Abstracts
This study was undertaken to estimate the antimicrobial resistance of Salmonella isolated from raw chicken. From November 2003 to April 2004 a total of 120 chicken carcasses were collected from 36 randomly selected sale points (supermarkets, traditional market, poultry slaughter house, flocks) in the urban and periurban zones of Dakar, Sénégal, and examined for the presence of Salmonella. Salmonella was isolated from 75 (62.5%) of the examined samples. Out of the 90 Salmonella isolates obtained, twenty one serotypes were identified, from which the most prevalent were S. Kentuchy 30%, S. Muenster (13.3%), S. Brancaster (8.8%), S. Enteritidis and S. Hadar (6.6%). All Salmonella isolates were tested for their susceptibility to 16 selected antimicrobial agents by the agar diffusion method. Seventy one (78.9%) isolates were resistant to one or more antimicrobials. Out of 71 resistant Salmonella isolates, 33 (46.5%) showed multiple resistance to five or more different antimicrobials. Resistance to ampicillin, trimethoprim, trimethoprim-sulphamethoxazole, tetracyclin and sulphonamides was the most frequent. We found 36 different patterns of multiresistant strains. The high level of antibiotic resistance of foodborne Salmonella isolates in the present study is an indication of indiscriminate and continuous use of subtherapeutic doses of antibiotics in animals. Furthermore, the results showed the possible significance of chicken meat as a source of multiple antimicrobial-resistant Salmonella for human infections and suggest the need for detailed epidemiological study.
Salmonella sp; poultry; antimicrobial resistance
O presente estudo foi realizado com o propósito de aviliar a resistência aos antibióticos dos sorovares da Salmonella isolados da carne de frango crua. De novembro de 2003 até abril de 2004, um total de 120 carcaças de frango foram compradas em 13 pontos de venda e 23 centros de criação de frangos. As Salmonella foram isoladas a partir de 75 (62,2%) carcaças analisadas. Vinte e um (21) sorotipos diferentes foram identificados, sendo os mais freqüentes S. Kentucky (30%), S. Muenster (13,3%), S. Brancaster (8,8%), S. Enteritidis e S. Hadar (6,6%). Todos os sorovares de salmonela foram examinados a fim de determinar a resistência à 16 antibióticos. Setenta e quatro (79,6%) foram resistentes à um antibiotico ou mais, das quais 33 (45,6%) mostraram resistência múltipla a cinco antibióticos (ampicilina, trimetoprim, trimetoprim-sulfametoxazol, tetraciclina e sulfonamidas). Foram encontrados 36 perfis diferentes de resistência múltipla. O nível elevado de resistência dos isolados de Salmonella encontrados na carne do presente estudo, é um indicador do uso indevido e contínuo de doses subterapêuticas de antibióticos nos animais. Por outro lado, os resultados do estudo demonstram a importância da carne de frango como fonte potencial de sorovares de Salmonella multiresistentes transmissíveis ao homem e sugerem um estudo epidemiólogico detalhado.
Salmonella sp; frangos; resistência à antimicrobianos
FOOD MICROBIOLOGY
Antimicrobial resistance of Salmonella isolated from poultry carcasses in Dakar (Senegal)
Resistência antimicrobiana de Salmonella isolada de carcaças de frango em Dakar, Sénégal
Rianatou Bada-AlambedjiI,* * Corresponding Author. Mailing address: Service de Microbiologie-Immunologie-Pathologie Infectieuse, Ecole Inter-Etats des Sciences et Mèdecine Vètèrinaires, BP 5077, Dakar, SÉNÉGAL. E-mail: riana@refer.sn ; Aïssatou FofanaI; Malang SeydiII; Ayayi Justin AkakpoI
IService de Microbiologie-Immunologie-Pathologie Infectieuse, Ecole Inter-Etats des Sciences et Médecine Vétérinaires, Dakar, Sénégal
IIService d'Hygiène et Industries des Denrées Alimentaires d'Origine Animale, Ecole Inter-Etats des Sciences et Médecine Vétérinaires, Dakar, Sénégal
ABSTRACT
This study was undertaken to estimate the antimicrobial resistance of Salmonella isolated from raw chicken. From November 2003 to April 2004 a total of 120 chicken carcasses were collected from 36 randomly selected sale points (supermarkets, traditional market, poultry slaughter house, flocks) in the urban and periurban zones of Dakar, Sénégal, and examined for the presence of Salmonella. Salmonella was isolated from 75 (62.5%) of the examined samples. Out of the 90 Salmonella isolates obtained, twenty one serotypes were identified, from which the most prevalent were S. Kentuchy 30%, S. Muenster (13.3%), S. Brancaster (8.8%), S. Enteritidis and S. Hadar (6.6%). All Salmonella isolates were tested for their susceptibility to 16 selected antimicrobial agents by the agar diffusion method. Seventy one (78.9%) isolates were resistant to one or more antimicrobials. Out of 71 resistant Salmonella isolates, 33 (46.5%) showed multiple resistance to five or more different antimicrobials. Resistance to ampicillin, trimethoprim, trimethoprim-sulphamethoxazole, tetracyclin and sulphonamides was the most frequent. We found 36 different patterns of multiresistant strains. The high level of antibiotic resistance of foodborne Salmonella isolates in the present study is an indication of indiscriminate and continuous use of subtherapeutic doses of antibiotics in animals. Furthermore, the results showed the possible significance of chicken meat as a source of multiple antimicrobial-resistant Salmonella for human infections and suggest the need for detailed epidemiological study.
Key words:Salmonella sp, poultry, antimicrobial resistance
RESUMO
O presente estudo foi realizado com o propósito de aviliar a resistência aos antibióticos dos sorovares da Salmonella isolados da carne de frango crua. De novembro de 2003 até abril de 2004, um total de 120 carcaças de frango foram compradas em 13 pontos de venda e 23 centros de criação de frangos. As Salmonella foram isoladas a partir de 75 (62,2%) carcaças analisadas. Vinte e um (21) sorotipos diferentes foram identificados, sendo os mais freqüentes S. Kentucky (30%), S. Muenster (13,3%), S. Brancaster (8,8%), S. Enteritidis e S. Hadar (6,6%). Todos os sorovares de salmonela foram examinados a fim de determinar a resistência à 16 antibióticos. Setenta e quatro (79,6%) foram resistentes à um antibiotico ou mais, das quais 33 (45,6%) mostraram resistência múltipla a cinco antibióticos (ampicilina, trimetoprim, trimetoprim-sulfametoxazol, tetraciclina e sulfonamidas). Foram encontrados 36 perfis diferentes de resistência múltipla. O nível elevado de resistência dos isolados de Salmonella encontrados na carne do presente estudo, é um indicador do uso indevido e contínuo de doses subterapêuticas de antibióticos nos animais. Por outro lado, os resultados do estudo demonstram a importância da carne de frango como fonte potencial de sorovares de Salmonella multiresistentes transmissíveis ao homem e sugerem um estudo epidemiólogico detalhado.
Palavras-chave:Salmonella sp, frangos, resistência à antimicrobianos
INTRODUCTION
The extensive use of antimicrobials in human and animals has led to an increase in bacterial multidrug resistant among several bacterial strains. This phenomenon of multiple resistance represents a worldwide problem both for veterinary and public health sectors. Bacterial resistance is observed especially when the antibiotics are abundantly used and that the bacteria can be transmitted easily between the individuals. Various antimicrobials in intensively managed food animals including chickens are often administered through feed or drinking water either for therapy, prophylaxis or growth promotion. Salmonella sp is one of the most frequently isolated bacteria in avian production units. The increasing single and multiple antimicrobial-resistant Salmonella strains isolated from human cases of salmonellosis has been associated with widespread use of antimicrobial agents in food animal production. This may represent a public health risk by transfer of resistant Salmonella strains to humans through the consumption of contaminated food and food products. Studies on antimicrobial resistant in Samonella have usually been undertaken in countries (1,10,15,20), that worked out excellent system to monitor resistance to antibiotics (3,9,11,17). In developing countries like Senegal, the situation of antimicrobial resistance is more complex and difficult. This is because Salmonella and other zoonotic bacterial pathogens are not routinely cultured and their resistance to commonly employed antimicrobials both in public health and veterinary practices is rarely determined. In Senegal, a new multiple resistant Salmonella serotype which emerged quasi simultaneously in food and human was identified (6). The purpose of the present study was to determine through the microbiological quality of chicken carcasses, the frequency and pattern of antimicrobial resistant Salmonella isolated from poultry in Dakar (Sénégal).
MATERIALS AND METHODS
Samples
A total of 120 whole chicken carcasses were purchased in 13 sale points and 23 flocks from November 2003 to April 2004 in the urban and periurban zones of Dakar. Sampling was done using a random sampling method. The number of carcasses purchased in poultry flocks was determined by the flock size. The samples were packed in iceboxes and transported immediately to the Hygiene and Animal Foodstuffs Inspection laboratory of Ecole Inter-Etats des Sciences et Médecine Vétérinaires of Dakar. The carcasses were frozen until their analysis.
Isolation and identification of Salmonella
The samples were examined for the presence of Salmonella according to the technique NF V 08-052 recommended by l'Association Française de Normalisation (AFNOR). Briefly, frozen samples were thawed at room temperature for about 3 hours. Twenty five grams of each sample were excised using a sterile scalpel. Each sample was put into a sterile stomacher bag containing 225 mL of buffered peptone water. The samples were then mixed for 2 minutes using a STOMACHER ND and incubated at 37ºC for 16 to 20 hours. After incubation, 0.1 mL of the pre-enriched sample was transferred into 10 mL of Rappaport Vassiliadis (Bio-Rad) broth and incubated at 42ºC for18 to 24 h. Another 1 mL aliquot was transferred into 10 ml of Selenite Cystine (Bio-Rad) broth and incubated at 37ºC for 24 hours. Following incubation, a loopful of each culture was streaked onto brillant green and Hecktoen agars (Bio-Rad) which were incubated at 37ºC for18 to 24 hours. Presumptive Salmonella colonies chosen from each plate were inoculated onto nutrient agar (Bio-Rad) and grown overnight at 37ºC. The cultures are then conveyed to the Microbiology-Immunology-Infectious Pathology Unit of Ecole Inter-Etats des Sciences et Médecine Vétérinaires of Dakar for identification. Salmonella isolates were screened biochemically using triple sugar iron, citrate, lysine decarboxylase, urease test, and indole test. Colonies that exhibited typical reactions were further biochemically characterised using API 20 E (Biomerieux) as recommended by the manufacturer. The isolates, which tested positive for Salmonella, were subcultured and sent for serotyping to the Medical Biology Unit, Pasteur Institute of Dakar.
Antimicrobial resistance test
The antimicrobial resistance of the isolates was determined by the agar diffusion method with Mueller Hinton agar and Bio-Rad disks (Marnes-La-Coquette, France).
The 16 antimicrobials tested were those commonly used in poultry herds or in human. There are: ampicillin (10 µg), nalidixic acid (30 µg), cephalotin (30 µg), cefoxitin (30 µg), chloramphenicol (30 µg), ciprofloxacin (5 µg), colistin (50 µg), flumequin (30 µg), furan (300 µg) gentamicin (15 µg), neomycin (30 UI), streptomycin (10 µg), sulphonamides (200 µg), tetracyclin (30 µg), trimethoprim (5µg) and trimethoprim-sulphamethoxazole (1.25 µg/25.75 µg). The categories susceptible or resistant were assigned on the basis of the critical points recommended by the French Committee on Guidelines for susceptibility testing (Comité de l'Antibiogramme de la Société Française de Microbiologie, C.A.-S.F.M.) (5).
RESULTS
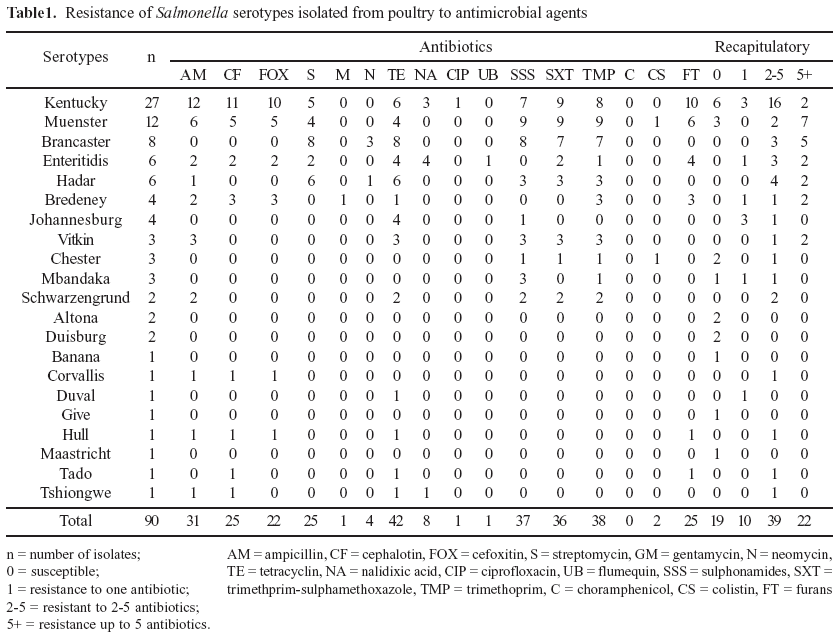
Salmonella were isolated from 75 (62.5%) of the 120 collected samples. Out of the 90 Salmonella isolates, twenty one different serotypes were identified of which S. Kentucky (30.0%), S. Muenster (13.3%), S. Brancaster (8.8%), S. Enteritidis and S. Hadar (6.6%) were the most prevalent ones. The distribution of serotypes is indicated in Table 1. From the total of 90 Salmonella isolates, 71 (78.9%) were resistant to one or more antimicrobials. Among the 71 resistant Salmonella isolates, 33 (45.6%) showed a multiple resistance pattern to 5 antimicrobials (ampicillin, trimethoprim, trimethoprim-sulfamethoxazole, sulphonamides and tetracyclin). The most significant percentages of resistance were obtained by decreasing order with tetracyclin (46.6%), trimethoprim (42.2%), sulphonamides (41.1%), Trimethoprim-Sulfamethoxazole (40%) and ampicillin (34.4%). Table 1 shows the frequency of resistance to one or more antimicrobials for the 21 identified serotypes.
Among the 21 serotypes identified, resistance was found in 16 of them. Fifteen multiresistant serotypes (to 2 ore more antimicrobials) were identified, including 7 resistant serotypes where resistance to more than 5 antimicrobials was noted. Thirty- six different antimicrobial multiresistance patterns were displayed by Table 2.
Five serotypes did not show any resistance with respect to antimicrobials tested: S. Altona (2 isolates), S. Duisburg (2 isolates), S. Banana (1 isolate), S. Give (1 isolate) and S. Tado (1 isolate).
DISCUSSION AND CONCLUSION
The level of contamination of chicken with Salmonella, observed in this study was higher than that obtained by Tall (unpublished data, 2003) in Senegal but comparable with those reported in some other African countries (13,19). Birds coming out of rearing units can be the principal source of contamination of the poultry meat by Salmonella. The results of the investigation showed that some stockbreeders do not respect prophylactic sanitary measures, which makes it possible for the bacteria to be maintained in the herd. The presence of 3 to 6 different serotypes in the same breeding reflects this lack of hygiene. However, plucking, evisceration and carcass washing, are all stages of the preparation which contribute to the whole process of contamination (Tall, unpublished data). The number of serotypes identified is significant: 21 from the 90 isolates. They are potentially pathogenic and different from those isolated by Tibaijuka et al. (19) in Ethiopia. They reported that from 244 poultry meat samples, 9 different serotypes were isolated from which the most prevalent were S. Braenderup, S. Anatum, S. Uganda and S. Saintpaul. In our study, serotypes Enteritidis and Hadar which play a significant role in collective food-borne diseases, were isolated. The antimicrobial susceptibility patterns of the Salmonella strains isolated indicated that a large proportion of the isolates were resistant to a variety of the drugs tested particularly tetracyclin, trimethoprim, sulphonamides, trimethoprime-sulfametoxazole, ampicillin, cephalotin and streptomycin. The percentages of resistance obtained with these antibiotics are comparable with those reported in other studies in France (17) and in Ethiopia (19). In Ethiopia, the level of resistance of 31 Salmonella isolates obtained from chicken meat samples was 60%.
In the present study, all Salmonella isolates were susceptible to chloramphenicol and only one was resistant to gentamicin. This may be explained by the limited availability and high cost of the above groups of antimicrobials that would reduce their frequent utilization in veterinary practice or public health practices in Senegal. Our results showed that Salmonella isolates were still largely susceptible to quinolones (9.7% of resistance to NA, 1.1% to flumequine). Although quinolones resistant Salmonella strains are seldom isolated (4,8), the reduction in the susceptibility must be regarded as an alarm signal, since quinolones are considered last resort antibiotics against multiple-drug resistant Salmonella strains. The evolutionary aspect of the resistance mechanisms calls for caution. Acquired resistances to furans are rare and develop slowly. The percentage of resistance to furans observed in our study was 29.0%. It is higher than the ones obtained by Poppe et al. (16) from Salmonella strains isolated from animals of which the percentages of resistance to furans were respectively 7.1%, 10.4%, 5.8%, and 5.1% over the years 1994 to 1997. This may be explained by the frequent use of furans (23.5%) for therapy in the poultry breeding and often by stockbreeders own decisions to medicate animals as reported by Bada-Alambedji et al. (2). Besides the increasing resistance to commonly used antibiotics in animals and humans, there is a concurrent increase in multiple resistant Salmonella isolates worldwide (7,14). The multiple resistance observed in the present study was to those antimicrobials commonly employed in veterinary practices and particularly to ampicillin, trimethoprim, trimethoprime-sulfametoxazole, tetracyclin, sulphonamides and streptomycin, which are also used for treatement of different human bacterial diseases. In a study of 54 Salmonella strains isolated from a total of 301 chicken meat and giblets in Ethiopia, 31 were resistant to one or more antimicrobials. Out of 31 resistant Salmonella isolates, 17 (54.8%) exhibited multiple resistance to up to 6 different antimicrobials (ampicillin, streptomycin, trimethoprim, sulfametoxazole, trimethoprim-sulfametoxazole and spectinomycin (19). In our study, all S. Brancaster and S. Hadar isolated showed multiple-drug resistance, followed by S. Enteritidis (83.3%), S. Muenster (75%), and S. Kentucky (66.6%). Salmonella Hadar isolates were resistant to streptomycin and tetracyclin. These frequencies of resistance are comparable with those reported by Poppe et al. (16). A study carried out in Ontario (Canada), showed that among strains of S. Hadar, isolated from poultry, 85.7% were resistant to these same antibiotics (12). On the other hand, S. Hadar isolates were susceptible to quinolones, contrary to an evolution to nalidixic acid resistance of S. Hadar isolated from animals in Belgium and reported by Chaslus-Dancla and Martel (8). Resistance to Nalidixic acid and tetracyclin was more frequent among S. Enteritidis isolates. Other studies covering several years that were done in the United Kingdom showed an increase in the frequency of nalidixic acid resistance among S. Enteritidis strains isolated from animals (18).
The results of the present study indicate single or multiple resistance of Salmonella strains isolated from poultry, that constitutes a potential source of transmission of these resistant strains to man and poses a problem in public health. This suggests the need for more prudent use of antibiotics by farmers, veterinarians and physicians. Further detailed epidemiological and molecular studies are essential on the frequency, sources of acquisition of resistant genes and distribution of antimicrobial resistant Salmonella among food animals, food products and humans in Sénégal.
ACKNOWLEDGMENTS
The authors would like to thank Inter-States School of veterinary Sciences and Medecine of Dakar for the financial support. They want also to thank Moussa Sène, Laboratory of microbiology and immunology and the technical staff of HIDAOA, veterinary school of Dakar for their excellent technical assistance in isolation and identification-antimicrobial resistance testing of Salmonella isolates; Dr Perrier Gros-Claude and the technical staff of the Medical Biology Unit, Pasteur Institute of Dakar for serotyping the Salmonella isolates.
Submitted: June 02, 2005; Returned to authors for corrections: March 16, 2006; Approved: July 18, 2006
- 1. Abouzeed, Y.M.; Hariharan, H.; Popp, C.; Kibenge, F.S.B. Characterization of Salmonella isolates from beef cattle, broiler chickens, and human sources on Prince Edward Island. Comp. Immun. Microbiol. Infect. Dis, 23, 253-266, 2000.
- 2. Bada-Alambédji, R.; Cardinale, E.; Biagui, C.; Akakpo, A.J.Recherche de résidus de substances à activité antibactérienne dans la chair de poulet consommée dans la région de Dakar (Sénégal). Bull. Acad. Vét. France, 157(2), 67-70, 2004.
- 3. Bager, F.; Aarestrup, F.M.; Jensen, N.E.; Madsen, M.; Meyling, A.; Wegener, H.C. Design of a system for monitoring antimicrobial resistance in pathogenic, zoonotic and indicator bacteria from food animals, Acta vetscand, 92(supplément): 77-86, 1999.
- 4. Boonmar, S.; Bangtrakulnonth, A.; Pornruangwong, S.; Samosornsuk, S.; Kaneko, K.; Ogawa, M. Significant increase in antibiotic resistance of Salmonella isolates from human beings and chicken meat in Thailand. Vet. Microbial., 62(1), 73-80, 1998.
- 5. Comité de l'Antibiogramme de la Société Française de Microbiologie (C.A.-S.F.M.). 1997. Communiqué 1997. Pathologie Biologie (Paris) 45: I-XII.
- 6. Cardinale, E.; Colbachini, P.; Perrier-Gros-Claude, J.D.; Gassama, A.; Aidara-Kane, A. Dual Emergence in Food and Human of a Novel multiresistant Serotype of Salmonella in Senegal: Salmonella enterica subsp. enterica Serotype 35: c: 1, 2. J. Clin. Microbiol, 39, 2373-2374, 2001.
- 7. Carraminana, J.J.; Rota, C.; Agustin, I.; Herrera, A. High prevalence of multiple resistance to antibiotics in Salmonella serotypes isolated from a poultry slaughterhouse in Spain. Vet. Microbial, 104(1-2), 133-9, 2004.
- 8. Chaslus-Dancla, E.; MARTEL, J.L. Résistance aux antibiotiques chez les animaux d'élevage. Bull. Soc. Fr. Microbiol, 12(2), 152-158, 1997.
-
9DANMAP 2000. Consumption of antimicrobial agents and occurrence of antimicrobial resistance in bacteria from food animals, foods, and humans in Denmark, Copenhague. DANMAP 56 p.2000.
- 10. Fedorka-Cray, J.; Dargatz, D.A.; Petersen, K.E.; Hollinger, K.; Wineland, N.E.; Headrick, M.; Tollefson, L.; Ferris, K. NARMS-EB veterinary isolates 1998 summary. Actes du Congrès sur le rôle de l'agriculture dans la gestion de la résistance aux antimicrobiens, Toronto, p.279, 1999.
- 11. Jones, Y.E.; Chapell, I.; McLaren, R.; Davies, R.; Wray, C. Antimicrobial resistance in Salmonella isolated from animals and their environment in England and Wales from 1988 to 1999. Vet. Rec, 150, 649-654, 2002.
- 12. Larkin, C.; Poppe, C.; McNab, B.; McEwen, B.; Mahdi, A.; Odumeru, J. Antibiotic resistance of Salmonella isolated from hog, beef and chicken carcass samples from provincially inspected abattoirs in Ontario. J. Food. Prot, 67, 448-455, 2004.
- 13. Molla, B.; Kleer, J.; Sinell, H.J. Antibiotic resistance pattern of foodborne Salmonella isolates in Addis Ababa (Ethiopia). Berl Munch Tierarztl Wochenschr, 112(2), 41-3, 1999.
- 14. Molla, B.; Mesfin, A.; Alemayehu, D. Multiple antimicrobial-resistant Salmonella serotypes isolated from chicken carcass and giblets in Debre Zeit and Addis Ababa, Ethiopia. Ethiop. J. Health Dev, 17(2), 131-149, 2003.
- 15. Nadeau, M.; Côté, G.; Higgins R. Surveillance de l'antibiorésistance chez des bactéries d'origine aviaire et porcine de 1993 à 1999 au Québec. Méd Vét Québec, 30, 195-199, 2000.
- 16. Poppe, C.; Ayroud, M.; Ollis, G.; Chirino-Trejo, M.; Smart, N.; Quessy, S.; Michel, P. Trends in antimicrobial resistance of Salmonella isolated from animals, foods of animal Origin, and the environment of animal production in Canada, 1994-1997. Microbial Drug resistance., 7(2), 197-212, 2001.
- 17. Sanders, P.; Gicquel, M.; Humbert, F.; Perrin-guyomard, A.; Salvat, G. Plan de surveillance de la résistance aux antibiotiques chez les bactéries indicatrices isolées de la flore intestinale des porcs et de la volaille 1999-2001. Bull. Acad. Vet. France, 155(3/4), 267-276, 2002.
- 18. Sojka, W.J.; Wray, C.; Mclaren, I. A survey of drug resistance in Salmonella isolated from animals in England and Wales in 1982 and 1983. Bc Vet. J, 142, 371-380, 1986.
- 19. Tibaijuka, B.; Molla, B.; Hildebrandt, G.; Kleer, J.; Salah, W.Antimicrobial resistance to Salmonellae isolated from retail raw chicken meat and giblets in Ethiopia. Bull. Anim. Hlth. Prod. Afr, 50(2), 86-95, 2002.
- 20. van Duijkeren, E.; Wannet, W.J.B.; Houwers, D.J.; Van Pelt, W. Antimicrobial susceptibilities of Salmonella strains isolated from humans, cattle, pig and chickens in the Netherlands from 1984 to 2001. J. Clin. Microbiol, 41(8), 3574-3578, 2003.
Publication Dates
-
Publication in this collection
01 Mar 2007 -
Date of issue
Dec 2006
History
-
Reviewed
16 Mar 2006 -
Received
02 June 2005 -
Accepted
18 July 2006