ABSTRACT
The karyotypes of three armored catfish species (Loricariidae) from the Iguaçu river, southern of the Brazil, were compared using different techniques: C-banding, Ag-NOR and fluorescence in situ hybridization (FISH), which used 5S and 18S rDNAs and total Cot-1 fraction as probes. Hypostomus commersoni and Hypostomus derbyi presented 2n = 68 chromosomes, with karyotype formulae 12m+12sm+14st+30a and 12m+12sm+10st+34a, respectively; whereas Hypostomus myersi presented 2n = 74 chromosomes and 12m+16sm+12st+34a. The chromosomal localization of the Ag-NORs, 5S and 18S rDNAs differed in number of sites and chromosomal localization among the studied species. The total Cot-1 probe permitted the visualization of the repetitive DNA fraction in karyotypes of each species. Cross-hybridizations using total Cot-1 probe revealed that these species have repetitive DNAs in common. However, this does not occur in H. commersoni in relation to the other species. The apparent karyotype similarity suggests a close relationship between the sympatric H. commersoni and H. derbyi species, but the small differences detected in the examined chromosomal markers indicate evolutionary divergence due to gene flow restriction among them. Hence, the present findings indicate different composition of repetitive sequences among studied species, which permit to infer its role in chromosomal differentiation of Hypostomus.
Keywords:
Armored catfish; FISH; Iguaçu river; karyotype evolution
INTRODUCTION
The armored catfishes are characterized by a body covered by bony plates and a ventral suckermouth 11 Ferraris Jr CJ. Checklist of catfishes, recent and fossil (Osteichthyes: Siluriformes), and catalogue of Siluriform primary types. Zootaxa. 2007; 1418: 1-682.. Loricariidae is the largest family of the Siluriformes, accounting 974 valid species 22 Eschmeyer WN, Fong JD (2018). Species of Fishes by family/subfamily. http://research.calacademy.org/research/ichthyology/catalog/Species>By Family.asp [accessed 29 July 2018].
http://research.calacademy.org/research/...
, and has phylogenetic studies that shown constant systematic reallocations in its subfamilies and genus 11 Ferraris Jr CJ. Checklist of catfishes, recent and fossil (Osteichthyes: Siluriformes), and catalogue of Siluriform primary types. Zootaxa. 2007; 1418: 1-682.,33 Reis RE, Pereira EHL, Armbruster JAW. Delturinae, a new loricariid catfish subfamily (Teleostei, Siluriformes), with revisions of Delturus and Hemipsilichthys. Zool J Linn Soc. 2006; 147: 277-299.
4 Chiachio MC, Oliveira C, Montoya-Burgos JI. Molecular systematic and historical biogeography of the armored Neotropical catfishes Hypoptopomatinae and Neoplecostominae (Siluriformes: Loricariidae). Mol Phylogenet Evol. 2008; 49: 606-617.
5 Cramer CA, Liedke AMR, Bonatto LS, Reis RE. The phylogenetic relationship of the Hypoptopomatinae and Neoplecostominae (Siluriformes: Loricariidae) as inferred from mitochondrial cytochrome c oxidase I sequences. Bull Fish Biol. 2008; 9: 51-59.-66 Cramer CA, Bonatto SL, Reis R. Molecular Phylogeny of the Neoplecostominae and Hypoptopomatinae (Siluriformes: Loricariidae) using multiple genes. Mol Phylogenet Evol. 2011; 59: 43-52.. The genus Hypostomus Lacépède, 1803 contains the greatest number of species in the Hypostominae subfamily (29 genera in total) 11 Ferraris Jr CJ. Checklist of catfishes, recent and fossil (Osteichthyes: Siluriformes), and catalogue of Siluriform primary types. Zootaxa. 2007; 1418: 1-682.,77 Hollanda Carvalho P, Lima FCT, Zawadzki CH. Two new species of the Hypostomus cochliodon group (Siluriformes: Loricariidae) from the rio Negro basin in Brazil. Neotrop Ichthyol. 2010; 8: 39-48.
8 Zawadzki CH, Weber C, Pavanelli CS. A new dark-saddled species of Hypostomus (Siluriformes: Loricariidae) from the upper rio Paraguay basin. Neotrop Ichthyol. 2010; 8: 719-725.-99 Zawadzki CH, Birindelli JLO, Lima FCT. A new armored catfish species of the genus Hypostomus Lacépède, 1803 (Siluriformes: Loricariidae) from the upper rio Xingu basin, Brazil. Neotrop Ichthyol. 2012; 10: 245-253., occurring in various freshwaters ecosystems throughout tropical and subtropical regions in South America 11 Ferraris Jr CJ. Checklist of catfishes, recent and fossil (Osteichthyes: Siluriformes), and catalogue of Siluriform primary types. Zootaxa. 2007; 1418: 1-682..
Hypostomus possesses apomorphic morphological and cytogenetic characters among loricariids 1010 Traldi JB, Blanco DR, Vicari MR, Martinez JF, Lui RL, Barros AV, et al. Chromosomal diversity in Hypostomus (Siluriformes, Loricariidae) with emphasis on physical mapping of 18S and 5S rDNA sites. Genet Mol Res. 2013; 12: 463-471., such as broad diploid number (2n) ranging (64 to 84 chromosomes) and, usually, multiple sites of rDNAs 1111 Cereali SS, Pompini E, Rosa R, Zawadzki CH, Froehlich O, Giuliano-Caetano L. Karyotype description of two species of Hypostomus (Siluriformes, Loricariidae) of the Planalto da Bodoquena, Brazil. Genet Mol Res. 2008; 7: 583-591.
12 Bueno V, Venere PC, Zawadzki CH, Margarido VP. Karyotypic diversification in Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae): biogeographical and phylogenetic perspective. Rev Fish Biol Fisher. 2013; 23: 103-112.
13 Bueno V, Zawadzki CH, Margarido VP. Trends in chromosome evolution in the genus Hypostomus Lacépède, 1803 (Osteichthyes, Loricariidae): a new perspective about the correlation between diploid number and chromosomes types. Rev Fish Biol Fisher. 2011; 22: 241-250.-1414 Bueno, V, Konerat JT, Zawadzki CH, Venere PC, Blanco DR, Margarido VP. Divergent chromosome evolution in Hypostominae tribes (Siluriformes: Loricariidae): Correlation of chromosomal data with morphological and molecular phylogenies. Zebrafish. 2018. doi: 10.1089/zeb.2018.1612
https://doi.org/10.1089/zeb.2018.1612...
, while some species have chromosomal characteristics considered to be plesiomorphic in the family, like sinteny in ribosomal genes families 1010 Traldi JB, Blanco DR, Vicari MR, Martinez JF, Lui RL, Barros AV, et al. Chromosomal diversity in Hypostomus (Siluriformes, Loricariidae) with emphasis on physical mapping of 18S and 5S rDNA sites. Genet Mol Res. 2013; 12: 463-471.. In Hypostomus, karyotype formulae, heterochromatin distribution, and number and localization of rDNAs sites are considered important evolutionary chromosome markers 1010 Traldi JB, Blanco DR, Vicari MR, Martinez JF, Lui RL, Barros AV, et al. Chromosomal diversity in Hypostomus (Siluriformes, Loricariidae) with emphasis on physical mapping of 18S and 5S rDNA sites. Genet Mol Res. 2013; 12: 463-471.,1515 Rubert M, Zawadzki CH, Giuliano-Caetano L. Cytogenetic characterization of Hypostomus nigromaculatus (Siluriformes: loricariidae). Neotrop Ichthyol. 2008; 6: 93-100.
16 Pansonato-Alves JC, Serrano EA, Utsunomia R, Scacchetti PC, Oliveira C, Foresti F. Mapping five repetitive DNA classes in sympatric species of Hypostomus (Teleostei: Siluriformes: Loricariidae): analysis of chromosomal variability. Rev Fish Biol Fisher. 2013; 23: 477-489.
17 Bueno V, Venere PC, Konerat JT, Zawadzki CH, Vicari MR, Margarido VP. Physical Mapping of the 5S and 18S rDNA in ten species of Hypostomus Lacépède 1803 (Siluriformes: Loricariidae). Evolutionary tendencies in the genus. Sci World J. 2014; 2014: 1-8.-1818 Rubert M, da Rosa R, Jerep FC, Bertollo LA, Giuliano-Caetano L. Cytogenetic characterization of four species of the genus Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae) with comments on its chromosomal diversity. Comp Cytogenet. 2011; 5: 397-410.. Chromosome variability at the intra- and interspecific level are detected in comparative cytogenetic studies 1616 Pansonato-Alves JC, Serrano EA, Utsunomia R, Scacchetti PC, Oliveira C, Foresti F. Mapping five repetitive DNA classes in sympatric species of Hypostomus (Teleostei: Siluriformes: Loricariidae): analysis of chromosomal variability. Rev Fish Biol Fisher. 2013; 23: 477-489.
17 Bueno V, Venere PC, Konerat JT, Zawadzki CH, Vicari MR, Margarido VP. Physical Mapping of the 5S and 18S rDNA in ten species of Hypostomus Lacépède 1803 (Siluriformes: Loricariidae). Evolutionary tendencies in the genus. Sci World J. 2014; 2014: 1-8.-1818 Rubert M, da Rosa R, Jerep FC, Bertollo LA, Giuliano-Caetano L. Cytogenetic characterization of four species of the genus Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae) with comments on its chromosomal diversity. Comp Cytogenet. 2011; 5: 397-410.. In addition, recent cytogenetic data showed that evolutionary breakpoint regions clustered in repetitive DNA regions promoted genomic reshuffle and chromosome evolution in Loricariidae species 1919 Blanco DR, Vicari MR, Lui RL, Bertollo LAC, Traldi JB, Moreira-Filho O. The role of the Robertsonian rearrangements in the origin of the XX/XY1Y2 sex chromosome system and in the chromosomal differentiation in Harttia species (Siluriformes, Loricariidae). Rev Fish Biol Fisher. 2013; 23: 127-134.
20 Blanco DR, Vicari MR, Lui RL, Artoni RF, Almeida MC, Traldi JB, et al. Origin of the X1X1X2X2/X1X2Y sex chromosome system of Harttia punctata (Siluriformes, Loricariidae) inferred from chromosome painting and FISH with ribosomal DNA markers. Genetica. 2014; 142: 119-126.
21 Blanco DR, Vicari MR, Lui RL, Traldi JB, Bueno V, Martinez JDF, et al. Karyotype diversity and evolutionary trends in armored catfish species of the genus Harttia (Siluriformes: Loricariidae). Zebrafish. 2017; 14: 169-176.
22 Barros AV, Wolski MAV, Nogaroto V, Almeida MC, Moreira-Filho O, Vicari MR. Fragile sites, dysfunctional telomere and chromosome fusions: what is 5S rDNA role? Gene. 2017; 608: 20-27.
23 Primo CC, Glugoski L, Almeida MC, Zawadzki CH, Moreira-Filho O, Vicari MR, et al. Mechanisms of chromosomal diversification in species of Rineloricaria (Actinopterygii: Siluriformes: Loricariidae). Zebrafish. 2017; 14: 161-168.
24 Primo CC, Glugoski L, Vicari MR, Nogaroto V. Chromosome mapping and molecular characterization of the Tc1/Mariner element in Rineloricaria (Siluriformes: Loricariidae). Braz Arch Biol Technol. 2018; 61: e18170623.-2525 Glugoski L, Giuliano-Caetano L, Moreira-Filho O, Vicari MR, Nogaroto V. Co-located hAT transposable element and 5S rDNA in an interstitial telomeric sequence suggest the formation of Robertsonian fusion in armored catfish. Gene. 2018; 650: 49-54..
Iguaçu River is an important tributary of Paraná River basin in the south region of Brazil and presents high endemism of its ichthyofauna due to the isolation caused by the formation of the Iguaçu waterfalls 22 Mya ago 2626 Zawadzki CH, Renesto E, Bini LM. Genetic and morphometric analysis of three species of the genus Hypostomus Lacépède, 1803 (Osteichthyes: Loricariidae) from the Rio Iguaçu basin (Brazil). Rev Suisse Zool. 1999; 106: 91-105.. Five species of Hypostomus are described in the Iguaçu River: Hypostomus albopunctatus (Regan, 1908), Hypostomus commersoni Valenciennes, 1836, Hypostomus derbyi (Haseman, 1911), Hypostomus myersi (Gosline, 1947) and Hypostomus nigropunctatus Garavello, Britski & Zawadzki, 2012, being H. derbyi and H. nigropunctatus restricted to this basin 2727 Garavelo JC, Britski HA, Zawadzki CH. The cascudos of the genus Hypostomus Lacépède (Ostariophysi: Loricariidae) from the rio Iguaçu basin. Neotrop Ichthyol. 2012; 10: 263-283.. The aim of the present study was to perform cytogenetical analysis in three Hypostomus species (H. commersoni, H. derbyi, and H. myersi) collected in in the middle Iguaçu River region (União da Vitória, Paraná State, Brazil - the type-localization of H. derbyi and H. myersi description), in order to establish chromosomal characterization among these sympatric and syntopic armored catfishes.
MATERIALS AND METHODS
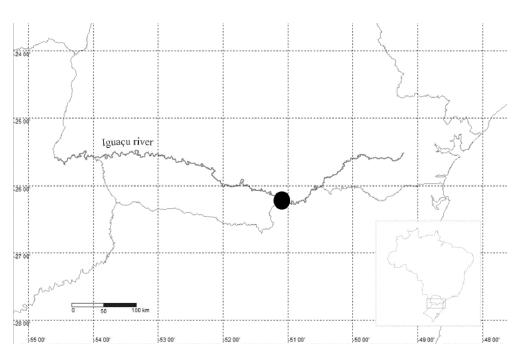
Specimens of H. commersoni (4 females and 4 males), H. derbyi (3 females and 6 males), and H. myersi (8 females and 6 males) were collected in the Iguaçu River (Fig. 1), Paraná River Basin, Brazil (26°15'1.11" S and 51°6'10.67" W). The specimens were identified and desposited in the ichthyologic collection of the Núcleo de Pesquisas em Limnologia, Ictiologia e Aquicultura (Nupélia; acronym = NUP) of Universidade Estadual de Maringá, Brazil (H. commersoni NUP 13581; H. derbyi NUP 13582; H. myersi NUP 13579). The procedures were performed according to the Ethics Committee in Animal Experimentation (Process CEUA 29/2016) of the Universidade Estadual de Ponta Grossa.
Map of Brazil, highlighting the state of Paraná and the Iguaçu river drainage. The circle refers to sampled collection region at União da Vitória.
Chromosome preparations were obtained by the air-drying method 2828 Bertollo LAC, Takahashi CS, Moreira-Filho O. Cytotaxonomic considerations on Hoplias lacerdae (Pisces, Erythrinidae). Braz J Genet. 1978; 2: 103-120., and then subjected to conventional Giemsa staining, C-banding for identification of heterochromatin regions 2929 Sumner AT. A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res. 1972; 75: 304-306.-3030 Lui RL, Blanco DR, Margarido VP, Moreira-Filho O. First description of B chromosomes in the family Auchenipteridae, Parauchenipterus galeatus (Siluriformes) of the São Francisco River basin (MG, Brazil). Mícron. 2009; 40: 552-559.and silver nitrate impregnation (AgNO3) for detection of nucleolar organizer regions (NORs) 3131 Howell WM, Black DA. Controlled silver-staining of nucleolus organizer regions with a protective colloidal developer: a 1-step method. Experientia. 1980; 36: 1014-1015.. Genomic DNA was obtained from liver using cetyltrimethylammonium bromide (CTAB) method 3232 Murray MG, Thompson WF. Rapid isolation of high molecular weight plant DNA. Nucleic Acid Res. 1980; 8: 4321-4326.. The DNA fraction containing moderately and highly repetitive DNAs was obtained by the Co t-1 DNA reassociation kinetic technique 3333 Zwick MS, Hanson RE, Mcknight TD, Nurul-Islam-Faridi M, Stelly DM. A rapid procedure for the isolation of Cot-1 DNA from plants. Genome. 1997; 40: 138-142.-3434 Wey WH, Zhao WP, Wang LJ, Chen B, Li YC, Song YC. Karyotyping of Brassica napus L. based on Cot-1 DNA banding by fluorescence in situ hybridization. J Integr Plant Biol. 2005; 47: 1479-1484., with modifications 3535 Vicari MR, Nogaroto V, Noleto RB, Cestari MM, Cioffi MB, Almeida MC, et al. Satellite DNA and chromosomes in Neotropical fishes: Methods, applications and perspectives. J Fish Biol. 2010; 76: 1094-1116.. Fragments obtained by Co t-1 ranged from 100 to 400 bp and were used as probe in cross-fluorescence in situ hybridization (cross-FISH) among the Hypostomus species. In addition, FISH experiments were performed using 18S 3636 Hatanaka T, Galetti Jr PM. Mapping of the 18S and 5S ribosomal RNA genes in the fish Prochilodus argenteus Agassiz, 1829 (Characiformes, Prochilodontidae). Genetica. 2004; 122: 239-244. and 5S rDNAs 3737 Martins C, Galetti Jr PM. Chromosomal localization of 5S DNA genes in Leporinus fish (Anostomidae, Characiformes). Chromosome Res. 1999; 7: 363-367. as probes, obtained from each of the three species studied. The probes were labeled by nick translation, using biotin-16-dUTP or digoxigenin-11-dUTP, following the manufacturer protocol (Roche Applied Science, Mannheim, Germany).
The general protocol for FISH procedure 3838 Pinkel D, Straume T, Gray JW. Cytogenetic analysis using quantitative, high-sensitivity, fluorescence hybridization. Proc Natl Acad Sci USA. 1986; 83: 2934-2938. was followed under stringency condition ~ 80 % (2.5 ng/µL probe, 50% formamide, 2x SSC - sodium saline citrate -, 10% dextran sulfate, for 18 h at 42 °C). Post-hybridization washes were performed in high stringency (50 % formamide at 42 °C for 20 min, 0.1x SSC at 60 °C for 15 min, and 4x SSC 0.05 % Tween at room temperature for 10 min). Signal detection was performed using antibodies Streptavidin Alexa Fluor 488 (Molecular Probes, Carlsbad, CA, USA) and anti-digoxigenin rhodamine (Roche Applied Science). Chromosomes were counterstained with 4´,6-diamidino-2-phenylindole (DAPI 0.2 μg/mL) in mounting medium Vectashield (Vector, Burlingame, CA, USA) and analyzed using an epifluorescence microscope. Chromosomes were classified according to Levan et al. 3939 Levan A, Fredga K, Sandberg AA. Nomenclature for centromeric position on chromosomes. Hereditas. 1964; 52: 201-220., like metacentric (m), submetacentric (sm), subtelocentric (st) and acrocentric (a) chromosomes. Fundamental number (FN) considered the m, sm and st chromosomes with two arms and acrocentric chromosomes with one arm.
RESULTS
Hypostomus commersoni
Hypostomus commersoni presented 2n = 68 chromosomes (12m+12sm+14st+30a), FN = 106 (Fig. 2A), and multiple Ag-NORs localized at the terminal region of the short arm of chromosome pair 15 and in the long arm of chromosome pair 28, both presenting size heteromorphism between the homologous (Fig 2). C-banding revealed positive heterochromatin predominantly at centromeric regions of most chromosomes, terminal bands in the short arm of the pair 13 and, one of the chromosome pairs 15 and 19, in the long arm of chromosome pairs 26 and 28, and one of the homologous of chromosome pair 4 (Fig. 2B). FISH mapping of 18S rDNA probe showed signals in the terminal regions of the short arm of chromosome pairs 13 and 15, and in terminal region of short arm in one chromosome of the pair 14, and the long arm of the pair 28. In addition, 5S rDNA sites were localized in the interstitial region of the short arm of chromosome pairs 5 and 19 and at the terminal region of the short arm of pair 9, 13 and 15 (Fig. 3A). Therefore, there was a synteny (18S/5S rDNA) at the terminal region in the short arm of chromosome pairs 13 and 15. Metaphases of H. commersoni subjected to FISH with H. commersoni total Co t-1 DNA probe showed low repetitive DNA accumulation at the terminal and interstitial regions of some chromosomes (Fig. 4A). Hypostomus commersoni metaphases hybridized with H. derbyi total Co t-1 DNA probe presented markers in many chromosomes (Fig. 4B). However, H. commersoni chromosomes did not show evident markers using H. myersi total Co t-1 DNA as probe (Fig. 4C).
Hypostomus derbyi
All specimens of H. derbyi analyzed presented 2n = 68 chromosomes (12m+12sm+10st+34a), FN = 102 (Fig. 2C) and presented single NOR at the terminal region of the short arm of pair 14 (Fig. 2). C-banding pattern showed the heterochromatin at the pericentromeric region of the short arm of the chromosome pair 2 and, at the terminal region of the short arm of pair 14 (Fig. 2D). 18S rDNA sites were localized in the chromosome pair 14, while the 5S rDNA sites were presented at the pericentromeric region of the short arm of the metacentric pair 2 (Fig. 3B). Hypostomus derbyi metaphases were subjected to hybridization with total Co t-1 probes of H. commersoni and H. derby total Co t-1 DNA probe revealing repetitive DNAs at the terminal and interstitial regions of some chromosomes in common (Fig. 4D and E, respectively). In addition, H. derbyi metaphases hybridized with H. myersi total Co t-1 probe presented signals predominantly on three chromosome pairs (Fig. 4F).
Hypostomus myersi
Specimens of H. myersi showed 2n = 74 chromosomes (12m+16sm+12st+34a) and FN = 114 (Fig. 2E). A single NOR site was detected at the terminal region of the long arm in chromosome pair 32 (acrocentric) and, frequently this site showed a size polymorphism between the homologous regions (Fig. 2). The heterochromatin was localized at centromeric regions of most chromosome pairs. We also detected heterochromatin at terminal regions of the long arm of the pairs 14, 20, 29, 30, 31, 32, 34, 36, and 37. In addition, the short arm of chromosome pair 7 and, and the interstitial region in the long arm of pairs 20, 21, and 23 had heterochromatic regions. A size heteromorphism was observed in heterochromatin block of the chromosome pair 20 in both, males and females. It was visualized an evident overlapping signal between heterochromatic sites and NORs on chromosome pair 32, that show a size heteromorphism (Fig. 2F). 18S rDNA site was localized in chromosome pair 32, while the 5S rDNA site was visualized in the m pair 2 (Fig. 3C). Metaphases of H. myersi were hybridized with H. commersoni total Co t-1 probe and not showed evident signals in the chromosomes (Fig. 4G). In addition, when metaphases of H. myersi were subjected to H. derbyi and H. myersi total Co t-1 probes, it was observed repetitive DNA accumulations at terminal and interstitial regions of many chromosomes (Fig. 4H and I, respectively).
Karyotypes of H. commersoni (A, B), H. derbyi (C, D) and H. myersi (E, F). submitted to Giemsa-stained (A, C, E) and C-banded chromosomes (B, D, F). Chromosome pairs carrying Ag-NORs are boxed. Bar = 10 µm.
Double FISH karyotypes with 5S rDNA (red markers) and 18S rDNA (green markers) probes. (A) Hypostomus commersoni, (B) H. derbyi and, (C) H. myersi. Bar = 10 µm.
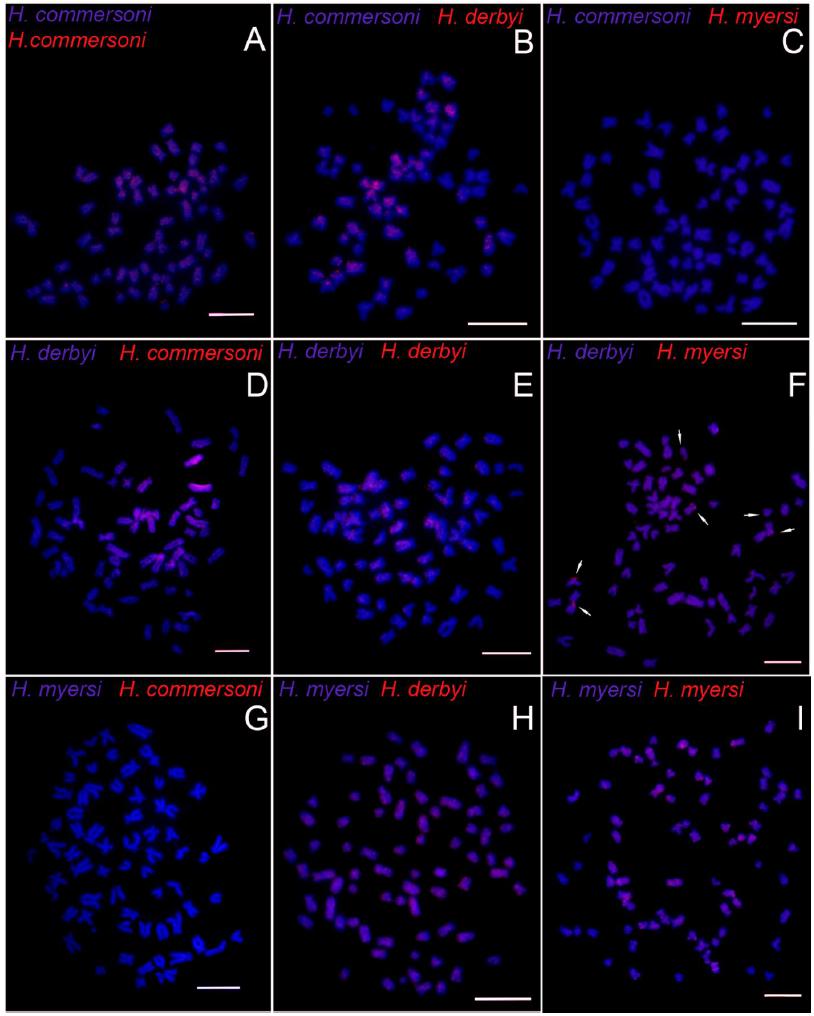
Metaphases of H. commersoni (A-C), H. derbyi (D-F) and H. myersi (G-I) after cross-FISH, using H. commersoni total Co t-1 DNA probe (A, D, G), H. derbyi total Co t-1 DNA probe (B, E, H) and, H. myersi total Co t-1 DNA probe (C, F, I). Red markers correspond to similar repeat DNA chromosomal regions among the species. Bar = 10 µm.
DISCUSSION
Several cytogenetic studies in Hypostomus revealed a great complexity and diversity of information 1010 Traldi JB, Blanco DR, Vicari MR, Martinez JF, Lui RL, Barros AV, et al. Chromosomal diversity in Hypostomus (Siluriformes, Loricariidae) with emphasis on physical mapping of 18S and 5S rDNA sites. Genet Mol Res. 2013; 12: 463-471.
11 Cereali SS, Pompini E, Rosa R, Zawadzki CH, Froehlich O, Giuliano-Caetano L. Karyotype description of two species of Hypostomus (Siluriformes, Loricariidae) of the Planalto da Bodoquena, Brazil. Genet Mol Res. 2008; 7: 583-591.
12 Bueno V, Venere PC, Zawadzki CH, Margarido VP. Karyotypic diversification in Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae): biogeographical and phylogenetic perspective. Rev Fish Biol Fisher. 2013; 23: 103-112.
13 Bueno V, Zawadzki CH, Margarido VP. Trends in chromosome evolution in the genus Hypostomus Lacépède, 1803 (Osteichthyes, Loricariidae): a new perspective about the correlation between diploid number and chromosomes types. Rev Fish Biol Fisher. 2011; 22: 241-250.
14 Bueno, V, Konerat JT, Zawadzki CH, Venere PC, Blanco DR, Margarido VP. Divergent chromosome evolution in Hypostominae tribes (Siluriformes: Loricariidae): Correlation of chromosomal data with morphological and molecular phylogenies. Zebrafish. 2018. doi: 10.1089/zeb.2018.1612
https://doi.org/10.1089/zeb.2018.1612...
15 Rubert M, Zawadzki CH, Giuliano-Caetano L. Cytogenetic characterization of Hypostomus nigromaculatus (Siluriformes: loricariidae). Neotrop Ichthyol. 2008; 6: 93-100.
16 Pansonato-Alves JC, Serrano EA, Utsunomia R, Scacchetti PC, Oliveira C, Foresti F. Mapping five repetitive DNA classes in sympatric species of Hypostomus (Teleostei: Siluriformes: Loricariidae): analysis of chromosomal variability. Rev Fish Biol Fisher. 2013; 23: 477-489.
17 Bueno V, Venere PC, Konerat JT, Zawadzki CH, Vicari MR, Margarido VP. Physical Mapping of the 5S and 18S rDNA in ten species of Hypostomus Lacépède 1803 (Siluriformes: Loricariidae). Evolutionary tendencies in the genus. Sci World J. 2014; 2014: 1-8.-1818 Rubert M, da Rosa R, Jerep FC, Bertollo LA, Giuliano-Caetano L. Cytogenetic characterization of four species of the genus Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae) with comments on its chromosomal diversity. Comp Cytogenet. 2011; 5: 397-410.,4040 Martinez ERM, Zawadzki CH, Foresti F, Oliveira C. Cytogenetic analysis of five Hypostomus species (Siluriformes, Loricariidae). Genet Mol Biol. 2011; 34: 562-568.
41 Mendes-Neto EO, Vicari MR, Artoni RF, Moreira-Filho O. Description of karyotype in Hypostomus regani (Ihering, 1905) (Teleostei, Loricariidae) from the Piumhi river in Brazil with comments on karyotype variation found in Hypostomus. Comp Cytogenet. 2011; 5: 133-142.
42 Bitencourt JA, Affonso PR, Giuliano-Caetano L, Dias AL. Identification of distinct evolutionary units in allopatric populations of Hypostomus cf. wuchereri Günther, 1864 (Siluriformes, Loricariidae): karyotypic evidence. Neotrop Ichthyol. 2011; 9: 317-324.
43 Bitencourt JA, Affonso PR, Giuliano-Caetano L, Dias AL. Heterochromatin heterogeneity in Hypostomus prope unae (Steindachner, 1878) (Siluriformes, Loricariidae) from Northeastern Brazil. Comp Cytogenet. 2011; 5: 329-344.
44 Bitencourt JA, Affonso PRAM, Giuliano-Caetano L, Carneiro PLS, Dias AL. Population divergence and peculiar karyoevolutionary trends in the loricariid fish Hypostomus aff. unae from northeastern Brazil. Genet Mol Res. 2012; 11: 933-943.
45 Endo KS, Martinez ERM, Zawadzki CH, Paiva LRS, Júlio Jr HF. Karyotype description of possible new species of the Hypostomus ancistroides complex (Teleostei: Loricariidae) and other Hypostominae. Acta Scient. 2012; 34: 181-189.-4646 Traldi JB, Vicari MR, Blanco DR, Martinez JF, Artoni RF, Moreira-Filho O. First karyotype description of Hypostomus iheringii (Regan, 1908): a case of heterochromatic polymorphism. Comp Cytogenet. 2012; 6: 115-125., showing extensive karyotype variation in this group. Cytogenetic analysis evidenced 2n = 54 chromosomes as an ancestral characteristic in Loricariidae, since it was reported in the basal genera and in the sister group Trichomycteridae 4747 Ziemniczak K, Barros AV, Rosa KO, Nogaroto V, Almeida MC, Cestari MM, et al. Comparative cytogenetics of Loricariidae (Actinopterygii: Siluriformes): emphasis in Neoplecostominae and Hypoptopomatinae. Ital J Zool. 2012; 79: 492-501.. Karyotype analyses in Hypostomus have allowed well-defined evolutionary trends, as the occurrence of high variation in the 2n, due to chromosomal rearrangements, like centric fissions 1212 Bueno V, Venere PC, Zawadzki CH, Margarido VP. Karyotypic diversification in Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae): biogeographical and phylogenetic perspective. Rev Fish Biol Fisher. 2013; 23: 103-112.
13 Bueno V, Zawadzki CH, Margarido VP. Trends in chromosome evolution in the genus Hypostomus Lacépède, 1803 (Osteichthyes, Loricariidae): a new perspective about the correlation between diploid number and chromosomes types. Rev Fish Biol Fisher. 2011; 22: 241-250.
14 Bueno, V, Konerat JT, Zawadzki CH, Venere PC, Blanco DR, Margarido VP. Divergent chromosome evolution in Hypostominae tribes (Siluriformes: Loricariidae): Correlation of chromosomal data with morphological and molecular phylogenies. Zebrafish. 2018. doi: 10.1089/zeb.2018.1612
https://doi.org/10.1089/zeb.2018.1612...
15 Rubert M, Zawadzki CH, Giuliano-Caetano L. Cytogenetic characterization of Hypostomus nigromaculatus (Siluriformes: loricariidae). Neotrop Ichthyol. 2008; 6: 93-100.
16 Pansonato-Alves JC, Serrano EA, Utsunomia R, Scacchetti PC, Oliveira C, Foresti F. Mapping five repetitive DNA classes in sympatric species of Hypostomus (Teleostei: Siluriformes: Loricariidae): analysis of chromosomal variability. Rev Fish Biol Fisher. 2013; 23: 477-489.
17 Bueno V, Venere PC, Konerat JT, Zawadzki CH, Vicari MR, Margarido VP. Physical Mapping of the 5S and 18S rDNA in ten species of Hypostomus Lacépède 1803 (Siluriformes: Loricariidae). Evolutionary tendencies in the genus. Sci World J. 2014; 2014: 1-8.
18 Rubert M, da Rosa R, Jerep FC, Bertollo LA, Giuliano-Caetano L. Cytogenetic characterization of four species of the genus Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae) with comments on its chromosomal diversity. Comp Cytogenet. 2011; 5: 397-410.
19 Blanco DR, Vicari MR, Lui RL, Bertollo LAC, Traldi JB, Moreira-Filho O. The role of the Robertsonian rearrangements in the origin of the XX/XY1Y2 sex chromosome system and in the chromosomal differentiation in Harttia species (Siluriformes, Loricariidae). Rev Fish Biol Fisher. 2013; 23: 127-134.-2020 Blanco DR, Vicari MR, Lui RL, Artoni RF, Almeida MC, Traldi JB, et al. Origin of the X1X1X2X2/X1X2Y sex chromosome system of Harttia punctata (Siluriformes, Loricariidae) inferred from chromosome painting and FISH with ribosomal DNA markers. Genetica. 2014; 142: 119-126.,4040 Martinez ERM, Zawadzki CH, Foresti F, Oliveira C. Cytogenetic analysis of five Hypostomus species (Siluriformes, Loricariidae). Genet Mol Biol. 2011; 34: 562-568.
41 Mendes-Neto EO, Vicari MR, Artoni RF, Moreira-Filho O. Description of karyotype in Hypostomus regani (Ihering, 1905) (Teleostei, Loricariidae) from the Piumhi river in Brazil with comments on karyotype variation found in Hypostomus. Comp Cytogenet. 2011; 5: 133-142.
42 Bitencourt JA, Affonso PR, Giuliano-Caetano L, Dias AL. Identification of distinct evolutionary units in allopatric populations of Hypostomus cf. wuchereri Günther, 1864 (Siluriformes, Loricariidae): karyotypic evidence. Neotrop Ichthyol. 2011; 9: 317-324.
43 Bitencourt JA, Affonso PR, Giuliano-Caetano L, Dias AL. Heterochromatin heterogeneity in Hypostomus prope unae (Steindachner, 1878) (Siluriformes, Loricariidae) from Northeastern Brazil. Comp Cytogenet. 2011; 5: 329-344.
44 Bitencourt JA, Affonso PRAM, Giuliano-Caetano L, Carneiro PLS, Dias AL. Population divergence and peculiar karyoevolutionary trends in the loricariid fish Hypostomus aff. unae from northeastern Brazil. Genet Mol Res. 2012; 11: 933-943.
45 Endo KS, Martinez ERM, Zawadzki CH, Paiva LRS, Júlio Jr HF. Karyotype description of possible new species of the Hypostomus ancistroides complex (Teleostei: Loricariidae) and other Hypostominae. Acta Scient. 2012; 34: 181-189.-4646 Traldi JB, Vicari MR, Blanco DR, Martinez JF, Artoni RF, Moreira-Filho O. First karyotype description of Hypostomus iheringii (Regan, 1908): a case of heterochromatic polymorphism. Comp Cytogenet. 2012; 6: 115-125., which could explain the increased 2n present in the H. commersoni (2n = 68 chromosomes), H. derbyi (2n = 68 chromosomes) and H. myersi (2n = 74 chromosomes).
Chromosome variability in Hypostomus may be assessed by intraspecific variation of 2n, karyotype formula, heterochromatin localization, and number and localization of NORs sites 1515 Rubert M, Zawadzki CH, Giuliano-Caetano L. Cytogenetic characterization of Hypostomus nigromaculatus (Siluriformes: loricariidae). Neotrop Ichthyol. 2008; 6: 93-100.,1717 Bueno V, Venere PC, Konerat JT, Zawadzki CH, Vicari MR, Margarido VP. Physical Mapping of the 5S and 18S rDNA in ten species of Hypostomus Lacépède 1803 (Siluriformes: Loricariidae). Evolutionary tendencies in the genus. Sci World J. 2014; 2014: 1-8.,4444 Bitencourt JA, Affonso PRAM, Giuliano-Caetano L, Carneiro PLS, Dias AL. Population divergence and peculiar karyoevolutionary trends in the loricariid fish Hypostomus aff. unae from northeastern Brazil. Genet Mol Res. 2012; 11: 933-943.. Hypostomus plecostomus (Linnaeus, 1758) was described with 2n = 54 chromosomes 4848 Muramoto JI, Ohmo S, Atkin NB. On the diploid state of the fish order Ostariophysi. Chromosoma. 1968; 24: 59-66.. However, it was considered a misidentification case, once the recent cytogenetic studies in this species has demonstrated 2n = 68 chromosomes 4949 Oliveira LC, Portela-Castro ALB, Ribeiro MO, Zawadski CH, Dutra ES, Martins-Santos IC. Karyotype structure of Hypostomus cf. plecostomus (Linnaeus, 1758) from Southem Amazon (MT): the occurrence of sex chromosomes (ZZ/ZW) and its evolutionary implications. Genet Mol Res. 2015; 14: 6625-6634.. Thus, in Hypostomus species the 2n is ranging from 64 chromosomes in Hypostomus sp. Xingu-1, Hypostomus faveolus Zawadzki, Birindelli & Lima, 2008 and Hypostomus
cochliodon Kner, 1854 to 84 chromosomes in Hypostomus perdido Zawadzki, Tencatt & Froehlich, 2014, cited as Hypostomus sp. 2 Perdido River 1111 Cereali SS, Pompini E, Rosa R, Zawadzki CH, Froehlich O, Giuliano-Caetano L. Karyotype description of two species of Hypostomus (Siluriformes, Loricariidae) of the Planalto da Bodoquena, Brazil. Genet Mol Res. 2008; 7: 583-591.
12 Bueno V, Venere PC, Zawadzki CH, Margarido VP. Karyotypic diversification in Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae): biogeographical and phylogenetic perspective. Rev Fish Biol Fisher. 2013; 23: 103-112.-1313 Bueno V, Zawadzki CH, Margarido VP. Trends in chromosome evolution in the genus Hypostomus Lacépède, 1803 (Osteichthyes, Loricariidae): a new perspective about the correlation between diploid number and chromosomes types. Rev Fish Biol Fisher. 2011; 22: 241-250.,5050 Zawadzki CH, Tencatt LFC, Froehlich O. A new unicuspid-toothed species of Hypostomus Lacépède, 1803 (Siluriformes: Loricariidae) from the rio Paraguai basin. Neotrop Ichthyol. 2014; 12: 97104.. The chromosomal data in this study showed that 2n = 68 and different karyotype formulae between H. commersoni and H. derbyi could be explained by pericentric inversions. Cytogenetic analysis in H. derbyi from the Iguaçu river in Curitiba, Paraná state, showed a different karyotype formula 5151 Maurutto FAM, Manvailer LFS, Sczepanski TS, Cestari MM, Artoni TF. Cytogenetic characterization of three allopatric species of Hypostomus Lacépède (1803) (Teleostei, Loricariidae). Caryologia. 2012; 65: 340-346. and can suggest chromosomal diversification and cryptic species for H. derbyi. In Hypostomus, the number of st/a chromosomes was postulated to be directly proportional to an increase in 2n 1313 Bueno V, Zawadzki CH, Margarido VP. Trends in chromosome evolution in the genus Hypostomus Lacépède, 1803 (Osteichthyes, Loricariidae): a new perspective about the correlation between diploid number and chromosomes types. Rev Fish Biol Fisher. 2011; 22: 241-250.. Although H.
myersi presented small numbers of st/a chromosomes and the highest 2n among the Hypostomus species analyzed in this work, the pericentric inversions occurred in the chromosomal diversification of the genus, as proposed by Bueno et al.1313 Bueno V, Zawadzki CH, Margarido VP. Trends in chromosome evolution in the genus Hypostomus Lacépède, 1803 (Osteichthyes, Loricariidae): a new perspective about the correlation between diploid number and chromosomes types. Rev Fish Biol Fisher. 2011; 22: 241-250..
In situ localization of ribosomal sites are considered important cytotaxonomic and evolutionary markers for understanding the karyotype differentiation in fishes 1010 Traldi JB, Blanco DR, Vicari MR, Martinez JF, Lui RL, Barros AV, et al. Chromosomal diversity in Hypostomus (Siluriformes, Loricariidae) with emphasis on physical mapping of 18S and 5S rDNA sites. Genet Mol Res. 2013; 12: 463-471.,2222 Barros AV, Wolski MAV, Nogaroto V, Almeida MC, Moreira-Filho O, Vicari MR. Fragile sites, dysfunctional telomere and chromosome fusions: what is 5S rDNA role? Gene. 2017; 608: 20-27.
23 Primo CC, Glugoski L, Almeida MC, Zawadzki CH, Moreira-Filho O, Vicari MR, et al. Mechanisms of chromosomal diversification in species of Rineloricaria (Actinopterygii: Siluriformes: Loricariidae). Zebrafish. 2017; 14: 161-168.
24 Primo CC, Glugoski L, Vicari MR, Nogaroto V. Chromosome mapping and molecular characterization of the Tc1/Mariner element in Rineloricaria (Siluriformes: Loricariidae). Braz Arch Biol Technol. 2018; 61: e18170623.-2525 Glugoski L, Giuliano-Caetano L, Moreira-Filho O, Vicari MR, Nogaroto V. Co-located hAT transposable element and 5S rDNA in an interstitial telomeric sequence suggest the formation of Robertsonian fusion in armored catfish. Gene. 2018; 650: 49-54.. Hypostomus commersoni, H. derbyi and H. myersi differed in localization and number of chromosome pairs bearing 18S rDNA sites. In H. derbyi and H. myersi, the presence of a single NOR indicated the maintenance of the plesiomorphic condition for Loricariidae 1717 Bueno V, Venere PC, Konerat JT, Zawadzki CH, Vicari MR, Margarido VP. Physical Mapping of the 5S and 18S rDNA in ten species of Hypostomus Lacépède 1803 (Siluriformes: Loricariidae). Evolutionary tendencies in the genus. Sci World J. 2014; 2014: 1-8.,4747 Ziemniczak K, Barros AV, Rosa KO, Nogaroto V, Almeida MC, Cestari MM, et al. Comparative cytogenetics of Loricariidae (Actinopterygii: Siluriformes): emphasis in Neoplecostominae and Hypoptopomatinae. Ital J Zool. 2012; 79: 492-501.,5252 Becker QMC, Castro RJ, Silva AM, Vizzotto PC. Cytogenetic characterization of two species of Hypostomus (Siluriformes, Loricariidae) from tributaries of the Vermelho river, Upper Paraguay river basin. Biodiversidade. 2014; 13: 2-13.-5353 Mariotto S, Centofante L, Vicari MR, Artoni RF, Moreira-Filho O. Chromosomal diversification in ribosomal DNA sites in Ancistrus Kner, 1854 (Loricariidae, Ancistrini) from three hydrographic basins of Mato Grosso, Brazil. Comp Cytogenet. 2011; 5: 289-300., while H. commersoni, with multiple NORs, showed a apomorphic condition 1010 Traldi JB, Blanco DR, Vicari MR, Martinez JF, Lui RL, Barros AV, et al. Chromosomal diversity in Hypostomus (Siluriformes, Loricariidae) with emphasis on physical mapping of 18S and 5S rDNA sites. Genet Mol Res. 2013; 12: 463-471.,1212 Bueno V, Venere PC, Zawadzki CH, Margarido VP. Karyotypic diversification in Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae): biogeographical and phylogenetic perspective. Rev Fish Biol Fisher. 2013; 23: 103-112.,1515 Rubert M, Zawadzki CH, Giuliano-Caetano L. Cytogenetic characterization of Hypostomus nigromaculatus (Siluriformes: loricariidae). Neotrop Ichthyol. 2008; 6: 93-100.-1616 Pansonato-Alves JC, Serrano EA, Utsunomia R, Scacchetti PC, Oliveira C, Foresti F. Mapping five repetitive DNA classes in sympatric species of Hypostomus (Teleostei: Siluriformes: Loricariidae): analysis of chromosomal variability. Rev Fish Biol Fisher. 2013; 23: 477-489.,1818 Rubert M, da Rosa R, Jerep FC, Bertollo LA, Giuliano-Caetano L. Cytogenetic characterization of four species of the genus Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae) with comments on its chromosomal diversity. Comp Cytogenet. 2011; 5: 397-410.,4040 Martinez ERM, Zawadzki CH, Foresti F, Oliveira C. Cytogenetic analysis of five Hypostomus species (Siluriformes, Loricariidae). Genet Mol Biol. 2011; 34: 562-568.,4545 Endo KS, Martinez ERM, Zawadzki CH, Paiva LRS, Júlio Jr HF. Karyotype description of possible new species of the Hypostomus ancistroides complex (Teleostei: Loricariidae) and other Hypostominae. Acta Scient. 2012; 34: 181-189.. A heterochromatin/rDNA association was found to be polymorphic in size between the homologous of the pairs 15 and 28 of H. commersoni and pair 32 of H. myersi, probable due an unequal crossing over and/or amplification of this region, like observed in other Hypostomus1212 Bueno V, Venere PC, Zawadzki CH, Margarido VP. Karyotypic diversification in Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae): biogeographical and phylogenetic perspective. Rev Fish Biol Fisher. 2013; 23: 103-112.,1515 Rubert M, Zawadzki CH, Giuliano-Caetano L. Cytogenetic characterization of Hypostomus nigromaculatus (Siluriformes: loricariidae). Neotrop Ichthyol. 2008; 6: 93-100.,1818 Rubert M, da Rosa R, Jerep FC, Bertollo LA, Giuliano-Caetano L. Cytogenetic characterization of four species of the genus Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae) with comments on its chromosomal diversity. Comp Cytogenet. 2011; 5: 397-410.,4141 Mendes-Neto EO, Vicari MR, Artoni RF, Moreira-Filho O. Description of karyotype in Hypostomus regani (Ihering, 1905) (Teleostei, Loricariidae) from the Piumhi river in Brazil with comments on karyotype variation found in Hypostomus. Comp Cytogenet. 2011; 5: 133-142.,4444 Bitencourt JA, Affonso PRAM, Giuliano-Caetano L, Carneiro PLS, Dias AL. Population divergence and peculiar karyoevolutionary trends in the loricariid fish Hypostomus aff. unae from northeastern Brazil. Genet Mol Res. 2012; 11: 933-943. and Loricariidae species 4747 Ziemniczak K, Barros AV, Rosa KO, Nogaroto V, Almeida MC, Cestari MM, et al. Comparative cytogenetics of Loricariidae (Actinopterygii: Siluriformes): emphasis in Neoplecostominae and Hypoptopomatinae. Ital J Zool. 2012; 79: 492-501.,5252 Becker QMC, Castro RJ, Silva AM, Vizzotto PC. Cytogenetic characterization of two species of Hypostomus (Siluriformes, Loricariidae) from tributaries of the Vermelho river, Upper Paraguay river basin. Biodiversidade. 2014; 13: 2-13.-5353 Mariotto S, Centofante L, Vicari MR, Artoni RF, Moreira-Filho O. Chromosomal diversification in ribosomal DNA sites in Ancistrus Kner, 1854 (Loricariidae, Ancistrini) from three hydrographic basins of Mato Grosso, Brazil. Comp Cytogenet. 2011; 5: 289-300..
The heterochromatin/rDNA association visualized in the analyzed species is a common feature in this genus. According to Vicari et al. 5454 Vicari MR, Artoni RF, Moreira-Filho O, Bertollo LAC. Colocalization of repetitive DNAs and silencing of major rDNA genes. A case report in the fish, Astyanax janeiroensis. Cytogenet Genome Res. 2008; 122: 67-72., the association between NORs and heterochromatin permit the dispersion of the rDNA sites throughout the genome, as visualized for H. commersoni, H. derby and H. myersi, which showed heterochromatin/rDNA associated chromosomal sites. In addition, H. commersoni and H. myersi presented additional heterochromatic blocks at centromeric, pericentromeric, and terminal regions, when compared to H. derby. Hypostomus myersi showed heterochromatic interstitial blocks in the long arms of some st and acrocentric chromosomes. The size heteromorphism of the heterochromatin block in pair 20 of the H. myersi is due to in cis accumulation of repeat sequences 4646 Traldi JB, Vicari MR, Blanco DR, Martinez JF, Artoni RF, Moreira-Filho O. First karyotype description of Hypostomus iheringii (Regan, 1908): a case of heterochromatic polymorphism. Comp Cytogenet. 2012; 6: 115-125.. Interstitial bands have also been observed in Hypostomus regani (Ihering, 1905) 1818 Rubert M, da Rosa R, Jerep FC, Bertollo LA, Giuliano-Caetano L. Cytogenetic characterization of four species of the genus Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae) with comments on its chromosomal diversity. Comp Cytogenet. 2011; 5: 397-410.,4141 Mendes-Neto EO, Vicari MR, Artoni RF, Moreira-Filho O. Description of karyotype in Hypostomus regani (Ihering, 1905) (Teleostei, Loricariidae) from the Piumhi river in Brazil with comments on karyotype variation found in Hypostomus. Comp Cytogenet. 2011; 5: 133-142., Hypostomus topavae (Godoy, 1969) 1313 Bueno V, Zawadzki CH, Margarido VP. Trends in chromosome evolution in the genus Hypostomus Lacépède, 1803 (Osteichthyes, Loricariidae): a new perspective about the correlation between diploid number and chromosomes types. Rev Fish Biol Fisher. 2011; 22: 241-250., Hypostomus unae (Steindachner, 1878) 4343 Bitencourt JA, Affonso PR, Giuliano-Caetano L, Dias AL. Heterochromatin heterogeneity in Hypostomus prope unae (Steindachner, 1878) (Siluriformes, Loricariidae) from Northeastern Brazil. Comp Cytogenet. 2011; 5: 329-344. and Hypostomus wuchereri (Günther, 1864) 4242 Bitencourt JA, Affonso PR, Giuliano-Caetano L, Dias AL. Identification of distinct evolutionary units in allopatric populations of Hypostomus cf. wuchereri Günther, 1864 (Siluriformes, Loricariidae): karyotypic evidence. Neotrop Ichthyol. 2011; 9: 317-324.. The pattern of heterochromatin bands at interstitial positions, which are equidistant in relation to the centromere and equilocal in non-homolog chromosomes of the same diploid group, may occur due to chromosome organization in nucleic interphase that would facilitate the transposition these segments5555 John B. The biology of heterochromatin. In: Verma RS, editor. Heterochromatin. Cambridge, Cambridge University Press, 1988.-5656 Schweizer D, Loidl J. A model for heterochomatin dispersion and the evolution of C-bands patterns. Chromosomes Today. 1987; 9: 61-74..
A syntenic chromosomal region of 18S and 5S rDNAs presenting a size heteromorphism between the homologous chromosomes was observed in H. commersoni. This size heteromorphism of 5S and 18S rDNA clusters may be resulted from unequal crossing-over, from the association of repetitive cis-sequences caused by mismatches of repeated DNA units 5757 Pendás AM, Morán P, Garcia-Vasquez E. Multi-chromosomal location of ribossomal RNA genes and heterochromatin association in brown trout. Chromosome Res. 1993; 1: 63-67. or, being mediated by the movement of transposable elements (TEs) associated with rDNA 5858 Symonová R, Majtánová S, Sember A, Staaks GBO, Bohlen J, Freyhof J, et al. Genome differentiation in a species pair of coregonine fishes: an extremely rapid speciation driven by stress-activated retrotransposons mediating extensive ribosomal DNA multiplications. BMC Evol Biol. 2013; 13: 1-11.. These mechanisms traditionally promote an increase or decrease of the heterochromatin block 5757 Pendás AM, Morán P, Garcia-Vasquez E. Multi-chromosomal location of ribossomal RNA genes and heterochromatin association in brown trout. Chromosome Res. 1993; 1: 63-67.. Syntenic 5S and 18S rDNAs observed on chromosome pairs 13 and 15 of H. commersoni may be considered a vestige of the ancestral condition of Loricariidae 4747 Ziemniczak K, Barros AV, Rosa KO, Nogaroto V, Almeida MC, Cestari MM, et al. Comparative cytogenetics of Loricariidae (Actinopterygii: Siluriformes): emphasis in Neoplecostominae and Hypoptopomatinae. Ital J Zool. 2012; 79: 492-501. or a recurrent feature in these species due to an intense movement of rDNA sites, especially when associated with TEs 1616 Pansonato-Alves JC, Serrano EA, Utsunomia R, Scacchetti PC, Oliveira C, Foresti F. Mapping five repetitive DNA classes in sympatric species of Hypostomus (Teleostei: Siluriformes: Loricariidae): analysis of chromosomal variability. Rev Fish Biol Fisher. 2013; 23: 477-489.,5858 Symonová R, Majtánová S, Sember A, Staaks GBO, Bohlen J, Freyhof J, et al. Genome differentiation in a species pair of coregonine fishes: an extremely rapid speciation driven by stress-activated retrotransposons mediating extensive ribosomal DNA multiplications. BMC Evol Biol. 2013; 13: 1-11.. In addition to the syntenic rDNA sites, H. commersoni presented multiple sites of 18S and 5S rDNAs, considered to be derived in Hypostomus, with origin by dispersion of theses sequences throughout the genome, which was confirmed by Traldi et al.1010 Traldi JB, Blanco DR, Vicari MR, Martinez JF, Lui RL, Barros AV, et al. Chromosomal diversity in Hypostomus (Siluriformes, Loricariidae) with emphasis on physical mapping of 18S and 5S rDNA sites. Genet Mol Res. 2013; 12: 463-471.
The repetitive fraction obtained by the Co t-1 reassociation kinetics has been considered an excellent methodology to recover repetitive DNA units, especially TEs, minisatellites, and microsatellites 6060 Zhang L, Xu C, Yu W. Cloning and characterization of chromosomal markers from a Cot-1 library of peanut (Arachis hypogaea L.). Cytogenet Genome Res. 2012; 137: 31-41.-6161 Schemberger MO, Oliveira JIN, Nogaroto V, Almeida MC, Artoni RF, Cestari MM, et al. Construction and characterization of a repetitive DNA library in Parodontidae (Actinopterygii: Characiformes): A genomic and evolutionary approach to the degeneration of the W sex chromosome. Zebrafish. 2014; 11: 518-527.. Except the satellite DNA, which is a highly in tandem repetitive sequence and the main heterochromatin marker 6262 Yunis JJ, Yasmineh WG. Heterochromatin, satellite DNA, and cell Function. Science. 1971; 174: 1200-1209.-6363 Plohl M, Luchetti A, Meštrovic N, Mantovani B. Satellite DNAs between selfishness and functionality: Structure, genomics and evolution of tandem repeats in centromeric (hetero) chromatin. Gene. 2008; 409: 72-82., the TEs, microsatellites, and minisatellites are disperse throughout the genomes, accumulating at heterochromatic regions 6161 Schemberger MO, Oliveira JIN, Nogaroto V, Almeida MC, Artoni RF, Cestari MM, et al. Construction and characterization of a repetitive DNA library in Parodontidae (Actinopterygii: Characiformes): A genomic and evolutionary approach to the degeneration of the W sex chromosome. Zebrafish. 2014; 11: 518-527.,6464 Ferreira IA, Martins C. Physical chromosome mapping of repetitive DNA sequences in Nile tilapia Oreochromis niloticus: Evidences for a differential distribution of repetitive elements in the sex chromosomes. Micron. 2008; 39: 411-418.-6565 Blanco DR, Lui RL, Vicari MR, Bertollo LAC, Moreira-Filho O. Comparative cytogenetics of giant trahiras Hoplias aimara and H. intermedius (Characiformes, Erythrinidae): chromosomal characteristics of minor and major ribosomal DNA and cross-species repetitive centromeric sequences mapping differ among morphologically identical karyotypes. Cytogenet Genome Res. 2010; 132: 71-78.. The cross-FISH with DNA Co t-1 probes of H. commersoni, H. derbyi, and H. myersi showed different distribution of repetitive DNAs among the species analyzed. We observed that H. derbyi shares sequences of repetitive DNAs with H. commersoni and H. myersi, whereas H. commersoni and H. myersi not possesses shared repeat DNA sequences. These data suggest a higher evolutionary dynamic for the repetitive fraction of the genome of the species H. commersoni and H. myersi, according to the evolutionary model combined with repetitive units of the genome 6666 Zimmer EA, Martin SL, Beverley SM, Kan YW, Wilson AC. Rapid duplication and loss of genes coding for the chains of hemoglobin. Proc Natl Acad Sci U S A. 1980; 77: 2158-2162.. Following this model, mutated copies of repetitive DNAs could spread throughout the genome by mechanisms as unequal cross-over 5959 Smith GP. Evolution of repeated DNA sequences by unequal crossing-over. Science. 1976; 191: 528-535. or hitchhiking with TEs, leading to sequence homogenization 6767 Dover GA. Molecular drive: A cohesive mode of species evolution. Nature. 1982; 299: 111-117. in the same genome. However, the effect of these mechanisms, when we compare different genomes, would be the differentiation of repetitive DNA copies, as verified for these species. Chromosome differentiation caused by the movement of repetitive sequences seems to be very important for speciation 5858 Symonová R, Majtánová S, Sember A, Staaks GBO, Bohlen J, Freyhof J, et al. Genome differentiation in a species pair of coregonine fishes: an extremely rapid speciation driven by stress-activated retrotransposons mediating extensive ribosomal DNA multiplications. BMC Evol Biol. 2013; 13: 1-11.,6868 Pucci MB, Barbosa P, Nogaroto V, Almeida MC, Artoni RF, Pansonato-Alves JC, et al. Population differentiation and speciation in the genus Characidium (Characiformes: Crenuchidae): effects of reproductive and chromosomal barriers. Biol J Linn Soc. 2014; 111: 541-553.
69 Barbosa P, Pucci MB, Nogaroto V, Almeida MC, Artoni RF, Vicari MR. Karyotype analysis of three species of Corydoras (Siluriformes: Callichthyidae) from southern Brazil: rearranged karyotypes and cytotaxonomy. Neotrop Ichthyol. 2017; 15: e160056.-7070 Pucci MB, Nogaroto V, Moreira-Filho O, Vicari MR. Dispersion of transposable elements and multigene families: Microstructural variation in Characidium (Characiformes: Crenuchidae) genomes. Genet Mol Biol. 2018; 41: 585-592.. Symonová et al.5858 Symonová R, Majtánová S, Sember A, Staaks GBO, Bohlen J, Freyhof J, et al. Genome differentiation in a species pair of coregonine fishes: an extremely rapid speciation driven by stress-activated retrotransposons mediating extensive ribosomal DNA multiplications. BMC Evol Biol. 2013; 13: 1-11. provided indirect evidence in which rDNA uses the spreading mechanism of retrotransposons, subsequently affecting recombination rates in genomes, leading to rapid genome divergence in close relationship species. Hence, these extensive genome rearrangements in Hypostomus could be associated with speciation event induced by retrotransposon genome spreading, leading to numeric and structural chromosome diversification.
Cytogenetic data obtained in the present study for the three sympatric and syntopic species of Hypostomus of the middle portion of the Iguaçu River revealed that although H. commersoni and H. derbyi presented the same diploid number, they diverged in relation to heterochromatic regions and 18S and 5S rDNA sites. Hypostomus derbyi and H. myersi, although presenting different diploid number and heterochromatin distribution, shared repetitive DNAs and 18S and 5S rDNA single sites. On the other hand, H. commersoni and H. myersi were more divergent in all cytogenetic characteristics analyzed.
ACKNOWLEDGEMENTS
The authors are grateful to the Ministério do Meio Ambiente/ Instituto Chico Mendes de Conservação da Biodiversidade (MMA/ICMBio, SISBIO nº: 15117) for authorization to collect the biological samples. This study was funded by CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior), Fundação Araucária (Fundação Araucária de Apoio ao Desenvolvimento Científico e Tecnológico do Estado do Paraná) and CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico).
REFERENCES
-
1Ferraris Jr CJ. Checklist of catfishes, recent and fossil (Osteichthyes: Siluriformes), and catalogue of Siluriform primary types. Zootaxa. 2007; 1418: 1-682.
-
2Eschmeyer WN, Fong JD (2018). Species of Fishes by family/subfamily. http://research.calacademy.org/research/ichthyology/catalog/Species>By Family.asp [accessed 29 July 2018].
» http://research.calacademy.org/research/ichthyology/catalog/Species>By Family.asp -
3Reis RE, Pereira EHL, Armbruster JAW. Delturinae, a new loricariid catfish subfamily (Teleostei, Siluriformes), with revisions of Delturus and Hemipsilichthys. Zool J Linn Soc. 2006; 147: 277-299.
-
4Chiachio MC, Oliveira C, Montoya-Burgos JI. Molecular systematic and historical biogeography of the armored Neotropical catfishes Hypoptopomatinae and Neoplecostominae (Siluriformes: Loricariidae). Mol Phylogenet Evol. 2008; 49: 606-617.
-
5Cramer CA, Liedke AMR, Bonatto LS, Reis RE. The phylogenetic relationship of the Hypoptopomatinae and Neoplecostominae (Siluriformes: Loricariidae) as inferred from mitochondrial cytochrome c oxidase I sequences. Bull Fish Biol. 2008; 9: 51-59.
-
6Cramer CA, Bonatto SL, Reis R. Molecular Phylogeny of the Neoplecostominae and Hypoptopomatinae (Siluriformes: Loricariidae) using multiple genes. Mol Phylogenet Evol. 2011; 59: 43-52.
-
7Hollanda Carvalho P, Lima FCT, Zawadzki CH. Two new species of the Hypostomus cochliodon group (Siluriformes: Loricariidae) from the rio Negro basin in Brazil. Neotrop Ichthyol. 2010; 8: 39-48.
-
8Zawadzki CH, Weber C, Pavanelli CS. A new dark-saddled species of Hypostomus (Siluriformes: Loricariidae) from the upper rio Paraguay basin. Neotrop Ichthyol. 2010; 8: 719-725.
-
9Zawadzki CH, Birindelli JLO, Lima FCT. A new armored catfish species of the genus Hypostomus Lacépède, 1803 (Siluriformes: Loricariidae) from the upper rio Xingu basin, Brazil. Neotrop Ichthyol. 2012; 10: 245-253.
-
10Traldi JB, Blanco DR, Vicari MR, Martinez JF, Lui RL, Barros AV, et al. Chromosomal diversity in Hypostomus (Siluriformes, Loricariidae) with emphasis on physical mapping of 18S and 5S rDNA sites. Genet Mol Res. 2013; 12: 463-471.
-
11Cereali SS, Pompini E, Rosa R, Zawadzki CH, Froehlich O, Giuliano-Caetano L. Karyotype description of two species of Hypostomus (Siluriformes, Loricariidae) of the Planalto da Bodoquena, Brazil. Genet Mol Res. 2008; 7: 583-591.
-
12Bueno V, Venere PC, Zawadzki CH, Margarido VP. Karyotypic diversification in Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae): biogeographical and phylogenetic perspective. Rev Fish Biol Fisher. 2013; 23: 103-112.
-
13Bueno V, Zawadzki CH, Margarido VP. Trends in chromosome evolution in the genus Hypostomus Lacépède, 1803 (Osteichthyes, Loricariidae): a new perspective about the correlation between diploid number and chromosomes types. Rev Fish Biol Fisher. 2011; 22: 241-250.
-
14Bueno, V, Konerat JT, Zawadzki CH, Venere PC, Blanco DR, Margarido VP. Divergent chromosome evolution in Hypostominae tribes (Siluriformes: Loricariidae): Correlation of chromosomal data with morphological and molecular phylogenies. Zebrafish. 2018. doi: 10.1089/zeb.2018.1612
» https://doi.org/10.1089/zeb.2018.1612 -
15Rubert M, Zawadzki CH, Giuliano-Caetano L. Cytogenetic characterization of Hypostomus nigromaculatus (Siluriformes: loricariidae). Neotrop Ichthyol. 2008; 6: 93-100.
-
16Pansonato-Alves JC, Serrano EA, Utsunomia R, Scacchetti PC, Oliveira C, Foresti F. Mapping five repetitive DNA classes in sympatric species of Hypostomus (Teleostei: Siluriformes: Loricariidae): analysis of chromosomal variability. Rev Fish Biol Fisher. 2013; 23: 477-489.
-
17Bueno V, Venere PC, Konerat JT, Zawadzki CH, Vicari MR, Margarido VP. Physical Mapping of the 5S and 18S rDNA in ten species of Hypostomus Lacépède 1803 (Siluriformes: Loricariidae). Evolutionary tendencies in the genus. Sci World J. 2014; 2014: 1-8.
-
18Rubert M, da Rosa R, Jerep FC, Bertollo LA, Giuliano-Caetano L. Cytogenetic characterization of four species of the genus Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae) with comments on its chromosomal diversity. Comp Cytogenet. 2011; 5: 397-410.
-
19Blanco DR, Vicari MR, Lui RL, Bertollo LAC, Traldi JB, Moreira-Filho O. The role of the Robertsonian rearrangements in the origin of the XX/XY1Y2 sex chromosome system and in the chromosomal differentiation in Harttia species (Siluriformes, Loricariidae). Rev Fish Biol Fisher. 2013; 23: 127-134.
-
20Blanco DR, Vicari MR, Lui RL, Artoni RF, Almeida MC, Traldi JB, et al. Origin of the X1X1X2X2/X1X2Y sex chromosome system of Harttia punctata (Siluriformes, Loricariidae) inferred from chromosome painting and FISH with ribosomal DNA markers. Genetica. 2014; 142: 119-126.
-
21Blanco DR, Vicari MR, Lui RL, Traldi JB, Bueno V, Martinez JDF, et al. Karyotype diversity and evolutionary trends in armored catfish species of the genus Harttia (Siluriformes: Loricariidae). Zebrafish. 2017; 14: 169-176.
-
22Barros AV, Wolski MAV, Nogaroto V, Almeida MC, Moreira-Filho O, Vicari MR. Fragile sites, dysfunctional telomere and chromosome fusions: what is 5S rDNA role? Gene. 2017; 608: 20-27.
-
23Primo CC, Glugoski L, Almeida MC, Zawadzki CH, Moreira-Filho O, Vicari MR, et al. Mechanisms of chromosomal diversification in species of Rineloricaria (Actinopterygii: Siluriformes: Loricariidae). Zebrafish. 2017; 14: 161-168.
-
24Primo CC, Glugoski L, Vicari MR, Nogaroto V. Chromosome mapping and molecular characterization of the Tc1/Mariner element in Rineloricaria (Siluriformes: Loricariidae). Braz Arch Biol Technol. 2018; 61: e18170623.
-
25Glugoski L, Giuliano-Caetano L, Moreira-Filho O, Vicari MR, Nogaroto V. Co-located hAT transposable element and 5S rDNA in an interstitial telomeric sequence suggest the formation of Robertsonian fusion in armored catfish. Gene. 2018; 650: 49-54.
-
26Zawadzki CH, Renesto E, Bini LM. Genetic and morphometric analysis of three species of the genus Hypostomus Lacépède, 1803 (Osteichthyes: Loricariidae) from the Rio Iguaçu basin (Brazil). Rev Suisse Zool. 1999; 106: 91-105.
-
27Garavelo JC, Britski HA, Zawadzki CH. The cascudos of the genus Hypostomus Lacépède (Ostariophysi: Loricariidae) from the rio Iguaçu basin. Neotrop Ichthyol. 2012; 10: 263-283.
-
28Bertollo LAC, Takahashi CS, Moreira-Filho O. Cytotaxonomic considerations on Hoplias lacerdae (Pisces, Erythrinidae). Braz J Genet. 1978; 2: 103-120.
-
29Sumner AT. A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res. 1972; 75: 304-306.
-
30Lui RL, Blanco DR, Margarido VP, Moreira-Filho O. First description of B chromosomes in the family Auchenipteridae, Parauchenipterus galeatus (Siluriformes) of the São Francisco River basin (MG, Brazil). Mícron. 2009; 40: 552-559.
-
31Howell WM, Black DA. Controlled silver-staining of nucleolus organizer regions with a protective colloidal developer: a 1-step method. Experientia. 1980; 36: 1014-1015.
-
32Murray MG, Thompson WF. Rapid isolation of high molecular weight plant DNA. Nucleic Acid Res. 1980; 8: 4321-4326.
-
33Zwick MS, Hanson RE, Mcknight TD, Nurul-Islam-Faridi M, Stelly DM. A rapid procedure for the isolation of Cot-1 DNA from plants. Genome. 1997; 40: 138-142.
-
34Wey WH, Zhao WP, Wang LJ, Chen B, Li YC, Song YC. Karyotyping of Brassica napus L. based on Cot-1 DNA banding by fluorescence in situ hybridization. J Integr Plant Biol. 2005; 47: 1479-1484.
-
35Vicari MR, Nogaroto V, Noleto RB, Cestari MM, Cioffi MB, Almeida MC, et al. Satellite DNA and chromosomes in Neotropical fishes: Methods, applications and perspectives. J Fish Biol. 2010; 76: 1094-1116.
-
36Hatanaka T, Galetti Jr PM. Mapping of the 18S and 5S ribosomal RNA genes in the fish Prochilodus argenteus Agassiz, 1829 (Characiformes, Prochilodontidae). Genetica. 2004; 122: 239-244.
-
37Martins C, Galetti Jr PM. Chromosomal localization of 5S DNA genes in Leporinus fish (Anostomidae, Characiformes). Chromosome Res. 1999; 7: 363-367.
-
38Pinkel D, Straume T, Gray JW. Cytogenetic analysis using quantitative, high-sensitivity, fluorescence hybridization. Proc Natl Acad Sci USA. 1986; 83: 2934-2938.
-
39Levan A, Fredga K, Sandberg AA. Nomenclature for centromeric position on chromosomes. Hereditas. 1964; 52: 201-220.
-
40Martinez ERM, Zawadzki CH, Foresti F, Oliveira C. Cytogenetic analysis of five Hypostomus species (Siluriformes, Loricariidae). Genet Mol Biol. 2011; 34: 562-568.
-
41Mendes-Neto EO, Vicari MR, Artoni RF, Moreira-Filho O. Description of karyotype in Hypostomus regani (Ihering, 1905) (Teleostei, Loricariidae) from the Piumhi river in Brazil with comments on karyotype variation found in Hypostomus. Comp Cytogenet. 2011; 5: 133-142.
-
42Bitencourt JA, Affonso PR, Giuliano-Caetano L, Dias AL. Identification of distinct evolutionary units in allopatric populations of Hypostomus cf. wuchereri Günther, 1864 (Siluriformes, Loricariidae): karyotypic evidence. Neotrop Ichthyol. 2011; 9: 317-324.
-
43Bitencourt JA, Affonso PR, Giuliano-Caetano L, Dias AL. Heterochromatin heterogeneity in Hypostomus prope unae (Steindachner, 1878) (Siluriformes, Loricariidae) from Northeastern Brazil. Comp Cytogenet. 2011; 5: 329-344.
-
44Bitencourt JA, Affonso PRAM, Giuliano-Caetano L, Carneiro PLS, Dias AL. Population divergence and peculiar karyoevolutionary trends in the loricariid fish Hypostomus aff. unae from northeastern Brazil. Genet Mol Res. 2012; 11: 933-943.
-
45Endo KS, Martinez ERM, Zawadzki CH, Paiva LRS, Júlio Jr HF. Karyotype description of possible new species of the Hypostomus ancistroides complex (Teleostei: Loricariidae) and other Hypostominae. Acta Scient. 2012; 34: 181-189.
-
46Traldi JB, Vicari MR, Blanco DR, Martinez JF, Artoni RF, Moreira-Filho O. First karyotype description of Hypostomus iheringii (Regan, 1908): a case of heterochromatic polymorphism. Comp Cytogenet. 2012; 6: 115-125.
-
47Ziemniczak K, Barros AV, Rosa KO, Nogaroto V, Almeida MC, Cestari MM, et al. Comparative cytogenetics of Loricariidae (Actinopterygii: Siluriformes): emphasis in Neoplecostominae and Hypoptopomatinae. Ital J Zool. 2012; 79: 492-501.
-
48Muramoto JI, Ohmo S, Atkin NB. On the diploid state of the fish order Ostariophysi. Chromosoma. 1968; 24: 59-66.
-
49Oliveira LC, Portela-Castro ALB, Ribeiro MO, Zawadski CH, Dutra ES, Martins-Santos IC. Karyotype structure of Hypostomus cf. plecostomus (Linnaeus, 1758) from Southem Amazon (MT): the occurrence of sex chromosomes (ZZ/ZW) and its evolutionary implications. Genet Mol Res. 2015; 14: 6625-6634.
-
50Zawadzki CH, Tencatt LFC, Froehlich O. A new unicuspid-toothed species of Hypostomus Lacépède, 1803 (Siluriformes: Loricariidae) from the rio Paraguai basin. Neotrop Ichthyol. 2014; 12: 97104.
-
51Maurutto FAM, Manvailer LFS, Sczepanski TS, Cestari MM, Artoni TF. Cytogenetic characterization of three allopatric species of Hypostomus Lacépède (1803) (Teleostei, Loricariidae). Caryologia. 2012; 65: 340-346.
-
52Becker QMC, Castro RJ, Silva AM, Vizzotto PC. Cytogenetic characterization of two species of Hypostomus (Siluriformes, Loricariidae) from tributaries of the Vermelho river, Upper Paraguay river basin. Biodiversidade. 2014; 13: 2-13.
-
53Mariotto S, Centofante L, Vicari MR, Artoni RF, Moreira-Filho O. Chromosomal diversification in ribosomal DNA sites in Ancistrus Kner, 1854 (Loricariidae, Ancistrini) from three hydrographic basins of Mato Grosso, Brazil. Comp Cytogenet. 2011; 5: 289-300.
-
54Vicari MR, Artoni RF, Moreira-Filho O, Bertollo LAC. Colocalization of repetitive DNAs and silencing of major rDNA genes. A case report in the fish, Astyanax janeiroensis. Cytogenet Genome Res. 2008; 122: 67-72.
-
55John B. The biology of heterochromatin. In: Verma RS, editor. Heterochromatin. Cambridge, Cambridge University Press, 1988.
-
56Schweizer D, Loidl J. A model for heterochomatin dispersion and the evolution of C-bands patterns. Chromosomes Today. 1987; 9: 61-74.
-
57Pendás AM, Morán P, Garcia-Vasquez E. Multi-chromosomal location of ribossomal RNA genes and heterochromatin association in brown trout. Chromosome Res. 1993; 1: 63-67.
-
58Symonová R, Majtánová S, Sember A, Staaks GBO, Bohlen J, Freyhof J, et al. Genome differentiation in a species pair of coregonine fishes: an extremely rapid speciation driven by stress-activated retrotransposons mediating extensive ribosomal DNA multiplications. BMC Evol Biol. 2013; 13: 1-11.
-
59Smith GP. Evolution of repeated DNA sequences by unequal crossing-over. Science. 1976; 191: 528-535.
-
60Zhang L, Xu C, Yu W. Cloning and characterization of chromosomal markers from a Cot-1 library of peanut (Arachis hypogaea L.). Cytogenet Genome Res. 2012; 137: 31-41.
-
61Schemberger MO, Oliveira JIN, Nogaroto V, Almeida MC, Artoni RF, Cestari MM, et al. Construction and characterization of a repetitive DNA library in Parodontidae (Actinopterygii: Characiformes): A genomic and evolutionary approach to the degeneration of the W sex chromosome. Zebrafish. 2014; 11: 518-527.
-
62Yunis JJ, Yasmineh WG. Heterochromatin, satellite DNA, and cell Function. Science. 1971; 174: 1200-1209.
-
63Plohl M, Luchetti A, Meštrovic N, Mantovani B. Satellite DNAs between selfishness and functionality: Structure, genomics and evolution of tandem repeats in centromeric (hetero) chromatin. Gene. 2008; 409: 72-82.
-
64Ferreira IA, Martins C. Physical chromosome mapping of repetitive DNA sequences in Nile tilapia Oreochromis niloticus: Evidences for a differential distribution of repetitive elements in the sex chromosomes. Micron. 2008; 39: 411-418.
-
65Blanco DR, Lui RL, Vicari MR, Bertollo LAC, Moreira-Filho O. Comparative cytogenetics of giant trahiras Hoplias aimara and H. intermedius (Characiformes, Erythrinidae): chromosomal characteristics of minor and major ribosomal DNA and cross-species repetitive centromeric sequences mapping differ among morphologically identical karyotypes. Cytogenet Genome Res. 2010; 132: 71-78.
-
66Zimmer EA, Martin SL, Beverley SM, Kan YW, Wilson AC. Rapid duplication and loss of genes coding for the chains of hemoglobin. Proc Natl Acad Sci U S A. 1980; 77: 2158-2162.
-
67Dover GA. Molecular drive: A cohesive mode of species evolution. Nature. 1982; 299: 111-117.
-
68Pucci MB, Barbosa P, Nogaroto V, Almeida MC, Artoni RF, Pansonato-Alves JC, et al. Population differentiation and speciation in the genus Characidium (Characiformes: Crenuchidae): effects of reproductive and chromosomal barriers. Biol J Linn Soc. 2014; 111: 541-553.
-
69Barbosa P, Pucci MB, Nogaroto V, Almeida MC, Artoni RF, Vicari MR. Karyotype analysis of three species of Corydoras (Siluriformes: Callichthyidae) from southern Brazil: rearranged karyotypes and cytotaxonomy. Neotrop Ichthyol. 2017; 15: e160056.
-
70Pucci MB, Nogaroto V, Moreira-Filho O, Vicari MR. Dispersion of transposable elements and multigene families: Microstructural variation in Characidium (Characiformes: Crenuchidae) genomes. Genet Mol Biol. 2018; 41: 585-592.
Publication Dates
-
Publication in this collection
2018
History
-
Received
07 Aug 2018 -
Accepted
26 Sept 2018