Abstracts
The aim of this work was to evaluate the physiological aspects of polyphosphate metabolism of Cunninghamella elegans grown in presence of copper. The growth profile was obtained by means of biomass yields, orthophosphate consumption, polyphosphate accumulation and phosphatases activities. The results revealed the influence of copper on the growth, observed by biomass yields. Orthophosphate consumption was faster in cells grown in the presence of copper. The presence of copper in the culture medium induced polyphosphate accumulation. The polyphosphate level was almost constant in the beginning of control culture growth, and could be related to the exponential growth phase. On the other hand, the copper treated cultures exhibited a significant reduction in the polyphosphate levels, indicating an active metabolization of the polymer. Acid phosphatase activity was not detected in the conditions studied, but alkaline phosphatase activity was significantly lower in the treated cultures. The results suggest the potential use of Cunninghamella elegans isolate in bioremediation and biosorption applied to environments polluted by copper.
Cunninghamella elegans; copper; polyphosphate; phosphatases
O presente trabalho teve como finalidade avaliar os aspectos fisiológicos do metabolismo do polifosfato em Cunninghamella. elegans cultivada em meio contendo cobre. O perfil de crescimento foi estabelecido em função da produção de biomassa, consumo de ortofosfato, acumulação de polifosfato e atividade das fosfatases. Os resultados obtidos indicaram a influência do metal pesado sobre o crescimento, como observado pelo rendimento da biomassa. O consumo da fonte de fósforo durante as primeiras 24 horas de crescimento na cultura tratada com cobre foi maior que na cultura controle. A acumulação de polifosfato permitiu verificar comportamentos distintos na ausência e presença do metal. A análise do polifosfato celular revelou que, nas amostras tratadas, o polímero é significativamente metabolizado durante o início do cultivo quando em presença do cobre. O isolado analisado não exibiu atividade para a fosfatase ácida. Contudo, o cultivo em presença de cobre induziu variações na expressão da enzima fosfatase alcalina. Uma diminuição significativa da atividade enzimática foi observada para a cultura tratada com o íon metálico. Os estudos demonstram o potencial de Cunninghamella elegans para biorremediação de ambientes contaminados com cobre.
Cunninghamella elegans; cobre; polifosfato; fosfatases
ENVIRONMENTAL AND SOIL MICROBIOLOGY
Copper influence on polyphosphate metabolism of Cunninghamella elegans
Influência de cobre no metabolismo de polifosfato de Cunninghamella elegans
Patrícia Mendes de SouzaI,II; Petrusk Homero MarinhoI,II; Marcos Antônio Barbosa de LimaI,II; Aline Elesbão do NascimentoII,III; Galba Maria de Campos TakakiI,II,IV* * Corresponding Author. Mailing address: Rua Nunes Machado, 42, Bloco J, UNICAP, Boa Vista. 50050-900, Recife, PE, Brasil. E-mail: takaki@unicap.br
IPrograma de Pós-Graduação em Biologia de Fungos, Universidade Federal de Pernambuco, Recife, PE, Brasil
IINúcleo de Pesquisas em Ciências Ambientais, Universidade Católica de Pernambuco, Recife, PE, Brasil
IIIDepartamento de Biologia, Universidade Católica de Pernambuco, Recife, PE, Brasil
IVDepartamento de Química - Universidade Católica de Pernambuco, Recife, PE Brasil
ABSTRACT
The aim of this work was to evaluate the physiological aspects of polyphosphate metabolism of Cunninghamella elegans grown in presence of copper. The growth profile was obtained by means of biomass yields, orthophosphate consumption, polyphosphate accumulation and phosphatases activities. The results revealed the influence of copper on the growth, observed by biomass yields. Orthophosphate consumption was faster in cells grown in the presence of copper. The presence of copper in the culture medium induced polyphosphate accumulation. The polyphosphate level was almost constant in the beginning of control culture growth, and could be related to the exponential growth phase. On the other hand, the copper treated cultures exhibited a significant reduction in the polyphosphate levels, indicating an active metabolization of the polymer. Acid phosphatase activity was not detected in the conditions studied, but alkaline phosphatase activity was significantly lower in the treated cultures. The results suggest the potential use of Cunninghamella elegans isolate in bioremediation and biosorption applied to environments polluted by copper.
Key words:Cunninghamella elegans, copper, polyphosphate, phosphatases
RESUMO
O presente trabalho teve como finalidade avaliar os aspectos fisiológicos do metabolismo do polifosfato em Cunninghamella. elegans cultivada em meio contendo cobre. O perfil de crescimento foi estabelecido em função da produção de biomassa, consumo de ortofosfato, acumulação de polifosfato e atividade das fosfatases. Os resultados obtidos indicaram a influência do metal pesado sobre o crescimento, como observado pelo rendimento da biomassa. O consumo da fonte de fósforo durante as primeiras 24 horas de crescimento na cultura tratada com cobre foi maior que na cultura controle. A acumulação de polifosfato permitiu verificar comportamentos distintos na ausência e presença do metal. A análise do polifosfato celular revelou que, nas amostras tratadas, o polímero é significativamente metabolizado durante o início do cultivo quando em presença do cobre. O isolado analisado não exibiu atividade para a fosfatase ácida. Contudo, o cultivo em presença de cobre induziu variações na expressão da enzima fosfatase alcalina. Uma diminuição significativa da atividade enzimática foi observada para a cultura tratada com o íon metálico. Os estudos demonstram o potencial de Cunninghamella elegans para biorremediação de ambientes contaminados com cobre.
Palavras-chave:Cunninghamella elegans, cobre, polifosfato, fosfatases
INTRODUCTION
Copper is an essential micronutrient for most, if not all, living organisms since it is the constituent of many metalloenzymes and proteins involved in electron transport, redox, and other important reactions. Copper is also required as a cofactor in a variety of proteins. Despite its importance, relatively little is understood about the molecular details of how organisms acquire this trace metal from the environment. Copper requirements by microorganisms are usually satisfied with very low concentrations of the metal (1 to 10 µM). In contrast, copper present at higher level in its free ionic form (Cu2+) is toxic to microbial cells. Microorganisms must possess delicate mechanisms to maintain intracellular copper within such a restricted level the neither interferes with normal metal homeostasis nor poses a risk of toxicity (9,11,12).
It is well recognized that microorganisms have a high affinity for metals and can accumulate both heavy and toxic metals by a variety of mechanisms. Microorganisms highly effective in sequestering heavy metals include bacteria, fungi, algae, and actinomycetes. These have been used to remove metals from polluted industrial and domestic effluents on a large scale. Microbial interactions with metals may have several implications for the environment. Microbes may play a large role in the biogeochemical cycling of toxic heavy metals, also in cleaning up or remedating metal-contaminated environments. There is also evidence of a correlation between tolerance to heavy metals and polyphosphate metabolism (6,13,15,25).
In microbial cells, inorganic polyphosphate (poly P) plays a significant role in increasing cell resistance to unfavorable environmental conditions and in regulating different biochemical processes. Many functions had been related to cellular poly P, such as ATP substitute and energy source, reserve for Pi, chelator of metal ions, buffer against alkali ions, regulator for stress and survival, channel for DNA entry, regulator of development and a component of the capsule cell (18,22).
The heavy metal tolerance has been related to poly P degradation and detoxification of heavy metals inside the cell. It has also been suggested that surface-associated poly P may be important in chelation of cations on the surface of cell (13,16,21).
Considering the role of Cunninghamella, a Mucoralean fungi, in xenobiotic metabolization/bioremediation, and the polyphosphate role in microbial tolerance and resistance to heavy metals, the present study was carried out to evaluate the physiological aspects of polyphosphate metabolism in Cunninghamella elegans during growth in presence of copper.
MATERIALS AND METHODS
Microorganisms and Culture Conditions
The isolate of Cunninghamella elegans (UCP 542), obtained from mangrove sediment, was kindly supplied by the Culture Collection of the Catholic University of Pernambuco, Brazil. The culture was mantained on Difco PDA (Potato Dextrose Agar), incubated at 28ºC. The SDA (Sabouraud Dextrose Agar) medium was used for spore production, during 5 days at 28ºC. A total of 107 spores/mL were collected and transferred to Erlenmeyer flasks containing 50 mL of SMM (Synthetic Medium for Mucoralean) and incubated during 120 hours, at 28ºC, at 150 Hz. A solution of 2 mM copper sulphate was prepared in distilled and deionized water, pH adjusted to 6.0 with 1N sodium hydroxide and 10% (v/v) acetic acid. All samples were prepared in five replicates.
Growth Curves
Samples collected at 12, 24, 36, 48, 72, 96 and 120 hours of culture were submitted to liophylization and mantained in a vacuum dissecator until constant weight. The final value corresponded to the arithmetic media of five replicates of each sample.
Analytical Procedure
The analytical procedures were performed using samples of culture supernatant collected at 12, 24, 36, 48, 72, 96 and 120 hours of cultivation. The final value corresponded to the arithmetic media of five replicates of each sample. Phosphate consumption was evaluated by spectrophotometric assay based on the Biosystems kit. A standard curve was produced using a potassium phosphate solution (0.5 to 5.0 g/L).
Polyphosphate Determination
The total cellular polyP was extracted and measured by the method described by Kornberg, 1995 (22). Cells were harvested at 12, 24, 36, 48, 72, 96 and 120 hours of culture, washed twice in 1.5M NaCl containing 0.01 M EDTA and 1mM NaF (wash buffer). The cell pellet was resuspended in 1.5 mL of wash buffer and sonicated on ice, for 24 minutes period with 2 minute intervals at 16 KHz. The resulting homogenate was centrifuged at 12,000 x g for 5 minutes to remove cell debris. To determine total intracellular polyP, 100 uL of concentrated HCl was added to 0.5 mL of cell extract and heated at 100ºC for 45 minutes. The liberated phosphate was measured spectrophotometrically in a Spectronic Genesys 2, at ultraviolet spectrum. The polyP concentrations were expressed in gram of phosphate per gram of biomass, as means of five replicas. An unhydrolized sample was used as a control of the background level of polyP.
Phosphatases Activity
The enzymes activities were evaluated by the method described by Joh et al., 1996 (19). Samples of 36 mg, collected at 12, 24, 36, 48, 72, 96 and 120 hours of cultivation, obtained from control and treated cultures, were washed in distilled water and suspended in 0.02M sodium acetate, pH 4.5, for the acid phosphatase detection and in 50 mM Tris-HCl buffer containing glycerol, pH 7.5, for the alkaline phosphatase detection. The samples were submitted to maceration during 5 minutes and homogenized during 2 minutes in ice bath. The extracts were centrifuged at 12,000 x g during 10 minutes at 4ºC to remove the cellular debris. The activity was determined by the use of Kit-Lab Test. The results corresponded to triplicates. The enzyme activity was expressed as International Units per Liter (UI/L), in which 1 UI/L represents the amount of enzyme that catalyses 1umol of substrate/minute/liter of sample.
RESULTS
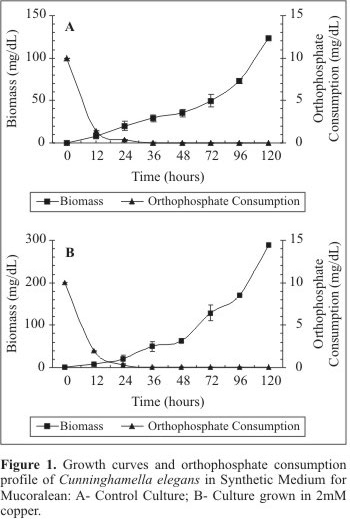
Figs. 1A and 1B exhibit the growth C. elegans in SMM culture medium in the absence and in the presence of copper, respectively. The cellular growth was evaluated by biomass production during 120 hours of cultivation. The control culture presented logarithmic growth during the experimental period. The maximum growth yield (122.9 mg/L) was obtained at 120 hours of growth (Fig. 1A). An orthophosphate consumption corresponding to 86.02% was observed during the first 12 hours of cultivation. The orthophosphate consumption was total at 48 hours of culture (Fig. 1A). When C. elegans was grown in the presence of copper the biomass yield was higher than that obtained in the control culture (288.2 mg/L). During the first 12 hours of cultivation, an orthophosphate consumption of 80.96% was detected. In the control culture, the total consumption of the phosphorus source was observed at 48 hours of culture (Fig. 1B). The results related to cellular biomass production revealed a significant difference between control and treated cultures. In the treated culture, the highest orthophosphate depletion occurred in the first 24 hours of culture and corresponded to 26.0% of the control culture.
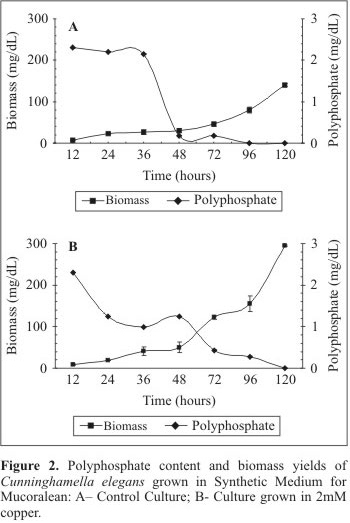
The cellular polyphosphate behavior is presented in Fig. 2. In the control culture, a progressive and slow decline in the polyphosphate content was detected during the experimental period, which could be related to increase in the biomass yield (Fig. 2A). The polyphosphate content in the mycelia was 2.3 mg/dL, 2.2 mg/dL and 2.15mg/dL, for 12 hours, 24 hours and 36 hours of growth, respectively. After 36 hours a significant decrease of polyphosphate content occurred. On the other hand, the analysis of copper treated cultures revealed the higher polyphosphate consumption during the first 36 hours of cultivation (Fig. 2B). The polyphosphate content in the mycelia of copper treated cultures was 2.3 mg/dL, 1.24 mg/dL and 0.99 mg/dL, for 12, 24 e 36 hours, respectively. The data revealed that poly P content decreased significantly in relation to the control sample. However, an increase in the polyphosphate content was observed at 48 hours of growth (1.24 mg/dL). After 48 hours, a new decline in polyphosphate content was observed. An analysis of the polyphosphate profile in the control and treated cultures revealed that in the presence of copper the polymer content decreases in a faster and continuous manner during the 120 hours of growth. On the other hand, in the control culture, a decrease of 92.1% of polyphosphate content was observed.
Fig. 3 exhibits the results for the alkaline phosphatase activity of the control and the treated samples. In the control samples, a continuous decrease in the enzyme activity was observed during the experimental period. Maximum activity was detected at 12 hours of cultivation, corresponding to 5.08 U.I/L. At 120 hours of culture, the activity was 2.22 U.I/L. The maximum enzymatic activity of C. elegans in cultures samples submitted to copper treatment was 1.28 U.I/L at 24 hours of cultivation. After 36 hours the values decreased progressively. The method described by Joh et al., 1996 (19) was inadequate to detect acid phosphatase activity in the control or treated cultures.
A comparative analysis between polyphosphate content and enzymatic activity for control and treated cultures is exhibited in Fig. 4. A progressive decrease of polyphosphate content and enzymatic activity during the experimental period was observed. A decrease of 56.29% in the alkaline phosphatase activity was observed for control cultures during the 120 hours of growth. The correlation between control and treated cultures revealed a decrease of 75.2% in the alkaline phosphatase activity in the first 12 hours of cultivation. On the other hand, the treated culture showed a decrease of 35.71% in the enzymatic activity during the experimental growth.
DISCUSSION
Microbial interactions with metals may have several implications for the environment and health. Microorganisms may play a large role in the biogeochemical cycling of toxic heavy metals, also in cleaning up or remediation of metal-contaminated environments. Other implications are not as beneficial, as the presence of metal tolerance mechanisms, which may contribute to the increase of natural resistance. It is very important to remember that any disturbance on the environment can affect humans, the environment it self and the microbial community on which other living cells depend (25,26,30).
The data presented in this work reveal the physiological aspects of the growth of C. elegans in presence of copper. The addition of 2mM copper to the growth medium has been shown to have a significant effect on the growth of C. elegans. As determined by biomass yields, the growth in control media was slower than in the presence of copper, when an increase of 57.3%, in the biomass production was observed.
Most results reported in the literature demonstate the toxic effects of heavy metals or the appearance of microbial resistance to these heavy metals. The intake and subsequent efflux of heavy metal ions in microorganisms usually include a redox reaction involving the metal, which some organisms can even use for energy and growth. This has an important implication on microbial tolerance to heavy metals because the solubility and toxicity of the metal depend on its oxidation state (14,17,28,29).
Microorganisms require metals in trace quantities for metabolism and growth, but higher concentrations can be toxic. The toxicity of metals is due to their ability to denature proteins. The blocking of functional groups, displacing an essential metal, or the modification of the active conformation of the molecule can cause this effect (11,12).
The copper inhibitory concentrations for bacteria are 1 mM, like Co2+ and Ni2+. Copper toxicity is based on the production of hydroperoxide radicals and on interaction with the cell membrane (5,6,10). Thus, the intracellular concentration of heavy metal cations, especially copper has to be tightly controlled. Copper metabolism was already studied in E. coli, some Pseudomonas-related species, Enterococus hirae and S. cerevisiae, bringing some light on copper metabolism (1,2,5,6,7,8,27,30).
The present work indicates that C. elegans is able to grown in copper containing medium and that the metal has a stimulatory effect on biomass prodution.
In fungi, metabolic changes associated to pH and composition of the culture medium, physical environmental aspects and growth phases are cited. Indeed, carbon, nitrogen and phosphorus sources are essential for the growth of microorganisms. The orthophosphate added to the culture media could regulate the cellular growth cycle, suggesting its utilization for molecular synthesis of cellular compounds (4,17).
In this work, it was shown that cultures of C. elegans submitted to copper treatment exhibited a faster orthophosphate remotion of the culture medium in the beginning of growth when compared to control cultures without the metal.
Some studies on relationship between medium components and culture development in presence of heavy metals, demonstrated that microorganisms exhibited a reduced biomass prodution even when the metal concentration was low (10,28).
The occurrence of polyphosphate has been reported for various eukaryote and prokaryote organisms. Some microrganisms can accumulate phosphate as poly P (21,22). Some studies revealed that the subsequent exposure to stress conditions promotes the poly P degradation. This mechanism has been coupled to bio-precipitation of heavy metals as cell-bound metal phosphates (13,21). However, the aspects of polyphosphate metabolism in fungi as a response to a metal presence in cultive media have not been reported yet.
In this report, the correlation between polyphosphate content in control and treated cells revealed that cultures submitted to copper treatment presented lower polyphosphate content in the initial stages of growth. In presence of copper, 57% of the polyP content was, possibly, degraded, if compared to control samples.
Some studies pointed out the role of polyP in metal precipitation. In prokariotic cells, Poly P biosynthesis and degradation, determine the appearance of metal resistance. In contrast, other results revealed that only intracellular polyP is related to this phenomenum (20,21).
The results obtained in this study indicate the association between polyP and biomass yield in cultures submitted to copper treatment. The data suggest that the polymer utilization for cellular growth increases in response to metal. Different enzymes are related to phosphate and polyP metabolism. Experimental models determine that phosphatases, exopyphosphatases, endopolyphosphatases and polyphosphate kinases are key enzymes, pointing out that these enzymes could be induced as a response to the phosphate and nitrogen concentration in the medium, its pH, and the growth phase (22,24).
The acid and alkaline phosphatase activities in response to copper presence demonstrated that C. elegans did not present acid phosphatase activity. On the other hand, alkaline phosphatase activity was inhibited by the presence of the metal in the culture medium. Compared to control cultures, a reduction of 75% in the enzyme activity was determined for treated cultures. Results in the literature for cadmium have shown that the activity of many enzymes associated to oxidative phosphorylation, photosynthesis, cell membrane permeability and integrity was impaired by the metal (29).
The present work revealed the influence of copper on the alkaline phosphatase activity in C. elegans. However, for survival in environments containing high concentrations of available metals, mechanisms to counter the inherent toxicity of the metal ions are required. The important role of fungi in biogeochemical cycles makes them candidates for studies of metal-microbe interactions. Fungal strains belonging to the class of Zygomycetes are of particular importance due to the presence in their cell walls of polymers such as chitin, chitosan, and glucan, which are known to be efficient metal ion biosorbents (3,13).
A better understanding of the factors responsible for metal resistance may help in the application of fungal biomass for the treatment of metal-contaminated water, and in enrichment or recycling of valuable metals.
ACKNOWLEDGEMENTS
The authors are greateful to PRONEX, PADCT, CNPq and UNICAP for the financial support.
Submitted: February 18, 2004; Returned to authors for corrections: April 25, 2005; Approved: November 22, 2005
- 1. Brown, N.L.; Barrett, S.R.; Camakaris, J.; Rouch, D.A.; Lee, B.T. Molecular genetics and transport analysis of the copper-resistance determinant (pco) from Escherichia coli plasmid pRJ1004. Mol. Microbiol, 17, 1153-1166, 1995.
- 2. Brown, N.L.; Rouch, D.A.; Lee, B.T. Copper resistance determinants in bacteria. Plasmid, 27, 41-51, 1992.
- 3. Campos-Takaki, G.M. Presence of inorganic polyphosphste in Mucorales fungi. Ann. Pub. Res. Cent. Path Fungi Microb. Toxic., Chiba University, Japão, 2000.
- 4. Carlile, M.J; Watkinson, S.C. The Fungi, Tokyo, Academic press., 1996.
- 5. Cervantes, C.; Gutierrez-Corona, F. Cooper resistance mechanisms in bacteria and fungi. FEMS Microbiol. Rev., 14, 121-138, 1994.
- 6. Cooksey, D.A. Copper uptake and resistance in bacteria. Mol. Microbiol., 7, 1-5, 1993.
- 7. Cooksey, D.A. Molecular mechanisms for copper resistance and accumulation in bacteria. FEMS Microbiol. Rev., 14, 381-386, 1994.
- 8. Dancis, A.; Haile, D.; Yuan, D.S.; Klausner, R.D. The Saccharomyces. cerevisiae copper transport protein (CTR1p) biochemical characterization, regulation by copper, and physiologic role in copper uptake. J. Biol. Biochem., 269, 25660-25667, 1994.
- 9. Dancis, A.; Yuan, D.S.; Haile, D.; Askwith, C.; Eide, D. Molecular characterization of copper transport protein in Saccharomyces. cerevisiae an unexpected role for copper in iron transport. Cell., 76, 393-402, 1994.
- 10. Donmez, G.; Aksu, Z. Bioaccumulation of Cooper (II) and Nickel (II) by the non-adapted and adapted growing Candida sp. Wat. Res., 35(6), 1425-1434, 2001.
- 11. Ford, T.; Mitchell, R. Microbial Transport of toxic metals, In: R. Mitchell (Ed.) Environmental Microbiology, Wiley-Liss, Inc., First ed., 83-101, 1992.
- 12. Ford, T.; Maki, J.; Mitchell, R. Metal-microbe interaction, In: Gaylarde, C.C. and H. A. Videla (Eds.) Bioextraction and Biodeterioration of metals, Cambridge University Press, First ed., 1-23, 1995.
- 13. Gadd, G. M. Interaction of fungi with toxic metals, New Phytology, 124, 25-60, 1993.
- 14. Gardea-Torresdey, J.L.; Cano-Aguilera; Webb, R.; Tiemann, K.J.; Gutierrez-Corona. Enhanced copper Adsorption and Morphological Alteration of cells of copper-stressed Mucor rouxii Environ. Toxicol. Chem., 16, 435-441, 1997.
- 15. Gomes, N.C.M.; Mendonça-Hagler, L.C.S.; Savaidis, I. Metal Biorremediation by Microorganisms. Rev. Microbiol., 29, 85-92, 1998.
- 16. Gonzalez, H.; Jensen, T.E. Nickel sequestering by polyphosphate bodies in Staphylococcus aureus. Microbios, 93, 179-185, 1998.
- 17. Griffin, D.H. Fungal Physiology. John Willey & Sons Incorporation Publishers., 458, 1994.
- 18. Harold, F.M. Inorganic Polyphosphate in biology: structure metabolism and function. Bacteriol. Rev., Dec., 772-794, 1966.
- 19. Joh, T.; Malick, D.H.; Yazaki, J.; Hayakawa, T. Purification and characterization of secreted phosphatase under phosphate - deficient condition in Pholiota nameko Mycoscience, 37, 65-70, 1996.
- 20. Keasling, J.D.; Hupf, G.A. Genetic Manipulation of Polyphosphate Metabolism affects Cadmium Tolerance in Escherichia coli Appl. Environ. Microbiol., 743-746, 1996.
- 21. Keasling, L.D.; Van Dien, S.J.; Trelstad, P.; Renninger, N.; Mcmahon, K. Application of polyphosphate metabolism to environmental and biotechnological problems. Biochemistry, Moscou, Mar., 65(3), 324-331, 1999.
- 22. Kornberg, A. Inorganic polyphosphate: toward marking a forgotten polymer unforgettable. J. Bacteriol., 177, 491-496, 1995.
- 23. Mcgrath, J.W.; Quinn, J.P. Intracellular Accumulation of Polyphosphate by the Candida humicola G-1 in response to acid pH. Appl. Environ. Microbiol., 66(9), 4068-4073, 2000.
- 24. Nesmeyanova, M.A. Review: Polyphosphates and Enzymes of Polyphosphate Metabolism in Escherichia coli. Skryabin Institute of Biochemistry and Physiology of Microorganisms, Russia Academy of Science, 1999.
- 25. Nies, D.H.; Silver, S. Ion efflux system involved in bacterial metal resistances. J. Ind. Microbiol., 14, 186-199, 1995.
- 26. Nies, D.H. Microbial heavy-metal resistance. Appl. Environ. Microbiol., 51, 730-750, 1999.
- 27. Rouch, D.A.; Brown, N.L. Copper-inducible transcriptional regulation at two promoters in the Escherichia coli copper resistance determination pco. Microbiology, 143, 1997.
- 28. Sing, C.; Yu, J. Cooper adsorption and removal from water by living mycelium of white-rot fungus Phanerochaete chrysosporium. Elsevier Sci., 9, 2746-2752, 1998.
- 29. Tobin, J.M.; White, C.; Gadd, G.M. Metal Accumulation by Fungi: Aplications in Environmental Biotechnology. J. Ind. Microbiol., 13, 126-130, 1994.
- 30. Trevors, J.T. Copper resistance in bacteria. Microbiol. Sci., 4, 29-31, 1987.
Publication Dates
-
Publication in this collection
10 May 2006 -
Date of issue
Dec 2005
History
-
Accepted
22 Nov 2005 -
Received
18 Feb 2004 -
Reviewed
25 Apr 2005