degradation after 20 days was in the range of 90-100 % for all except benzene that was degraded only to 7 % 3636. Mallakin A, Ward OP, Degradation of BTEX compounds in liquid media and in peat biofilters. J. Ind. Microbiol. 1996; 16: 309-318..
Catechols are formed during biodegradation of a variety of aromatic compounds by aerobic microorganisms1616. Tsai SC, Li YK. Purification and characterization of catechol 1,2- dioxygenase from a phenol degrading Candida albicans TL3. Arch Microbiol. 2007; 187: 199-206.. C12D found in Psuedomonas sp. was the first enzyme proved to be a dioxygenase that catalyses the conversion of catechol to cis-cis muconic acid3737. Hayaishi O, Hashimoto K. Pyrocatechase- A new enzyme catalyzing oxidative breakdown of pyrocatechin. J. biochem. 1950; 37: 371-374.. Under aerobic conditions BTEX compounds are initiated for ring cleavage by the introduction of hydroxyl groups that are initially catalyzed by oxygenases, hence it typically proceeds through catechol intermediates3838. Kim JM, Jeon CO. Isolation and characterization of a new benzene, toluene and ethylbenzene degrading bacterium , Acinetobacter sp.B113. Curr Microbiol. 2009; 58: 70-75.. Aromatic compounds undergo degradation through oxygenetic ring fission by C12D (EC 1.13.11.1). It catalyzes the intradiol cleavage of the aromatic ring at 1,2 (ortho) position of catechol that leads to production of cis-cis muconic acid. In this study of enzymatic activity, ortho pathway for degradation of BTEX were identified and the activity of the enzyme C12D was quantified. The environmental conditions that affect the activity of C12D from Bacillus pumilus were determined in cell free extract. According to the results obtained, pH affected the activity of the enzyme so did the temperature and metal ions. In our findings the SDS PAGE releaved a band at 35 kDa. Similarly the subunit molecular weight of C12D was estimated to be 31,558 and 34,500 Da from Rhodochrous NCIMB 13259 3939. Philip DS, Andrew AF, Charles AF. Purification and characterization of catechol 1,2-dioxygenase from Rhodococcus rhodochrous NCIMB 13259 and cloning and sequencing of its catA gene. Biochem. J. 1998; 333: 741-747. and Stenotrophomonas maltophilia strain KB24040. Guzik U, Kocurek KH, Stinik M, et al. High activity catechol 1,2-dioxygenase from Stenotrophomonas mattophilia strain KB2 as a useful tool in cis,cis- muconic acid production. Antonie van Leeuwenhoek. 2013; 103: 1297- 1307., receptively. Three bands were observed in Zymography which were in agreement with the findings of three isoenzymes of C12D that have been reported from Pseudomonas arvilla C-1, which were formed by the combination of two nonidentical subunits4141. Nakai, C, Horiike K, Kuramitsu S, et al. Three isozymes of Catechol 1,2- Dioxygenase (Pyrocatechase),((, (β and ββ, from Pseudomonas arvilla C-1. J. Biol. Chem. 1990; 265: 660-65.. Since only the presence of the isoforms were studied, further studies on their kinetic properties and cellular distributions have to be investigated to make clear the physiological significance of these isoforms. In conclusion C12D, from Bacillus pumilus MVSV3 could serve as a better tool in degrading BTEX to catechol and production of muconic acid. The wide spectrum of the enzyme over temperature and pH range and on different metal ions may be further used for efficient degradation of BTEX.
ACKNOWLEDGEMENTS
The authors gratefully acknowledge the Anna University, Chennai, India for providing the financial support (Anna Centenary Research Fellowship) to Ms. S. Sheeba Varma. We are also thankful to Dr. Hirul Islam, Pondicherry Center for Biological Sciences for supporting a part of this work.
REFERENCES
-
1Jindrova E, Chocova M, Demnerova K, et al. Bacterial aerobic degradation of benzene, toluene, ethylbenzene and Xylene. Folia Microbiol. 2002; 47: 83-93.
-
2Cao B, Nagarajan K, Loh KC. Biodegradaion of aromatic compounds: current status and opportunities for biomolecule approaches. Appl Microbiol Biotechnol. 2009; 85: 207-228.
-
3Payel S, Andrew RR, Shilpi G. Degradation of Aromatic Hydrocarbons (BTEX) by a solvent tolerant bacterial consortium. Journal of Urban and Environmental Engineering. 2013; 7: 274-279.
-
4Georg F, Matthias B, Johann H. Microbial degradation of aromatic compounds- from one strategy to four. Nature Reviews, Microbiology. 2011; 9: 803-816.
-
5Machnicka A, Suschka J. Activity of selected microorganism and mixtures in BTX biodegradation. Pol. J. Environ. Stud. 2001; 10: 341-346.
-
6Muftah HEN, Janice AA, Ayat EET. Aerobic biodegradation of BTEX: progresses and prospects. Journal of Environmental Chemical Engineering. 2012; 2: 1104-1122.
-
7Emmanuel VPP, Dayananda S. Purification and characterization of catechol 1,2 dioxygenase from Acinetobacter sp DS002 and cloning , sequencing of partial cat A gene. Indian J. Microbiol. 2011; 51: 312-318.
-
8Tancsics A, Scoboszlay S, Kriszt, B, et al. Applicability of the functional gene catechol 1,2- dioxygenase as a biomarker in the detection of BTEX degrading Rhodococcus species J. Appl. Microbiol. 2008; 105: 1026-1033.
-
9Sheeba VS, Brinda LM, Velan M. Isolation and characterization of non-adapted BTEX degrading bacterial strains from petroleum contaminated environment.. J. Pure, Appl. Microbio. 2015; 9: 3161-3169.
-
10Brinda LM, Velan, M. Biodegradation of the toxic polycyclic aromatic hydrocarbons, Phenanthrene by an indigenously isolated Alkaligenes feacalis MVMB1 strain. IPCBEE, 2nd international conference in environmental science and technology, IACSIT PRESS, Singapore; 2011.
-
11Lowry OH, Rosebrough NJ, Farr L, et al. Protein measurement with the folin phenol reagent. J.Biol.Chem. 1951; 193: 265-275.
-
12Laemmli UK. Cleavage of structural protein during the assembly of the head of bacteriophage T4. Nature. 1970; 227: 682-685.
-
13Matsumura E, Ooi S, Murakami S, Takenaka S, et al. Constitutive synthesis, purification and characterization of catechol 1,2-dioxygenase from the aniline- assimilating bacterium Rhodococcus spAN-22. J. Biosci. Bioeng. 2004; 98: 71-76.
-
14Piyali M, Pranab R, Copper enhanced monooxygenase activity and FT-IR spectroscopic characterization of biotransformation products in trichloroethylene degrading bacterium : Stenotrophomonas maltophilia PM102, Hindawi Publishing corporation, BioMed Research International. 2013; Article ID 723680, Available from: http://dx.doi.org/10.1155/2013/723680
» http://dx.doi.org/10.1155/2013/723680 -
15Parke D.Application of p-toluidine in chromogenic detection of catechol and protocatechuate, diphenolic intermediates in catabolism of aromatic compounds. Applied And Environmental Microbiology, 1992; 58;2694-2697.
-
16Tsai SC, Li YK. Purification and characterization of catechol 1,2- dioxygenase from a phenol degrading Candida albicans TL3. Arch Microbiol. 2007; 187: 199-206.
-
17Silva AS, Jacques RJS, Andreazza R, et al. Properties of catechol 1,2-dioxygenase in the cell free extract and immobilized extract of Mycobacterium fortuitum Braz. J. Microbiol. 2013; 44: 291-297.
-
18Goudar CT, Strevett KA. Comparision of relative rates of BTEX biodegradation using respirometry. J. Ind. Microbiol. Biotechnol. 1998; 21: 11-18.
-
19You Y, Shim J, Cho CH, et al. Biodegradation of BTEX mixture by Psuedomonas putida YNS1 isolated from oil contaminated soil. J. Basic Microbiol. 2013; 53: 469-475.
-
20Hou CT, Jackson MA, Bagby MO, et al. Microbial oxidation of cumene by octane grown cells. Appl Microbiol Biotechnol. 1994; 1: 178-182.
-
21Martin HVA, Sytze K, Dick BJ. Handbook on Biodegradation and Biological treatment of Hazardous Organic Compounds; Kluwer academic publishers; 1998, 2, p-189-200.
-
22Kou X, Li Q. The preparation and properties of catechol-1,2- dioxygenase from Psuedomonas putida Acta Microbiologica Sinica. 1990; 30: 397-399.
-
23Guzik U, Gren I, Kocurek KH, et al. Catechol 1,2-dioxygenase from the new aromatic compounds - degrading Psuedomonas putida strain N6. Int. Biodeterior. Biodegrad. 2011; 65: 504-512.
-
24Miguzez CB, Greer CW, Ingram JM. Purification and properties of chlorocatechol 1,2-dioxygenase from Alcaligenes denitrifcans BRI 6011. Can. J. Microbiol. 1993; 39: 1-5.
-
25Chen YP, Lovell R. Purification and properties of catechol 1,2- dioxygenase from Rhizobium leguminosarum biovar viceae USDA 2370. Appl. Environ. Microbiol. 1990; 56: 1971-1973.
-
26Dorn E, knackmuss HJ. Chemical structure and biodegradability of halogenated aromatic compounds - Two catechol 1,2-dioxygenases from a 3-Chlorobenzoate- grown Pseudomonas Biochem. J. 1978; 174: 73-84.
-
27Murakami A, Nakanishi Y, Shinke R, et al. Catechol 1,2- dioxygenase isoenzymes in soil bacteria metabolizing aromatic compounds. Soil. Biol. Biochem. 1991; 23: 815-819.
-
28Kalcheva EO, Faiziev MM, Shanskaya VO, et al. Regulation of two aspartokinase isoenzymes in Streptococcus bovis Can. J. Microbiol. 1994; 40: 224-227.
-
29Naessens M, Vandamme EJ. Multiple forms of microbial enzymes. Biotechnol. Lett. 2003; 25: 1119-1124
-
30Dokic L, Narancic T, Runic JN, et al. Four Bacillus sp. Soil isolates capable of degrading phenol, toluene, biphenyl, naphthlene and other aromatic compounds exhibit different aromatic catabolic potentials. Arch. Biol. Sci 2011; 63: 1057-1067.
-
31Gurujeyalakshmi G, Mahadevan A. Degradation of guaiacol glyceryl ether (GCE) by Bacillus subtilis Appl Microbiol Biotechnol. 1987; 26: 289-293.
-
32El-Naas M, Acio JA, El-Telib AE. Aerobic biodegradation of BTEX: progresses and prospects, Journal of Environmental Chemical Engineering , 2014, 2: 1104-1122.
-
33Lee EH, Cho KS. Effect of substrate interaction on the degradation of methyl tert- butyl ether, benzene, toluene, ethylbenzene and xylene by Rhodococcus sp J. Hazard. Mater. 2009; 167: 669-674.
-
34Shim H, Yang ST. Biodegrad ation of benzene, toluene, ethylbenzene and o-xylene by a coculture of Pseudomonas putida and Pseudomonas fluorescence immobilized in a fibrous- bed bioreactor. J. Biotechnol. 1999; 67: 99-112.
-
35Alvarez PJJ, Vogel TM, Degradation of BTEX and their aerobic metabolites by indigenous microorganisms under nitrate reducing conditions. Wat. Sci. Tech. 1995; 31 : 15-28.
-
36Mallakin A, Ward OP, Degradation of BTEX compounds in liquid media and in peat biofilters. J. Ind. Microbiol. 1996; 16: 309-318.
-
37Hayaishi O, Hashimoto K. Pyrocatechase- A new enzyme catalyzing oxidative breakdown of pyrocatechin. J. biochem. 1950; 37: 371-374.
-
38Kim JM, Jeon CO. Isolation and characterization of a new benzene, toluene and ethylbenzene degrading bacterium , Acinetobacter spB113. Curr Microbiol. 2009; 58: 70-75.
-
39Philip DS, Andrew AF, Charles AF. Purification and characterization of catechol 1,2-dioxygenase from Rhodococcus rhodochrous NCIMB 13259 and cloning and sequencing of its catA gene. Biochem. J. 1998; 333: 741-747.
-
40Guzik U, Kocurek KH, Stinik M, et al. High activity catechol 1,2-dioxygenase from Stenotrophomonas mattophilia strain KB2 as a useful tool in cis,cis- muconic acid production. Antonie van Leeuwenhoek. 2013; 103: 1297- 1307.
-
41Nakai, C, Horiike K, Kuramitsu S, et al. Three isozymes of Catechol 1,2- Dioxygenase (Pyrocatechase),((, (β and ββ, from Pseudomonas arvilla C-1. J. Biol. Chem. 1990; 265: 660-65.
Publication Dates
-
Publication in this collection
2017
History
-
Received
05 May 2016 -
Accepted
21 July 2016