Abstracts
In this study, the presence of putative periodontal organisms, Porphyromonas gingivalis, Prevotella intermedia, Tannerella forsythensis, Fusobacterium nucleatum,Dialister pneumosintes,Actinobacillus actinomycetemcomitans,Campylobacter rectus,Eikenella corrodens and Treponema denticola were examined from subgingival samples of 40 dogs of different breeds with (25) and without (15) periodontitis, by using the PCR method. The PCR products of each species showed specific amplicons. Of the 25 dogs with periodontitis, P. gingivalis was detected in 16 (64%) samples, C. rectus in 9 (36%), A. actinomycetemcomitans in 6 (24%), P. intermedia in 5 (20%), T. forsythensis in 5 (20%), F. nucleatum in 4 (16%) and E. corrodens in 3 (12%). T. denticola and D. pneumosintes were not detected in clinical samples from dogs with periodontitis. Moreover, P. gingivalis was detected only in one (6.66%) crossbred dog without periodontitis. Our results show that these microorganisms are present in periodontal microbiota of dogs with periodontitits, and it is important to evaluate the role of these putative periodontal microorganisms play in the periodontitis in household pets particularly, dogs in ecologic and therapeutic terms, since these animals might acquire these periodontopahogens from their respective owners.
Periodontal microorganisms; dogs; periodontitis; oral anaerobic bacteria
Neste estudo, a presença de patógenos periodontais, Porphyromonas gingivalis, Prevotella intermedia, Tannerella forsythensis, Fusobacterium nucleatum, Dialister pneumosintes, Actinobacillus actinomycetemcomitans, Campylobacter rectus, Eikenella corrodens e Treponema denticola foi determinada por PCR, em amostras subgengivais de 40 cães com (25) e sem (15) doença periodontal. Os produtos amplificados pelo PCR para cada espécie bacteriana mostraram amplicons específicos. Dos 25 cães apresentando doença periodontal, P. gingivalis foi detectado em 16 amostras (64%), C. rectus em 9 (36%), A. actinomycetemcomitans em 6 (24%), P. intermedia em 5 (20%), T. forsythensis em 5 (20%), F. nucleatum em 4 (16%) e E. corrodens em 3 (12%). Em nenhuma amostra clínica periodontal foi observada a presença de T. denticola ou D. pneumosintes. Adicionalmente, P. gingivalis foi detectado em apenas um (6,66%) cão saudável, sem raça definida. Nossos resultados mostram que esses microrganismos estão presentes na microbiota periodontal de cães com periodontitis, e isto torna importante a avaliação do papel que esses microrganismos periodontais desempenham na periodontite de animais domésticos, particularmente cães, em termos ecológicos e terapêuticos, desde que esses cães podem adquirir esses periodontopatógenos de seus respectivos proprietários.
Microrganismos periodontais; cães; periodontite; bactéria anaeróbia oral
VETERINARY MICROBIOLOGY
Detection of putative periodontal pathogens in subgingival specimens of dogs
Detecção de patógenos periodontais em amostras subgengivais de cães
Sheila Alexandra Belini NishiyamaI; Gerusa Neyla Andrade SenhorinhoI; Marco Antônio GiosoII; Mario Julio Avila-CamposI,* * Corresponding Author. Mailing address: USP - ICB - Dep. de Microbiologia - Lab. Anaeróbios - Av. Prof. Lineu Prestes, 1374 Cid. Universitária 05508-900 são Paulo, SP - Brasil. Tel. (11) 3091-7344 ou (11) 3091-7354. E-mail: mariojac@usp.br
IAnaerobe Laboratory, Department of Microbiology, Institute of Biomedical Science
IIVeterinary and Zootechny School, University of São Paulo, São Paulo, SP, Brazil
ABSTRACT
In this study, the presence of putative periodontal organisms, Porphyromonas gingivalis, Prevotella intermedia, Tannerella forsythensis, Fusobacterium nucleatum,Dialister pneumosintes,Actinobacillus actinomycetemcomitans,Campylobacter rectus,Eikenella corrodens and Treponema denticola were examined from subgingival samples of 40 dogs of different breeds with (25) and without (15) periodontitis, by using the PCR method. The PCR products of each species showed specific amplicons. Of the 25 dogs with periodontitis, P. gingivalis was detected in 16 (64%) samples, C. rectus in 9 (36%), A. actinomycetemcomitans in 6 (24%), P. intermedia in 5 (20%), T. forsythensis in 5 (20%), F. nucleatum in 4 (16%) and E. corrodens in 3 (12%). T. denticola and D. pneumosintes were not detected in clinical samples from dogs with periodontitis. Moreover, P. gingivalis was detected only in one (6.66%) crossbred dog without periodontitis. Our results show that these microorganisms are present in periodontal microbiota of dogs with periodontitits, and it is important to evaluate the role of these putative periodontal microorganisms play in the periodontitis in household pets particularly, dogs in ecologic and therapeutic terms, since these animals might acquire these periodontopahogens from their respective owners.
Key words: Periodontal microorganisms; dogs; periodontitis; oral anaerobic bacteria
RESUMO
Neste estudo, a presença de patógenos periodontais, Porphyromonas gingivalis, Prevotella intermedia, Tannerella forsythensis, Fusobacterium nucleatum, Dialister pneumosintes, Actinobacillus actinomycetemcomitans, Campylobacter rectus, Eikenella corrodens e Treponema denticola foi determinada por PCR, em amostras subgengivais de 40 cães com (25) e sem (15) doença periodontal. Os produtos amplificados pelo PCR para cada espécie bacteriana mostraram amplicons específicos. Dos 25 cães apresentando doença periodontal, P. gingivalis foi detectado em 16 amostras (64%), C. rectus em 9 (36%), A. actinomycetemcomitans em 6 (24%), P. intermedia em 5 (20%), T. forsythensis em 5 (20%), F. nucleatum em 4 (16%) e E. corrodens em 3 (12%). Em nenhuma amostra clínica periodontal foi observada a presença de T. denticola ou D. pneumosintes. Adicionalmente, P. gingivalis foi detectado em apenas um (6,66%) cão saudável, sem raça definida. Nossos resultados mostram que esses microrganismos estão presentes na microbiota periodontal de cães com periodontitis, e isto torna importante a avaliação do papel que esses microrganismos periodontais desempenham na periodontite de animais domésticos, particularmente cães, em termos ecológicos e terapêuticos, desde que esses cães podem adquirir esses periodontopatógenos de seus respectivos proprietários.
Palavras-chave: Microrganismos periodontais, cães, periodontite, bactéria anaeróbia oral
INTRODUCTION
The periodontal disease is an inflammatory condition of the dental periodontal tissues and it is often observed in small domestic animals, such as dogs (2,20). Depending on the severity, the periodontal diseases can be broadly classified into two stages: gingivitis, characterized by tissue inflammation, and periodontitis, associated with attachment loss, alveolar bone resorption, and tooth loss (10).
These processes are caused by accumulation of subgingival biofilm and the severity is mediated by the presence of specific microorganisms and the host immunity status (16). The pathogenesis of the periodontal disease in dogs has been little studied and these animals are used as research models for human periodontal disease (1,17,20). Studies have suggested that the periodontal etiologic agents are similar to those of human disease (30,36). Moreover, it has been demonstrated a strong correlation between the presence of putative periodontal organisms and the destruction of periodontal tissues. Organisms such as Porphyromonas gingivalis, Prevotella intermedia,Actinobacillus actinomycetemcomitan,Treponema denticola, Fusobacterium nucleatum, Dialister pneumosintes,Tannerella forsythensis,Treponema denticola, Eikenella corrodens and Campylobacter rectus have been related to several forms of periodontal disease in humans and animals (2,3,5,29).
Molecular methods are currently available for detecting periodontopathogens, since techniques of bacterial culture are used to identify putative anaerobic pathogens from destructive periodontal disease, but it can be somewhat cumbersome and expensive (5). Polymerase chain reaction (PCR) has been used in the direct identification of periodontal pathogens from subgingival specimens due to ability to accurately detect microbial species in mixed populations (3,4).
The aim of this study was to detect putative periodontal organisms from adult dogs with naturally developed periodontitis and without periodontitis.
MATERIAL AND METHODS
Dogs and clinical samples
Twenty-five dogs with periodontitis and 15 healthy dogs were selected from a Private Dental Veterinary Clinic (Odontovet, Sao Paulo, SP, Brazil) and from the School of Veterinary Medicine and Zootechny at the University of Sao Paulo (Sao Paulo, SP, Brazil). The selected animals belonged to four Crossbred and 36 to different breeds, as follow: 10 Poodle, 2 Schnauzer, 4 Yorkshire, 3 Beagle, 3 Cocker Spaniel, 3 Lhasa Apso, 1 Chow Chow, 1 Basset Hound, 1 German Shepherd, 1 Golden, 2 Maltese, 2 Rottweiller, 1 West Highland and 2 Daschund. They were 18 dogs and 22 bitches and from 1-year to 6-years of age. All the dogs with periodontitis showed clinical evidences of bleeding on probing, alveolar loss and periodontal pockets depth equal or exceeding 5 mm. Animals with systemic diseases, as well as those which had received antimicrobial therapy within the previous three months were excluded. The Ethic Committee for Animal Experimentation at the ICB/USP approved this study.
Initially, animals were anesthetized with propofol (2 mg/kg) and diazepam (5.5 mg/kg) by an intramuscular injection; isoflurane and oxygen were used by endotracheal way. Then, dental supra-gingival biofilm was removed by using a sterile gauze and the subgingival samples were obtained from only one periodontal site (canine, incisive or pre-molars) by using two fine sterile paper points (No. 40, Tanariman Ind. Ltd. AM, Brazil) inserted to the depth of the periodontal pocket or the gingival crevices and left in place for 60s. Samples were transported in 300 µl of sterilized ultra-pure water.
DNA extraction and analysis
DNA extraction was performed according to Avila-Campos (6). Samples were homogenized and washed twice at 12,000 x g for 10 minutes. The pellet was ressuspended in 300 µl of ultra-pure water and boiled for 10 minutes. After centrifugation (14,000 x g, 10 minutes), the supernatant (DNA) was saved and transferred to a new tube and used as template. The quality of DNA samples was verified by using an agarose gel electrophoresis, and all of them were stored at -20ºC.
PCR amplification
All the clinical samples were included in the PCR analysis. Species-specific primer pairs based on the sequence of the gene 16S rDNA were used according to Ashimoto et al. (3). The amplification reaction was performed in 25 µl containing 2.5 µl of 10 X PCR buffer, 1.25 µl of MgCl2 (50 mM), 1.0 µl of dNTP (0.2 mM; Invitrogen do Brasil, Sao Paulo, SP, Brazil), 1.0 µl of each primer (0.4 M), 0.25 µl of Platinum Taq DNA polymerase (0.5U; Invitrogen), 8 µl of sterilized ultra-pure water and 10 ng of DNA. The amplifications were performed in a DNA thermal cycler (Perkin Elmer, GeneAmp PCR System 9700, Norwalk, CT, USA) programmed for 94ºC (5 minutes), followed by 30 cycles at 94ºC for 30 seconds; annealing temperature adequate for each primer pairs (Table 1) for 30 seconds; 72ºC for 30 seconds, then 72ºC (5 minutes) to allow the completion of the DNA extension. In each respective PCR amplification DNA from P. gingivalis ATCC 33277, A. actinomycetemcomitans ATCC 29523, F. nucleatum ATCC 10953, D. pneumosintes ATCC 33048, P. intermedia ATCC 25611, T. denticola ATCC 33520, T. forsythensis ATCC 43037, E. corrodens ATCC 23834 and C. rectus ATCC 33238 were used as positive controls. A negative control without template DNA was included in each PCR run.
Amplified products were compared by electrophoresis in 1% of agarose gel in 1 X TBE buffer (1M Tris, 0.9M boric acid, 0.01M EDTA, pH 8.4) (Invitrogen), at 60 V, for 2.5 hours, stained with ethidium bromide (0.5 mg/ml) and photographed on an UV light transilluminator (Kodak Digital Science System 120TM). A 1-kb plus DNA ladder (Invitrogen) was used as a molecular marker.
RESULTS
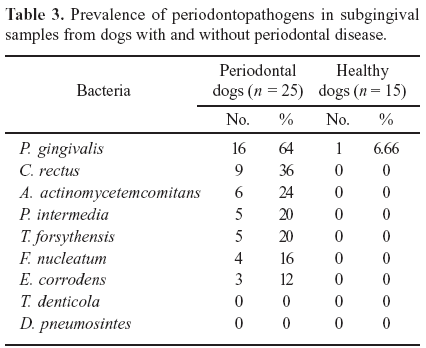
The total bacterial distribution in 25 dogs with periodontitis and 15 healthy dogs is described in Table 2. It can also be observed that Poodles showed the presence of the six studied microorganisms followed by Yorkshire, Beagle and Basset Hound dogs. P. gingivalis (64%) and C. rectus (36%) were present in most of the periodontal animals. Moreover, A. actinomycetemcomitans was present in 24%, P. intermedia and T. forsythensis in 20%, F. nucleatum in 16% and E. corrodens in 12% of the periodontal dogs (Table 3). Of the 25 dogs with periodontitis only German Shepherd did not show the presence of any organism. No dog without periodontitis showed the presence of any studied periodontopathogen except one (6.66%) crossbred dog harboring P. gingivalis. On the other hand, none of the periodontal or healthy dogs showed T. denticola or D. pneumosintes. Moreover, all the evaluated dogs positive to any periodontal microorganism were over one year of age, but none association between the presence of organisms and analyzed breeds was observed.
DISCUSSION
In veterinary dentistry, it is estimated that more than 80% of adult dogs have periodontal disease (18). Since these organisms have an important role in oral infection, their isolation and identification are required (12,18). Periodontal bacteria such as E. corrodens, A. actinomycetemcomitans, P. gingivalis, P. intermedia and F. nucleatum are recognized as important opportunistic pathogens in the development of periodontal and non-oral diseases of humans and dogs (7,16,19).
Studies have shown the presence of different organisms of the subgingival samples from dogs with gingivitis or induced periodontitis (1,17,27). In our study, we detected putative periodontal organisms from subgingival samples of dogs with naturally occurring periodontitis. P. gingivalis was detected in 64% of the evaluated periodontal dogs, according to Allaker et al. (2), who identified this organism in 68% of the evaluated dogs.
It is well known that the presence of plaque at the gingival margin and in the sulcus induces a gingival inflammation in dogs, since it is irritating and causes gingivitis (19). Animals over two years old, mainly dogs, suffer in some degree of this chronic inflammatory disease (26) and the bacteria often implicated in that process are P. gingivalis and P. intermedia, as well as in human periodontitis (2). In this study, none association between the presence of the microorganisms and breed dogs analyzed was observed. It was also interesting to note that P. gingivalis was detected only in one (6.66%) of the healthy crossbred dogs, and it is explained due to an accelerated disease progression observed in companion animals compared to humans may be due to a relative lack of routine dental care.
The evaluated periodontal microorganisms were observed 12% to 64% of the dogs with periodontitis and it suggests that the detected microorganisms can play an important role in progression of periodontitis. In this study, we analyzed different dog breeds and certainly the dietary regimen, age, sex and other factors such as the animal behavior could interfere with the prevalence and in the progression of disease (35).
Our results show that C. rectus was the second most often bacteria detected in the evaluated periodontal dogs (36%). Similarly, studies have shown that in humans this organism is an active pathogen in periodontitis (3,33). On the other hand, C. rectus has been detected in low levels from samples of ligature-induced periodontitis in beagle dogs (27).
A. actinomycetemcomitans is considered an important periodontal pathogen in the human aggressive periodontitis (13). In this study, this organism was detected in 24% of the periodontal samples. In addition, none healthy dog harbored A. actinomycetemcomitans. Data have shown that A. actinomycetemcomitans is recovered from healthy human without periodontal disease in approximately 20% to 25% of the examined subgingival samples, and it has been suggested that the presence of this organism in healthy sites can be an exogenous contamination (3,25).
F. nucleatum is a Gram negative anaerobe often recovered from subgingival samples, particularly from human periodontitis (25). In our study, F. nucleatum was present in 16% of the periodontal dogs but none subgingival sample from healthy dogs showed DNA of this organism. Moreover, studies have shown the high prevalence of F. nucleatum in gingivitis and in advanced periodontitis, particularly, in beagle dogs (27,31).
P. intermedia/P. nigrescens isolates are commonly found in the oral cavity of periodontal healthy or diseased individuals as well as those with extra-oral infections (23). These two species, P. intermedia and P. nigrescens are not distinguishable by routine phenotypic methods (15). The identification to species level is based on DNA homology studies, serotyping using monoclonal antibodies, sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), hybridization with species specific oligonucleotide probes or polymerase chain reaction (24,29). In our study, 20% of the periodontal dogs harbored P. intermedia. Studies have shown that the same genotypes of P. intermedia can colonize spouses or parents and children (14); the same can be true to owners and household pets, due to these animals have a close relationship with members of a family, particularly those with pedigree.
T. forsythensis is found in the subgingival and periodontal pockets of humans and it is known to cause periodontal disease (30,32). This organism has been isolated from the oral cavity of primates but not from other animals. Moreover, T. forsythensis has been isolated from human wounds produced by cat and dog bite, suggesting that these strains were originated in the oral cavity of dogs and cats (22). In our study, T. forsythensis was detected in 20% of the evaluated dogs and it can suggest a possible role in periodontal microbiota.
E. corrodens appears to be an opportunistic pathogen in man and dogs (1), but its prevalence in the dog oral cavity is still unknown. Our data showed the occurrence of E. corrodens in 12% of the evaluated periodontal dogs. On the other hand, this organism has been observed in 60% of healthy adult humans (9).
It is well known that several oral spirochetes are part of the oral microbiota in household pets such as dogs and cats; however, their role in pathogenesis and in the progression of periodontal disease is not still clear (21,28). Moreover, the role and the diversity of the Treponema spp. in human oral microbiota or in periodontal disease are still unclear (30,33). T. denticola has been related to subgingival biofilm from dogs, suggesting that some household pets may be susceptible to the bacterial infection similarly from those found in humans (28). In our study, none of the analyzed dogs harbored this organism.
D. pneumosintes has been reported in literature as a putative periodontopathogen and it can be recovered from oral and extra-oral infections (11). This organism appears to be related to the increase of the alveolar bone loss in man (8). Nevertheless, this periodontal organism has not been described in oral samples of dogs or other household pets.
Because these animals can acquire these periodontal pathogens from their respective owners, our results suggest the need to determine the role of these putative periodontal microorganisms in the periodontal disease of household pets, particularly dogs.
ACKNOWLEDGEMENTS
The authors are grateful to Zulmira Alves de Souza for her technical support. This study was partly supported by Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP Grant 02/02568-0). During the course of this work, SBN and GNS were supported by the fellowship from CAPES, and MJA-C fellowships are partly supported by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq).
Submitted: April 07, 2006; Returned to authors for corrections: July 13, 2006; Approved: October 13, 2006
- 1. Allaker, R.P.; Langlois, T.; Hardie, J.M. (1994). Prevalence of Eikenella corrodens and Actinobacillus actinomycetemcomitans in the dental plaque of dogs. Vet. Rec., 14, 519-520.
- 2. Allaker, R.P.; Young, K.A.; Langlois, T.; Rosayro, R.; Hardie, J.M. (1997). Dental plaque flora of the dog with reference to fastidious and anaerobic bacteria associated with bites. J. Vet. Dent, 14, 127-130.
- 3. Ashimoto, A.; Chen, C.; Bakker, I.; Slots, J. (1996). Polymerase chain reaction detection of 8 putative periodontal pathogens in subgingival plaque of gingivitis and advanced periodontitis lesions. Oral Microbiol. Immunol, 11, 266-273.
- 4. Avila-Campos, M.J.; Sacchi, C.T.; Whitney, A.M.; Steigerwalt, A.G.; Mayer, L.W. (1999). Arbitrarily primed-polymerase chain reaction for identification and epidemiologic subtyping of oral isolates of Fusobacterium nucleatum.J. Periodontol., 70, 1202-1208.
- 5. Avila-Campos, M.J.; Velásquez-Meléndez, G. (2002). Prevalence of putative periodontopathogens from periodontal patients and healthy subjects in São Paulo, SP, Brazil. Rev. Inst. Med. Trop. São Paulo, 44, 1-5.
- 6. Avila-Campos, M.J. (2003). PCR detection of four periodontopathogens from subgingival clinical samples. Braz. J. Microbiol., 34, 34-81.
- 7. Braga, C.A.S.B.; Resende, C.M.F.; Pestana, A.C.N.R.; Carmo, L.S.; Costa, J.E.; Silva, L.A.F.; Assis, L.N.; Lima, L.A.; Farias, L.M.; Carvalho, M.A.R. (2005). Isolamento e identificação da microbiota periodontal de cães da raça Pastor Alemão. Ciência Rural., 35, 385-390.
- 8. Contreras, A.; Doan, N.; Chen, C.; Rusitanonta, T.; Flynn M.J.; Slots, J. (2000). Importance of Dialister pneumosintes in human periodontitis. Oral Microbiol. Immunol, 15, 269-272.
- 9. Chen, C.K.C.; Dunford, R.G.; Reynolds, H.S.; Zambon, J.J. (1989). Eikenella corrodens in the human oral cavity. J. Periodontol., 60, 611-616.
- 10. Darveu, R.P.; Belton, C.M.; Reife, R.A.; Lamont, R.J. (1998). Local chemokine paralysis, a novel pathogenic mechanism for Porphyromonas gingivalis. Infect. Immun., 66, 1660-1665.
- 11. Doan, N.; Contreras, A.; Flynn, J.; Slots, J.; Chen, C. (2000). Molecular identification of Dialister pneumosintes in subgingival plaque of humans. J. Clin. Microbiol 38, 3043-3047.
- 12. Dzink, J.L.; Socransky, S.S.; Haffajee, A.D. (1988). The predominant cultivable microbiota of active and inactive lesions of destructive periodontal diseases. J. Clin. Periodontol., 15, 316-323.
- 13. Fives-Taylor, P.; Meyer, D.; Mintz, K. (1996). Virulence factors of the periodontopathogen Actinobacillus actinomycetemcomitans.J. Periodontol., 96, 291-297.
- 14. Fukui, K.; Kato, N.; Kato, H.; Watanabe, K.; Tatematsu, N. (1999). Incidence of Prevotella intermedia and Prevotella nigrescens carriage among family members with subclinical periodontal disease. J. Clin. Microbiol., 37, 3141-3145.
- 15. Frandsen, E.V.G.; Poulsen, K.; Kilian, M. (1995). Confirmation of the species Prevotella intermedia and Prevotella nigrescens. Int. J. Syst. Bacteriol., 45, 429-435.
- 16. Genco, C.A. (1998). Animal models for Porphyromonas gingivalis-mediated periodontal disease. Trends Microbiol, 6, 445-449.
- 17. Hale, F.A. (2003). The owner-animal-environment triad in the treatment of canine periodontal disease. J. Vet. Dent., 20, 118-122.
- 18. Hardan, J.; Dreier, K.; Wong, J.; Sfintescu, C.; Evans, R.T. (2005). Pigmented-anaerobic bacteria associated with canine periodontitis. Vet. Microbiol, 106, 119-128.
- 19. Harvey, C.E.; Thornsberry, C.; Miller, B.R. (1995). Subgingival bacteria-comparison of culture results in dogs and cats with gingivitis. J. Vent. Dent., 12, 147-150.
- 20. Harvey, C.E. (1998). Periodontal disease in dogs. Etiopathogenesis, prevalence, and significance. Vet. Clin. North Am. Small Anim. Pract, 28, 1111-1128.
- 21. Hennet, P.R.; Harvey, C.E. (1991). Anaerobes in periodontal disease in the dog: a review. J. Vet. Dent, 8, 18-21.
- 22. Hudspeth, M.K.; Hunt G.S.; Maiden, M.F.J.; Citron, D.M.; Goldstein, E.J.C. (1999). Characterization of Bacteroides forsythus strains from cat and dog bite wounds in humans and comparison with monkey and human oral strains. J. Clin. Microbiol., 37, 2003-2006.
- 23. Jousimies-Somer, H.; Savolainen, S.; Makitie, A.; Ylikoshi, J. (1993). Bacteriologic finding in peritonsillar abscesses in young adults. Clin. Infect. Dis., 16, 292-298.
- 24. Mättö, J.; Saarela, M.; von Toil-Lindén, B.; Alaluusua, S.; Jousimies-Somer, H.; Asikainen, S. (1996). Similarity of salivary and subgingival Prevotella intermedia and Prevotella nigrescens isolates by arbitrarily primed polymerase chain reaction. Oral Microbiol Immunol., 11, 395-401.
- 25. Moore, W.E.C.; Moore L.V.H. (1994). The bacteria of periodontal diseases. Periodontol, 2000. 5, 66-77.
- 26. Penman, S.; Harvey, C.E. (1990). Periodontal disease. In: Harvey, C.E.; Orr, H. S. (Eds.), Manual of Small Animal Dentistry B.S.A.V. A., Cheltenham, British Small Animal Veterinary Association, p.37-39.
- 27. Renvert, S.; Wikström, M.; Mugrabi, M.; Claffey, N. (1996). Histological and microbiological aspects of ligature-induced periodontitis in beagle dogs. J. Clin. Periodontol., 23, 310-319.
- 28. Riviere, G.R.; Thompson, A.J.; Brannan, R.D.; McCoy, D.E.; Simonson, L.G. (1996). Detection of pathogen-related oral spirochetes, Treponema denticola and Treponema socranskii in dental plaque from dogs. J. Vet. Dent., 13, 135-138.
- 29. Saarela, M.; Asikainen, S.; Chen, C.; Alaluusua, S.; Slots, J. (1995). Comparison of arbitrarily primed polymerase chain reaction and ribotyping for subtyping Actinobacillus actinomycetemcomitans. Anaerobe, 1, 97-102.
- 30. Socransky, S.S.; Haffajee, A.D.; Cugini, M.A.; Smith, C.; Kent Jr., R.L. (1998). Microbial complexes in subgingival plaque. J. Clin. Microbiol., 25, 134-144.
- 31. Syed, S.A.; Svanberg, M.; Svanberg, G. (1981). The predominant cultivable dental plaque flora of beagle dogs with periodontitis. J. Clin. Periodontol, 8, 45-56.
- 32. Tang, K.S.; Song, K-P.; Ong, G. (2001). Bacteroides forsythus prtH genotype in periodontitis patients: occurrence and association with periodontal disease. J. Periodont. Res, 36, 398-403.
- 33. Tanner, A.; Maiden, M.F.J.; Macuch, P.J.; Murray, L.L.; Kent Jr., R.L. (1998). Microbiota of health, gingivitis and initial periodontitis. J. Clin. Periodontol, 25, 85-98.
- 34. Valdez, M.; Haines, R.; Riviere, K.H.; Riviere, G.R.; Thomas, D. (2000). Isolation of oral spirochetes from dogs and cats and provisional identification using a polymerase chain reaction (PCR) analysis specific for human plaque Treponema spp. J. Vet. Dent, 17, 23-26.
- 35. Watson, A.D.J. (1994). Diet and periodontal disease in dogs and cats. Australian Vet. J., 71, 313-318.
Publication Dates
-
Publication in this collection
17 Apr 2007 -
Date of issue
Mar 2007
History
-
Accepted
13 Oct 2006 -
Reviewed
13 July 2006 -
Received
07 Apr 2006