Abstracts
Aggregatibacter actinomycetemcomitans is an important etiologic agent of the periodontitis and is associated with extra-oral infections. In this study, the detection of the ltxA gene as well as the ltx promoter region from leukotoxic A. actinomycetemcomitans isolated from 50 Brazilian patients with periodontitis and 50 healthy subjects was performed. The leukotoxic activity on HL-60 cells was also evaluated. Leukotoxic activity was determined using a trypan blue exclusion method. The 530 bp deletion in the promoter region was evaluated by PCR using a PRO primer pair. A. actinomycetemcomitans was detected by culture and directly from crude subgingival biofilm by PCR using specific primers. By culture, A. actinomycetemcomitans was detected in nine (18%) of the periodontal patients and one (2%) healthy subject. However, by PCR, this organism was detected in 44% of the periodontal patients and in 16% of the healthy subjects. It was verified a great discrepancy between PCR detection of the ltx operon promoter directly from crude subgingival biofilm and from bacterial DNA. Only one periodontal sample harbored highly leukotoxic A. actinomycetemcomitans. Moreover, biotype II was the most prevalent and no correlation between biotypes and leukotoxic activity was observed. The diversity of leukotoxin expression by A. actinomycetemcomitans suggests a role of this toxin in the pathogenesis of periodontal disease and other infectious diseases.
biotype; leukotoxin; periodontitis
Aggregatibacter actinomycetemcomitans é um importante agente etiológico da periodontite e produz infecções extra-bucais. Neste estudo, foram detectados os biótipos, o gene ltxA associado à produção de leucotoxina e o promotor ltx em A. actinomycetemcomitans de pacientes com e sem periodontite. A atividade leucotóxica sobre células HL-60 também foi avaliada. A atividade leucotóxica foi determinada através do método de exclusão do azul de tripam. A deleção de 530 bp no promotor ltx foi avaliada usando-se o par de iniciadores PRO. A. actinomycetemcomitans foi detectado por cultura e por PCR. Por cultura, A. actinomycetemcomitans foi detectado em nove pacientes com periodontite (18%) e em um indivíduo sadio (2%). Por PCR esse microrganismo foi detectado em 44% dos pacientes com periodontite e em 16% dos saudáveis. Verificou-se diferença estatística entre a detecção do promotor do operon ltx, por PCR, diretamente do biofilme subgengival e do DNA bacteriano. Somente uma amostra clínica apresentou A. actinomycetemcomitans altamente leukotóxico. O biótipo II foi o mais prevalente e não foi observada correlação biótipo-atividade leucotóxica. A expressão da leucotoxina por A. actinomycetemcomitans na doença periodontal e outras doenças infecciosas necessita ser avaliado.
biótipo; leucotoxina; periodontite
MEDIC AL MICROBIOLOGY
Distribution of biotypes and leukotoxic activity of Aggregatibacter actinomycetemcomitans isolated from Brazilian patients with chronic periodontitis
Distribuição de biótipos e atividade leucotóxica de Aggregatibacter actinomycetemcomitans isolados de pacientes brasileiros com periodontite crônica
Elerson Gaetti-Jardim Jr.I; Thais Cristiane WahasuguiIII; Paulo Henrique TomazinhoIII; Márcia Martins MarquesII; Viviane NakanoIII; Mario Julio Avila-CamposIII,* * Corresponding Author. Mailing address: Anaerobe Laboratory, Department of Microbiology, ICB, University of São Paulo. Av. Prof. Lineu Prestes 1374, São Paulo, SP, 05508-900, Brazil. Phone/fax: +55 11 3091-7344/3091-7354. E-mail: mariojac@usp.br
ILaboratório de Patologia, Faculdade de Odontologia, Universidade Estadual Paulista, Araçatuba, SP, Brasil
IIFaculdade de Odontologia, Universidade de São Paulo, SP, Brasil
IIILaboratório de Anaeróbios, Departamento de Microbiologia, Universidade de São Paulo, São Paulo, SP, Brasil
ABSTRACT
Aggregatibacter actinomycetemcomitans is an important etiologic agent of the periodontitis and is associated with extra-oral infections. In this study, the detection of the ltxA gene as well as the ltx promoter region from leukotoxic A. actinomycetemcomitans isolated from 50 Brazilian patients with periodontitis and 50 healthy subjects was performed. The leukotoxic activity on HL-60 cells was also evaluated. Leukotoxic activity was determined using a trypan blue exclusion method. The 530 bp deletion in the promoter region was evaluated by PCR using a PRO primer pair. A. actinomycetemcomitans was detected by culture and directly from crude subgingival biofilm by PCR using specific primers. By culture, A. actinomycetemcomitans was detected in nine (18%) of the periodontal patients and one (2%) healthy subject. However, by PCR, this organism was detected in 44% of the periodontal patients and in 16% of the healthy subjects. It was verified a great discrepancy between PCR detection of the ltx operon promoter directly from crude subgingival biofilm and from bacterial DNA. Only one periodontal sample harbored highly leukotoxic A. actinomycetemcomitans. Moreover, biotype II was the most prevalent and no correlation between biotypes and leukotoxic activity was observed. The diversity of leukotoxin expression by A. actinomycetemcomitans suggests a role of this toxin in the pathogenesis of periodontal disease and other infectious diseases
Key-words: Aggregatibacter actinomycetemcomitans, biotype, leukotoxin, periodontitis.
RESUMO
Aggregatibacter actinomycetemcomitans é um importante agente etiológico da periodontite e produz infecções extra-bucais. Neste estudo, foram detectados os biótipos, o gene ltxA associado à produção de leucotoxina e o promotor ltx em A. actinomycetemcomitans de pacientes com e sem periodontite. A atividade leucotóxica sobre células HL-60 também foi avaliada. A atividade leucotóxica foi determinada através do método de exclusão do azul de tripam. A deleção de 530 bp no promotor ltx foi avaliada usando-se o par de iniciadores PRO. A. actinomycetemcomitans foi detectado por cultura e por PCR. Por cultura, A. actinomycetemcomitans foi detectado em nove pacientes com periodontite (18%) e em um indivíduo sadio (2%). Por PCR esse microrganismo foi detectado em 44% dos pacientes com periodontite e em 16% dos saudáveis. Verificou-se diferença estatística entre a detecção do promotor do operon ltx, por PCR, diretamente do biofilme subgengival e do DNA bacteriano. Somente uma amostra clínica apresentou A. actinomycetemcomitans altamente leukotóxico. O biótipo II foi o mais prevalente e não foi observada correlação biótipo-atividade leucotóxica. A expressão da leucotoxina por A. actinomycetemcomitans na doença periodontal e outras doenças infecciosas necessita ser avaliado.
Palavras-chave: Aggregatibacter actinomycetemcomitans, biótipo, leucotoxina, periodontite.
INTRODUCTION
Aggregatibacter actinomycetemcomitans is a capnophilic gram-negative microorganism, member of the indigenous oral microbiota, and it is found in periodontal lesions, especially in young adults. This organism has been associated with several infectious diseases, such as septic endocarditis, brain and lung abscesses, osteomyelitis, and cardiovascular pathologies (8,27), and aggressive and chronic periodontal diseases (25,32,33).
This microorganism is grouped into 10 different biotypes, and their distribution depends on several factors, such as, geographic location and clinical condition of the patients (3,28). However, there is a need to confirm this statement in Brazilian population, which presents several peculiarities, such as racial miscegenation and cultural aspects. In addition, there are no available data to support the relationship virulence-biotype in this microorganism.
The virulence of A. actinomycetemcomitans is not well understood, but this organism produces virulence factors including a heat-labile leukotoxin (7,9) that belongs to the repeat-in-toxin (RTX) family. The gene ltxA encodes the leukotoxin, genes ltxB and ltxD encode proteins presumably required for the toxin secretion, and gene ltxC encodes the acyl-transferase production that is responsible for the toxin transformation, from protoxin to the active form (20).
Genetic determinants for the synthesis, activation, and secretion of leukotoxin are localized on ltx operon, which seems to be present in all A. actinomycetemcomitans strains (16,31), however levels of toxin expression vary considerably in different clones. It has been observed that the leukotoxin expression is related to the presence of a 530 bp deletion in the promoter region, although physiological factors may also be involved in this regulation (18).
The presence of leukotoxin has been associated with the ability of A. actinomycetemcomitans to evade the main defense line into periodontal pocket and it may contribute to the pathogenesis of periodontal disease (17). The leukotoxic activity is determined by a cytolytic action that kills human polymorphonuclear leukocytes, T lymphocytes and macrophages. In contrast, epithelial and endothelial cells, fibroblasts, and platelets are resistant to this action (8,17).
Due to the variation in the leukotoxin genes transcriptional regulation, A. actinomycetemcomtians have been classified into high and low leukotoxin-producing strains. The occurrence of high leucotoxin-producing A. actinomycetemcomtians strains has showed variations in different ethnic populations (12,16) and the presence of low leukotoxin-producing strains in patients with aggressive periodontal disease, as well as, in healthy subjects (5,26) suggest that genetic and environmental factors may interfere with the leukotoxin expression and the host's response (10). Therefore, in this study, the biotypes distribution and the ltx gene presence in A. actinomycetemcomitans isolated from Brazilian patients with advanced periodontitis were determined.
MATERIAL AND METHODS
Patients and subgingival samples
Fifty patients with chronic periodontitis visiting the Clinic of Periodontology of the Dental School of the University of São Paulo, São Paulo, Brazil, and 50 periodontally healthy subjects were selected. Patients with periodontitis were 27 females and 23 males aged between 18 to 50 years old (mean age 37.72 ± 10.63), while healthy subjects were 30 females and 20 males aged from 18 to 35 years old (mean age 23.76 ± 5.79). All patients displayed > 25 teeth, did not require pre-medication with antibiotics for a periodontal examination, and were not under any periodontal therapy during the last 6 months. Periodontal patients and healthy subjects were submitted to a full-mouth periapical radiographic examination with a Kodak film (Ektaspeed plus). Patients with chronic periodontitis showed clinical and radiographic evidences of bone loss and periodontal pocket depth exceeding 5mm, while healthy subjects did not show evidences of bone loss and gingival inflammation. Exclusion criteria included pregnancy, history of self-medication, nursing, diabetes, autoimmune diseases and other systemic pathology.
Subgingival samples from both groups were obtained by using two sterile paper points (Dentsplay, Ind. Co. Ltd., RJ, Brazil) inserted to the apical portion of the periodontal pocket or gingival crevice for 60 s, and transported in VMGA III medium (23). Samples were plated, in duplicate, onto selective trypticase soy-serum-bacitracin-vancomycin agar (30) and after 72 h of incubation in anaerobiosis (90% N2 + 10% CO2) at 37ºC, characteristic colonies of A. actinomycetemcomitans were identified by biochemical methods (29). Ethic Committee of the Institute of Biomedical Sciences, University of São Paulo (ICB 087/CEP), approved this study.
Biotyping
The biotyping of all A. actinomycetemcomitans isolated was based on dextrin, maltose, mannitol and xylose fermentation (28).
DNA extraction
Bacterial DNA was obtained from clinical samples and from colonies. Subgingival samples were homogenized and 500 µl were transferred to tubes containing 500 µl of sterile Milli-Q water. After centrifugation at 14,000 x g for 10 min, the pellet was suspended in 300 µl of sterile Milli-Q water and boiled for 10 min. In addition, two colonies were picked from brain heart infusion agar supplemented with 0.5% yeast extract and 5% horse blood and resuspended in 500 µl of sterile Milli-Q water, homogenized and boiled for 10 min (2). Finally, both clinical samples and colonies after boiling were harvested by centrifugation at 14,000 x g for 10 min, and the supernatants were stored at -20ºC.
PCR assay
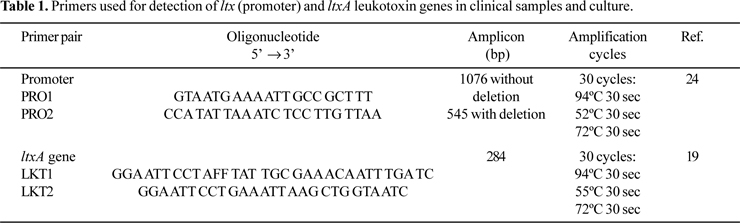
PCR amplification for detecting A. actinomycetemcomitans from clinical samples was performed in final volumes of 25 µl, containing 10 X PCR buffer, 2.5 mM MgCl2, 200 µM dNTP (Invitrogen do Brasil, São Paulo, Brazil), 1.5 U Platinum Taq DNA polymerase (Invitrogen), 0.4 µM each primer and 10 ng of DNA template. In all the PCR reactions this mixture was used. Table 1 shows the primer pairs used in PCR reactions. Amplifications were performed in a Perkin Elmer Amp PCR System 9700. The temperature profile included an initial step of 94ºC for 5 min, followed by 30 cycles of 94ºC for 30 s, annealing temperature of primer pair for 30 s, and 72ºC for 30 s and a final step of 72ºC for 5 min (Table 1).
Twenty microliters of the amplified products were analyzed by electrophoresis in 1% agarose gel. Gels were stained with 0.5 µg/ml ethidium bromide and photographed under UV transiluminator with a Kodak Eletrophoresis Documentation and Analyses System-120. A. actinomycetemcomitans JP2 with a 530-bp deletion was used a positive control, and A. actinomycetemcomitans ATCC 33383 as negative control.
Leukotoxin assay
Leukotoxin activity was determined by using trypan blue exclusion method, with modifications (14). Briefly, promyelocytic leukemic cell line HL-60 was grown in RPMI 1640 medium with 10% fetal bovine serum (FBS), penicillin G (100 UI/ml), and streptomycin (100 µg/ml) in an atmosphere of 5% CO2 at 37ºC. 100 µl of bacterial inoculum (106 UFC) or supernatant from A. actinomycetemcomitans cultures were added and incubated at 37ºC for 2 h. The proportion of HL-60 cells and bacterial cells was estimated in 1/100.
HL-60 cells without bacteria or supernatant were used as negative controls. The assay was performed in microtubes without antimicrobials. After incubation, 100 µl of 0.4% trypan blue were added to mixtures, placed on hemocytometer and then observed under a light microscope. The cells taking up the trypan blue (death cells) and cells excluding the dye (viable cells) were counted in a Neubauer chamber. Percentages of lyses were determined by dividing the number of surviving cells by the number of cells in the negative control, and values were given as the average of assays. The percentage of viable cells as the average of five viability measurements, each requiring the scoring of 100 cells was determined. Measurements were repeated five times to assess the reproducibility of viability measurement.
Highly leukotoxic strains were able to lyses > 70%; strains showing medium toxicity produced cell lyses values from 70% to 50%; strains with low toxicity produced cell lyses values from 50% to 25%, and non-toxic strains < 25%. Highly producer A. actinomycetemcomitans JP2 was used as positive control, A. actinomycetemcomitans ATCC 43718 (Y4), a medium leukotoxin-producing strain, and A. actinomycetemcomitans SUNY AB 67 or ATCC 33383, non-leukotoxic strains, used as negative control.
Statistical analysis
The results were expressed as median, standard deviation and percentages. Chi-square (χ2) and multiple comparison tests were performed to check differences in detection of A. actinomycetemcomitans, with a significant level at 5%.
RESULTS
Bacterial detection and leukotoxin-producing strains
Probing depth in patients with periodontitis was 6.24 ± 1.39 mm, while healthy subjects presented probing depth of 1.78 ± 0.54. A total of 91 A. actinomycetemcomitans strains were obtained from 9 (18%) patients with periodontitis and three isolates were obtained from one (2%) healthy subject.
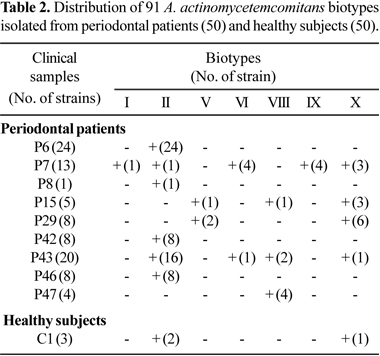
Seven biotypes from periodontal patients were identified, I, II, V, VI, VIII, IX and X. Biotype II was found in 12% of the periodontal patients and it was the most prevalent, followed by biotype X found in 8% of the patients. Only one healthy subject harbored three strains (two biotypes II and one X). Most of strains isolated from clinical samples generally belonged to the same biotype, but some clinical samples harbored from two to five biotypes as observed in Table 2.
This microorganism was detected in 44% of the periodontitis clinical samples and in 16% of the healthy subjects by using the LKT primer pairs. Moreover, by using the PRO primers, A. actinomycetemcomitans was detected in 6% of the clinical samples, but it was not observed in samples from healthy individuals. The presence of the highly leukotoxic strains showing the 530 bp deletion was verified in 13 (14.2%) of the 91 A. actinomycetemcomitans obtained from periodontitis patients. The three strains recovered from healthy subjects produced a 1.0 kb amplicon characteristic of low leukotoxin-producing strains. Fig. 1 shows highly leukotoxic and medium to low leukotoxic strains.
Leukotoxin assay
As presented in Table 3, some patients were colonized by strains producing different levels of leukotoxin. The toxic activity in most of the isolates showed to be cell bound, since supernatants of the highly leukotoxic strains did not produce significant damage on HL-60 cells.
DISCUSSION
Human subgingival microbiota displays a great heterogeneity in populations from different geographic locations (11). In this study, by culture, A. actinomycetemcomitans was isolated from 18% of the periodontal patients and 2% of the healthy subjects, in accordance with Asikainen et al. (1) and Malheiros and Avila-Campos (22). Moreover, studies have shown that the occurrence of A. actinomycetemcomitans varied from 20% to 80% in adult periodontitis patients (3,22,33). On the other hand, by using a PCR method, this microorganism was detected in 44% of the periodontal patients, and in 16% of the healthy subjects. Yang et al. (33) detected this microorganism in 64.4% and 64% of the patients with chronic periodontitis and healthy subjects, respectively. In Brazil, is observed a multi-ethnic population, with strong race mixture and peculiar habits, and the occurrence of different oral pathogens in dental or subgingival biofilms may reflects this characteristic.
In this study, A. actinomycetemcomitans strains were grouped in seven biotypes, but no relationship among any biotype and leukotoxin-producing strains or presence or absence of 530-bp deletion was observed (P = 0.278). Haubek et al. (13) have suggested that highly leukotoxic A. actinomycetemcomitans comprise a single clone, and in our study, only 13 strains isolated from a single patient were considered highly leukotoxic strains, which belonged to five different biotypes I, II, VI, IX or X.
In addition, biotype II and X were the most prevalent colonizing 12% and 8% of the periodontal patients, respectively. In healthy subjects, biotypes II, VI and X were observed. Few studies have shown the presence of biotypes in A. actinomycetemcomitans and the biotype II appears to be the most prevalent in Brazilian periodontal patients (3,22).
Some findings have suggested that highly leukotoxic strains may be important in the pathogenesis of the periodontal disease in certain populations (16). Previous reports have demonstrated a strong correlation between leukotoxic strains and periodontal breakdown (12), especially in younger patients with localized early bone loss (6,21).
Different populations seem to show highly variable frequency of the 530-bp leukotoxin promoter deletion (6,12,14,16), which is associated with the maximum leukotoxin expression. The frequency of this genetic deletion in the promoter is especially relevant, since subjects harboring highly leukotoxic A. actinomycetemcomitans are 22.5 times more likely to convert from health status to localized aggressive periodontitis than those colonized by strains harboring full-length leukotoxin promoter region (4). A strong correlation between highly leukotoxic strains of A. actinomycetemcomitans and periodontitis has been reported in Brazilian population (6).
Leukotoxin produced by A. actinomycetemcomitans is bound to bacterial and vesicle surfaces by weak interactions mediated by electrostatic forces or by hydrophobic protein epitopes (31). Data of the cytotoxicity assay on HL-60 cells are in accordance with this statement, since supernatant of cultures showed reduced cytolytic activities (15).
In summary, the diversity of leukotoxin expression by A. actinomycetemcomitans strains suggests the need to a better evaluation about the role of this leukotoxin in periodontal diseases and other infections, obtaining biological data to be considered in epidemiological studies.
ACKNOWLEDGEMENTS
The authors thank Mrs. Zulmira Alves de Souza. This study was supported by grants of Fundação do Amparo à Pesquisa do Estado de São Paulo (FAPESP) Proc. No. 00/07784-7 and 00/07582-5.
REFERENCES
1. Asikainen, S.; Alaluusua, S.; Kari, K.; Kleemola-Kujala, E. (1986). Subgingival microflora and periodontal conditions in healthy teenagers. J. Periodontol., 57, 505-509.
2. Avila-Campos, M.J.; Velásquez-Meléndez, G. (2002). Prevalence of putative periodontopathogens from periodontal patients and healthy subjects in São Paulo, SP, Brazil. Rev. Inst. Med. Trop. S. Paulo, 44, 1-5.
3. Avila-Campos, M.J.; Carvalho, M.A.R.; Zelante, F. (1995). Distribution of biotypes and antimicrobial susceptibility of Actinobacillus actinomycetemcomitans. OralMicrobiol. Immunol., 10, 382-384.
4. Bueno, I.C.; Mayer, M.P.A.; DiRienzo, J.M. (1998). Relationship between conversion of localized juvenile periodontitis-susceptible children from health to disease and Actinobacillus actinomycetemcomitans leukotoxin promoter structure. J.Periodontol., 70, 998-1007.
5. Contreras, A.; Rusitananta, T.; Chen, C.; Wagner, W.G.; Michalowicz, B.S.; Slots, J. (2000). Frequency of 530-bp deletion in Actinobacillusactinomycetemcomitans leukotoxin promoter region. Oral Microbiol. Immunol., 15, 338-340.
6. Cortelli, J.R.; Cortelli, S.C.; Jordan, S.; Haraszthy, V.I.; Zambon, J.J. (2005). Prevalence of periodontal pathogens in Brazilians with aggressive or chronic periodontitis. J. Clin. Periodontol., 32, 860-866.
7. Diaz, R.; Al Ghofaily, L.; Patel, J. Balashova, N.V.; Freitas, A.C.; Labib, I.; Kachlany, S.C. (2006). Characterization of leukotoxin from a clinical strain of Actinobacillusactinomycetemcomitans. Microb. Pathogen., 40, 48-55.
8. Fine, D.H.; Kaplan, J.B.; Kachlany, S.C.; Schreiner, H.C. (2006). How we got attached to Actinobacillus actinomycetemcomitans: a model for infectious diseases. Periodontology2000, 42, 114-157.
9. Fives-Taylor, P.; Meyer, D.; Mintz, K. (1996). Virulence factors of the periodontopathogen Actinobacillus actinomycetemcomitans. J. Periodontol., 67, 291-297.
10. Fong, K.P.; Chung, W.O.; Lamont, R.J.; Demuth, D.R. (2001) Intra- and interspecies regulation of gene expression by Actinobacillus actinomycetemcomitans. Infect. Immun., 69, 7625-7634.
11. Haffajee, A.D.; Bogren, A.; Hasturk, H.; Feres, M.; Lopez, N.J.; Socransky, S.S. (2004). Subgingival microbiota of chronic periodontitis subjects from different geographic locations. J. Clin. Periodontol., 31, 996-1002.
12. Haubek, D.; Dirienzo, J.M.; Tinoco, E.M.B.; Westergaard, J.; Lopez, N.J.; Chung, C.P.; Poulsen, K.; Kilian, M. (1997). Racial tropism of a highly toxic clone of Actinobacillus actinomycetemcomitans associated with juvenile periodontitis. J. Clin. Microbiol., 35, 3037-3042.
13. Haubek, D.; Havemose-Poulsen, A.; Westergaard, J. (2006). Aggressive periodontitis in a 16-year-old Ghanaian adolescent, the original source of Actinobacillus actinomycetemcomitans strain HK1651 - a 10-year follow up. Int. J. PediatricDent., 16, 370-375.
14. He, T.; Nishihara, T.; Demuth, D.R.; Ishikawa, J. (1999). A novel insertion sequence increases the expression of leukotoxin in Actinobacillus actinomycetemcomitans clinical isolates. J. Periodontol., 70, 1261-1268.
15. Johansson, A.; Claesson, R.; Hänström, L.; Kalfas, S. (2003). Serum-mediated release of leukotoxin from the cell surface of the periodontal pathogen Actinobacillusactinomycetemcomitans. Eur. J. Oral Sci., 111, 209-215.
16. Johansson, A.; Hänström, L.; Kalfas, S. (2000). Inhibition of Actinobacillusactinomycetemcomitans leukotoxicity by bacteria from the subgingival flora. Oral Microbiol. Immunol., 15, 218-225.
17. Kaplan, J.B.; Schreiner, H.C.; Furgang, D.; Fine, D.H. (2002) Population structure and genetic diversity of Actinobacillus actinomycetemcomitans strains isolated from localized juvenile periodontitis patients. J. Clin. Microbiol., 40, 1181-1187.
18. Kelk, P.; Johansson, A.; Claesson, R.; Hänström, L.; Kalfas, S. (2003). Caspase 1 involvement in human monocyte lyses induced by Actinobacillus actinomycetemcomitans leukotoxin. Infect. Immun., 71, 4448-4455.
19. Kraig, E.; Dailey, T.; Kolodrubetz, D. (1990). Nucleotide sequence of the leukotoxin gene from Actinobacillus actinomycetemcomitans: homology to the alpha-hemolysin/leukotoxin gene family. Infect. Immun., 58, 920-929.
20. Lally, E.T.; Hill, R.B.; Kieba, I.R.; Korostoff, J. (1999). The interaction between RTX toxins and target cells. TRENDS Microbiol., 7, 357-361.
21. Leung, W.K.; Ngai, V.K.S.; Yau, J.Y.Y.; Cheung, B.P.K.; Tsang, P.W.K.; Corbet, E.F. (2005). Characterization of Actinobacillus actinomycetemcomitans isolated from young Chinese aggressive periodontitis patients. J. Periodont. Res., 40, 258-268.
22. Malheiros, V.J.; Avila-Campos, M.J. (2004). Detection of pathogens from periodontal lesions. Rev. Saúde Públ., 38, 723-728.
23. Möller, A.J. (1966). Microbiol examination of root canals and periapical tissues of human teeth: methodological studies. Odontol. Tidskr., 74, 1-138.
24. Mombelli, A.; Gmür, R.; Lang, N.P.; Corbet, E.; Frey, F. (1999). Actinobacillus actinomycetemcomitans in Chinese adults. Serotype distribution and analysis of the leukotoxin gene promoter locus. J. Clin. Periodontol., 26, 505-510.
25. Mullally, B.H.; Dace, B.; Shelburne, C.E.; Wolf, L.F.; Coulter, W.A. (2000). Prevalence of periodontal pathogens in localized and generalized forms of early-onset periodontitis. J. Periodont. Res., 35, 232-241.
26. Müller, H.P.; Heinecke, A.; Fuhrmann, A.; Eger, T.; Zöller, L. (2001). Intraoral distribution of Actinobacillus actinomycetemcomitans in young adults with minimal periodontal disease. J. Periodont.Res., 36, 114-123.
27. Nakano, K.; Inaba, H.; Nomura, R.; Nemoto, H.; Tamura, K.; Miyamoto, E.; Yoshioka, H.; Taniguchi, K.; Amano, A.; Ooshima, T. (2007). Detection and serotype distribution of Actinobacillusactinomycetemcomitans in cardiovascular specimens from Japanese patients. Oral Microbiol. Immunol., 22, 136-139.
28. Slots, J.; Reynolds, H.S.; Genco, R.J. (1980). Actinobacillus actinomycetemcomitans in human periodontal disease: a cross-sectional microbiological investigation. Infect. Immun., 29, 1013-1020.
29. Slots, J. (1982). Salient biochemical characteristic of Actinobacillus actinomycetemcomitans. Arch. Microbiol., 131, 60-67.
30. Slots, J. (1982). Selective medium for Actinobacillus actinomycetemcomitans. J. Clin. Microbiol., 15, 606-609.
31. Tonjun, T.; Haas, R. (1993). Identification of Actinobacillusactinomycetemcomitans by leukotoxin gene-specific hybridization and polymerase chain reaction assays. J. Clin. Microbiol., 31, 1856-1859.
32. Wu, Y.M.; Yan, J.; Chen, L.L.; Gu, Z.Y. (2007). Association between infection of different strains of Porphyromonas gingivalis and Actinobacillusactinomycetemcomitans in subgingival plaque and clinical parameters in chronic periodontitis. J. Zhejiang Univ. Science B, 8, 121-131.
33. Yang, H.W.; Huang, Y.F.; Chan, Y.; Chou, M.Y. (2005). Relationship of Actinobacillusactinomycetemcomitans serotypes to periodontal condition: prevalence and proportions in subgingival plaque. Eur. J. Oral Sci., 113, 28-33.
Submitted: November 04, 2007; Returned to authors for corrections: February 25, 2007; Approved: October 22, 2008
- 1. Asikainen, S.; Alaluusua, S.; Kari, K.; Kleemola-Kujala, E. (1986). Subgingival microflora and periodontal conditions in healthy teenagers. J. Periodontol., 57, 505-509.
- 2. Avila-Campos, M.J.; Velásquez-Meléndez, G. (2002). Prevalence of putative periodontopathogens from periodontal patients and healthy subjects in São Paulo, SP, Brazil. Rev. Inst. Med. Trop. S. Paulo, 44, 1-5.
- 3. Avila-Campos, M.J.; Carvalho, M.A.R.; Zelante, F. (1995). Distribution of biotypes and antimicrobial susceptibility of Actinobacillus actinomycetemcomitans OralMicrobiol. Immunol., 10, 382-384.
- 4. Bueno, I.C.; Mayer, M.P.A.; DiRienzo, J.M. (1998). Relationship between conversion of localized juvenile periodontitis-susceptible children from health to disease and Actinobacillus actinomycetemcomitans leukotoxin promoter structure. J.Periodontol, 70, 998-1007.
- 5. Contreras, A.; Rusitananta, T.; Chen, C.; Wagner, W.G.; Michalowicz, B.S.; Slots, J. (2000). Frequency of 530-bp deletion in Actinobacillusactinomycetemcomitans leukotoxin promoter region. Oral Microbiol. Immunol., 15, 338-340.
- 6. Cortelli, J.R.; Cortelli, S.C.; Jordan, S.; Haraszthy, V.I.; Zambon, J.J. (2005). Prevalence of periodontal pathogens in Brazilians with aggressive or chronic periodontitis. J. Clin. Periodontol., 32, 860-866.
- 7. Diaz, R.; Al Ghofaily, L.; Patel, J. Balashova, N.V.; Freitas, A.C.; Labib, I.; Kachlany, S.C. (2006). Characterization of leukotoxin from a clinical strain of Actinobacillusactinomycetemcomitans Microb. Pathogen., 40, 48-55.
- 8. Fine, D.H.; Kaplan, J.B.; Kachlany, S.C.; Schreiner, H.C. (2006). How we got attached to Actinobacillus actinomycetemcomitans: a model for infectious diseases. Periodontology2000, 42, 114-157.
- 9. Fives-Taylor, P.; Meyer, D.; Mintz, K. (1996). Virulence factors of the periodontopathogen Actinobacillus actinomycetemcomitans J. Periodontol., 67, 291-297.
- 10. Fong, K.P.; Chung, W.O.; Lamont, R.J.; Demuth, D.R. (2001) Intra- and interspecies regulation of gene expression by Actinobacillus actinomycetemcomitans. Infect. Immun, 69, 7625-7634.
- 11. Haffajee, A.D.; Bogren, A.; Hasturk, H.; Feres, M.; Lopez, N.J.; Socransky, S.S. (2004). Subgingival microbiota of chronic periodontitis subjects from different geographic locations. J. Clin. Periodontol., 31, 996-1002.
- 12. Haubek, D.; Dirienzo, J.M.; Tinoco, E.M.B.; Westergaard, J.; Lopez, N.J.; Chung, C.P.; Poulsen, K.; Kilian, M. (1997). Racial tropism of a highly toxic clone of Actinobacillus actinomycetemcomitans associated with juvenile periodontitis. J. Clin. Microbiol., 35, 3037-3042.
- 13. Haubek, D.; Havemose-Poulsen, A.; Westergaard, J. (2006). Aggressive periodontitis in a 16-year-old Ghanaian adolescent, the original source of Actinobacillus actinomycetemcomitans strain HK1651 - a 10-year follow up. Int. J. PediatricDent., 16, 370-375.
- 14. He, T.; Nishihara, T.; Demuth, D.R.; Ishikawa, J. (1999). A novel insertion sequence increases the expression of leukotoxin in Actinobacillus actinomycetemcomitans clinical isolates. J. Periodontol., 70, 1261-1268.
- 15. Johansson, A.; Claesson, R.; Hänström, L.; Kalfas, S. (2003). Serum-mediated release of leukotoxin from the cell surface of the periodontal pathogen Actinobacillusactinomycetemcomitans Eur. J. Oral Sci, 111, 209-215.
- 16. Johansson, A.; Hänström, L.; Kalfas, S. (2000). Inhibition of Actinobacillusactinomycetemcomitans leukotoxicity by bacteria from the subgingival flora. Oral Microbiol. Immunol., 15, 218-225.
- 17. Kaplan, J.B.; Schreiner, H.C.; Furgang, D.; Fine, D.H. (2002) Population structure and genetic diversity of Actinobacillus actinomycetemcomitans strains isolated from localized juvenile periodontitis patients. J. Clin. Microbiol., 40, 1181-1187.
- 18. Kelk, P.; Johansson, A.; Claesson, R.; Hänström, L.; Kalfas, S. (2003). Caspase 1 involvement in human monocyte lyses induced by Actinobacillus actinomycetemcomitans leukotoxin. Infect. Immun, 71, 4448-4455.
- 19. Kraig, E.; Dailey, T.; Kolodrubetz, D. (1990). Nucleotide sequence of the leukotoxin gene from Actinobacillus actinomycetemcomitans: homology to the alpha-hemolysin/leukotoxin gene family. Infect. Immun., 58, 920-929.
- 20. Lally, E.T.; Hill, R.B.; Kieba, I.R.; Korostoff, J. (1999). The interaction between RTX toxins and target cells. TRENDS Microbiol., 7, 357-361.
- 21. Leung, W.K.; Ngai, V.K.S.; Yau, J.Y.Y.; Cheung, B.P.K.; Tsang, P.W.K.; Corbet, E.F. (2005). Characterization of Actinobacillus actinomycetemcomitans isolated from young Chinese aggressive periodontitis patients. J. Periodont. Res., 40, 258-268.
- 22. Malheiros, V.J.; Avila-Campos, M.J. (2004). Detection of pathogens from periodontal lesions. Rev. Saúde Públ., 38, 723-728.
- 23. Möller, A.J. (1966). Microbiol examination of root canals and periapical tissues of human teeth: methodological studies. Odontol. Tidskr., 74, 1-138.
- 24. Mombelli, A.; Gmür, R.; Lang, N.P.; Corbet, E.; Frey, F. (1999). Actinobacillus actinomycetemcomitans in Chinese adults. Serotype distribution and analysis of the leukotoxin gene promoter locus. J. Clin. Periodontol, 26, 505-510.
- 25. Mullally, B.H.; Dace, B.; Shelburne, C.E.; Wolf, L.F.; Coulter, W.A. (2000). Prevalence of periodontal pathogens in localized and generalized forms of early-onset periodontitis. J. Periodont. Res., 35, 232-241.
- 26. Müller, H.P.; Heinecke, A.; Fuhrmann, A.; Eger, T.; Zöller, L. (2001). Intraoral distribution of Actinobacillus actinomycetemcomitans in young adults with minimal periodontal disease. J. Periodont.Res, 36, 114-123.
- 27. Nakano, K.; Inaba, H.; Nomura, R.; Nemoto, H.; Tamura, K.; Miyamoto, E.; Yoshioka, H.; Taniguchi, K.; Amano, A.; Ooshima, T. (2007). Detection and serotype distribution of Actinobacillusactinomycetemcomitans in cardiovascular specimens from Japanese patients. Oral Microbiol. Immunol., 22, 136-139.
- 28. Slots, J.; Reynolds, H.S.; Genco, R.J. (1980). Actinobacillus actinomycetemcomitans in human periodontal disease: a cross-sectional microbiological investigation. Infect. Immun., 29, 1013-1020.
- 29. Slots, J. (1982). Salient biochemical characteristic of Actinobacillus actinomycetemcomitans Arch. Microbiol., 131, 60-67.
- 30. Slots, J. (1982). Selective medium for Actinobacillus actinomycetemcomitans. J. Clin. Microbiol., 15, 606-609.
- 31. Tonjun, T.; Haas, R. (1993). Identification of Actinobacillusactinomycetemcomitans by leukotoxin gene-specific hybridization and polymerase chain reaction assays. J. Clin. Microbiol., 31, 1856-1859.
- 32. Wu, Y.M.; Yan, J.; Chen, L.L.; Gu, Z.Y. (2007). Association between infection of different strains of Porphyromonas gingivalis and Actinobacillusactinomycetemcomitans in subgingival plaque and clinical parameters in chronic periodontitis. J. Zhejiang Univ. Science B, 8, 121-131.
- 33. Yang, H.W.; Huang, Y.F.; Chan, Y.; Chou, M.Y. (2005). Relationship of Actinobacillusactinomycetemcomitans serotypes to periodontal condition: prevalence and proportions in subgingival plaque. Eur. J. Oral Sci, 113, 28-33.
Publication Dates
-
Publication in this collection
06 Apr 2009 -
Date of issue
Dec 2008
History
-
Accepted
25 Feb 2007 -
Received
04 Nov 2007